Abstract
The emergence of SARS-CoV-2 variants of concern with progressively increased transmissibility between humans is a threat to global public health. The Omicron variant of SARS-CoV-2 also evades immunity from natural infection or vaccines1, but it is unclear whether its exceptional transmissibility is due to immune evasion or intrinsic virological properties. Here we compared the replication competence and cellular tropism of the wild-type virus and the D614G, Alpha (B.1.1.7), Beta (B.1.351), Delta (B.1.617.2) and Omicron (B.1.1.529) variants in ex vivo explant cultures of human bronchi and lungs. We also evaluated the dependence on TMPRSS2 and cathepsins for infection. We show that Omicron replicates faster than all other SARS-CoV-2 variants studied in the bronchi but less efficiently in the lung parenchyma. All variants of concern have similar cellular tropism compared to the wild type. Omicron is more dependent on cathepsins than the other variants of concern tested, suggesting that the Omicron variant enters cells through a different route compared with the other variants. The lower replication competence of Omicron in the human lungs may explain the reduced severity of Omicron that is now being reported in epidemiological studies, although determinants of severity are multifactorial. These findings provide important biological correlates to previous epidemiological observations.
Similar content being viewed by others
Main
Since the emergence of COVID-19 in December 2019, the causative agent SARS-CoV-2 has continued to evolve in humans, generating variants of progressively increased transmissibility between humans. The World Health Organization (WHO) has classified several lineages as variants of concern (VOCs) on the grounds of their high transmission rate, potential for immune evasion, unusual epidemiological properties, or adverse impact on diagnostics and therapeutics2. These VOCs include the Alpha, Beta, Gamma, Delta and Omicron virus lineages.
The first virus variant emerged in February 2020 carrying the D614G amino acid substitution in the S protein and this quickly became the dominant virus variant globally. It was shown to replicate more rapidly in primary airway epithelial cells and in the nasal cavity of experimentally infected hamsters, explaining its greater transmissibility in humans3. The Alpha lineage, which was first reported in the United Kingdom in September 2020, had 13 mutations in the spike (S) protein, and the N501Y and P681H mutations in the spike protein contributed to its increased transmissibility4,5. The deletion at position 60–70 in the spike protein led to a loss of detection of the S gene target in some diagnostic tests (by PCR with reverse transcription (RT–PCR)), termed S-gene target failure6. The Beta lineage first detected in South Africa in August 2020, contained 10 mutations in the S protein and mutations N501Y and K417N enhanced viral transmission while E484K contributed to immune evasion7,8. The Delta lineage was first identified in the state of Maharashtra, India in October 2020, had multiple amino acid substitutions in the S protein—including L452R and E484Q, which enhance angiotensin-converting enzyme 2 (ACE2) binding, transmission and immune evasion9,10. The Omicron variant was identified in Botswana and South African in November 2021 and was designated as VOC by the WHO in the same month. It has 37 amino acid substitutions in the S protein, 15 of which are in the receptor-binding domain. Thus, it was probable that the phenotype of the virus would be markedly affected for transmission, escape from previous immunity or both. As of 14 December 2021, the Omicron variant has been detected in 76 countries globally11. Differences in disease severity between variants has been more subtle. A modest increase in disease severity has been reported for the Delta VOC, with higher hospital admissions compared with the Alpha VOC12,13. The extensive amino acid substitutions in the spike protein of Omicron14 are likely to have major implications on transmissibility, disease severity and immune evasion from serum neutralizing antibodies from both infection- and vaccine-elicited and therapeutic monoclonal antibodies. However, there is limited information on the biological or virological characteristics of this variant. Omicron replication is ACE2 dependent and it has a considerable ability to evade neutralizing antibodies elicited by past infection or vaccination1.
The correlation of the phenotype of a SARS-CoV-2 variant in vitro with epidemiology is well illustrated by the experimental observations with the D614G mutation. An isogenic wild-type (WT) virus engineered to carry the spike D614G substitution had enhanced viral replication in human lung epithelial cells and primary human airway tissues by increasing the infectivity and stability of virions15. Hamsters infected with this virus produced higher infectious titres in nasal washes and the trachea, but not in the lungs, supporting clinical evidence showing that the mutation enhances viral loads in the upper respiratory tract of patients with COVID-19 and may increase transmission. However, studies using physiologically relevant experimental models investigating the phenotype of the Omicron variant in comparison to previous variants of SARS-CoV-2 are lacking.
We have previously used ex vivo explant cultures of the human bronchi and lung parenchyma to study the competence of viral replication and the cellular tropism of avian influenza viruses16,17, MERS18,19,20 and SARS-CoV-2 (ref. 21). This provided a suitable platform to rapidly compare the replication profile and tropism of the Omicron variant with others and to provide insights into the observed epidemiology of this variant. We initially compared the viral replication kinetics of WT SARS-CoV-2 with the D614G variant and VOCs from Alpha and Beta lineages in ex vivo explant cultures of human bronchi and lungs. In a subsequent set of experiments, we compared the viral replication profiles of the WT with the Delta and B.1.1.529/Omicron lineages. We also investigated the cellular tropism of the virus using immunohistochemistry and compared the dependency of WT, Delta and Omicron variants on transmembrane serine protease 2 (TMPRSS2) and cathepsins for their replication.
