Abstract
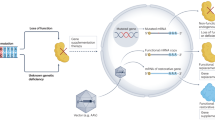
Gene therapy is a potentially curative medicine for many currently untreatable diseases, and recombinant adeno-associated virus (rAAV) is the most successful gene delivery vehicle for in vivo applications1,2,3. However, rAAV-based gene therapy suffers from several limitations, such as constrained DNA cargo size and toxicities caused by non-physiological expression of a transgene4,5,6. Here we show that rAAV delivery of a suppressor tRNA (rAAV.sup-tRNA) safely and efficiently rescued a genetic disease in a mouse model carrying a nonsense mutation, and effects lasted for more than 6 months after a single treatment. Mechanistically, this was achieved through a synergistic effect of premature stop codon readthrough and inhibition of nonsense-mediated mRNA decay. rAAV.sup-tRNA had a limited effect on global readthrough at normal stop codons and did not perturb endogenous tRNA homeostasis, as determined by ribosome profiling and tRNA sequencing, respectively. By optimizing the AAV capsid and the route of administration, therapeutic efficacy in various target tissues was achieved, including liver, heart, skeletal muscle and brain. This study demonstrates the feasibility of developing a toolbox of AAV-delivered nonsense suppressor tRNAs operating on premature termination codons (AAV-NoSTOP) to rescue pathogenic nonsense mutations and restore gene function under endogenous regulation. As nonsense mutations account for 11% of pathogenic mutations, AAV-NoSTOP can benefit a large number of patients. AAV-NoSTOP obviates the need to deliver a full-length protein-coding gene that may exceed the rAAV packaging limit, elicit adverse immune responses or cause transgene-related toxicities. It therefore represents a valuable addition to gene therapeutics.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
Amplicon next-generation sequencing, ribosome profiling, tRNA sequencing and RNA-seq data can be found in the NCBI’s Gene Expression Omnibus (GEO) using GEO Series accession number GSE179275. Source data are provided with this paper.
Code availability
Bioinformatics analysis was performed using publicly available programs and parameters described in the Methods. The code for Ribo-seq is available at https://github.com/dolphinnext/ribosome-profiling. The code for tRNA-seq is available at https://github.com/dolphinnext/trnaseq.
Change history
12 May 2022
In the version of this article initially published, the name of John Leuck was not included in the Peer review information for this article, which has now been amended.
References
Wang, D., Tai, P. W. L. & Gao, G. Adeno-associated virus vector as a platform for gene therapy delivery. Nat. Rev. Drug Discov. 18, 358–378 (2019).
Li, C. & Samulski, R. J. Engineering adeno-associated virus vectors for gene therapy. Nat. Rev. Genet. 21, 255–272 (2020).
Mendell, J. R. et al. Current clinical applications of in vivo gene therapy with AAVs. Mol. Ther. 29, 464–488 (2020).
Hordeaux, J. et al. Adeno-associated virus-induced dorsal root ganglion pathology. Hum. Gene Ther. 31, 808–818 (2020).
Van Alstyne, M. et al. Gain of toxic function by long-term AAV9-mediated SMN overexpression in the sensorimotor circuit. Nat. Neurosci. 24, 930–940 (2021).
Golebiowski, D. et al. Direct intracranial injection of AAVrh8 encoding monkey β-N-acetylhexosaminidase causes neurotoxicity in the primate brain. Hum. Gene Ther. 28, 510–522 (2017).
Wang, D., Zhang, F. & Gao, G. CRISPR-based therapeutic genome editing: strategies and in vivo delivery by AAV vectors. Cell 181, 136–150 (2020).
Chang, J. C., Temple, G. F., Trecartin, R. F. & Kan, Y. W. Suppression of the nonsense mutation in homozygous β0 thalassaemia. Nature 281, 602–603 (1979).
Temple, G. F., Dozy, A. M., Roy, K. L. & Kan, Y. W. Construction of a functional human suppressor tRNA gene: an approach to gene therapy for β-thalassaemia. Nature 296, 537–540 (1982).
Porter, J. J., Heil, C. S. & Lueck, J. D. Therapeutic promise of engineered nonsense suppressor tRNAs. Wiley Interdiscip. Rev. RNA. 12, e1641 (2021).
