Abstract
Studying tissue composition and function in non-human primates (NHPs) is crucial to understand the nature of our own species. Here we present a large-scale cell transcriptomic atlas that encompasses over 1 million cells from 45 tissues of the adult NHP Macaca fascicularis. This dataset provides a vast annotated resource to study a species phylogenetically close to humans. To demonstrate the utility of the atlas, we have reconstructed the cell–cell interaction networks that drive Wnt signalling across the body, mapped the distribution of receptors and co-receptors for viruses causing human infectious diseases, and intersected our data with human genetic disease orthologues to establish potential clinical associations. Our M. fascicularis cell atlas constitutes an essential reference for future studies in humans and NHPs.
Similar content being viewed by others
Main
A major obstacle to building a comprehensive human cell atlas1 is obtaining a full range of ‘high-quality’ samples of sufficient size. Given their evolutionary proximity, non-human primates (NHPs) represent the nearest-to-human alternative. Generating an NHP cell atlas (NHPCA) would produce a catalogue of features that could be used to study human physiology, disease and ageing. It would also provide insights into the evolutionary mechanisms underlying different body function between NHPs and humans.
NHPs comprise a large and diverse group of species with major ecological, dietary, locomotor and behavioural differences2. Because of their characteristics, including a more frequent reproductive cycle and wide availability, macaques, in particular Macaca fascicularis (also known as cynomolgus, crab-eating or long-tailed monkey), are now used for research purposes worldwide3. Here we used adult M. fascicularis tissues to generate the largest NHP cell transcriptomic dataset thus far. To facilitate exploration of this resource, we have created the NHPCA website, an open and interactive database (https://db.cngb.org/nhpca/).
Generation of an adult monkey cell atlas
We isolated cells/nuclei for 45 different tissue samples from five male and three female 6-year-old monkeys (Fig. 1a, Supplementary Fig. 1 and Supplementary Table 1a). Most tissues were profiled by single-nucleus RNA sequencing (snRNA-seq), which circumvents complications associated with dissociation protocols and allowed us to profile frozen samples, but for some tissues we used single-cell RNA sequencing (scRNA-seq). The lymph node was profiled using both scRNA-seq and snRNA-seq for comparison. All experiments used the DNBelab C4 droplet-based platform for library generation4. After filtering, we retained transcriptomic data for a total of 1,144,706 cells/nuclei (Fig. 1a), with numbers ranging from 84,619 in the cerebellum to 2,694 in the vagina (Supplementary Table 1a). For lymph node, the comparison between scRNA-seq and snRNA-seq identified a similar number of genes and unique molecular identifiers (UMIs) (Supplementary Fig. 2a–f). Likewise, cell cluster integration showed a good match between the two methods, although snRNA-seq was more efficient at capturing less abundant cell types. These results confirm the utility of snRNA-seq for generating large-scale cell atlases5,6.
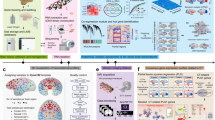
a, Left, schematic representation of the monkey tissues analysed in this study. The cartoons used to generate this schematic diagram were purchased from BioRender.com. A total of 45 tissues were collected from 3 female and 5 male 6-year-old monkeys. A UMAP visualization is shown of global clustering of all cells from the dataset coloured by tissue (middle), and bar plots show the number of cells/nuclei profiled for each tissue after quality control (right). n = 1,144,706 individual cells/nuclei analysed. b, UMAP visualization of all clusters coloured by major cell types. A total of 113 cell clusters were identified in the dataset. Cell type annotation for all major clusters is provided in the legend to the right; NKT, natural killer T; OPC, oligodendrocyte progenitor cell. Source data
In global visualization of cell clustering using uniform manifold approximation and projection (UMAP), each tissue tended to cluster separately, with those from the same system generally clustering more closely to each other (Fig. 1a and Supplementary Figs. 3–7). On the basis of the expression levels of specific markers (Supplementary Fig. 8), we defined 113 cell clusters in the global UMAP view of all tissues (Fig. 1b and Supplementary Table 1b, c). On average, we detected 1,445 genes and 2,583 UMIs per cell/nucleus (Supplementary Fig. 9). The number of cells for each of these 113 cell types ranged from 76,602 for granule cells in the cerebellum to 21 for oligodendrocytes in the pineal gland (Supplementary Fig. 10). Reassuringly, many of the 113 clusters were largely composed of a cell type belonging to a specific tissue (Fig. 1b and Supplementary Fig. 11a). However, cell types such as endothelial, stromal and various immune cells were shared between different tissues, as expected (Supplementary Fig. 11b). We next generated individual UMAP representations for each tissue and applied unbiased graph-based Seurat clustering, identifying 463 cell clusters across all tissues. A detailed annotation of the cell populations detected in each tissue is provided in Supplementary Figs. 12–15 and Supplementary Table 1d, e. Our M. fascicularis atlas can be searched interactively by tissue, cell type and gene through the NHPCA website.
To demonstrate the potential for cross-species comparisons, we selected a total of 12 NHP tissues overlapping with single-cell mouse (Mouse Cell Atlas, MCA) and human (Human Cell Landscape, HCL) cell atlases7,8 (Supplementary Figs. 16–19 and Supplementary Table 1f). Cell numbers as well as gene and UMI capture rates were higher in NHPCA for all 12 tissues. We observed good correlation of tissue marker genes with both the mouse and human datasets in all cases. Likewise, the number of detected main cell types was roughly comparable in the three species (111 in monkeys, 110 in mice and 106 in humans), but with differences in the proportions. For example, over 80% of liver cells detected in monkeys corresponded to hepatocytes, in line with the normal proportion of 60–80% in this tissue9,10, but only 3% and 6.7% of corresponding cells were hepatocytes in human and mouse liver, respectively. This discrepancy might be related to a bias in cell population capture when using different platforms or the use of nuclei versus whole cells. We performed immunostaining of monkey liver sections for the hepatocyte marker albumin, observing as expected that most cells were positive (Supplementary Fig. 20a). Differentially expressed genes (DEGs) between specific tissue cell populations in the three datasets can be examined using our website. As proof of principle of the application for studying body-wide cell–cell interactions, we examined the distribution of insulin and glucagon receptors throughout the 12 tissues (Supplementary Fig. 20b). Although the patterns were similar, species-specific differences were observed. Additional ligand–receptor interactions in each of the 45 monkey tissues and the comparison between species for the 12 shared tissues can also be explored using our website, and we have provided an option for uploading individual tissue datasets to enable customized comparisons.
Common cell types across tissues
We inspected common cell types populating different tissues throughout the monkey body8,11,12,13. First, we selectively combined and reclustered stromal cells, macrophages (including microglia), endothelial cells and smooth muscle cells from all analysed tissues. Although considerable diversity was observed, many cell clusters grouped together on the basis of tissue origin (Supplementary Fig. 21a–d). We also performed DEG analysis to obtain tissue-specific signatures, identifying substantial heterogeneity (Supplementary Fig. 21e–h and Supplementary Table 2a–d).
Notably, our snRNA-seq data offer the possibility of studying cell populations that cannot be characterized by conventional scRNA-seq analysis, such as myonuclei from multinucleated skeletal muscle fibres. We combined and reclustered cells from tissues in our atlas known to contain skeletal muscle cells. This approach identified distinct populations in the abdominal wall, diaphragm and tongue, whereas nuclei from the oesophagus were more homogenous (Fig. 2a). Myonuclei in the abdominal wall, diaphragm and tongue comprised MYH7+ type I (slow-twitch) and MYH2+ type II (fast-twitch) myofibres14 (Fig. 2b, c and Supplementary Table 2e–g). Differential thresholds of MYH2 and GPD2 expression further subdivided type II myonuclei into type IIa (MYH2high) and type IIb (MYH2lowGPD2+) myonuclei. In line with previous reports, we did not detect type IIb myonuclei in the tongue15. Moreover, type I and type IIa tongue myonuclei clustered in close proximity, which may be related to the tongue being a highly innervated muscle.
a, UMAP visualization of global clustering of skeletal muscle cells. Clusters are coloured by tissue (abdominal wall, diaphragm, oesophagus and tongue). b, UMAP representation of all reclustered skeletal muscle cells coloured by subtype. c, UMAP visualization of specific markers used to identify type I (MYH7), type IIa (MYH2) and type IIb (GPD2) myonuclei, FAPs (LVRN), MTJ nuclei (NAV3 and COL22A1), NMJ nuclei (ETV5 and MUSK) and satellite cells (PAX7), as shown in b. Because of their small proportions, the latter four populations are indicated by a red arrow. d, Stacked bar plots representing the proportions of skeletal muscle nuclei (myonucleus subtypes type I, type IIa and type IIb, MTJ and NMJ nuclei, and satellite cells and FAPs) in the indicated tissues. e, Heat map showing DEGs among the skeletal muscle populations highlighted in d. f, Bubble plots showing DEGs for each of the myonucleus subtypes comparing different tissues. g, UMAP visualization of mesothelial (meso) cells from selected tissues (adrenal gland, bladder, diaphragm, fallopian tube, ovary and visceral adipose tissue). Four different clusters of mesothelial cells belonging to the visceral adipose tissue are indicated by the dashed red line. h, Violin plots showing the differential expression of mesothelial and immune markers in the visceral adipose tissue clusters highlighted by the dashed red line in g. i, UMAP visualization of three different clusters of mesothelial cells from the ovary. Mesothelial cells, surface epithelial (surface epi) cells and progenitor-like epithelial (prog-like epi) cells are highlighted in red, blue and yellow, respectively. j, UMAP visualization of LGR5 expression in ovarian mesothelial cells. k, Violin plots showing DEGs among the three populations of ovarian mesothelial cells highlighted in the UMAP visualization. Source data
Differential thresholds of MYH2 and GPD2 expression further subdivided type II myonuclei into type IIa (MYH2high) and type IIb (MYH2lowGPD2+). In addition, we discriminated, albeit at low proportions, NAV3+ neuromuscular junction (NMJ) nuclei in the diaphragm and ETV5+ myotendinous junction (MTJ) nuclei in both the tongue and diaphragm (Fig. 2b–d). Moreover, we detected PAX7 + nuclei from satellite cells in the tongue and diaphragm, while a small cluster of LVRN+ fibroadipogenic progenitors (FAPs) could be annotated in the diaphragm, abdominal wall and oesophagus. Skeletal muscle nuclei exhibited subtype-specific and tissue-specific gene expression signatures and Gene Ontology (GO) terms (Fig. 2e, f and Supplementary Fig. 22a–c). We also observed substantial myonucleus heterogeneity within the same subtype and tissue (Fig. 2f).