Higher replication of Omicron in bronchi
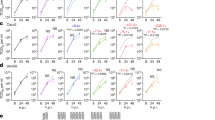
We first compared the replication kinetics of WT, D614G, Alpha and Beta strains that were isolated from returning travellers or the community in Hong Kong in ex vivo cultures of human bronchi and lungs by titrating infectious virus using 50% tissue culture infectious dose (TCID50) titrations. The only significant difference observed was a higher replication of the Beta variant in human bronchi at 72 hours post-inferction (hpi); no significant difference was noted between viruses at 24 and 48 hpi (Fig. 1a). When WT, Delta and Omicron variants were compared, the Omicron variant replicated to significantly higher titres compared with the WT or Delta at 24 and 48 hpi—a difference of greater than 70-fold (Fig. 1b). At 72 hpi, both Delta and Omicron viruses replicated significantly more than the WT strain in human bronchi, but there was no significant difference between the Delta and Omicron viruses. When these experiments were performed at 33 °C, the viral titres were similar to those at 37 °C for each virus (data not shown). In ex vivo cultures of human lung tissue, the only significant difference observed was a reduction in replication of Omicron compared with the WT strain at 24, 48 and 72 hpi (Fig. 1c, d). These findings in bronchus and lung tissues were confirmed in area under the curve (AUC) analysis of aggregate virus titres at 24–72 hpi (Fig. 1e, f). The individual donor datasets of viral titres for the comparison of WT, Delta and Omicron are shown in Extended Data Fig. 1. The same consistent trend was observed for each individual donor. Similar trends were observed between ORF1b gene quantification and TCID50 results—a higher level of viral gene was detected in bronchi infected with the Omicron variant compared with the WT and Delta strains, while a higher number of viral gene copies was measured in lung tissues infected with the WT strain compared with the Delta and Omicron variants (Extended Data Fig. 2).
a–d, Human ex vivo cultures of bronchi and lungs were infected with 5 × 105 TCID50 per ml at 37 °C. Viruses released in the culture supernatants were measured over time using the TCID50 assay. a, c, Viral replication kinetics of SARS-CoV-2 WT, D614G, Alpha and Beta in human ex vivo cultures of bronchi (a) and lungs (c). b, d, Viral replication kinetics of SARS-CoV-2 WT, Delta and Omicron in human ex vivo cultures of bronchi (b) and lungs (d). The horizontal dotted line denotes the limit of detection in the TCID50 assay. Data are geometric mean ± s.d. n = 6. Statistical analysis was performed using two-way analysis of variance (ANOVA) followed by Tukey’s test; P < 0.05 was considered to be statistically significant, and exact P values are presented. e, f, Viral titres from a (e) to d (f) are depicted as AUC. Data are the geometric mean ± s.d. n = 6. Statistical analysis was performed using one-way ANOVA followed by Tukey’s test; P < 0.05 was considered to be statistically significant, and exact P values are presented.
The tropism of each virus variant in the bronchi and lungs was visualized through immunohistochemical staining of the SARS-CoV-2 nucleoprotein (Fig. 2). The virus variants did not appear to differ in cell tropism in the bronchus tissues that were infected ex vivo. For Omicron, there was evidence of extensive virus infection—the ciliated epithelium, goblet and club cells, in which infection of WT, Alpha, Beta, Delta and Omicron viruses was observed on the basis of immunofluorescence staining (Figs. 2a and 3). Transmission electron microscopy also showed the presence of Omicron viral particles in membrane-bound vesicles in the cytoplasm, as well as on the surface attached to microvilli of ciliated cells (Extended Data Fig. 4). The virus variants did not appear to differ in cell tropism in the lung tissue (Fig. 2b). Immunohistochemical staining showed positive staining in the lung tissues, with the spindle-shaped antigen-positive cells morphologically resembling type 1 pneumocytes. In lung tissues infected with Omicron, there were decreased staining intensities of viral antigen identified (Extended Data Fig. 3). By contrast, greater numbers of cells with positive staining for viral antigen were observed in the bronchial epithelium infected with Omicron compared with all of the other variants and the WT virus.
a, b, Ex vivo cultures of human bronchi (a) and lungs (b) infected with WT and various variants of SARS-CoV-2 with 5 × 105 TCID50 per ml or mock at 37 °C. The tissues were fixed with formalin at 72 hpi Paraffin-embedded sections were analysed using immunohistochemical staining with a polyclonal antibody against the SARS-CoV-2 nucleoprotein. Positive cells are red-brown. Scale bars, 100 μm. The images are representative of two individual donors.
Ex vivo cultures of human bronchi were infected with SARS-CoV-2 WT, Alpha, Beta, Delta, Omicron or mock. At 72 hpi, the tissues were fixed with formalin, embedded in paraffin and stained for immunofluorescence (green) analysis of the indicated cell markers: acetyl-α-tubulin+ (ciliated cells), mucin 5AC+ (secretory goblet cells) and club cell protein 10+ (club cells); monoclonal antibodies against the SARS-CoV-2 nucleoprotein (red) and 4′,6-diamidino-2-phenylindole-positive for nuclei (blue) were used. The images are representative of two individual donors. Scale bars, 100 μm.
ACE2 and TMPRSS2 expression in the respiratory tract
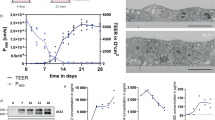
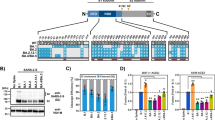
ACE2 and transmembrane serine protease 2 (TMPRSS2) contribute to the entry of SARS-CoV-2 into cells. The latter cleaves the spike S2 domain (S2′), enabling the virus to enter the cell through cell fusion at the outer cell membrane, distinct from viral entry through the endocytic pathway22. ACE2 expressed at the cell surface exists as short and long forms, with the short form lacking the domain that binds to SARS-CoV-2 spike23. We showed that there was significantly higher expression of both long- and short-form ACE2 in human bronchi compared with in the lungs (Fig. 4a). Immunohistochemical staining also showed clear evidence of more extensive ACE2 staining in the bronchi compared with in the lungs (Fig. 4b and 4c). We next investigated the TMPRSS2 expression and found significantly higher expression of TMPRSS2 mRNA in the bronchi compared with in the lungs (Fig. 4a).