Wang, D. et al. Characterization of an MPS I-H knock-in mouse that carries a nonsense mutation analogous to the human IDUA-W402X mutation. Mol. Genet. Metab. 99, 62–71 (2010).
Bigger, B. W., Begley, D. J., Virgintino, D. & Pshezhetsky, A. V. Anatomical changes and pathophysiology of the brain in mucopolysaccharidosis disorders. Mol. Genet. Metab. 125, 322–331 (2018).
Hampe, C. S. et al. Mucopolysaccharidosis type I: current treatments, limitations, and prospects for improvement. Biomolecules 11, 189 (2021).
Ingolia, N. T., Ghaemmaghami, S., Newman, J. R. & Weissman, J. S. Genome-wide analysis in vivo of translation with nucleotide resolution using ribosome profiling. Science 324, 218–223 (2009).
Wangen, J. R. & Green, R. Stop codon context influences genome-wide stimulation of termination codon readthrough by aminoglycosides. eLife. 9, e52611 (2020).
Behrens, A., Rodschinka, G. & Nedialkova, D. D. High-resolution quantitative profiling of tRNA abundance and modification status in eukaryotes by mim-tRNAseq. Mol. Cell 81, 1802–1815.e7 (2021).
Koukuntla, R., Ramsey, W. J., Young, W. B. & Link, C. J. U6 promoter-enhanced GlnUAG suppressor tRNA has higher suppression efficacy and can be stably expressed in 293 cells. J. Gene Med. 15, 93–101 (2013).
Keeling, K. M., Xue, X., Gunn, G. & Bedwell, D. M. Therapeutics based on stop codon readthrough. Annu. Rev. Genomics Hum. Genet. 15, 371–394 (2014).
Manuvakhova, M., Keeling, K. & Bedwell, D. M. Aminoglycoside antibiotics mediate context-dependent suppression of termination codons in a mammalian translation system. RNA 6, 1044–1055 (2000).
Phillips-Jones, M. K., Hill, L. S., Atkinson, J. & Martin, R. Context effects on misreading and suppression at UAG codons in human cells. Mol. Cell. Biol. 15, 6593–6600 (1995).
Roy, B. et al. Ataluren stimulates ribosomal selection of near-cognate tRNAs to promote nonsense suppression. Proc. Natl Acad. Sci. USA 113, 12508–12513 (2016).
Xue, X. et al. Identification of the amino acids inserted during suppression of CFTR nonsense mutations and determination of their functional consequences. Hum. Mol. Genet. 26, 3116–3129 (2017).
Lueck, J. D. et al. Engineered transfer RNAs for suppression of premature termination codons. Nat. Commun. 10, 822 (2019).
Giege, R., Sissler, M. & Florentz, C. Universal rules and idiosyncratic features in tRNA identity. Nucleic Acids Res. 26, 5017–5035 (1998).
Bunge, S. et al. Genotype–phenotype correlations in mucopolysaccharidosis type I using enzyme kinetics, immunoquantification and in vitro turnover studies. Biochim. Biophys. Acta 1407, 249–256 (1998).
Oussoren, E. et al. Residual α-l-iduronidase activity in fibroblasts of mild to severe mucopolysaccharidosis type I patients. Mol. Genet. Metab. 109, 377–381 (2013).
Parker, D. J. et al. Growth-optimized aminoacyl-tRNA synthetase levels prevent maximal tRNA charging. Cell Syst. 11, 121–130.e6 (2020).
Hinnebusch, A. G. Translational regulation of GCN4 and the general amino acid control of yeast. Annu. Rev. Microbiol. 59, 407–450 (2005).
Buvoli, M., Buvoli, A. & Leinwand, L. A. Suppression of nonsense mutations in cell culture and mice by multimerized suppressor tRNA genes. Mol. Cell. Biol. 20, 3116–3124 (2000).
Xie, J. et al. Short DNA hairpins compromise recombinant adeno-associated virus genome homogeneity. Mol. Ther. 25, 1363–1374 (2017).
Davidoff, A. M., Ng, C. Y., Zhou, J., Spence, Y. & Nathwani, A. C. Sex significantly influences transduction of murine liver by recombinant adeno-associated viral vectors through an androgen-dependent pathway. Blood 102, 480–488 (2003).