Next, to study the heterogeneity among adipocytes, we combined and reclustered cells from subcutaneous and visceral adipose tissues, resulting in nine major clusters (Supplementary Fig. 23a–d). We noticed a marked distinction between mature adipocytes and putative adipocyte progenitors, as reflected by differential expression of ADIPOQ and CD34. Subcutaneous mature adipocytes and adipocyte progenitors were enriched for FOS expression. Likewise, SLC11A1 and SPOCK3 marked mature subcutaneous and visceral adipocytes, respectively. Adipocyte progenitors were composed of two populations for visceral tissue (WT1+ITLN1+ and CFDhighWT1lowITLN1–), three populations for subcutaneous tissue (ESR1+, CXCL14+APOD+ and DPP4+) and one population shared by both tissues (NOX4+). These results are consistent with markers described in previous reports16,17,18,19. We validated coexpression of CD34 and NOX4 in a subset of adipocyte progenitors of both subcutaneous and visceral adipose tissue by immunostaining (Supplementary Fig. 24a, b). Pseudotime analysis characterized the trajectory of adipocyte maturation from progenitors in both subcutaneous and visceral adipose tissue (Supplementary Fig. 24c, d). We did not detect substantial proliferation in any of the progenitor populations on the basis of expression of the pan-cycling marker MKI67 (Supplementary Fig. 23c), suggesting that these populations are not transitory.
Finally, we combined and reclustered all tissues that contained mesothelial cells, a type of specialized epithelial cell. Mesothelial cells from the bladder, ovary and fallopian tube were in close proximity, whereas those from other tissues clustered more separately (Fig. 2g). We also detected within-tissue mesothelial cell heterogeneity, in particular for visceral adipose tissue and ovary. In the former, we observed a cluster of immune-like mesothelial cells that, apart from expression of the typical mesothelial markers (MSLN, ITLN1 and PKHD1L1), also expressed high levels of immune cell markers (for example, PTPRC, IL7R and TRAC) (Fig. 2h). This is in agreement with the emerging concept that structural cells have immune properties8,11 and the known immunomodulatory role of the visceral adipose tissue in responses to gut bacteria20. In the ovary, we identified a classical mesothelial population and two close PAX8+ (ref. 21) epithelial-like populations (one mature and one progenitor-like) of mesothelial origin (Fig. 2i–k). Progenitor-like ovarian epithelial cells have previously been reported22. In line with previous work, we observed that they expressed well-known stem cell markers such as LGR5 (ref. 22) and CD44 (ref. 23). Immunostaining for CD44 and single-molecule fluorescence in situ hybridization (smFISH) for LGR5 confirmed their coexpression in a subset of monkey surface epithelial cells (Supplementary Fig. 24e). Pseudotime analysis reconstructed the trajectory from progenitor-like cells to ovarian epithelial cells (Supplementary Fig. 24f). As in adipose tissue, we did not detect substantial proliferation in progenitor-like ovarian epithelial cells on the basis of expression of MKI67.
These findings substantially add up with previous studies of common cell type heterogeneity and tissue-specific molecular signatures8,11,12,13. Our dataset provides a new interactive resource for further dissecting these, clarifying the underlying mechanisms and studying interspecies differences.
Wnt signalling components in tissues
A cell body atlas of large dimensions is ideal for investigating multifaceted cell–cell interactions, including those occurring in cytokine or growth factor-mediated signalling pathways. Apart from having essential roles in embryonic development, Wnt factors control growth and maintenance of numerous tissues throughout life. We thus performed a survey of Wnt pathway24 components throughout the monkey body to thoroughly dissect target cells and potentially identify previously unappreciated populations.
LGR proteins (LGR4, LGR5 and LGR6) act as amplifiers of Wnt signals by inhibiting negative regulators25. Accordingly, LGR5 and LGR6 often mark and regulate cells with homeostatic or adult stem cell function in specific mammalian tissues, whereas LGR4 has a less well-understood function26. We observed expression of LGR5 across multiple monkey tissues, with the highest levels in type I skeletal muscle myonuclei, epithelial cells of the uterus and fallopian tube, oligodendrocyte progenitor cells (OPCs) and kidney tubule cells (Fig. 3a). To the best of our knowledge, with the exception of epithelial cells in the uterus and fallopian tube25, these tissues have not previously been reported to contain substantial numbers of LGR5+ cells in adult mammals. In this regard, it is worth noting that the majority of reports of LGR5+ cells thus far have been in genetically engineered mouse models owing to the lack of specific tools and reagents to study other mammals25. The expression of LGR6 was more restricted (Supplementary Fig. 25a), with higher abundance in cardiomyocytes, thyroid follicular cells, folliculostellate cells of the pituitary gland and, as previously reported, smooth muscle cells27. We also detected LGR5+ or LGR6+ cells in other tissues, including in both previously reported (for example, ovary epithelial cells22, hepatocytes28 and colon enterocytes29) and unreported (for example, LGR5+ cells in bipolar cells of the neurosensory retina) tissues (Supplementary Figs. 26–30 and Supplementary Table 3). In general, expression of LGR5 and LGR6 did not overlap, apart from in fallopian tube epithelial cells and gallbladder smooth muscle cells (Supplementary Fig. 25b). Moreover, there was little overlap between LGR5+ or LGR6+ cells with those expressing MKI67, apart from epithelial cells of the fallopian tube and uterus and basal cells from the salivary gland. In contrast to LGR5 and LGR6, LGR4 was ubiquitously expressed across most tissues (Supplementary Fig. 25c).
a, Top, global UMAP visualization of LGR5 expression across all tissues. Bottom, bubble plot showing the LGR5 expression level and ratio in the indicated cell types. b, Co-embedding of kidney snRNA-seq (blue) and scATAC-seq (red) datasets. c, Integrated kidney snRNA-seq and scATAC-seq data. Cell clusters are coloured by cell type. DCTC, distal convoluted tubule cell; Endo, endothelial cell; LOH, loop of Henle; Myofibro, myofibroblast. d, UMAP visualization of LGR5 across kidney cell types (top) and ArchR track visualization of aggregated scATAC-seq signal at the LGR5 locus in each cell type (bottom). The bar plots on the right indicate the ratio of LGR5+ cells in kidney cells. e, Representative images from smFISH detection of LGR5 (red) and SLC12A3 (yellow) expression in kidney. Scale bars, 50 μm (left and middle) and 20 μm (right). The right panel represents a magnification of the area indicated by the white boxes in the left and middle panels. f, UMAP visualization of muscle cells clustered by tissue (abdominal wall, aorta, bladder, carotid, diaphragm, fallopian tube, heart, oesophagus, ovary, prostate, salivary gland, spermaduct, testis, tongue, uterus and vagina). The dashed lines encompass clusters of cells belonging to a specific muscle type (cardiac, skeletal or smooth muscle). g, UMAP visualizations of LGR5, MYH2 and MYH7 across skeletal muscle cell types. The dashed line in the left panel indicates clusters belonging to the diaphragm; the one in the right panel indicates LGR5+MYH7 + cells. h, Representative images from smFISH detection of LGR5, MYH7 and their coexpression in skeletal myonuclei of the diaphragm. Scale bar, 20 μm. The panel at the bottom is a magnification of the area indicated by the white box in the adjacent panel; scale bar, 40 μm. Source data
In the kidney, LGR5+ cells were mostly enriched in the distal convoluted tubule (DCT) and, to a less extent, in the descending and ascending loop of Henle (Fig. 3a). To support this observation, we performed single-cell assay for transposase-accessible chromatin with sequencing (scATAC-seq) of monkey kidney and integrated the results with our snRNA-seq dataset (Fig. 3b, c and Supplementary Fig. 31a, b). The analysis showed peaks of open chromatin at both the LGR5 promoter and a putative enhancer in cell types expressing LGR5 (Fig. 3d). Double smFISH for LGR5 and the DCT cell (DCTC) marker SLC12A3 confirmed coexpression of both genes in a substantial proportion of DCTCs, but showed little or no expression in other cell types (Fig. 3e). To study potential interspecies differences in the Wnt pathway, we merged our monkey kidney data with adult human8,30,31 and mouse7,32,33 kidney snRNA-seq and scRNA-seq datasets. Interestingly, there was lower LGR5 expression in adult human and mouse kidneys, including in DCTCs, than in monkey (Supplementary Fig. 32a–c). The finding in mice is consistent with the low levels of Lgr5 detected in adult mouse kidney using reporter mice or FISH probes34. We also performed a head-to-head comparison of DCTC gene expression, which showed that interspecies differences extend beyond LGR5 (Supplementary Fig. 32d, e and Supplementary Table 4).