a, The mRNA expression of both the long and short forms, the short form and the long form of ACE2, and TMPRSS2 was measured by using quantitative PCR with reverse transcription (RT–qPCR). The fold change in expression in bronchi (n = 6) and lungs (n = 8) was normalized to the average gene expression of lung tissues. Data are mean ± s.d. Statistical analysis was performed using unpaired two-sided Student’s t-tests. b, c, Immunohistochemical staining of paraffin-embedded sections with antibodies against non-specific forms (b) and the long form (c) of ACE2 in bronchi and lungs. Positive cells are red-brown. Scale bars, 100 μm. The images are representative of two individual donors. d, Viral replication kinetics of WT, Delta and Omicron in Vero E6 (E6) and Vero E6-TMPRSS2 (T2) cells determined using the TCID50 assay. Cells were infected with SARS-CoV-2 viruses at a multiplicity of infection (MOI) of 0.01. Data are the geometric mean ± s.d. of the virus titre. n = 3. The horizontal dotted line denotes the detection limit of the TCID50 assay. Statistical analysis was performed using two-way ANOVA followed by Tukey’s test. e, The viral titres from d are shown as AUC. Data are mean ± s.d. n = 3. Statistical analysis was performed using one-way ANOVA followed by Tukey’s test. f, g, The effects of treatment with camostat mesylate (Cam) and E64d on viral replication. T2 cells were pretreated with camostat mesylate, E64d or both for 1 h. Infection of SARS-CoV-2 WT, Delta and Omicron was performed at a MOI of 0.01. After infection, the cells were replenished with fresh medium with the drug. ORF1b gene copies were measured in culture supernatants at 24 hpi using RT–qPCR. Data are the mean ± s.d. percentages of ORF1b gene copies using vehicle treatment as reference. n = 6. Statistical analysis was performed using one-way ANOVA followed by Tukey’s test. P < 0.05 was considered to be statistically significant, and exact P values are presented.
Omicron preference for endocytic entry
We investigated WT, Delta and Omicron virus replication for their dependence on TMPRSS2. We compared the viral replication kinetics of the WT, Delta and Omicron strains in Vero E6 and Vero E6-TMPRSS2 (E6/T2) cells. Although virus titres at 24 hpi were higher in E6/T2 cells for the WT and Delta, the magnitude of the difference was 100–1,000-fold for Delta, 3–100-fold for Omicron and around 10-fold for the WT (Fig. 4d). The AUC levels of the viral titres at 24–48 hpi confirmed these findings (Fig. 4e). Omicron replicated to lower titres compared with the WT and Delta strains at 24 hpi in E6/T2 cells (Extended Data Fig. 5), which was also shown in their AUC levels (Fig. 4e). We next infected Vero E6/T2 cells with the WT, Delta and Omicron strains in the presence of camostat mesylate (an inhibitor of serine proteases, including TMPRSS2)22 or E64d (an inhibitor of cathepsins)24. The viability of Vero E6/T2 cells was above 80% after treatment with camostat mesylate at concentrations up to 300 µM or E64d at concentrations up to 120 µM (Extended Data Fig. 6). Although both WT and Omicron infection was partially reduced by camostat mesylate, Delta was more sensitive to the inhibitor (Fig. 4f, g). By contrast, while both WT and Delta infection was partially suppressed by E64d, Omicron infection was reduced more substantially to around 5% of the control (Fig. 4g). The combined treatment of camostat mesylate and E64d completely blocked the infection of all three viruses (Fig. 4f). This suggests that, in contrast to Delta, Omicron may be more dependent on the endocytic pathway to enter cells.
Discussion
Preliminary epidemiological data from the United Kingdom suggest that the risk of household transmission with Omicron is 3.2 times (95% confidence interval 2.0–5.0) higher than with Delta, which in turn was a highly transmissible virus compared with the WT virus. Omicron has a reproduction number (R) of 3.7 (ref. 25). It has been unclear whether this increased efficiency of transmission is driven by mutations that confer immune escape from antibodies in the population elicited by previous infection or vaccination, by intrinsic viral factors or a combination of the two. It is increasingly clear that high neutralizing antibody titres elicited by previous infection or immunization are markedly compromised by Omicron1. Our results suggest that the Omicron variant has a substantial (over 70-fold increase) and significantly higher replication competence in the human bronchi compared with both WT and Delta viruses at 24 hpi The extent to which this faster replication in the bronchi might contribute to transmissibility is unclear, but higher infectious virus load in conducting airways might result in increased amounts of infectious virus released while breathing or speaking, therefore enhancing transmission through the airborne route. There are also recent reports suggesting that Omicron has increased viral replication compared with the Delta and WT virus in differentiated human nasal epithelial cultures in vitro26. Infectious SARS-CoV-2 has been detected in fine aerosol particles in air exhaled by patients with COVID-1927. The mechanistic reasons underlying the increased replication competence in the bronchi remain to be elucidated. The Omicron variant has 37 amino acid substitutions in the spike protein, 15 of which are in the receptor-binding domain14. Infection with Omicron is ACE2 dependent1 and the binding of the spike of Omicron spike to ACE2 is enhanced compared with that of the WT virus28. We showed that there is more extensive ACE2 expression in human bronchi compared with in the lungs, which may explain the enhanced replication of SARS-CoV-2 in the bronchi. Omicron also has amino acid substitutions in the nucleocapsid protein (R203K and G204R) that are associated with enhanced virus replication29.