Keeling, K. M. et al. Leaky termination at premature stop codons antagonizes nonsense-mediated mRNA decay in S. cerevisiae. RNA 10, 691–703 (2004).
Kim, Y. K., Furic, L., Desgroseillers, L. & Maquat, L. E. Mammalian Staufen1 recruits Upf1 to specific mRNA 3′UTRs so as to elicit mRNA decay. Cell 120, 195–208 (2005).
Maquat, L. E., Tarn, W. Y. & Isken, O. The pioneer round of translation: features and functions. Cell 142, 368–374 (2010).
van Tol, H. & Beier, H. All human tRNATyr genes contain introns as a prerequisite for pseudouridine biosynthesis in the anticodon. Nucleic Acids Res. 16, 1951–1966 (1988).
Dong, J., Qiu, H., Garcia-Barrio, M., Anderson, J. & Hinnebusch, A. G. Uncharged tRNA activates GCN2 by displacing the protein kinase moiety from a bipartite tRNA-binding domain. Mol. Cell 6, 269–279 (2000).
Fechter, P., Rudinger-Thirion, J., Theobald-Dietrich, A. & Giege, R. Identity of tRNA for yeast tyrosyl-tRNA synthetase: tyrosylation is more sensitive to identity nucleotides than to structural features. Biochemistry 39, 1725–1733 (2000).
Kurosaki, T., Popp, M. W. & Maquat, L. E. Quality and quantity control of gene expression by nonsense-mediated mRNA decay. Nat. Rev. Mol. Cell Biol. 20, 406–420 (2019).
Colombo, M., Karousis, E. D., Bourquin, J., Bruggmann, R. & Muhlemann, O. Transcriptome-wide identification of NMD-targeted human mRNAs reveals extensive redundancy between SMG6- and SMG7-mediated degradation pathways. RNA 23, 189–201 (2017).
Huang, L. et al. Targeting translation termination machinery with antisense oligonucleotides for diseases caused by nonsense mutations. Nucleic Acid Ther. 29, 175–186 (2019).
Wang, J. et al. In vivo delivery of suppressor tRNA overcomes a pathogenic nonsense mutation in mice. Mol. Ther. 29, S128 (2021).
Chan, K. Y. et al. Engineered AAVs for efficient noninvasive gene delivery to the central and peripheral nervous systems. Nat. Neurosci. 20, 1172–1179 (2017).
Moreno, A. M. et al. Immune-orthogonal orthologues of AAV capsids and of Cas9 circumvent the immune response to the administration of gene therapy. Nat. Biomed. Eng. 3, 806–816 (2019).
Li, A. et al. AAV-CRISPR gene editing is negated by pre-existing immunity to Cas9. Mol. Ther. 28, 1432–1441 (2020).
Gadalla, K. K. et al. Improved survival and reduced phenotypic severity following AAV9/MECP2 gene transfer to neonatal and juvenile male Mecp2 knockout mice. Mol. Ther. 21, 18–30 (2013).
Kramarski, L. & Arbely, E. Translational read-through promotes aggregation and shapes stop codon identity. Nucleic Acids Res. 48, 3747–3760 (2020).
Hashimoto, S., Nobuta, R., Izawa, T. & Inada, T. Translation arrest as a protein quality control system for aberrant translation of the 3′-UTR in mammalian cells. FEBS Lett. 593, 777–787 (2019).
Arribere, J. A. et al. Translation readthrough mitigation. Nature 534, 719–723 (2016).
Lombardi, S. et al. Translational readthrough of GLA nonsense mutations suggests dominant-negative effects exerted by the interaction of wild-type and missense variants. RNA Biol. 17, 254–263 (2020).
Kuzmin, D. A. et al. The clinical landscape for AAV gene therapies. Nat. Rev. Drug Discov. 20, 173–174 (2021).
Keller, A., Nesvizhskii, A. I., Kolker, E. & Aebersold, R. Empirical statistical model to estimate the accuracy of peptide identifications made by MS/MS and database search. Anal. Chem. 74, 5383–5392 (2002).