In the neocortex, cell cluster integration of available human35 and mouse snRNA-seq datasets with the monkey data indicated differential LGR5 expression patterns between species. LGR5 was highest in OPCs in monkeys and in oligodendrocytes in humans, whereas in mice it was higher in inhibitory neurons than in OPCs and oligodendrocytes (Supplementary Fig. 33a–c). Pseudotime analysis showed high LGR5 abundance along the OPC maturation trajectory towards oligodendrocytes in monkey OPCs (Supplementary Fig. 33d, e). Double immunostaining for the OPC marker PDGFRA and LGR5 confirmed their coexpression in OPCs from monkey neocortex (Supplementary Fig. 33f). We also combined and reclustered all types of muscle cells in our atlas (Fig. 3f). LGR5 was more enriched in MYH7 + slow-twitch myonuclei of the abdominal wall and diaphragm (Fig. 3g), whereas LGR6 was higher in cardiomyocytes and smooth muscle cells (aorta, ovary, carotid and vagina) (Supplementary Fig. 34a). LGR5 and LGR6 expression in slow-twitch skeletal myonuclei and cardiomyocytes, respectively, was validated by smFISH (Fig. 3h and Supplementary Fig. 34b). In mice, Lgr5 is known to be expressed in NMJ myonuclei36 and a subset of satellite cells activated following injury37, but we did not detect enrichment of LGR5 in either cell type in our monkey dataset (Supplementary Figs. 26 and 27). The lack of enrichment in satellite cells is unsurprising given that we did not apply any injury before obtaining the skeletal muscle tissues. Yet, we could detect LGR6 in mouse and human cardiomyocytes using previously reported snRNA-seq datasets38,39 (Supplementary Fig. 34c, d). Similarly, LGR6 was enriched in several monkey pituitary cell populations, with the highest expression in folliculostellate cells, which have been reported to be pituitary gland stem cells40 (Supplementary Fig. 34e). In line with this, these cells also showed expression of other progenitor markers such as SOX2, PAX6, CD44 and CXCR4 (Supplementary Fig. 34f). Moreover, DEGs specific to this LGR5+ population in comparison with other pituitary cells were enriched in GO terms related to development (Supplementary Fig. 34g).
Next, we examined the genes encoding Wnt factors and the R-spondin family (RSPO1–RSPO4) of ligands for LGR proteins25 in a panel of monkey tissues containing cells with high LGR5 and LGR6 (Supplementary Figs. 35a, b and 36–39). RSPO cytokine expression was widely distributed among tissues, but higher levels were found in mesenchymal-like cells (for example, smooth muscle cells of the epididymis, hepatic stellate cells and folliculostellate cells from the pituitary gland) and mesothelial cells (for example, of the diaphragm, fallopian tube and ovary). Of note, RSPO2 expression was high in inhibitory neurons from the neocortex (Supplementary Fig. 38a). The expression of Wnt factors was more limited and in general lower than that of RSPO cytokines, but we noticed high levels of WNT9B in principal cells and principal-like cells from the collecting duct in the kidney (Supplementary Fig. 35a, c), WNT2B in mesothelial cells from the fallopian tube (Supplementary Fig. 37a) and ovary (Supplementary Fig. 38c), and, as expected, WNT2 in endothelial cells from the liver41 (Supplementary Fig. 37c). WNT9B expression was lower in mouse7,32,33 and in particular human8,30,31 kidney snRNA-seq datasets than in monkey (Supplementary Fig. 35e). Supporting the monkey snRNA-seq data, scATAC-seq analysis of the WNT9B locus showed increased enhancer accessibility in monkey principal and principal-like cells (Supplementary Fig. 35d). High levels of WNT9B in these cells may be responsible for inducing LGR5 (a Wnt pathway target) in monkey DCTCs. In fact, Wnt factors are known to act predominantly on neighbouring cells24,42, and cells from the collecting duct and DCT are in closer physical proximity than other nephron structures (Supplementary Fig. 35f). We further analysed Wnt receptors and other co-receptors43 as well as the TCF family of transcription factors bound by β-catenin44 as a resource for exploration (Supplementary Figs. 35a, b and 36–39). Thus, Wnt and other signalling pathways can be explored in monkey tissues and compared between species using our NHPCA website.
Cell type vulnerability to viruses
To examine the utility of our atlas for advancing knowledge of disease pathogenesis, we first mapped the expression of the main viral receptors and co-receptors for a panel of 126 viruses, including respiratory pathogens, across all monkey tissues. As expected, NCAM1 (encoding the rabies virus receptor) was enriched in astrocytes, oligodendrocytes and neurons, in line with knowledge of this virus attacking the central nervous system45. CD46 (encoding the receptor for measles and herpes viruses) was enriched in epithelial cells from the bladder, cells from the female and male reproductive system, and liver endothelial cells (Fig. 4a, Supplementary Fig. 40 and Supplementary Table 5a).
a, Heat map showing the expression of entry receptors and related molecules for a selection of viruses (indicated on the right) in all cell types (indicated at the bottom). Definitions for abbreviations are provided in the Supplementary Note. b, UMAP visualizations of ACE2 (left) and TMPRSS2 (right) expression in all cell types. The bubble plot next to each UMAP plot shows the expression level and ratio of ACE2 and TMPRSS2 in the indicated cell types. c, UMAP projection of ACE2+TMPRSS2+ cells (highlighted in yellow). The bar plots on the right show the ratio of cells expressing both genes. d, Bubble plots showing the ratio and expression levels of ACE2 and TMPRSS2 in gallbladder, kidney, liver and lung in monkeys and humans. The colour of each bubble represents the level of expression, and the size indicates the proportion of expressing cells. e, Left, ArchR track visualization of aggregated scATAC-seq signal at the ACE2 locus in each of the annotated kidney cell types. Predicted binding of human transcription factors based on DNA sequence is shown in the corresponding open chromatin regions of ACE2. Right, bar plots indicating the ratio (%) of ACE2+ cells in each annotated cell type of the monkey kidney. Inset, UMAP visualization of ACE2 in the integrated scATAC-seq and snRNA-seq data from monkey kidney. The red dashed line demarcates the separation of proximal tubule S1 and S3 cells. Asc LOH, ascending loop of Henle cell; AT1, alveolar type 1 cell; CTC, connecting tubule cell; Desc LOH, descending loop of Henle cell; Hep, hepatocyte; Mono, monocyte; PTC, proximal tubule cell; SMC, smooth muscle cell. Source data
Given the current coronavirus disease 2019 (COVID-19) pandemic caused by severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2; ref. 46), we focused on the receptor for this virus, ACE2, and the serine protease TMPRSS2 (ref. 47) to assess their expression in monkey tissues. This knowledge offers the major advantage of studying COVID-19 pathogenesis in a species that is often used for modelling the disease48. Although lung is the predominantly affected tissue in humans, other tissues such as the kidney (especially proximal tubule cells) and liver are also affected, and clarifying the mechanisms of tissue targeting would improve understanding of disease course and transmissibility5,49. TMPRSS2 showed broad expression across multiple monkey tissues, whereas ACE2 was more restricted (Fig. 4b, Supplementary Figs. 41 and 42, and Supplementary Table 5b). The highest ACE2 expression was found in the gallbladder (mucous, endothelial, glandular and smooth muscle cells), Sertoli cells from the testis, kidney epithelial cells (mostly proximal tubule cells), the lung (ciliated, club and, in particular, alveolar type 2 (AT2) cells) and the liver (hepatocytes and especially cholangiocytes). ACE2 in these tissues was notably heterogeneous, suggesting that regulatory mechanisms fine-tune its expression levels. Double-positive (ACE2+TMPRSS2+) cells have a higher risk of infection by SARS-CoV-2 (refs. 5,47,50,51), and we noticed the largest numbers of these cells among monkey gallbladder cells, in agreement with reports of patients with COVID-19 developing acute cholecystitis52. Considerable coexpression was also observed in cells from the lung and kidney, with less overlap observed in other cell types such as bladder epithelial cells and pancreatic ductal and islet cells (Fig. 4c). We next performed a comparative analysis of ACE2 and TMPRSS2 expression in monkeys and humans8. Similar patterns were seen in liver in the two species, whereas more distinct patterns were observed in the gallbladder, kidney and lung (Fig. 4d).
For a representative tissue with substantial ACE2 levels and a substantial proportion of ACE2+TMPRSS2+ cells, we looked at integrated snRNA-seq and scATAC-seq data from monkey kidney. This analysis identified discrete peaks of open chromatin in the ACE2 promoter and enhancer regions, with the greatest signal in a population of proximal tubule cells containing the highest proportion of ACE2-expressing cells (Fig. 4e). Motif analysis of these peaks demonstrated enrichment in binding sites for STAT1, STAT3, FOXA1, JUNB and several interferon response factor (IRF) proteins. These transcription factors are targets of tissue-protective and innate immune responses mediated by interleukin (IL)-6, IL-1 and interferons53. In this regard, dysregulation of both IL-6 and IL-1β has been implicated in the pathogenesis of severe COVID-19 disease54. Thus, we investigated the coexpression of their receptors (IL6R, IL1R1 and IL1RAP) with ACE2 in monkey kidney, only observing good correlation with ACE2 expression in proximal tubule cells for IL6R (Supplementary Fig. 43a). This observation suggests a potential link between IL-6, STAT transcription factors and enhanced ACE2 levels that may either facilitate viral reservoirs or exacerbate COVID-19 disease progression owing to increased viral dissemination (Supplementary Fig. 43b). In addition to ACE2 and TMPRSS2, numerous other molecules have been implicated in facilitating SARS-CoV-2 binding to the cell surface or in COVID-19 pathogenesis55. Their expression or coexpression in monkey tissues, other associations and virus–host interactions, as well as interspecies differences, can be studied using our NHPCA website.
