Abstract
The brain is the seat of body weight homeostasis. However, our inability to control the increasing prevalence of obesity highlights a need to look beyond canonical feeding pathways to broaden our understanding of body weight control1,2,3. Here we used a reverse-translational approach to identify and anatomically, molecularly and functionally characterize a neural ensemble that promotes satiation. Unbiased, task-based functional magnetic resonance imaging revealed marked differences in cerebellar responses to food in people with a genetic disorder characterized by insatiable appetite. Transcriptomic analyses in mice revealed molecularly and topographically -distinct neurons in the anterior deep cerebellar nuclei (aDCN) that are activated by feeding or nutrient infusion in the gut. Selective activation of aDCN neurons substantially decreased food intake by reducing meal size without compensatory changes to metabolic rate. We found that aDCN activity terminates food intake by increasing striatal dopamine levels and attenuating the phasic dopamine response to subsequent food consumption. Our study defines a conserved satiation centre that may represent a novel therapeutic target for the management of excessive eating, and underscores the utility of a ‘bedside-to-bench’ approach for the identification of neural circuits that influence behaviour.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Materials availability
This study did not generate any new unique reagents. Mouse lines used in this study are on deposit at Jackson Laboratories and are listed under ‘Mice’.
Data and code availability
The sequencing datasets generated in this study are accessible at Gene Expression Omnibus under accession GSE184385. This manuscript contains all other datasets except the processed sequencing data, raw fibre photometry datasets and codes for analysis which have been uploaded to Mendeley Data (https://data.mendeley.com//datasets/j2mgy5486k/2). Source data are provided with this paper.
References
Berthoud, H. R. & Morrison, C. The brain, appetite, and obesity. Annu. Rev. Psychol. 59, 55–92 (2008).
Speakman, J. R. A nonadaptive scenario explaining the genetic predisposition to obesity: the “predation release” hypothesis. Cell Metab. 6, 5–12 (2007).
Rossi, M. A. & Stuber, G. D. Overlapping brain circuits for homeostatic and hedonic feeding. Cell Metab. 27, 42–56 (2018).
Woods, S. C. & Ramsay, D. S. Food intake, metabolism and homeostasis. Physiol. Behav. 104, 4–7 (2011).
Berthoud, H. R. The neurobiology of food intake in an obesogenic environment. Proc. Nutr. Soc. 71, 478–487 (2012).
Gautron, L., Elmquist, J. K. & Williams, K. W. Neural control of energy balance: translating circuits to therapies. Cell 161, 133–145 (2015).
Angulo, M. A., Butler, M. G. & Cataletto, M. E. Prader–Willi syndrome: a review of clinical, genetic, and endocrine findings. J. Endocrinol. Invest. 38, 1249–1263 (2015).
Holsen, L. M. et al. Importance of reward and prefrontal circuitry in hunger and satiety: Prader-Willi syndrome vs simple obesity. Int. J. Obes. 36, 638–647 (2012).
Barrachina, M. D., Martinez, V., Wang, L., Wei, J. Y. & Tache, Y. Synergistic interaction between leptin and cholecystokinin to reduce short-term food intake in lean mice. Proc. Natl Acad. Sci. USA 94, 10455–10460 (1997).
Kebschull, J. M. et al. Cerebellar nuclei evolved by repeatedly duplicating a conserved cell-type set. Science 370, eabd5059 (2020).
Betley, J. N., Cao, Z. F., Ritola, K. D. & Sternson, S. M. Parallel, redundant circuit organization for homeostatic control of feeding behavior. Cell 155, 1337–1350 (2013).
Carta, I., Chen, C. H., Schott, A. L., Dorizan, S. & Khodakhah, K. Cerebellar modulation of the reward circuitry and social behavior. Science 363, eaav0581 (2019).
Salamone, J. D., Correa, M., Mingote, S. & Weber, S. M. Nucleus accumbens dopamine and the regulation of effort in food-seeking behavior: implications for studies of natural motivation, psychiatry, and drug abuse. J. Pharmacol. Exp. Ther. 305, 1–8 (2003).
Palmiter, R. D. Is dopamine a physiologically relevant mediator of feeding behavior? Trends Neurosci. 30, 375–381 (2007).