Nesvizhskii, A. I., Keller, A., Kolker, E. & Aebersold, R. A statistical model for identifying proteins by tandem mass spectrometry. Anal. Chem. 75, 4646–4658 (2003).
Ingolia, N. T., Brar, G. A., Rouskin, S., McGeachy, A. M. & Weissman, J. S. The ribosome profiling strategy for monitoring translation in vivo by deep sequencing of ribosome-protected mRNA fragments. Nat. Protoc. 7, 1534–1550 (2012).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Yukselen, O., Turkyilmaz, O., Ozturk, A. R., Garber, M. & Kucukural, A. DolphinNext: a distributed data processing platform for high throughput genomics. BMC Genomics 21, 310 (2020).
Clarke, L. A. et al. Murine mucopolysaccharidosis type I: targeted disruption of the murine α-l-iduronidase gene. Hum. Mol. Genet. 6, 503–511 (1997).
Wang, D. et al. The designer aminoglycoside NB84 significantly reduces glycosaminoglycan accumulation associated with MPS I-H in the Idua-W392X mouse. Mol. Genet. Metab. 105, 116–125 (2012).
Wang, D. et al. Cas9-mediated allelic exchange repairs compound heterozygous recessive mutations in mice. Nat. Biotechnol. 36, 839–842 (2018).
Crowe, A. R. & Yue, W. Semi-quantitative determination of protein expression using immunohistochemistry staining and analysis: an integrated protocol. Bio Protoc. 9, e3465 (2019).
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30, 2114–2120 (2014).
Evans, M. E., Clark, W. C., Zheng, G. & Pan, T. Determination of tRNA aminoacylation levels by high-throughput sequencing. Nucleic Acids Res. 45, e133 (2017).
Acknowledgements
We are grateful to P. D. Zamore, O. J. Rando, D. D. Nedialkova and members of the Wang laboratory and the Gao laboratory for helpful discussions. We thank staff at the Viral Vector Core at University of Massachusetts Chan Medical School for producing the AAV vectors used in this study. The Wang laboratory is supported by grants from the National Institutes of Health (NIH) (P01HL158506), the Grace Science Foundation, the Pitt–Hopkins Research Foundation, and Believe in a Cure. The Gao laboratory is supported by grants from the NIH (R01NS076991, P01AI100263, P01HL131471, R01AI121135, UG3HL147367, UH3HL147367, R01HL152723, R01HL097088 and U19AI149646) and the Cystic Fibrosis Foundation. O.Y. and A.K. are supported by the National Center for Advancing Translational Sciences (UL1TR001453). Illustrations were created with BioRender.com.
Author information
Authors and Affiliations
Contributions
bioinformatics analysis. Z.J. performed the histological analysis. J.X. analysed the AAV vector integrity data. J.W., K.M. and S.A.S. analysed the mass spectrometry data. G.G. and D.W. supervised the study. J.W., Y.Z. and D.W. wrote the original draft with input from all authors.
Corresponding authors
Ethics declarations
Competing interests
J.W., Y.Z., C.A.M., G.G. and D.W. are inventors of a patent application filed by the University of Massachusetts Chan Medical School concerning the design and applications of sup-tRNAs described in this study. G.G. is a scientific co-founder of Voyager Therapeutics, Adrenas Therapeutics and Aspa Therapeutics and holds equity in these companies. G.G. and D.W. are inventors on patents related to AAV-based gene therapy, some of which were licensed to commercial entities. The other authors declare no competing interests.
Peer review
Peer review information
Nature thanks Steven Gray, Lynne Maquat, John Leuck and Christopher Trotta for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Sup-tRNAs suppressed UAG PTCs in HEK293 cells.