Mapping traits and diseases to cell types
We next assessed the potential effect of genetic variation linked to complex human traits and diseases in specific monkey body cell types by applying a large panel of genome-wide association studies (GWAS) to our NHPCA. We linked human single-nucleotide polymorphisms from 163 GWAS taken from the UK Biobank (https://nealelab.github.io/UKBB_ldsc/downloads.html) to orthologous coordinates in the monkey transcriptome to calculate the enrichment of traits across the genes expressed in each cell cluster annotated in our dataset (Fig. 5, Supplementary Fig. 44 and Supplementary Table 6a). As a general trend, we observed enriched heritability for neurological traits such as ‘schizophrenia’ and ‘depression’ in clusters corresponding to neural cells. Alzheimer’s disease traits were enriched in immune cells, in line with the knowledge that immune dysfunction contributes to the pathogenesis of this disease56. In line with expectations, we also observed enrichment of immunological-related traits (‘lymphocyte count’, ‘monocyte count’ and traits related to immune disorders) in myeloid cells and B and T lymphocytes. Likewise, blood-related traits such as ‘mean sphered cell volume’ and ‘red blood cell distribution width’ were enriched in erythroid cells. Notably, we observed enrichment for traits such as ‘body mass index’ or ‘waist–hip ratio’ in lower digestive tract epithelial cells and somatotrope cells from the pituitary gland. Similarly, type 2 diabetes- and cholesterol-related traits showed not only the expected association with pancreatic cells (acinar, ductal and islet cells) and hepatocytes, but also associations with several kidney cell populations57. Our analysis also indicated enrichment of attention deficit and hyperactivity disorder, which often presents with motor abnormalities58, in skeletal muscle type II myonuclei in addition to neural cells (Fig. 5). To evaluate differences in target cell specificity among species, we further compared a selected panel of GWAS traits to cell types within the neocortex (our own dataset), heart and kidney in mice33,39, humans30,35,38 and monkey (Supplementary Fig. 45a). Neurological and neuropathological traits were more strongly linked to neurons in humans and monkeys than to those in mice. Notably, migraine had a higher score in human and monkey excitatory neurons than in mice but was more highly enriched in kidney intercalated cells of these two species.
The heat map shows the association of selected common human traits and diseases (indicated on the right) with the monkey cell types (indicated at the bottom) annotated in our dataset. The coloured boxes indicate selected enriched patterns. Definitions for abbreviations are provided in the Supplementary Note.Source data
We also generated a correlation map of specific mutant genes causing human diseases (Supplementary Fig. 46 and Supplementary Table 6b). As expected, genes related to retinitis pigmentosa were specifically expressed in monkey photoreceptors, while genes related to porphyria were associated with erythroblasts. In addition, we compared the interspecies distribution of a selection of genes related to human neurological diseases in mouse, human35 and monkey neocortex. As with the GWAS, we observed a generally higher correlation of the expression in specific cell types between humans and monkeys than between either of these species and mice (Supplementary Fig. 45b). However, some genes were linked to different cell types in monkeys and humans. For instance, spinocerebellar ataxia caused by mutations in PLEKHG4 (ref. 59) and ataxia telangiectasia caused by mutations in ATM were enriched in astrocytes and oligodendrocytes60, respectively, in humans, while they were enriched in distinct types of inhibitory neurons in monkeys and mice. Further scrutiny of these and other GWAS datasets and disease-related genes as well as wider interspecies comparisons using our website should provide additional insights.
Discussion
Despite the enormous potential, few NHP tissues have been profiled thus far at the single-cell/nucleus level, and use of different species, experimental conditions and platforms makes comparisons challenging13,61. To address this, we have generated the first version of a large-scale cell transcriptomic atlas for an NHP widely used in research studies, M. fascicularis, and an open, expandable and interactive NHPCA database to facilitate its exploration.
In addition to the study of NHP physiology, our dataset will be valuable for understanding tissues that either have not been profiled at all at the single-cell/nucleus level in humans or lack sufficient cell numbers, enabling interspecies adaptive comparisons and predicting disease susceptibility. With respect to the latter, the observed association between IL-6, STAT transcription factors and ACE2 in the kidney could explain the reported positive effects of tocilizumab, a humanized monoclonal antibody against IL-6R, for the treatment of patients with severe COVID-19 disease62. Although it is currently under debate whether the human kidney is infected by SARS-CoV-2 (ref. 63), this positive feedback loop may exist in other tissues. Notably, we have also shown that the distribution of ACE2 and TMPRSS2 expression across different cell types is not identical between monkeys and humans. This could influence SARS-CoV-2 pathogenesis and may for example explain why drugs such as hydroxychloroquine, despite providing promising results in monkey cell lines in vitro, are not effective in humans64. The analysis of human genetic disease susceptibility confirmed clinical associations between motor symptoms and attention deficit and hyperactivity disorder58 as well as between migraine and the kidney65. Interspecies comparison for a panel of genes showed that differences in target cell susceptibility exist between humans and monkeys, further demonstrating that a cautious approach is required when modelling human diseases in NHPs.
Notably, in the survey for Wnt pathway components, we identified an unexpected enrichment of LGR5+ cells in the monkey DCT in comparison with mice and humans. The maintenance of high levels of LGR5 in DCTCs and of WNT9B in cells from the collecting duct suggests that the monkey DCT could have different properties than in mice and humans, but this remains to be studied. Similarly, LGR5+ cells in the neocortex correspond mainly to OPCs in monkeys, oligodendrocytes (and, to a lesser extent, OPCs) in humans and inhibitory neurons in mice. This is consistent with the knowledge that Wnt activity regulates oligodendrocyte function and OPC to oligodendrocyte differentiation66, but points to interspecies differences in the mode of action. Likewise, the expression of LGR5 in skeletal slow-twitch myofibres and LGR6 in the pituitary gland and heart is intriguing. During development, Wnt activity regulates skeletal myogenesis and myofibre typing, cardiomyocyte progenitor proliferation and pituitary gland growth67,68, but little is known about its role in adults.
Apart from these analyses and comparisons, our NHPCA website provides a platform for interactive comparisons with manually uploaded datasets. When doing this, the type of sequencing platform and use of single-nucleus versus single-cell analysis should be considered, as these factors can influence the number of captured genes as well as the cell populations detected and their relative proportions. In the future, the NHPCA database will be extended with additional omics layers and datasets from disease modelling studies and ageing. It will also be relevant to compare our M. fascicularis atlas with future cell atlases from humans and other non-endangered NHPs. Altogether, this information will be instrumental for advancing knowledge of primates.
Methods
Ethics statement
All experimental protocols in this study were reviewed and approved by the Institutional Animal Care and Use Committee of Huazhen Bioscience (permit no. HZ2019027) and the Institutional Review Board on Ethics Committee of Beijing Genomics Institute (BGI; permit nos BGI-IRB 19125-T2 and BGI-IRB 21136). The study was also implemented in compliance with the US National Institutes of Health Guide for the Care and Use of Laboratory Animals (8th edition, 2011).
Collection of animal tissues
A total of three female and five male cynomolgus monkeys, approximately 6 years old, were obtained from Huazhen Laboratory Animal Breeding Centre (Guangzhou, China). Monkeys were anaesthetized with an injection of ketamine hydrochloride (10 mg per kg) and sodium pantabarbital (40 mg per kg) before being euthanized by exsanguination. Wild-type C57BL/6J male mice, approximately 8 weeks old, were purchased from Guangdong Medical Lab Animal Center. Mice were provided with food and water ad libitum and maintained on a regular 12-h day/12-h night cycle. Ambient temperature was set to 18–23 °C, and relative humidity was set to 40–60%. One mouse was euthanized by neck dislocation. Monkey and mouse tissues were isolated and placed on an ice-cold board for dissection. Each tissue (except for bone marrow, peripheral blood and tissues on which enzymatic digestion was performed) was cut into 5–10 pieces of roughly 50–200 mg each. Samples were transferred to cryogenic vials (Corning, 430488) and then snap frozen in liquid nitrogen and stored in liquid nitrogen until nuclear extraction was performed. Peripheral blood mononuclear cells (PBMCs) from heparinized venous blood and bone marrow cells were isolated using Lymphoprep medium (STEMCELL Technologies, 07851) according to a standard density gradient centrifugation protocol. Cells from these two tissues were resuspended in freezing medium composed of 90% FBS (Thermo Fisher, 1921005PJ) and 10% DMSO (Sigma-Aldrich, D2650) and frozen using a Nalgene Mr. Frosty Cryo 1 °C Freezing Container (Thermo Fisher Scientific, 5100-0001) in a −80 °C freezer for 24 h before being transferred to liquid nitrogen for long-term storage.