Epidemiological studies suggested that the Delta variant was substantially more transmissible than Alpha variant30, which itself was more transmissible than the earlier virus strains4. Thus, our data indicating that Delta variant has higher titres of infectious virus than WT virus in the bronchi is also compatible with the epidemiological observations. It has been shown that pseudoviruses expressing Delta spike have a greater ability to infect ACE2low human bronchial epithelial cells compared with the previous variants31. Moreover, the spike protein of Delta is in a predominantly cleaved state, which may promote its replication efficiency in human airway32. P681R enhanced the cleavage of furin-cleavage sequences, which contributes to the enhanced fitness of Delta over Alpha in a competition assay in Calu-3 and human airway epithelium in vitro models33,34.
Our data showing that the Omicron variant has lower viral replication competence in the lungs compared with the bronchi are of particular interest. This difference is also confirmed by the immunohistochemistry studies showing less virus infected cells in human lung explant cultures ex vivo. The biological determinants of this divergence in comparative replication competence of Omicron and Delta variants in the bronchi and lungs remain to be examined. These observations may suggest that Omicron may have reduced clinical severity, but such interpretations need to be qualified because the disease severity of COVID-19 is determined not only by virus replication but also by dysregulated innate immune responses. Recent epidemiological studies from South Africa and the United Kingdom suggest that Omicron has caused less hospitalization compared with Delta35,36. Thus, the findings from the ex vivo cultures are in concordance with recent epidemiological assessments both in regard to transmission and disease severity.
Our finding that Omicron infection was less dependent on TMPRSS2 activities but more sensitive to a cathepsin inhibitor compared with Delta suggests that Omicron may enter cells primarily through the endocytic pathway, wheras Delta preferentially enters cells through fusion at the cell surface. The adoption of a ubiquitous endocytic pathway potentially broadens the cellular spectrum for Omicron infection and enables Omicron to infect cells with ACE2 expression irrespective of the presence of TMPRSS2. Single-cell sequencing results indicate that cells with co-expression of ACE2 and cathepsins are more abundant in the upper airway than cells that co-express ACE2 and TMPRSS2, which may explain the increased replication competence of Omicron in the bronchi26. Our findings that Omicron preferentially enters cells through the endosomal pathway while Delta is more reliant on the cell surface fusion is consistent with the reports on alveolar epithelial cells and nasal epithelial cells24,26. The use of therapeutic inhibitors of TMPRSS2 may be of limited benefit in the management of clinical infections with the Omicron variant. Although there have been single-cell studies on the distribution of TMPRSS2 in cell cultures, the data on the extent of its distribution are hampered by the sensitivity and specificity of antibodies in fixed tissues.
One of the limitations of this study is that only one virus strain from each lineage was tested. Furthermore, all six virus variants were not tested in parallel in the same experiment, but this is very difficult to carry out logistically as there is limited tissue available to test six viruses with replicates.
In summary, our findings reveal that Omicron has faster and enhanced viral replication efficiency in the human bronchi compared with the previous lineages, suggesting that it has an intrinsic capacity for enhanced transmission. The lower replication competence of Omicron in human lungs is compatible with lower disease severity compared with Delta. Both of these observations are concordant with epidemiological data. Even if disease severity is modestly reduced, the very efficient transmissibility of Omicron will pose a major threat to global public health and health care systems. Investigations on preventing Omicron infection through vaccination boosters and therapeutic options are urgently needed.
Methods
SARS-CoV-2 isolation
Vero E6 cells (ATCC) were used for virus isolation and propagation of the WT virus, D614G strain and Alpha variant, and Vero E6-TMPRSS2 overexpressed cells (provided by M. Takeda)37 were used for the Beta, Delta and Omicron variants. Both cell lines were cultured in DMEM with 10% FBS. The original clinical samples were collected from patients with a confirmed SARS-CoV-2 infection in Hong Kong from January 2020 to November 2021 (Extended Data Table 1) and isolated as previously described38. Viruses were isolated from clinical samples of the nasopharyngeal and throat swab from patients infected with SARS-CoV-2 in virus transport medium. Vero E6-TMPRSS2 (E6/T2) cells seeded at 1 × 105 cells in 24-well plate were inoculated with 50 µl of sample and topped up with 2% FBS DMEM medium to 1 ml for 1 h at 37 °C. The cells were washed once with PBS, replenished with fresh medium and observed daily for cytopathic effect (CPE). Culture supernatants were collected when the CPE reached around 50% and was defined as P1. The virus was further propagated in E6/T2 cells. The virus stock was aliquoted and stored frozen at −80 °C. Aliquots were titrated to TCID50 in respective cell lines. All cell lines used were mycoplasma free. The experiments were carried out in a bio-safety level 3 (BSL-3) facility at the School of Public Health, LKS Faculty of Medicine, The University of Hong Kong. Informed consent was obtained from all of the participants and approval was granted by the Institutional Review Board (IRB) of the University of Hong Kong and the Hospital Authority (Hong Kong West) (IRB approval no. UW 20-862 and UW20-863).
Ex vivo cultures and infection of human respiratory tract
Fresh non-tumour bronchi (n = 12) and lungs (n = 12) tissues were obtained from patients aged 51–78 years undergoing elective surgery in Department of Surgery at Queen Mary Hospital from May to December 2021 and were removed as part of routine clinical care but surplus for routine diagnostic requirements as described previously39,40. The donor information is listed in the Extended Data Table 2. The virus infection procedures were performed as previously described38. In brief, similarly sized pieces of human bronchus and lung tissues were infected with each virus at 5 × 105 TCID50 per ml for 1 h at 37 °C. Each tissue fragment was washed three times in culture medium to remove residual virus inoculum, topped up with fresh medium and incubated at 37 °C as indicated. Mock-infected tissues were used as negative controls. Aliquots of culture medium were removed at the indicated times and stored at −80 °C until titration. Infectious viral titres in culture supernatants were assayed by TCID50 in Vero E6 or Vero E6-TMPRSS2 cells, respectively, depending on the cells used for virus isolation and passage. Infected tissues were fixed in 10% formalin and processed for immunostaining at 72 hpi. Bronchus tissues were also fixed in formalin and then processed for transmission electron microscopy as previously described41.