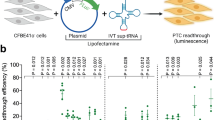
a, Workflow of a fluorescence reporter assay to examine sup-tRNA-induced readthrough of the Y39X premature termination codon (PTC) in the enhanced green fluorescence protein (EGFP) gene. Representative fluorescence images and quantification of fluorescent cells by flow cytometry are shown. Scale bar = 250 µm. b, Dual-luciferase reporter assay to quantify G418 (0.1 mg/mL) or sup-tRNA-induced readthrough at the Idua-W401X PTC. Readthrough efficiency is calculated as the normalized ratio of Gaussia luciferase (Gluc) activity to Cypridina luciferase (Cluc) activity. Data are mean ± s.d. of three biological replicates. c, Dual-luciferase reporter assay using escalating amounts of sup-tRNATyr plasmid in co-transfection. d, Dual-luciferase reporter assay using a series of constructs harboring different nucleotides immediately downstream of the W401X PTC (+4 nucleotide), in the absence or presence of G418 (0.1 mg/mL) or sup-tRNATyr plasmid co-transfection. e, Representative Western blot images and quantification of protein expression from mouse Idua cDNA variants that encode different amino acid residues at codon 401. Data are mean ± s.d. of three biological replicates. Statistical analysis was performed by one-way ANOVA followed by two-sided Dunnett’s multiple comparisons test (b and e). **p < 0.01, ***p < 0.001, ns: not significant. Nucleotide sequences of EGFP-Y39X and dual-luciferase reporters are shown in Supplementary Table 4. For gel source data, see Supplementary Figure 1.
Extended Data Fig. 2 Comparable expression level and enzymatic activity among Idua variants.
a, Representative images and quantification of EGFP+ human fibroblasts by flow cytometry following Lenti.EGFP infection. Scale bar = 250 µm. b, Workflow to examine mouse Idua cDNA variants expression and IDUA enzyme activity in IDUAW402X/W402X patient fibroblasts. The lentiviral construct expressing FLAG-tagged mouse IDUA variants is shown. c, Representative Western blot images and quantification of relative protein expression of mouse IDUA variants with different amino acid residues at codon 401. d, Absolute IDUA enzyme activity of different mouse IDUA variants. e, IDUA enzyme activity of different mouse IDUA variants normalized to expression level. Data are mean ± s.d. of three technical replicates (a), or of three biological replicates (c, d and e). Statistical analysis was performed by one-way ANOVA followed by two-sided Dunnett’s multiple comparisons (e). *p < 0.05, **p < 0.01, ***p < 0.001, ns: not significant. For gel source data, see Supplementary Figure 1.
Extended Data Fig. 3 Good correlation between replicates in Ribo-seq.
Density of coding sequence (CDS) is shown comparing two biological replicates of MPS-I patient fibroblasts treated under indicated conditions. Pearson correlations (two-sided) of log-transformed CDS densities and p values are shown. Histogram shows the CDS distribution sorted by binned densities.
Extended Data Fig. 4 Optimizing sup-tRNA expression cassette for in vivo rAAV delivery.
a, Schematic showing AAV9 vector constructs expressing 1x, 2x, or 4x sup-tRNATyr. Nucleotide sequences are shown in Supplementary Table 4. b, Workflow to study in vivo rAAV9 delivery of various sup-tRNATyr expression cassettes in the IduaW401X/W401X knock-in mice (KI/KI). c, IDUA enzymatic activity in liver (n = 5, 4, 6, 4 as indicated by individual circles) and heart (n = 5, 5, 5, 4 as indicated by individual circles) tissue lysates derived from mice treated by indicated AAV9 vectors. Data are mean ± s.d. Statistical analysis was performed by one-way ANOVA followed by two-sided Dunnett’s multiple comparisons test against the untreated group. d, Representative denaturing gel electrophoresis of extracted rAAV vector genomes of the 1x, 2x, 4x constructs of three technical replicates. Predicated vector genome sizes are labeled at the top. Black arrowheads: intact vector genomes; red arrows: truncated vector genomes. e, Characterization of serum IDUA activity (n = 4 per group), urine GAGs (n = 7 or 4 per group), liver IDUA activity (n = 3 or 5 per group), and liver GAGs (n = 3 or 5 per group) in KI/+ and wildtype (+/+) mice. Statistical analysis was performed by two-sided Student t-test. *p < 0.05, **p < 0.01, ns: not significant.