Single-nucleus/cell suspension preparation
Single-nucleus isolation was performed as previously described69. In brief, tissues were thawed, minced and transferred to a 1-ml Dounce homogenizer (TIANDZ) with 1 ml of homogenization buffer A containing 250 mM sucrose (Ambion), 10 mg ml–1 BSA (Ambion), 5 mM MgCl2 (Ambion), 0.12 U μl–1 RNasin Plus (Promega, N2115), 0.12 U μl–1 RNasein (Promega, N2115) and 1× cOmplete Protease Inhibitor Cocktail (Roche, 11697498001). Frozen tissues were kept in an ice box and homogenized by 25–50 strokes of the loose pestle (pestle A), after which the mixture was filtered using a 100-µm cell strainer into a 1.5-ml tube (Eppendorf). The mixture was then transferred to a clean 1-ml Dounce homogenizer to which 750 μl of buffer A containing 1% Igepal (Sigma, CA630) was added, and the tissue was further homogenized by 25 strokes of the tight pestle (pestle B). After this, the mixture was filtered through a 40-µm strainer into a 1.5-ml tube and centrifuged at 500g for 5 min at 4 °C to pellet the nuclei. The pellet was resuspended in 1 ml of buffer B containing 320 mM sucrose, 10 mg ml–1 BSA, 3 mM CaCl2, 2 mM magnesium acetate, 0.1 mM EDTA, 10 mM Tris-HCl, 1 mM DTT, 1× cOmplete Protease Inhibitor Cocktail and 0.12 U μl–1 RNasein. This was followed by a centrifugation at 500g for 5 min at 4 °C to pellet the nuclei. Nuclei were then resuspended with cell resuspension buffer at a concentration of 1,000 nuclei per μl for library preparation.
Because of technical limitations in obtaining high-quality nuclei, scRNA-seq was performed for colon, duodenum, spleen, stomach, skin and testis. To do this, cells were obtained from fresh tissue by enzymatic digestion. Tissues were first rinsed with PBS, minced into small pieces by mechanical dissociation and incubated for 1 h in 10 ml DS-LT buffer (0.2 mg ml–1 CaCl2, 5 μM MgCl2, 0.2% BSA and 0.2 mg ml–1 Liberase in HBSS) at 37 °C. After this, the tissue digestion was stopped by adding 3 ml of FBS, followed by filtration through a 100-µm cell strainer and centrifugation at 500g for 5 min at 4 °C. Cells from lymph node and spleen were obtained from fresh tissue by mechanical dissociation. Cells from bone marrow and PBMCs were obtained as described in the ‘Collection of animal tissues’ section. Samples were filtered through a 40-µm cell strainer and centrifuged at 500g for 5 min at 4 °C. Pellets were resuspended in cell resuspension buffer at 1,000 cells per μl for library preparation.
scRNA-seq and snRNA-seq sample preparation
The DNBelab C Series Single-Cell Library Prep Set (MGI, 1000021082) was used as previously described4. In brief, single-nucleus/cell suspensions were used for droplet generation, emulsion breakage, bead collection, reverse transcription and cDNA amplification to generate barcoded libraries. Indexed libraries were constructed according to the manufacturer’s protocol. Concentrations were measured with a Qubit ssDNA Assay Kit (Thermo Fisher Scientific, Q10212). Libraries were sequenced on a DNBSEQ-T1 or DNBSEQ-T7 sequencer at the China National GeneBank (Shenzhen, China) with the following sequencing strategy: 41-bp read length for read 1 and 100-bp read length for read 2.
scATAC-seq sample preparation
scATAC-seq libraries were prepared using the DNBelab C Series Single-Cell ATAC Library Prep Set70 (MGI, 1000021878). In brief, nuclei were extracted from tissue using the same protocol as describe above. After Tn5 tagmentation, transposed single-nucleus suspensions were converted to barcoded scATAC-seq libraries through droplet encapsulation, pre-amplification, emulsion breakage, captured bead collection, DNA amplification and purification. Indexed libraries were prepared according to the manufacturer’s protocol. Concentrations were measured with a Qubit ssDNA Assay Kit. Libraries were sequenced on a BGISEQ-500 sequencer at the China National GeneBank (Shenzhen, China) with the following sequencing strategy: 50-bp read length for read 1 and 76-bp read length for read 2.
scRNA-seq and snRNA-seq data processing
Raw data processing
Raw sequencing reads from DNBSEQ-T1 or DNBSEQ-T7 were filtered and demultiplexed using PISA (v0.2; https://github.com/shiquan/PISA). Reads were aligned to the Macaca_fascicularis_5.0 genome using STAR (v2.7.4a)71 and sorted by sambamba (v0.7.0)72. For tissues sequenced with scRNA-seq, reads were aligned to the exons of mRNA as normal. For tissues sequenced with snRNA-seq, a custom ‘pre-mRNA’ reference was created for alignment of count reads to introns as well as to exons because of the large amount of unspliced pre-mRNA in the cell nucleus. Thus, each gene’s transcript in snRNA-seq was counted by including exon and intron reads together73. Next, a cell/nucleus versus gene UMI count matrix was generated with PISA.
Ambient RNA removal
Ambient RNA noise was reduced using SoupX (v1.4.8; https://github.com/constantAmateur/SoupX)74 with default settings apart from the contamination fraction (represented as rho). The rho value was automatically parameterized using the autoEstCont function in tissues where rho was lower than 0.05 or higher than 0.2. In other tissues, the rho value was manually set to 0.2 using the setContaminationFraction function if the autoEstCont value was between 0.05 and 0.2.
Doublet removal
For each library, we performed doublet removal using DoubletFinder75. DoubletFinder first averages the transcriptional profile of randomly chosen cell pairs to create pseudo-doublets and then predicts doublets according to each real cell’s similarity in gene expression to the pseudo-doublets. Doublet removal was performed with the default parameter of DoubletFinder, and the 5% of cells most similar to the pseudo-doublets were excluded.
Cell clustering and cell type identification in scRNA-seq and snRNA-seq data
Cells or nuclei were preprocessed and filtered on the basis of a minimal expression threshold of 500 genes and genes being expressed by at least three cells or nuclei. Cells or nuclei fulfilling these criteria were kept for downstream analysis. In addition, cells or nuclei with more than 10% mitochondrial gene counts were removed. Global clustering of the complete cynomolgus monkey tissue dataset was performed using Scanpy (v1.6.0)76 in a Python environment (v3.6). Filtered data were transformed by ln(counts per million (CPM)/100 + 1). Three thousand highly variable genes were selected according to their average expression and dispersion. The number of UMIs and the percentage of mitochondrial genes were regressed out, and each gene was scaled with default options. Parameters used in each function were manually curated to obtain the optimal clustering of cells. Dimension reduction started with principal-component analysis, and the number of principal components used for UMAP visualization depended on the importance of the embeddings. The Louvain method was then used to detect subgroups of cells. For individual clustering, each tissue dataset was visualized using the Seurat package (v4.0.3)77 in the R environment (v4.0.2). Data from different replicates were normalized using the NormalizeData function with default options, and the top 2,000 most variable genes of each replicate were then calculated by FindVariableFeatures with the vst method. The replicable variable genes across replicates were selected to perform the FindIntegrationAnchors function for batch correction and then used to created an integrated data assay. The standard workflow for clustering and visualization was performed on the basis of the integrated data assay with default parameters according to the guidance of Seurat (https://satijalab.org/seurat/articles/integration_introduction.html). For kidney data, replicates were aligned to the monkey FM1 data with the FindIntegrationAnchors function using option reference = 1. Finally, each cluster was annotated by extensive literature review and searches for specific gene expression patterns.
DEGs and GO term enrichment
In the global clustering, we performed DEG analysis using the sc.pl.rank_genes_groups function in Scanpy. In other analyses, we used the FindMarkers or FindAllMarkers function in Seurat. Analysis of DEGs among different cell types within one tissue was performed with the FindAllMarkers function. DEGs were defined as genes with a fold change > 2 and adjusted P < 0.01. GO enrichment analysis was performed using the CompareCluster function of ChIPseeker (v1.22.1)78. Only GO terms with Q value < 0.05 were retained.
Cross-species comparisons
Between-atlas comparisons
For interspecies cell atlas analysis, data were retrieved from the HCL8 and MCA7. The count matrix for each tissue in the three species was preprocessed in three steps: (1) orthologous gene lists were downloaded from Ensembl79 and only genes that were orthologous for all three species were kept; (2) only genes expressed in at least one cell in each of the three species were kept; and (3) gene names for the human and mouse count matrix were converted into orthologues in M. fascicularis. After preprocessing, the count matrices of the three species were integrated and subjected to clustering using the standard integrated pipeline of Seurat with one additional criterion that only cells expressing more than 200 genes were kept. Seurat clusters were then annotated into different cell types using cell-type-specific markers defined in this paper.
Cross-species comparisons for other tissues
To obtain more accurate comparisons, we specifically chose three tissues, namely kidney7,8,30,31,32,33, neocortex35 (mouse neocortex data from our own samples) and heart38,39. Apart from the MCA and HCL kidney data, we downloaded the following data from public databases: human kidney, GSE121862 and GSE151302; mouse kidney (Tabula Muris), GSE107585; human neocortex, GSE97942; human heart, ERP123138; mouse heart, E-MTAB-7869; we also used our own mouse neocortex data (https://db.cngb.org/nhpca/). All data, except those from the MCA, HCL and Tabula Muris, were processed using our pipeline described above in the ‘scRNA-seq and snRNA-seq data processing’ section. Data were integrated using the same preprocessing, clustering and annotation method described above. Clusters with cell numbers lower than 200 were excluded. After annotation, we performed DEG analysis by comparing our dataset and each of the downloaded datasets within the same cell type. We used a critical cut-off in this analysis: fold change > 2 and adjusted P < 0.01. Only DEGs shared by three human datasets or three mouse datasets were considered to be species-specific DEGs.