Viral titration by TCID50 assay
A confluent 96-well tissue culture plates of Vero-E6 or Vero E6-TMPRSS2 cells was prepared one day before the virus titration (TCID50) assay. Cells were washed once with PBS and replenished with DMEM (Gibco) with 2% foetal bovine serum (Gibco) supplemented with 100 U ml−1 penicillin and 100 µg ml−1 streptomycin (Gibco). Serial dilutions of virus supernatant, from 0.5 log to 7 log, were performed and each virus dilution was added to the plates in quadruplicate. The plates were observed for cytopathic effect daily. The end point of viral dilution leading to CPE in 50% of inoculated wells was estimated using the Karber method42. AUC was calculated from the viral titres from different time points indicated in the y axis.
Immunohistochemistry staining of paraffin-embedded tissues
Human respiratory tract tissues (lungs and bronchi) were fixed with 10% formalin overnight at 4 °C and the fixed tissues were embedded in paraffin blocks. For immunohistochemistry, the 4 µm sliced sections were microwaved for 15 min for antigen retrieval. Endogenous peroxidase activity was stopped by quenching the tissue sections with 3% H2O2 for 20 min. The slides were then blocked with 10% normal horse serum at room temperature and incubated with primary antibodies (anti-SARS-CoV-2 nucleoprotein (NP) (40143-T62, Sino Biological) or ACE2 (long form, ab108252; short form, ab15348, Abcam)) for 90 min at room temperature followed by horseradish peroxidase-conjugated anti-rabbit antibodies (Vector Laboratory). The sections were developed using NovaRED Substrate Kit (Vector Laboratory). The cell nuclei were counterstained with Mayer’s Hematoxylin.
To characterize the SARS-CoV-2-infected cells, double-antibody immunofluorescence staining against SARS-CoV-2 NP and different cellular markers was performed. The tissue sections were first stained with SARS-CoV-2 NP antibody similar to as mentioned above except, after SARS-CoV-2 NP antibody incubation, the sections were incubated with alkaline phosphatase-conjugated anti-rabbit or anti-mouse antibodies (Vector Laboratory) and developed using the Vector Red (VR) Substrate Kit (Vector Laboratory). The sections were then microwaved, incubated with SCGB1A1/CC10 (Protein-Tech), acetylated α-tubulin (Santa Cruz), MUC5AC (Thermo-Fisher), for 90 min at room temperature followed by goat anti rabbit-AF488 or goat anti mouse-AF488. The cell nuclei were counterstained with DAPI (blue). The sections were imaged using the Nikon Eclipse Ti-S microscope.
Quantification of immunohistochemistry
Immunohistochemically stained samples were analysed using ImageJ Fiji (v.2.1.0) as previously described43,44. In brief, each original IHC (RGB) image was split into single-coloured images and Nova-red (SARS-CoV-2 NP) stained image was selected for the quantification of SARS-CoV-2 NP expression. The Nova-red (SARS-CoV-2 NP) stained signal was quantified after adjusting the threshold, which is set at the same value for all the images. The results are presented as frequency of expression and was calculated as percentage of positively stained area divided by the total image area of lung tissues or by the epithelium area of bronchial tissues.
RT–qPCR
The viral RNA in culture supernatants was extracted using the QIAamp Viral RNA Mini Kit (Qiagen). RNA was reverse-transcribed by using random 6-mer primers with the PrimeScript RT Reagent Kit (Takara). mRNA expression of target genes was detected using the corresponding primers (Extended Data Table 3) using an ABI ViiA 7 real-time PCR system (Applied Biosystems). All of the procedures were performed according to the manufacturers’ instructions. The viral gene and gene expression profiles of cell lysates were quantified and normalized to β-actin as previously described45,46,47,48.
Replication of SARS-CoV-2 in Vero-E6 and Vero-E6-TMPRSS2 cells and drug treatment
Vero-E6 and Vero-E6-TMPRSS2 cells in 48-well plates were infected with SARS-CoV-2 at a MOI of 0.01 for viral replication kinetics. Viral titres in culture supernatants (1–48 hpi) were determined using TCID50 assay (see above). To assess the effect of TMPRSS2 inhibitor and cathepsins inhibitor, the Vero-E6-TMPRSS2 cells were infected with SARS-CoV-2 at a MOI of 0.01. Camostat mesylate (Sigma-Aldrich) or E64d (Sigma-Aldrich) at 30 µM was added 1 h before, during and after infection. Vehicle was used as a negative control. Viral replication was evaluated by quantification of SARS-CoV-2 ORF1b copy number in the culture supernatants by using qPCR at 24 hpi (ref. 48).
Quantification of cell viability
Cell viability of Vero-E6-TMPRSS2 after drug treatment was evaluated using the Cell Counting Kit 8 (WST-8/CCK8) (Abcam). In brief, cells grown to 70% confluency in 48-well plates were incubated for 24 h in the absence or presence of various concentrations (0.41–300 µM) of camostat mesylate and (1.88–120 µM) of E64d. Next, the culture medium was aspirated and washed once with PBS, incubated with CCK8 working solution (CCK8:medium, 1:10) at 37 °C, 5% CO2 for 2 h. Subsequently, optical density values at 450 nm were measured using a multiplate reader (BMG FLUOstar OPTIMA). Cell viability was expressed as a percentage of that of the control cells.