Extended Data Fig. 5 rAAV9.2xstRNATyr alleviated lysosomal abnormalities in MPS-I mice and yielded long-term (>6 months) therapeutic effect.
a-c, Western blot images and quantification of mouse LAMP1 protein expression (a), glucuronidase activity (b), and hexosaminidase activity (c) in the liver and heart of KI/KI mice with (+) or without (-) rAAV9.2xsup-tRNATyr treatment (n = 4 per group), KI/+ mice without (-) rAAV9.2xsup-tRNATyr treatment (n = 3). Data are mean ± s.d. of individual animals (circles). d, Serum IDUA enzyme activity in untreated KI/+ mice (green line, n = 8), untreated KI/KI mice (red line, n = 7) and KI/KI mice treated with rAAV9.2xsup-tRNATyr (blue line, n = 7) at various timepoints after treatment at 6 weeks old. IDUA activity in untreated KI/+ mice was normalized to 50% of that in WT (+/+) mice at each timepoint, and IDUA activity in treated KI/KI mice as percentage of that in WT (+/+) mice is labeled. e, Urine GAG levels in KI/KI with (+) or without (-) rAAV9.2xsup-tRNATyr treatment (n = 5 per group), KI/+ mice without (-) rAAV9.2xsup-tRNATyr treatment (n = 6) determined at 28 weeks post treatment. Statistical analysis was performed by one-way ANOVA followed by two-sided Dunnett’s multiple comparisons test (a-c and e). **p < 0.01, ***p < 0.001, ns: not significant. For gel source data, see Supplementary Figure 1.
Extended Data Fig. 6 rAAV9.2xstRNATyr rescued MPS-I phenotype in female mice.
a, Workflow to assess in vivo rAAV9.2xsup-tRNATyr treatment efficacy in female IduaW401X/W401X knock-in mice (KI/KI). b, IDUA enzymatic activity in liver and heart tissue lysates derived from KI/KI mice with (+) or without (-) rAAV9.2xsup-tRNATyr treatment (n = 4 and 3, respectively). Statistical analysis was performed by two-sided Welch’s t-test. c-e, Tissue GAG (c), glucuronidase activity (d), and hexosaminidase activity (e) levels in KI/KI mice with (+) or without (-) rAAV9.2xsup-tRNATyr treatment (n = 4 or 3 per group), KI/+ mice without (-) rAAV9.2xsup-tRNATyr treatment (n = 4). Data are mean ± s.d. of individual animals (circles). Statistical analysis was performed by one-way ANOVA followed by two-sided Dunnett’s multiple comparisons test (c-e). *p < 0.05, **p < 0.01, ***p < 0.001, ns: not significant.
Extended Data Fig. 7 No gross toxicity was observed in mice treated with rAAV9.2xsup-tRNATyr.
a, Representative H&E staining images of liver, brain, spleen, and kidney sections in untreated (n = 3) or sup-tRNATyr-treated (n = 4) male KI/+ mice. Mice were treated at 6 weeks old, and euthanized at 16 weeks old. Scale bar = 200 µm. b, Blind pathological assessment of the liver, brain, spleen, and kidney H&E slides as described in (a). c, Summary of endpoint serum clinical biochemistry of untreated (n = 3) or sup-tRNATyr-treated (n = 4) KI/+ mice. Data are mean ± s.d. of biological replicates. Statistical analysis was performed by two-sided Student t-test. *p < 0.05, **p < 0.01. Note that for the endpoints showing statistical significance, values of both groups are within the normal ranges (https://phenome.jax.org/projects/CGDpheno2).
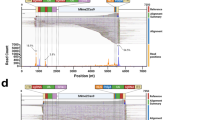
Extended Data Fig. 8 Ribosome profiling revealed that global readthrough is largely restricted to UAG in the liver.
a, Metagene plot showing normalized reads of ribosome protected fragments (RPFs) relative to the distance from the normal stop codon at position 0. RPFs from untreated (blue, n = 3 mice), or rAAV9.2xsup-tRNATyr-treated KI/KI male mice (red, n = 3 mice) are overlaid. b, Magnified view of the 3′UTR showing increased RPFs in this region in the rAAV9.2xsup-tRNATyr treated mouse livers. c, Box plot of ribosome readthrough score (RRTS) derived from ribosome profiling of KI/KI mouse livers with (+) or without (-) rAAV9.2xsup-tRNATyr treatment (n = 3 per group). RRTS values were calculated for transcripts harboring different normal stop codons, namely UAA (orange), UAG (blue), and UGA (green), respectively. Center line indicates the median, the box ends indicate the first and third quartiles, and the whiskers indicate the range of the remaining data excluding outliers. d, Scatter plot showing RPF densities at each codon (codon occupancy) in KI/KI mouse livers with (+) or without (-) rAAV9.2xsup-tRNATyr treatment (n = 3 per group). Two tyrosine (Y) codons are highlighted in red.