Hinton, E. C. et al. Neural representations of hunger and satiety in Prader–Willi syndrome. Int. J. Obes. 30, 313–321 (2006).
Chen, Y., Lin, Y. C., Kuo, T. W. & Knight, Z. A. Sensory detection of food rapidly modulates arcuate feeding circuits. Cell 160, 829–841 (2015).
Betley, J. N. et al. Neurons for hunger and thirst transmit a negative-valence teaching signal. Nature 521, 180–185 (2015).
Alhadeff, A. L. et al. Natural and drug rewards engage distinct pathways that converge on coordinated hypothalamic and reward circuits. Neuron 103, 891–908.e896 (2019).
Somana, R. & Walberg, F. Cerebellar afferents from the nucleus of the solitary tract. Neurosci. Lett. 11, 41–47 (1979).
Zhu, J. N. & Wang, J. J. The cerebellum in feeding control: possible function and mechanism. Cell Mol. Neurobiol. 28, 469–478 (2008).
Sodersten, P., Bergh, C., Leon, M. & Zandian, M. Dopamine and anorexia nervosa. Neurosci. Biobehav. Rev. 60, 26–30 (2016).
Daberkow, D. P. et al. Amphetamine paradoxically augments exocytotic dopamine release and phasic dopamine signals. J. Neurosci. 33, 452–463 (2013).
Alhadeff, A. L., Rupprecht, L. E. & Hayes, M. R. GLP-1 neurons in the nucleus of the solitary tract project directly to the ventral tegmental area and nucleus accumbens to control for food intake. Endocrinology 153, 647–658 (2012).
Keiflin, R., Pribut, H. J., Shah, N. B. & Janak, P. H. Ventral tegmental dopamine neurons participate in reward identity predictions. Curr. Biol. 29, 93–103.e103 (2019).
Butler, M. G., Bittel, D. C., Kibiryeva, N., Talebizadeh, Z. & Thompson, T. Behavioral differences among subjects with Prader–Willi syndrome and type I or type II deletion and maternal disomy. Pediatrics 113, 565–573 (2004).
Holsen, L. M. et al. Neural mechanisms underlying food motivation in children and adolescents. Neuroimage 27, 669–676 (2005).
Holsen, L. M. et al. Neural mechanisms underlying hyperphagia in Prader–Willi syndrome. Obesity 14, 1028–1037 (2006).
Holsen, L. M. et al. Genetic subtype differences in neural circuitry of food motivation in Prader-Willi syndrome. Int. J. Obes. 33, 273–283 (2009).
LaBar, K. S. et al. Hunger selectively modulates corticolimbic activation to food stimuli in humans. Behav. Neurosci. 115, 493–500 (2001).
Gelman, N. et al. MR imaging of human brain at 3.0 T: preliminary report on transverse relaxation rates and relation to estimated iron content. Radiology 210, 759–767 (1999).
Whitfield-Gabrieli, S. Region of interest extraction (REX) toolbox (Boston, MA). p497 (2009).
Holmes, C. J. et al. Enhancement of MR images using registration for signal averaging. J. Comp. Assist. Tomography 22, 324–333 (1998).
Vong, L. et al. Leptin action on GABAergic neurons prevents obesity and reduces inhibitory tone to POMC neurons. Neuron 71, 142–154 (2011).
Backman, C. M. et al. Characterization of a mouse strain expressing Cre recombinase from the 3' untranslated region of the dopamine transporter locus. Genesis 44, 383–390 (2006).
Allen, W. E. et al. Thirst-associated preoptic neurons encode an aversive motivational drive. Science 357, 1149–1155 (2017).
Madisen, L. et al. A toolbox of Cre-dependent optogenetic transgenic mice for light-induced activation and silencing. Nat. Neurosci. 15, 793–802 (2012).
Borgius, L., Restrepo, C. E., Leao, R. N., Saleh, N. & Kiehn, O. A transgenic mouse line for molecular genetic analysis of excitatory glutamatergic neurons. Mol. Cell Neurosci. 45, 245–257 (2010).