Statistical analysis
Experiments with the human ex vivo cultures were performed independently with six different donors. The results shown in the figures are the geometric mean ± s.d. AUC was calculated by integrating infectious virus titres at 24–72 hpi in ex vivo bronchus or lung tissues or at 24–48 hpi in cell lines. The differences in log10-transformed viral titres and quantitative viral RNA of ORF1b between viruses and over time were compared using two-way ANOVA followed by Tukey’s multiple-comparison test using GraphPad Prism v.9.1.2. Comparisons of AUC and quantitative viral RNA of ORF1b between viruses were calculated using one-way ANOVA followed by Tukey’s multiple-comparison test. Differences were considered to be significant at P < 0.05.
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this paper.
Data availability
All data generated or analysed during this study are included in the Article. The sequences of all SARS-CoV-2 viruses used in this study are available in Genbank , and a list of the accession IDs is provided in Extended Data Table 1. Source data are provided with this paper.
References
Cele, S. et al. Omicron extensively but incompletely escapes Pfizer BNT162b2 neutralization. Nature https://doi.org/10.1038/s41586-021-04387-1 (2021).
Tracking SARS-CoV-2 Variants (WHO, 2021); https://www.who.int/en/activities/tracking-SARS-CoV-2-variants/
Baric, R. S. Emergence of a highly fit SARS-CoV-2 variant. N. Engl. J. Med. 383, 2684–2686 (2020).
Davies, N. G. et al. Estimated transmissibility and impact of SARS-CoV-2 lineage B.1.1.7 in England. Science 372, eabg3055 (2021).
Lubinski, B. et al. Functional evaluation of the P681H mutation on the proteolytic activation of the SARS-CoV-2 variant B.1.1.7 (Alpha) spike. iScience 25, 103589 (2022).
Srivastava, S., Banu, S., Singh, P., Sowpati, D. T. & Mishra, R. K. SARS-CoV-2 genomics: an Indian perspective on sequencing viral variants. J. Biosci. 46, 22 (2021).
Zhou, D. et al. Evidence of escape of SARS-CoV-2 variant B.1.351 from natural and vaccine-induced sera. Cell 184, 2348–2361 (2021).
Khan, A. et al. Higher infectivity of the SARS-CoV-2 new variants is associated with K417N/T, E484K, and N501Y mutants: an insight from structural data. J. Cell. Physiol. 236, 7045–7057 (2021).
Ozono, S. et al. SARS-CoV-2 D614G spike mutation increases entry efficiency with enhanced ACE2-binding affinity. Nat. Commun. 12, 848 (2021).
Augusto, G. et al. In vitro data suggest that Indian Delta variant B.1.617 of SARS-CoV-2 escapes neutralization by both receptor affinity and immune evasion. Allergy 77, 111–117 (2021).
Weekly Epidemiological Update on COVID-19—14 December 2021 (WHO, 2021).
Sheikh, A., McMenamin, J., Taylor, B. & Robertson, C. SARS-CoV-2 Delta VOC in Scotland: demographics, risk of hospital admission, and vaccine effectiveness. Lancet 397, 2461–2462 (2021).
Twohig, K. A. et al. Hospital admission and emergency care attendance risk for SARS-CoV-2 Delta (B.1.617.2) compared with Alpha (B.1.1.7) variants of concern: a cohort study. Lancet Infect. Dis. 22, 35–42 (2021).
Gu, H. et al. Probable transmission of SARS-CoV-2 Omicron variant in quarantine hotel, Hong Kong, China, November 2021. Emerg. Infect. Dis. 28, 460–462 (2021).
Plante, J. A. et al. Spike mutation D614G alters SARS-CoV-2 fitness. Nature 592, 116–121 (2021).
Hui, K. P. et al. Tropism and innate host responses of influenza A/H5N6 virus: an analysis of ex vivo and in vitro cultures of the human respiratory tract. Eur. Respir. J. 49, 1601710 (2017).
Chan, M. C. et al. Tropism and innate host responses of a novel avian influenza A H7N9 virus: an analysis of ex-vivo and in-vitro cultures of the human respiratory tract. Lancet Respir. Med. 1, 534–542 (2013).
Chan, R. W. et al. Tropism and replication of Middle East respiratory syndrome coronavirus from dromedary camels in the human respiratory tract: an in-vitro and ex-vivo study. Lancet Respir. Med. 2, 813–822 (2014).
Zhou, Z. et al. Phenotypic and genetic characterization of MERS coronaviruses from Africa to understand their zoonotic potential. Proc. Natl Acad. Sci. USA 118, e2103984118 (2021).
Chu, D. K. W. et al. MERS coronaviruses from camels in Africa exhibit region-dependent genetic diversity. Proc. Natl Acad. Sci. USA 115, 3144–3149 (2018).
Hui, K. P. Y. et al. Tropism, replication competence, and innate immune responses of the coronavirus SARS-CoV-2 in human respiratory tract and conjunctiva: an analysis in ex-vivo and in-vitro cultures. Lancet Respir. Med. 8, 687–695 (2020).
Hoffmann, M. et al. SARS-CoV-2 cell entry depends on ACE2 and TMPRSS2 and is blocked by a clinically proven protease inhibitor. Cell 181, 271–280 (2020).
Blume, C. et al. A novel ACE2 isoform is expressed in human respiratory epithelia and is upregulated in response to interferons and RNA respiratory virus infection. Nat. Genet. 53, 205–214 (2021).
Willett, B. J. et al. The hyper-transmissible SARS-CoV-2 Omicron variant exhibits significant antigenic change, vaccine escape and a switch in cell entry mechanism. Preprint at https://doi.org/10.1101/2022.01.03.21268111 (2022).
SARS-CoV-2 Variants of Concern and Variants Under Investigation (UK Health Security Agency, 2021).
Peacock, T. P. et al. The SARS-CoV-2 variant, Omicron, shows rapid replication in human primary nasal epithelial cultures and efficiently uses the endosomal route of entry. Preprint at https://doi.org/10.1101/2021.12.31.474653 (2022).