Extended Data Fig. 9 mRNA-seq analysis on human fibroblasts.
Cells were infected with Lenti.sup-tRNATyr.EGFP or Lenti.EGFP as control for seven days as described in Fig. 2a. a, Volcano plot showing differentially expressed transcripts in Lenti.sup-tRNATyr.EGFP-treated cells compared to control cells. Red and blue dots denote significantly upregulated and downregulated transcripts (adjusted p < 0.01 and fold change > 2), respectively. b, Venn diagram showing the upregulated transcripts in a and published set of 1,000 human putative NMD substrates.
Extended Data Fig. 10 rAAV.2xsup-tRNATyr treatment efficacy correlates with gene delivery efficiency.
a, IDUA activity in the tibialis anterior (TA) muscle lysates derived from KI/KI mice ± rAAV9.2xsup-tRNATyr treatment via either intravenous (IV) injection of 2x1012 GC (left panel; n = 4 per group) or from KI/KI mice ± rAAV9.2xsup-tRNATyr treatment via intramuscular (IM) injection of 3x1011 GC to the TA muscle (right panel; n = 4 or 6 per group). Tissues were harvested at 10 weeks (blue), 4 weeks (yellow), or 12 weeks (orange) post treatment. The right y-axis denotes absolute IDUA specific activity, and the left y-axis denotes relative activity normalized to untreated heterozygous mice as 50% of WT (+/+) level. b, Quantification of AAV vector genome copy numbers in the TA muscle of the mice described in a. c, IDUA enzymatic activity in the TA muscle lysates derived from KI/KI mice ± rAAVMYO.2xsup-tRNATyr treatment via IV injection at 2x1012 GC (n = 4 or 3 per group), and euthanized at 4 weeks post treatment. d, IDUA activity in the brain lysates derived from KI/KI mice ± rAAV.2xsup-tRNATyr treatment via intravenous (IV) injection. Mice were treated with either rAAV9 (left panel; n = 4) at 2x1012 GC or rAAV.PHPeB (right panel; n = 5 per group) at 1x1011 GC (yellow) or 2x1011 GC (orange). e, Quantification of AAV vector genome copy numbers in the brain of the mice described in d. f, IDUA activity in the hippocampus lysates derived from KI/KI mice ± rAAV9.2xsup-tRNATyr treatment via intrahippocampal injection of 2.2x1010 GC (n = 4 or 3 per group). Mice were euthanized at 4 weeks post treatment. Data are mean ± s.d. of individual animals (circles). Statistical analysis was performed by one-way Brown-Forsythe and Welch ANOVA followed by two-sided Dunnett T3’s multiple comparisons test (a, c, d, f), or ANOVA followed by two-sided Dunnett’s multiple comparison test (b, e). *p < 0.05, **p < 0.01, ***p < 0.001, ns: not significant.
Supplementary information
Supplementary Information
This file contains Supplementary Figs. 1 and 2.
Supplementary Tables
This file contains Supplementary Tables 1–5.
Source data
Rights and permissions
About this article
Cite this article
Wang, J., Zhang, Y., Mendonca, C.A. et al. AAV-delivered suppressor tRNA overcomes a nonsense mutation in mice. Nature 604, 343–348 (2022). https://doi.org/10.1038/s41586-022-04533-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-022-04533-3
This article is cited by
-
Translation velocity determines the efficacy of engineered suppressor tRNAs on pathogenic nonsense mutations
Nature Communications (2024)
-
Adeno-associated virus as a delivery vector for gene therapy of human diseases
Signal Transduction and Targeted Therapy (2024)
-
Hijacking endogenous mRNA for genetic code expansion
Nature Chemical Biology (2024)
-
Deep generative design of RNA family sequences
Nature Methods (2024)
-
Tracking endogenous proteins based on RNA editing-mediated genetic code expansion
Nature Chemical Biology (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.