Common cell analysis
For each common cell type, we extracted cells from all tissues in our dataset according to the cell type annotation presented in Supplementary Figs. 12–15. For the downstream analysis, we excluded common cell clusters from each individual tissue if the cell number of the cell cluster was less than 200. Data from different replicates were integrated following a standard integration pipeline using Seurat. To reduce the influence of ambient RNA and technical differences between snRNA-seq and scRNA-seq, the analysis of tissue-specific DEGs in Fig. 2 and Supplementary Fig. 21 was stringently defined. We first performed DEG analysis by comparing a selected cell type and other cell types within an individual tissue to define selected cell-type-specific genes in each tissue. We computed Pi, j,k as the fraction of cells in tissue i expressing gene j in cell population k. A given cell-type-specific gene j in tissue i (SCSGi) was defined using the following cut-off: log2(fold change) > 2, adjusted P < 0.01 and (Pi, j,c1 – Pi, j,c2)/Pi, j,c1 > 0.8 (where c1 represents a given cell type in tissue i and c2 represents other cell types in tissue i). After this, we tested whether SCSGi genes were differentially expressed in a given cell type in tissue i as compared to other tissues. Genes were finally determined to be tissue-specific DEGs of a given cell type in tissue i if they met the following conditions: log2(fold change) > 0.5 and adjusted P < 0.01.
Pseudotime trajectory analysis
The cell lineage trajectory was inferred using Monocle 2 (ref. 80) according to the tutorial. After the cell trajectory was constructed, DDRtree was used to visualize it in two-dimensional space.
Cell–cell interaction networks
To assess the cellular cross-talk between different cell types in each tissue, we used CellPhoneDB, a public repository of ligand–receptor interactions81. Cell-type‐specific receptor–ligand interactions between cell types were identified on the basis of specific expression of a receptor by one cell type and a ligand by another cell type. The interaction score refers to the mean total of the average expression values for all individual ligand–receptor partners in the corresponding interacting pairs of cell types. Before analysis, cells from the same cell type were aggregated in groups of 20 to make pseudo-cells in each organ. For this analysis, we applied a statistical method to ensure that only receptors or ligands expressed by more than 10% of the cells in the given cluster were considered. The total mean of the average expression values for individual partners in the corresponding interacting pairs of cell types was calculated.
Association of human GWAS and genetic disease data with monkey cell types
To test the enrichment of genes related to human diseases and traits for each cluster of cells based on global clustering, we applied linkage disequilibrium (LD) score regression analysis as previously described (https://github.com/bulik/ldsc/wiki/LD-Score-Estimation-Tutorial)82. For this, we only considered DEGs with an adjusted P < 0.01 and fold change > 2 in the tested cell types. Then, we converted the genome coordinates of Macaca_fascicularis_5.0 into hg19 genome coordinates by orthologous gene list download from Ensembl. The summary statistics file for each trait was downloaded from the UK Biobank database or published studies (Supplementary Table 6a). To calculate cell-type-specific LD scores, we first created annotation files for 22 chromosomes in each cell type with script make_annot.py using options --bed-file --bimfile 1000G.EUR.QC.bim --annot-file. Then, the annotation files were used as input to compute LD scores with the ldsc.py script using options --l2 --bfile 1000G.EUR.QC --ld-wind-cm 1 --annot --thin-annot --print-snps. Next, we ran the ldsc.py script with the --h2-cts flag to perform regressions following the standard workflow (https://github.com/bulik/ldsc/wiki/Cell-type-specific-analyses). We report the coefficient P value as a measure of the association of each cell type with the traits. All plots show the −log10-transformed P-value z-score of partitioned LD score regression. The cross-species GWAS analysis was performed on the basis of the integrated Seurat object.
scATAC-seq data processing
Raw sequencing reads from BGISEQ-500 were filtered, demultiplexed and aligned to the Macaca_fascicularis_5.0 genome using PISA. Fragment files for each library were generated for downstream analysis. The transcription start site enrichment score and fragment number for each nucleus were calculated using ArchR83. Cells with transcription start site enrichment scores lower than 5 and fragment numbers lower than 1,000 were removed. We then calculated the doublet score with the addDoubletScores function in ArchR and filtered using the filterDoublets function with parameter filterRatio = 2. Clustering analysis was performed using ArchR by first identifying a robust set of peak regions followed by iterative latent semantic indexing (LSI) clustering. In brief, we created 500-bp tiles across the genome and determined whether each cell was accessible within each tile. Next, we performed an LSI dimensionality reduction on these tiles with the addIterativeLSI function in ArchR. We then performed Seurat clustering (FindClusters) on the LSI dimensions at a resolution of 0.8. Anchors between the scATAC-seq and scRNA-seq/snRNA-seq datasets were identified and used to transfer cell type labels identified from the scRNA-seq/snRNA-seq data. Data were co-embedded using the TransferData function of Seurat.
Transcription factor motif enrichment analysis
To predict the motif footprint in peaks within the ACE2 promoter and enhancer sequences, we extracted the genome sequence in each peak region with Seqkit (v0.7.0)84. Sequences were matched to all Homo sapiens motifs form JASPAR2018 using the matchMotifs function in motifmatchr (v1.8.0) with the default parameter.
Immunofluorescence staining
Staining of monkey liver, subcutaneous and visceral adipose tissue, ovary and neocortex samples was conducted following a standard protocol. In brief, paraffin-embedded sections were deparaffinized, incubated with primary antibody for albumin (1:250 dilution; Abcam, ab207327) in liver, with primary antibody for CD34 (1:50 dilution; BioLegend, 34063) and NOX4 (1:100 dilution; Invitrogen, MA5-32090) in both types of adipose tissue, with primary antibody for CD44 (1:50 dilution; Proteintech, 60224-1-lg) in ovary, and with primary antibodies for PDGFRα (1:500 dilution; Cell Signaling, 3174S) and LGR5 (1:50 dilution; Abcam, ab273092) in neocortex overnight at 4 °C, followed by incubation with a secondary antibody conjugated to Alexa Fluor 488 (1:250 dilution; Jackson ImmunoResearch, 715-545-150) or Cy3 (1:250 dilution; Jackson ImmunoResearch, 711-165-152) for 30 min at room temperature. Slides were mounted with Slowfade Mountant+DAPI (Life Technologies, S36964) and sealed.
smFISH
smFISH of monkey kidney, diaphragm and heart tissues was performed using RNAscope Fluorescent Multiplex and RNAscope Multiplex Fluorescent v2 (Advanced Cell Diagnostics) according to the manufacturer’s instructions. The following alterations were made: the thickness of the paraffin section was adjusted to 5 μm, the target retrieval boiling time was adjusted to 15 min, and the incubation time with Protease plus at 40 °C was adjusted to 30 min. The following fluorescence channels were used for RNAscope probes: LGR5 (C1), SLC12A3 (C2), LGR6 (C2) and MYH7 (C2). For ovary, LGR5 (C1) probe was used before staining with primary antibody for CD44 (Proteintech, 60224-1-Ig) and subsequent incubation with secondary antibody (Alexa Fluor, Jackson ImmunoResearch) for 30 min at room temperature. Slides were mounted with Slowfade Mountant+DAPI (Life Technologies, S36964) and sealed.
Statistics and reproducibility
For smFISH and immunofluorescence staining experiments, each in situ hybridization probe or antibody staining was repeated with similar results on at least three separate samples and on at least two sections per sample. The experiments were not randomized, and the investigators were not blinded to allocation during experiments and outcome assessment. No statistical methods were used to predetermine sample size.
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this paper.
Data availability
All raw data produced in this study (including NHPCA and mouse neocortex data) have been deposited to the CNGB Nucleotide Sequence Archive (accession code CNP0001469). All NHPCA count matrix data are available from https://db.cngb.org/nhpca/download. We have also provided the NHPCA website (https://db.cngb.org/nhpca/), an open and interactive database for exploration. The public datasets used in this study can be accessed as described below: the HCL count matrix is available at https://figshare.com/articles/dataset/HCL_DGE_Data/7235471, the MCA count matrix is available at https://figshare.com/articles/dataset/MCA_DGE_Data/5435866 and the count matrix for the Tabula Muris dataset is available at https://figshare.com/projects/Tabula_Muris_Transcriptomic_characterization_of_20_organs_and_tissues_from_Mus_musculus_at_single_cell_resolution/27733. The Gene Expression Omnibus (GEO) accession numbers for the two human kidney datasets are GSE121862 and GSE151302. The GEO accession number for the mouse kidney data is GSE107585. The GEO accession number for the human neocortex data is GSE97942. The human heart data can be accessed at the European Nucleotide Archive (https://www.ebi.ac.uk/ena/) using accession number ERP123138. The mouse heart data can be found through accession number E-MTAB-7869 in the database of the European Bioinformatics Institute (https://www.ebi.ac.uk/arrayexpress/experiments/E-MTAB-7869/). Summary statistics files for each human trait were downloaded from the UK Biobank database or published studies (data links in Supplementary Table 6a). Source data are provided with this paper.
Code availability
Computer code used for processing the snRNA-seq, scRNA-seq and scATAC-seq data is available at https://github.com/single-cell-BGI/NHPCA.
References
Rozenblatt-Rosen, O., Stubbington, M. J. T., Regev, A. & Teichmann, S. A. The Human Cell Atlas: from vision to reality. Nature 550, 451–453 (2017).