Tong, Q., Ye, C. P., Jones, J. E., Elmquist, J. K. & Lowell, B. B. Synaptic release of GABA by AgRP neurons is required for normal regulation of energy balance. Nat. Neurosci. 11, 998–1000 (2008).
Su, Z., Alhadeff, A. L. & Betley, J. N. Nutritive, post-ingestive signals are the primary regulators of AgRP neuron activity. Cell Rep. 21, 2724–2736 (2017).
Alhadeff, A. L., Park, O., Hernandez, E. & Betley, J. N. Inhibition of itch by hunger and AgRP neuron activity. Neuroscience 450, 126–134 (2020).
Sun, F. et al. A genetically encoded fluorescent sensor enables rapid and specific detection of dopamine in flies, fish, and mice. Cell 174, 481–496.e419 (2018).
Goebel, M., Stengel, A., Wang, L. & Tache, Y. Central nesfatin-1 reduces the nocturnal food intake in mice by reducing meal size and increasing inter-meal intervals. Peptides 32, 36–43 (2011).
Lerner, T. N. et al. Intact-brain analyses reveal distinct information carried by SNc dopamine subcircuits. Cell 162, 635–647 (2015).
Zalocusky, K. A. et al. Nucleus accumbens D2R cells signal prior outcomes and control risky decision-making. Nature 531, 642–646 (2016).
Weir, J. B. New methods for calculating metabolic rate with special reference to protein metabolism. 1949. Nutrition 6, 213–221 (1990).
Betley, J. N. et al. Stringent specificity in the construction of a GABAergic presynaptic inhibitory circuit. Cell 139, 161–174 (2009).
Young, M. D. & Behjati, S. SoupX removes ambient RNA contamination from droplet-based single-cell RNA sequencing data. Gigascience 9, giaa151 (2020).
Stuart, T. et al. Comprehensive integration of single-cell data. Cell 177, 1888–1902.e1821 (2019).
Wolf, F. A., Angerer, P. & Theis, F. J. SCANPY: large-scale single-cell gene expression data analysis. Genome Biol. 19, 15 (2018).
Levine, J. H. et al. Data-driven phenotypic dissection of AML reveals progenitor-like cells that correlate with prognosis. Cell 162, 184–197 (2015).
McGinnis, C. S., Murrow, L. M. & Gartner, Z. J. DoubletFinder: doublet detection in single-cell RNA sequencing data using artificial nearest neighbors. Cell Syst. 8, 329–337.e324 (2019).
Hafemeister, C. & Satija, R. Normalization and variance stabilization of single-cell RNA-seq data using regularized negative binomial regression. Genome Biol. 20, 296 (2019).
Becht, E. et al. Dimensionality reduction for visualizing single-cell data using UMAP. Nat Biotechnol 37, 38–44 (2019).
Marques, S. et al. Oligodendrocyte heterogeneity in the mouse juvenile and adult central nervous system. Science 352, 1326–1329 (2016).
Saunders, A. et al. Molecular diversity and specializations among the cells of the adult mouse brain. Cell 174, 1015–1030.e1016 (2018).
Vanlandewijck, M. et al. A molecular atlas of cell types and zonation in the brain vasculature. Nature 554, 475–480 (2018).
Zeisel, A. et al. Molecular architecture of the mouse nervous system. Cell 174, 999–1014.e1022 (2018).
Carter, R. A. et al. A single-cell transcriptional atlas of the developing murine cerebellum. Curr. Biol. 28, 2910–2920.e2912 (2018).
Wizeman, J. W., Guo, Q., Wilion, E. M. & Li, J. Y. Specification of diverse cell types during early neurogenesis of the mouse cerebellum. eLife 8, e42388 (2019).
Wang, F. et al. RNAscope: a novel in situ RNA analysis platform for formalin-fixed, paraffin-embedded tissues. 14, 22–29 (2012).
Goldstein, N. et al. Hypothalamic detection of macronutrients via multiple gut–brain pathways. Cell Metab. 33, 676–687.e5 (2021).
Andermann, M. L. & Lowell, B. B. Toward a wiring diagram understanding of appetite control. Neuron 95, 757–778 (2017).