Adenaiye, O. O. et al. Infectious SARS-CoV-2 in exhaled aerosols and efficacy of masks during early mild infection. Clin. Infect. Dis. https://doi.org/10.1093/cid/ciab797 (2021).
Cameroni, E. et al. Broadly neutralizing antibodies overcome SARS-CoV-2 Omicron antigenic shift. Nature https://doi.org/10.1038/s41586-021-04386-2 (2021).
Wu, H. et al. Nucleocapsid mutations R203K/G204R increase the infectivity, fitness, and virulence of SARS-CoV-2. Cell Host Microbe 29, 1788–1801 (2021).
Allen, H. et al. Household transmission of COVID-19 cases associated with SARS-CoV-2 Delta variant (B.1.617.2): national case-control study. Lancet Reg. Health. Eur. 12, 100252 (2022).
Li, H., Liu, T., Wang, L., Wang, M. & Wang, S. SARS-CoV-2 Delta variant infects ACE2(low) primary human bronchial epithelial cells more efficiently than other variants. J. Med. Virol. https://doi.org/10.1002/jmv.27372 (2021).
Mlcochova, P. et al. SARS-CoV-2 B.1.617.2 Delta variant replication and immune evasion. Nature 599, 114–119 (2021).
Liu, Y. et al. Delta spike P681R mutation enhances SARS-CoV-2 fitness over Alpha variant. Preprint at https://doi.org/10.1101/2021.08.12.456173 (2021).
Lubinski, B. et al. Spike protein cleavage-activation mediated by the SARS-CoV-2 P681R mutation: a case-study from its first appearance in variant of interest (VOI) A.23.1 identified in Uganda. Preprint at https://doi.org/10.1101/2021.06.30.450632 (2021).
Wolter, N. et al. Early assessment of the clinical severity of the SARS-CoV-2 omicron variant in South Africa: a data linkage study. Lancet 399, 437–446 (2022).
SARS-CoV-2 Variants of Concern and Variants Under Investigation in England Technical briefing: Update on Hospitalisation and Vaccine Effectiveness for Omicron VOC-21NOV-01 (B.1.1.529) (UK Health Security Agency, 2021).
Matsuyama, S. et al. Enhanced isolation of SARS-CoV-2 by TMPRSS2-expressing cells. Proc. Natl Acad. Sci. USA 117, 7001–7003 (2020).
Hui, K. P. Y. et al. Tropism, replication competence, and innate immune responses of the coronavirus SARS-CoV-2 in human respiratory tract and conjunctiva: an analysis in ex-vivo and in-vitro cultures. Lancet Respi. Med. 8, 687–695 (2020).
Chan, M. C. et al. Tropism and innate host responses of the 2009 pandemic H1N1 influenza virus in ex vivo and in vitro cultures of human conjunctiva and respiratory tract. Am. J. Pathol. 176, 1828–1840 (2010).
Hui, K. P. et al. Tropism and innate host responses of influenza A/H5N6 virus: an analysis of ex vivo and in vitro cultures of the human respiratory tract. Eur. Respir. J. 49, 1601710 (2017).
Yen, H. L. et al. Cellular tropism of SARS-CoV-2 in the respiratory tract of Syrian hamsters and B6.Cg-Tg(K18-ACE2)2Prlmn/J transgenic mice. Vet. Pathol. https://doi.org/10.1177/03009858211043084 (2021).
Kärber, G. Beitrag zur kollektiven behandlung pharmakologischer reihenversuche. Naunyn Schmiedebergs Arch. Exp. Pathol. Pharmakol. 162, 480–483 (1931).
Crowe, A. R. & Yue, W. Semi-quantitative determination of protein expression using immunohistochemistry staining and analysis: an integrated protocol. Bio Protoc. 9, e3465 (2019).
Berglin, L. et al. In situ characterization of intrahepatic non-parenchymal cells in PSC reveals phenotypic patterns associated with disease severity. PLoS ONE 9, e105375 (2014).
Chan, R. W. et al. DAS181 inhibits H5N1 influenza virus infection of human lung tissues. Antimicrob. Agents Chemother. 53, 3935–3941 (2009).
Hui, K. P. et al. H5N1 influenza virus-induced mediators upregulate RIG-I in uninfected cells by paracrine effects contributing to amplified cytokine cascades. J. Infect. Dis. 204, 1866–1878 (2011).
Hui, K. P. et al. Induction of proinflammatory cytokines in primary human macrophages by influenza A virus (H5N1) is selectively regulated by IFN regulatory factor 3 and p38 MAPK. J. Immunol. 182, 1088–1098 (2009).
Bui, C. H. T. et al. Tropism of SARS-CoV-2, SARS-CoV, and influenza virus in canine tissue explants. J. Infect. Dis. 224, 821–830 (2021).
Acknowledgements
T. Shack, H.-W. Yeung, C. Lin and Kevin Fung provided technical support. We acknowledge research funding from the National Institute of Allergy and Infectious Diseases, National Institutes of Health, Department of Health and Human Services (contract no. 75N93021C00016) and the Theme-Based Research Scheme (T11-705/21N and T11-712/19-N) under University Grants Committee of Hong Kong Special Administrative Region. The funding providers had no role in study design, data collection, analysis or interpretation of the data, or in the writing of the report or in the decision to submit it for publication. The authors have not been paid to write this Article by a pharmaceutical company or other agency. The authors had full access to all the data in the study and had final responsibility for the decision to submit for publication. The corresponding author had full access to all of the data and the final responsibility to submit for publication.