Carbone, L. et al. Gibbon genome and the fast karyotype evolution of small apes. Nature 513, 195–201 (2014).
Taylor, K. Clinical veterinarian’s perspective of non-human primate (NHP) use in drug safety studies. J. Immunotoxicol. 7, 114–119 (2010).
Zhu, L. et al. Single-cell sequencing of peripheral mononuclear cells reveals distinct immune response landscapes of COVID-19 and influenza patients. Immunity 53, 685–696 (2020).
Delorey, T. M. et al. COVID-19 tissue atlases reveal SARS-CoV-2 pathology and cellular targets. Nature 595, 107–113 (2021).
Ding, J. et al. Systematic comparison of single-cell and single-nucleus RNA-sequencing methods. Nat. Biotechnol. 38, 737–746 (2020).
Han, X. et al. Mapping the mouse cell atlas by Microwell-seq. Cell 172, 1091–1107 (2018).
Han, X. et al. Construction of a human cell landscape at single-cell level. Nature 581, 303–309 (2020).
Brazovskaja, A. et al. Cell atlas of the regenerating human liver after portal vein embolization. Preprint at bioRxiv https://doi.org/10.1101/2021.06.03.444016 (2021).
Bram, Y. et al. Cell and tissue therapy for the treatment of chronic liver disease. Annu. Rev. Biomed. Eng. 23, 517–546 (2021).
Krausgruber, T. et al. Structural cells are key regulators of organ-specific immune responses. Nature 583, 296–302 (2020).
Kalucka, J. et al. Single-cell transcriptome atlas of murine endothelial cells. Cell 180, 764–779 (2020).
Geirsdottir, L. et al. Cross-species single-cell analysis reveals divergence of the primate microglia program. Cell 179, 1609–1622 (2019).
Petrany, M. J. et al. Single-nucleus RNA-seq identifies transcriptional heterogeneity in multinucleated skeletal myofibers. Nat. Commun. 11, 6374 (2020).
Stål, P., Marklund, S., Thornell, L. E., De Paul, R. & Eriksson, P. O. Fibre composition of human intrinsic tongue muscles. Cells Tissues Organs 173, 147–161 (2003).
Vijay, J. et al. Single-cell analysis of human adipose tissue identifies depot- and disease-specific cell types. Nat. Metab. 2, 97–109 (2020).
Merrick, D. et al. Identification of a mesenchymal progenitor cell hierarchy in adipose tissue. Science 364, eaav2501 (2019).
Ribeiro, R. et al. Human periprostatic white adipose tissue is rich in stromal progenitor cells and a potential source of prostate tumor stroma. Exp. Biol. Med. 237, 1155–1162 (2012).
Schröder, K., Wandzioch, K., Helmcke, I. & Brandes, R. P. Nox4 acts as a switch between differentiation and proliferation in preadipocytes. Arter. Thromb. Vasc. Biol. 29, 239–245 (2009).
Ha, C. W. Y. et al. Translocation of viable gut microbiota to mesenteric adipose drives formation of creeping fat in humans. Cell 183, 666–683 (2020).
Adler, E., Mhawech-Fauceglia, P., Gayther, S. A. & Lawrenson, K. PAX8 expression in ovarian surface epithelial cells. Hum. Pathol. 46, 948–956 (2015).
Ng, A. et al. Lgr5 marks stem/progenitor cells in ovary and tubal epithelia. Nat. Cell Biol. 16, 745–757 (2014).
Parte, S. C., Batra, S. K. & Kakar, S. S. Characterization of stem cell and cancer stem cell populations in ovary and ovarian tumors. J. Ovarian Res. 11, 69 (2018).
Nusse, R. & Clevers, H. Wnt/β-catenin signaling, disease, and emerging therapeutic modalities. Cell 169, 985–999 (2017).
Leung, C., Tan, S. H. & Barker, N. Recent advances in Lgr5+ stem cell research. Trends Cell Biol. 28, 380–391 (2018).
Barker, N. & Clevers, H. Leucine-rich repeat-containing G-protein-coupled receptors as markers of adult stem cells. Gastroenterology 138, 1681–1696 (2010).
Lee, J.-H. et al. Anatomically and functionally distinct lung mesenchymal populations marked by Lgr5 and Lgr6. Cell 170, 1149–1163 (2017).
Huch, M. et al. In vitro expansion of single Lgr5+ liver stem cells induced by Wnt-driven regeneration. Nature 494, 247–250 (2013).
Barker, N. et al. Identification of stem cells in small intestine and colon by marker gene Lgr5. Nature 449, 1003–1007 (2007).
Lake, B. B. et al. A single-nucleus RNA-sequencing pipeline to decipher the molecular anatomy and pathophysiology of human kidneys. Nat. Commun. 10, 2832 (2019).
Muto, Y. et al. Single cell transcriptional and chromatin accessibility profiling redefine cellular heterogeneity in the adult human kidney. Nat. Commun. 12, 2190 (2021).
Tabula Muris Consortium. et al. Single-cell transcriptomics of 20 mouse organs creates a Tabula Muris. Nature 562, 367–372 (2018).
Park, J. et al. Single-cell transcriptomics of the mouse kidney reveals potential cellular targets of kidney disease. Science 360, 758–763 (2018).
Barker, N. et al. Lgr5+ve stem/progenitor cells contribute to nephron formation during kidney development. Cell Rep. 2, 540–552 (2012).
Lake, B. B. et al. Integrative single-cell analysis of transcriptional and epigenetic states in the human adult brain. Nat. Biotechnol. 36, 70–80 (2018).
Nakashima, H. et al. R-spondin 2 promotes acetylcholine receptor clustering at the neuromuscular junction via Lgr5. Sci. Rep. 6, 28512 (2016).
Leung, C. et al. Lgr5 marks adult progenitor cells contributing to skeletal muscle regeneration and sarcoma formation. Cell Rep. 33, 108535 (2020).
Litvinukova, M. et al. Cells of the adult human heart. Nature 588, 466–472 (2020).
Vidal, R. et al. Transcriptional heterogeneity of fibroblasts is a hallmark of the aging heart. JCI Insight 4, e131092 (2019).
Vankelecom, H. Non-hormonal cell types in the pituitary candidating for stem cell. Semin. Cell Dev. Biol. 18, 559–570 (2007).
Klein, D. et al. Wnt2 acts as a cell type-specific, autocrine growth factor in rat hepatic sinusoidal endothelial cells cross-stimulating the VEGF pathway. Hepatology 47, 1018–1031 (2008).
Nusse, R. Wnt signaling and stem cell control. Cell Res. 18, 523–527 (2008).
Niehrs, C. The complex world of WNT receptor signalling. Nat. Rev. Mol. Cell Biol. 13, 767–779 (2012).
Zhang, M. et al. β-catenin safeguards the ground state of mouse pluripotency by strengthening the robustness of the transcriptional apparatus. Sci. Adv. 6, eaba1593 (2020).
Devakumar, D. et al. Infectious causes of microcephaly: epidemiology, pathogenesis, diagnosis, and management. Lancet Infect. Dis. 18, e1–e13 (2018).
Zhu, N. et al. A novel coronavirus from patients with pneumonia in China, 2019. N. Engl. J. Med. 382, 727–733 (2020).
Hoffmann, M. et al. SARS-CoV-2 cell entry depends on ACE2 and TMPRSS2 and is blocked by a clinically proven protease inhibitor. Cell 181, 271–280 (2020).
Rockx, B. et al. Comparative pathogenesis of COVID-19, MERS, and SARS in a nonhuman primate model. Science 368, 1012–1015 (2020).
Diao, B. et al. Human kidney is a target for novel severe acute respiratory syndrome coronavirus 2 infection. Nat. Commun. 12, 2506 (2021).
Muus, C. et al. Single-cell meta-analysis of SARS-CoV-2 entry genes across tissues and demographics. Nat. Med. 27, 546–559 (2021).
Sungnak, W. et al. SARS-CoV-2 entry factors are highly expressed in nasal epithelial cells together with innate immune genes. Nat. Med. 26, 681–687 (2020).
Ying, M. et al. COVID-19 with acute cholecystitis: a case report. BMC Infect. Dis. 20, 437 (2020).
Tosi, M. F. Innate immune responses to infection. J. Allergy Clin. Immunol. 116, 241–249 (2005).
Bell, L. C. K. et al. Transcriptional response modules characterize IL-1β and IL-6 activity in COVID-19. iScience 24, 101896 (2021).
Nie, X. et al. Multi-organ proteomic landscape of COVID-19 autopsies. Cell 184, 775–791 (2021).
Gate, D. et al. Clonally expanded CD8 T cells patrol the cerebrospinal fluid in Alzheimer’s disease. Nature 577, 399–404 (2020).
Zhong, J., Yang, H. & Kon, V. Kidney as modulator and target of "good/bad" HDL. Pediatr. Nephrol. 34, 1683–1695 (2019).
Athanasiadou, A. et al. Early motor signs of attention-deficit hyperactivity disorder: a systematic review. Eur. Child Adolesc. Psychiatry 29, 903–916 (2020).
Amino, T. et al. Redefining the disease locus of 16q22.1-linked autosomal dominant cerebellar ataxia. J. Hum. Genet. 52, 643–649 (2007).
Tse, K. H. & Herrup, K. DNA damage in the oligodendrocyte lineage and its role in brain aging. Mech. Ageing Dev. 161, 37–50 (2017).