Brakeman, P. R. et al. Homer: a protein that selectively binds metabotropic glutamate receptors. Nature 386, 284–288 (1997).
Berridge, K. C. 'Liking' and 'wanting' food rewards: brain substrates and roles in eating disorders. Physiol. Behav. 97, 537–550 (2009).
Wise, R. A. Role of brain dopamine in food reward and reinforcement. Philos. Trans. R. Soc. B 361, 1149–1158 (2006).
Becker, M. I. & Person, A. L. Cerebellar control of reach kinematics for endpoint precision. Neuron 103, 335–348.e335 (2019).
Dacre, J. et al. A cerebellar-thalamocortical pathway drives behavioral context-dependent movement initiation. Neuron 109, 2326–2338.e2328 (2021).
Darmohray, D. M., Jacobs, J. R., Marques, H. G. & Carey, M. R. Spatial and temporal locomotor learning in mouse cerebellum. Neuron 102, 217–231.e214 (2019).
Frontera, J. L. et al. Bidirectional control of fear memories by cerebellar neurons projecting to the ventrolateral periaqueductal grey. Nat. Commun. 11, 5207 (2020).
Gao, Z. et al. A cortico-cerebellar loop for motor planning. Nature 563, 113–116 (2018).
Kelly, E. et al. Regulation of autism-relevant behaviors by cerebellar-prefrontal cortical circuits. Nat. Neurosci. 23, 1102–1110 (2020).
Xiao, L., Bornmann, C., Hatstatt-Burkle, L. & Scheiffele, P. Regulation of striatal cells and goal-directed behavior by cerebellar outputs. Nat. Commun. 9, 3133 (2018).
Giovannucci, A. et al. Cerebellar granule cells acquire a widespread predictive feedback signal during motor learning. Nat. Neurosci. 20, 727–734 (2017).
Locke, T. M. et al. Dopamine D1 receptor-positive neurons in the lateral nucleus of the cerebellum contribute to cognitive behavior. Biol. Psychiatry 84, 401–412 (2018).
Fujita, H., Kodama, T. & du Lac, S. Modular output circuits of the fastigial nucleus mediate diverse motor and nonmotor functions of the cerebellar vermis. eLife 9, e58613 (2020).
DiFeliceantonio, A. G. et al. Supra-additive effects of combining fat and carbohydrate on food reward. Cell Metab. 28, 33–44.e33 (2018).
Kralj-Hans, I., Baizer, J. S., Swales, C. & Glickstein, M. Independent roles for the dorsal paraflocculus and vermal lobule VII of the cerebellum in visuomotor coordination. Exp. Brain Res. 177, 209–222 (2007).
Sobel, N. et al. Odorant-induced and sniff-induced activation in the cerebellum of the human. J. Neurosci. 18, 8990–9001 (1998).
Wagner, M. J., Kim, T. H., Savall, J., Schnitzer, M. J. & Luo, L. Cerebellar granule cells encode the expectation of reward. Nature 544, 96–100 (2017).
Nieoullon, A., Cheramy, A. & Glowinski, J. Release of dopamine in both caudate nuclei and both substantia nigrae in response to unilateral stimulation of cerebellar nuclei in the cat. Brain Res. 148, 143–152 (1978).
Miller, J. L. et al. Enhanced activation of reward mediating prefrontal regions in response to food stimuli in Prader–Willi syndrome. J. Neurol. Neurosurg. Psychiatry 78, 615–619 (2007).
Shapira, N. A. et al. Satiety dysfunction in Prader–Willi syndrome demonstrated by fMRI. J. Neurol. Neurosurg. Psychiatry 76, 260–262 (2005).
Brady, R. O., Jr et al. Cerebellar-prefrontal network connectivity and negative symptoms in schizophrenia. Am. J. Psychiatry 176, 512–520 (2019).
Halko, M. A., Farzan, F., Eldaief, M. C., Schmahmann, J. D. & Pascual-Leone, A. Intermittent theta-burst stimulation of the lateral cerebellum increases functional connectivity of the default network. J. Neurosci. 34, 12049–12056 (2014).
Miterko, L. N. et al. Neuromodulation of the cerebellum rescues movement in a mouse model of ataxia. Nat. Commun. 12, 1295 (2021).