Author information
Authors and Affiliations
Contributions
K.P.Y.H. was responsible for study design, coordination, analysis, interpretation of results, performed experiments and writing of the manuscript. J.C.W.H., M.-c.C., K.-c.N., R.H.H.C., K.-l.L. and T.T.K. performed experiments, analysis and interpretation of results. H.G. performed analysis and interpretation of sequencing results. K.-Y.S., M.K.Y.H. and T.W.K.A. provided human lung and bronchus tissue and read the manuscript. L.L.M.P. analysed and interpreted sequencing results and read the manuscript. M.P. analysed and interpreted results, and writing and revision of the manuscript. J.M.N. performed analysis, coordinated sample collection and interpreted results from immunohistochemical staining, and read the manuscript. M.C.W.C. was responsible for study design, overall coordination, interpretation of results and writing of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature thanks Peter Openshaw and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Viral replication kinetics of SARS-CoV-2 variants in ex vivo cultures of human respiratory tract from individual donors.
Human ex vivo cultures of bronchus and lung were infected with 5x105 TCID50/mL at 37 °C. Virus released in the culture supernatants were measured over time by TCID50 assay. Viral replication kinetics of wild-type (WT), Delta and Omicron in human ex vivo cultures of a. bronchus and b. lung from individual donors are shown. The horizontal dotted line denotes the limit of detection in the TCID50 assay. Bar-charts show the geometric mean.
Extended Data Fig. 2 Viral RNA of SARS-CoV-2 variants in ex vivo cultures of human respiratory tract.
Human ex vivo cultures of bronchus and lung were infected with 5x105 TCID50/mL at 37 °C. Viral RNA of ORF1b gene released in the culture supernatants were measured over time by real time PCR. Viral ORF1b gene copies of wild-type (WT), Delta and Omicron in human ex vivo cultures of bronchus and lung are shown. Bar-charts show the mean (n = 4) (+/−SD). Statistics were performed using Two-way ANOVA followed by a Tukey’s multiple-comparison test. p<0.05 was considered as statistically significant and exact p values are shown.
Extended Data Fig. 3 Quantification of viral antigen staining of SARS-CoV-2 variants in ex vivo cultures of human bronchus and lung.
Ex vivo cultures of human bronchus and human lung were infected with wild-type (WT) and various variants of SARS-CoV-2 with 5x105 TCID50/mL or mock at 37 °C and the tissues were fixed with formalin at 72 h.p.i. Paraffin-embedded sections were subjected to immunohistochemical staining with a polyclonal antibody against the SARS-CoV-2 nucleoprotein (SARS-2 NP). The expression of SARS-2 NP in a. bronchus for wild-type (WT) (n = 4), D614G (n = 3), Alpha (n = 5), Beta, (n = 2), Delta (n = 2), Omicron (n = 3) and mock (n = 2) and b. lung for wild-type (WT) (n = 5), D614G (n = 5), Alpha (n = 2), Beta, (n = 2), Delta (n = 3), Omicron (n = 3) and mock (n = 2) is presented. Bar-charts show the mean (+/−SD). Statistics were performed using One-way ANOVA followed by a Tukey’s multiple-comparison test and samples with sample number < 3 were excluded in the test. p<0.05 was considered as statistically significant and exact p values are shown.
Extended Data Fig. 4 Omicron viral particles in human bronchial mucosa.
Transmission electron micrograph of human bronchial mucosa infected with SARS-CoV-2 Omicron variant. Tissues were fixed after 72 h in formalin, followed by processing into Epon, ultrathin sectioning and staining. Ciliated cells are present and there are virus particles present in membrane bound vesicles in the cytoplasm indicated by red boxes, as well as on the surface attached to microvilli indicated by blue boxes. Micrograph is a representative photo of three individual donors. Scale bar: 1 μm.
Extended Data Fig. 5 Viral replication kinetics of SARS-CoV-2 variants in Vero E6-TMPRSS2 cells.
Vero E6-TMPRSS2 cells were infected with SARS-CoV-2 wild-type (WT), Delta and Omicron at MOI 0.01. Virus released in culture supernatants were measured over time by TCID50 assay. The horizontal dotted line denotes the limit of detection in the TCID50 assay. Bar-charts show the geometric mean of three independent experiments (n = 3) (+/−SD). Statistics were performed using Two-way ANOVA followed by a Tukey’s multiple-comparison test. p<0.05 was considered as statistically significant and exact p values are shown.
Extended Data Fig. 6 Viability of Vero E6-TMPRSS2 cells in the presence of camostat mesylate and E64d.
Vero E6-TMPRSS2 cells were treated with a. camostat mesylate (Cam) from 0.41 to 300 µM or b. E64d from 1.88 to 120 µM for 48 h. Cell viability was measured using Cell Counting Kit 8. Untreated cells were used as 100 % viability. Results are mean of three independent experiments (n = 3) (+/−SD). The half-maximal inhibitory concentration (IC50) was calculated by using Prism.
Supplementary information
Rights and permissions
About this article
Cite this article
Hui, K.P.Y., Ho, J.C.W., Cheung, Mc. et al. SARS-CoV-2 Omicron variant replication in human bronchus and lung ex vivo. Nature 603, 715–720 (2022). https://doi.org/10.1038/s41586-022-04479-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-022-04479-6
This article is cited by
-
SARS-CoV-2 infection increases airway bleeding risk in patients after tracheostomies
Virology Journal (2024)
-
Postoperative morbidity and mortality of patients with COVID-19 undergoing cardiovascular surgery: an inverse propensity-weighted study
BMC Anesthesiology (2024)
-
Latent and incubation periods of Delta, BA.1, and BA.2 variant cases and associated factors: a cross-sectional study in China
BMC Infectious Diseases (2024)
-
Inhalation of ACE2-expressing lung exosomes provides prophylactic protection against SARS-CoV-2
Nature Communications (2024)
-
Omicron Spike confers enhanced infectivity and interferon resistance to SARS-CoV-2 in human nasal tissue
Nature Communications (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.