Wang, S. et al. Single-cell transcriptomic atlas of primate ovarian aging. Cell 180, 585–600 (2020).
Villiger, P. M. et al. Tocilizumab for induction and maintenance of remission in giant cell arteritis: a phase 2, randomised, double-blind, placebo-controlled trial. Lancet 387, 1921–1927 (2016).
Le Stang, M.-B., Desenclos, J., Flamant, M., Chousterman, B. G. & Tabibzadeh, N. The good treatment, the bad virus, and the ugly inflammation: pathophysiology of kidney involvement during COVID-19. Front. Physiol. 12, 209 (2021).
Cavalcanti, A. B. et al. Hydroxychloroquine with or without azithromycin in mild-to-moderate Covid-19. N. Engl. J. Med. 383, 2041–2052 (2020).
Verghese, D., Alrifai, T., Nimmagadda, M. & Upadhyay, M. It could be in the kidneys: fibromuscular dysplasia and the association with headaches and mood disorders. BMJ Case Rep. 12, e231322 (2019).
Chavali, M. et al. Wnt-dependent oligodendroglial–endothelial interactions regulate white matter vascularization and attenuate injury. Neuron 108, 1130–1145 (2020).
Heallen, T. et al. Hippo pathway inhibits Wnt signaling to restrain cardiomyocyte proliferation and heart size. Science 332, 458–461 (2011).
Osmundsen, A. M., Keisler, J. L., Taketo, M. M. & Davis, S. W. Canonical WNT signaling regulates the pituitary organizer and pituitary gland formation. Endocrinology 158, 3339–3353 (2017).
Bakken, T. E. et al. Single-nucleus and single-cell transcriptomes compared in matched cortical cell types. PLoS ONE 13, e0209648 (2018).
Yu, Y. et al. Single-nucleus chromatin accessibility landscape reveals diversity in regulatory regions across distinct adult rat cortex. Front. Mol. Neurosci. 14, 651355 (2021).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Tarasov, A., Vilella, A. J., Cuppen, E., Nijman, I. J. & Prins, P. Sambamba: fast processing of NGS alignment formats. Bioinformatics 31, 2032–2034 (2015).
Del-Aguila, J. L. et al. A single-nuclei RNA sequencing study of Mendelian and sporadic AD in the human brain. Alzheimers Res. Ther. 11, 71 (2019).
Young, M. D. & Behjati, S. SoupX removes ambient RNA contamination from droplet-based single-cell RNA sequencing data. Gigascience 9, giaa151 (2020).
McGinnis, C. S., Murrow, L. M. & Gartner, Z. J. DoubletFinder: doublet detection in single-cell RNA sequencing data using artificial nearest neighbors. Cell Syst. 8, 329–337 (2019).
Wolf, F. A., Angerer, P. & Theis, F. J. SCANPY: large-scale single-cell gene expression data analysis. Genome Biol. 19, 15 (2018).
Hao, Y. et al. Integrated analysis of multimodal single-cell data. Cell 184, 3573–3587 (2021).
Yu, G., Wang, L. G. & He, Q. Y. ChIPseeker: an R/Bioconductor package for ChIP peak annotation, comparison and visualization. Bioinformatics 31, 2382–2383 (2015).
Yates, A. D. et al. Ensembl 2020. Nucleic Acids Res. 48, D682–D688 (2020).
Qiu, X. et al. Reversed graph embedding resolves complex single-cell trajectories. Nat. Methods 14, 979–982 (2017).
Efremova, M., Vento-Tormo, M., Teichmann, S. A. & Vento-Tormo, R. CellPhoneDB: inferring cell–cell communication from combined expression of multi-subunit ligand–receptor complexes. Nat. Protoc. 15, 1484–1506 (2020).
Bryois, J. et al. Genetic identification of cell types underlying brain complex traits yields insights into the etiology of Parkinson’s disease. Nat. Genet. 52, 482–493 (2020).
Granja, J. M. et al. ArchR is a scalable software package for integrative single-cell chromatin accessibility analysis. Nat. Genet. 53, 403–411 (2021).
Shen, W., Le, S., Li, Y. & Hu, F. SeqKit: a cross-platform and ultrafast toolkit for FASTA/Q file manipulation. PLoS ONE 11, e0163962 (2016).
Acknowledgements
We thank W. Liu and L. Xu from the Huazhen Laboratory Animal Breeding Centre for helping in the collection of monkey tissues, D. Zhu and H. Li from the Bioland Laboratory (Guangzhou Regenerative Medicine and Health Guangdong Laboratory) for technical help, G. Guo and H. Sun from Zhejiang University for providing HCL and MCA gene expression data matrices, G. Dong and C. Liu from BGI Research, and X. Zhang, P. Li and C. Qi from the Guangzhou Institutes of Biomedicine and Health for experimental advice or providing reagents. This work was supported by the Shenzhen Basic Research Project for Excellent Young Scholars (RCYX20200714114644191), Shenzhen Key Laboratory of Single-Cell Omics (ZDSYS20190902093613831), Shenzhen Bay Laboratory (SZBL2019062801012) and Guangdong Provincial Key Laboratory of Genome Read and Write (2017B030301011). In addition, L.L. was supported by the National Natural Science Foundation of China (31900466), Y. Hou was supported by the Natural Science Foundation of Guangdong Province (2018A030313379) and M.A.E. was supported by a Changbai Mountain Scholar award (419020201252), the Strategic Priority Research Program of the Chinese Academy of Sciences (XDA16030502), a Chinese Academy of Sciences–Japan Society for the Promotion of Science joint research project (GJHZ2093), the National Natural Science Foundation of China (92068106, U20A2015) and the Guangdong Basic and Applied Basic Research Foundation (2021B1515120075). M.L. was supported by the National Key Research and Development Program of China (2021YFC2600200).
Author information
Authors and Affiliations
Contributions
L.H., Y. Hou, X.X., M.A.E. and L.L. conceived the idea; Y. Hou, X.X., M.A.E. and L.L. supervised the work; L.H., Xiaoyu Wei, Y. Yuan, M.A.E. and L.L. designed the experiments; L.H., Xiaoyu Wei, G.V., Y. Yuan, X. Zhang, P.F., P.G., Xingyuan Liu, F.Y., S.S., G.L., J.A., Y. Lei, Y. Lai, M.C., C.-W. Wong, X.G., S.L. and J.M. collected tissue samples; C.L., G.V., Zhifeng Wang, Y. Yuan, X. Zhang, P.F., Q.D., Ya Liu, Y. Huang, H.L., B.W., M.C., J.X., M.W., C. Wang, Y.Z., Y. Yu, H. Zheng, Y.W. and S.X. performed the experiments. L.H., Xiaoyu Wei, G.V., Z. Zhuang, X. Zou, T.P., Y. Lai, L.W., Q. Shi, H. Yu, Yang Liu, D.X., F.H., Z. Zhu and C. Ward performed data analysis. L.H., Xiaoyu Wei, C.L., G.V., Z. Zhuang, X. Zou, Z. Wang, T.P., Y. Yang, J.L. and L.L. prepared the figures. H. Yu, Xiaofeng Wei, F.C., T.Y., W.D. and J.C. prepared the website. Zongren Wang, Z.P., C.-W.W., B.Q., A.S., J.I., L.F., Yan Liu, Z.L., Xiaolei Liu, H. Zhang, M.L., Q. Sun, P.H.M., N.B., P.M.-C., Y.G., J.M., M.U., T.T., S.L., H. Yang and J.W. provided relevant advice and reviewed the manuscript. L.H., G.V., M.A.E. and L.L. wrote the manuscript with input from all authors. All other authors contributed to the work. All authors read and approved the manuscript for submission.
Corresponding authors
Ethics declarations
Competing interests
Employees of BGI have stock holdings in BGI. All other authors declare no competing interests.
Peer review
Peer review information
Nature thanks Benjamin Humphreys, Itai Yanai and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
This file contains the Supplementary Note, the legends for Supplementary Figs. 1–46 and the legends for Supplementary Tables 1–6.
Supplementary Table 1
Description of all profiled monkey tissues, cell types and markers used for cluster annotation.
Supplementary Table 2
Global analysis of monkey common cell types and tissue-specific signatures.
Supplementary Table 3
Global distribution of LGR5, LGR6 and MKI67 expression in monkey tissues.
Supplementary Table 4
Species-specific genes in kidney DCTCs between monkey, human and mouse.
Supplementary Table 5
Expression of virus receptors, ACE2 and TMPRSS2 in monkey tissues.
Supplementary Table 6
Association of GWAS traits and human genetic diseases with monkey cell types.
Rights and permissions
About this article
Cite this article
Han, L., Wei, X., Liu, C. et al. Cell transcriptomic atlas of the non-human primate Macaca fascicularis. Nature 604, 723–731 (2022). https://doi.org/10.1038/s41586-022-04587-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-022-04587-3
This article is cited by
-
VGLL1 cooperates with TEAD4 to control human trophectoderm lineage specification
Nature Communications (2024)
-
A spatiotemporal atlas of mouse liver homeostasis and regeneration
Nature Genetics (2024)
-
An organism-wide atlas of hormonal signaling based on the mouse lemur single-cell transcriptome
Nature Communications (2024)
-
Single-nucleus transcriptome inventory of giant panda reveals cellular basis for fitness optimization under low metabolism
BMC Biology (2023)
-
Traditional Chinese Medicine Shi-Bi-Man regulates lactic acid metabolism and drives hair follicle stem cell activation to promote hair regeneration
Chinese Medicine (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.