Stoodley, C. J. et al. Altered cerebellar connectivity in autism and cerebellar-mediated rescue of autism-related behaviors in mice. Nat. Neurosci. 20, 1744–1751 (2017).
Acknowledgements
We thank H. Fujita, K. Gallagher and A. Thanawalla for comments on the manuscript. N.G. is supported by the National Science Foundation Graduate Research Fellowship Program (DGE-1845298). E.A. is supported by the National Institutes of Health (NIH) (DP2NS105555, R01NS111479 and U19NS112959), the Searle Scholars Program, The Pew Charitable Trusts, and the McKnight Foundation. O.M.S. is supported by the National University of Singapore. M.A.H. is supported by the NIH (R01MH111868; R56MH125995). R.O.B. is supported by the NIH (R01MH116170). L.M.H. is supported by the NIH (R01DK104772). A.L.A. is supported by the Monell Chemical Senses Center, NIH (R00DK119574), the Klingenstein-Simons Fellowship Award, the American Heart Association (857082) and the Penn Institute for Diabetes, Obesity, and Metabolism. J.N.B. is supported by the University of Pennsylvania School of Arts and Sciences, the American Diabetes Association (118IBS116), the American Heart Association (AHA 17SDG33400158), the Whitehall Foundation, the Klingenstein-Simons Fellowship Award and the NIH (1R01DK114104). A.I.C. and A.Y.T.L. were supported by the Warwick-NTU Neuroscience Programme. A.I.C., O.M.S. and T.H.C. are supported by the Singapore Ministry of Education (MOE2018-T2-1-065). A.I.C. and T.H.C. are supported by the Singapore Ministry of Education (MOE2017-T3-1-002). J.N.B. and A.I.C. are supported by the NIH (1R01DK124801).
Author information
Authors and Affiliations
Contributions
A.Y.T.L., A.I.C. and J.N.B. initiated the project and prepared the manuscript with comments from all authors. A.Y.T.L., N.G., J.R.G., K.-P.H., N.Z., A.K.K.Y., J.R.E.C., J.Y.C., A.M.M., H.S.T.H., C.L., L.M.H. and A.L.A. performed experiments. A.Y.T.L., N.G., J.R.G., E.A., O.M.S., T.H.C., A.S.B., L.E.M., M.A.H., R.O.B., L.M.H., A.L.A., A.I.C. and J.N.B. designed experiments and analysed data.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature thanks Dana Small, Larry Zweifel and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
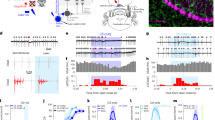
Extended Data Fig. 1 fMRI paradigm for response to food cue.
Subjects with Prader-Willi syndrome (PWS) and controls underwent two separate scanning sessions (top left, group): either during fasting or post-meal (bottom left, session). During each scanning session, participants were presented with visual cues that alternate between food (muffin) and non-food (dog) categories (right, stimulus)8.
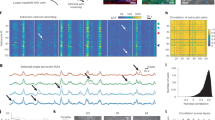
Extended Data Fig. 2 Neural activation pattern following food infusion and refeeding in mice.
(a-c) Experimental design for Targeted Recombination of Activated Populations (TRAP) labelling of neurons activated by water infusion (IG water), Ensure infusion (IG Ensure) or refeeding (Refed) in Fos2A::iCreER; Ai9 mice. (d-i) tdTomato expression in the DCN after water infusion (d; e, magnified of box in d), 1 kcal Ensure infusion (f; g, magnified of box in f), and chow refeeding (h; i, magnified of box in h). Scale bar, 500 µm (d, f, h), 100 µm (e, g, i). (j) Heatmap depicting the activated cells recombined in DCN subregions following infusion and refeeding (n = 9). (k-m) tdTomato expression in the nucleus tractus solitarius (NTS) 3 weeks after water infusion (k), 1 kcal Ensure infusion (l), and chow refeeding (m). Scale bar, 500 µm. (n-p) tdTomato expression in the paraventricular hypothalamic nucleus (PVH) 3 weeks after water infusion (n), 1 kcal Ensure infusion (o), and chow refeeding (p). Scale bar, 250 µm. (q-s) tdTomato expression in the arcuate hypothalamic nuclei (ARC) 3 weeks after water infusion (q), 1 kcal Ensure infusion (r), and chow refeeding (s). Scale bar, 250 µm. (t-v) tdTomato expression in the lateral parabrachial nucleus (LPBN) 3 weeks after water infusion (t), 1 kcal Ensure infusion (u), and chow refeeding (v). Scale bar, 250 µm. (w-y) tdTomato expression in the central amygdaloid nucleus (CEA) 3 weeks after water infusion (w), 1 kcal Ensure infusion (x), and chow refeeding (y). Scale bar, 500 µm. (z-bb) tdTomato expression in the bed nucleus of the stria terminalis (BNST) 3 weeks after water infusion (z), 1 kcal Ensure infusion (aa), and chow refeeding (bb). Scale bar, 250 μm. (cc) Heatmap depicting relative density of cells recombined following water and calorie intake (top), and 1 kcal Ensure infusion and refeeding (bottom, n = 9). (dd) Schematic depicting the DCN and key feeding brain regions that sense food cues and nutrients62. Statistical analysis in Supplementary Table 1
Extended Data Fig. 3 Mapping the DCN subregions that suppress food intake.
(a) Schematic of the deep cerebellar nuclei. The lateral subnuclei of the anterior deep cerebellar nuclei (aDCN) are depicted in maroon (aDCN-LAT, consisting of Lat and LatPC, bregma -5.68 to -5.88 mm), interposed subnuclei of the aDCN are depicted in pink (aDCN-INT, consisting of IntA, IntDL, IntP, and IntPPC, bregma -6.00 to -6.35 mm). The posterior DCN is in grey (pDCN, consisting of IntP, IntPPC, Med, MedDL, and MedL, bregma -6.36 to -6.64 mm, see also e and g). (b) Distribution of cells expressing hM3D(Gq) across 9 DCN subnuclei in mice with hM3D(Gq) targeted to the lateral nucleus (aDCN-LAT) and mice with hM3D(Gq) targeted to the interposed nucleus (aDCN-INT). Mice with targeting to the LAT show a reduction in food intake following DREADD activation (aDCN-LAT in maroon, n = 5 mice, and aDCN-INT in pink, n=3 mice). (c) Schematised serial coronal sections depicting regions where hM3D expression results in food intake reduction (magenta). (d) Representative images of the entire DCN in a aDCN-LAT hM3D(Gq) mouse with hM3D(Gq) expression in the lateral nucleus. Scale bar, 2000 µm. (e) Expression of mCherry (as control viral vector) in the aDCN (red: mCherry). Scale bar, 500 µm. (f) Chow intake in mice with mCherry expression in the aDCN following vehicle or CNO treatment (n = 8, paired t-test, P = 0.539). (g) hM3D(Gq) expression in the pDCN (red: hM3D(Gq)). Scale bar, 500 µm. (h) Chow intake in mice with hM3D(Gq) expression in the pDCN following vehicle or CNO treatment (n = 12, paired t-test, P = 0.548). (i) Chow intake in mice with mCherry expression in the pDCN following vehicle or CNO treatment (n=8, paired t-test, P = 0.722). Data are expressed as mean ± SEM. Lat, lateral; LatPC, lateral parvicellular; IntDL, interposed dorsolateral; IntA, interposed anterior nucleus; IntP, interposed posterior; IntPPC, interposed posterior parvicellular; MedDL, medial dorsolateral; Med, medial; MedL, medial lateral. Statistical analysis in Supplementary Table 1
Extended Data Fig. 4 Neural activity in the aDCN suppresses food intake independent of hunger state with no compensatory metabolic changes.
(a) Experimental design: meal pattern measurements of 24-h food-deprived mice following vehicle or CNO i.p. administration. (b) Latency to first bite in food-deprived mice with hM3D(Gq) expression in the aDCN-LAT (n = 9), aDCN-INT (n = 16) or mCherry control in the aDCN (n = 8) following vehicle or CNO treatment (two-way ANOVA interaction P < 0.001, main effect P = 0.005; Holm-Sidak’s, P < 0.001). (c) Average meal duration during a 1-h chow intake assay following 24-h food deprivation in mice with hM3D(Gq) expression in the aDCN-LAT (n = 9), aDCN-INT (n = 16 mice), or control mCherry expression in the aDCN (n = 8) following vehicle or CNO treatment (two-way ANOVA interaction P = 0.005, main effect P < 0.001; Holm-Sidak’s, P < 0.001). (d) Rate of food intake during a 1-h chow intake assay following 24-h food deprivation in mice with hM3D expression in the aDCN-LAT (n = 9), aDCN-INT (n = 16), or control mCherry expression in the aDCN (n = 8) following vehicle or CNO treatment (two-way ANOVA interaction P = 0.748). (e) Experimental design: meal pattern measurements of ad libitum fed mice following vehicle or CNO i.p. administration. (f) Chow intake in ad libitum fed mice with hM3D(Gq) expression following vehicle or CNO treatment (aDCN-LAT: n = 9, aDCN-INT: n = 16, mCherry control: n = 8; two-way ANOVA, interaction P = 0.001, main effect P = 0.006; Holm-Sidak’s, P < 0.001). (g) Latency to first bite in ad libitum fed mice with hM3D(Gq) expression following vehicle or CNO treatment (aDCN-LAT: n = 9, aDCN-INT: n = 16, mCherry control: n = 8; two-way ANOVA interaction P < 0.001, main effect P < 0.001; Holm-Sidak’s, P<0.001). (h) Chow intake in ad libitum fed mice with mCherry control or hM3D(Gq) expression in the pDCN following vehicle or CNO treatment (pDCN mCherry: n = 8, pDCN hM3D(Gq): n = 12; two-way ANOVA interaction P=0.358). (i) Schematic of the metabolic monitoring experiment. (j) Energy expenditure (kcal) over a 48-h period in mice with mCherry control (n = 8) or hM3D(Gq) (n = 7) expression in the aDCN-LAT (unpaired t-test, P = 0.004). (k) Energy intake (EI) and energy expenditure (EE) over 48-h period in mice with mCherry control or hM3D(Gq) expression in the aDCN-LAT following CNO treatment normalized to vehicle treatment (n = 7 control, 8 aDCN-LAT-hM3D(Gq), repeated measures two-way ANOVA interaction P < 0.001, main effect P<0.001; Holm-Sidak’s, P < 0.001, P = 0.009 (EE); P<0.001, P < 0.001, P < 0.001, P = 0.006 (EI)). Data are expressed as mean ± SEM, two-sided P values, t-tests and post-hoc comparisons: **P<0.01, ***P < 0.001, ANOVA interaction: ∞∞∞P < 0.01, ∞∞∞P<0.001; ANOVA main effect of group: ¤¤P < 0.01, ¤¤¤P<0.001. Statistical analysis in Supplementary Table 1
Extended Data Fig. 5 aDCN activity suppresses food intake regardless of hedonic value of food.
(a) Experimental timeline: 24-h food deprivation followed by measurements of chow intake over 12-h (top). Cumulative kcal of chow intake in food-deprived mice with hM3D(Gq) expression in the aDCN-LAT following vehicle or CNO treatment (bottom; n = 9 hM3D(Gq); repeated measures two-way ANOVA interaction P < 0.001, main effect P < 0.001; Holm-Sidak’s, P = 0.094 (30 min), P=0.008 (1 h), P < 0.001 (2 h), P < 0.001 (4h), P < 0.001 (6h), P < 0.001 (8 h), P<0.001 (10 h), P < 0.001 (12 h)). (b) 12-h food intake in food-deprived mice expressing hM3D(Gq) in the aDCN-LAT (n = 9 mice, paired t-test, P = 0.008). (c) 12-h food intake in food-deprived mice expressing mCherry in the aDCN-LAT (n = 8 mice, paired t-test, P = 0.391). (d) Food intake during the first 2-h of refeeding in food-deprived mice expressing hM3D(Gq) in the aDCN-LAT (n = 9 mice, paired t-test, P < 0.001). (e) Food intake during the first 2-h of refeeding in food-deprived mice expressing mCherry in the aDCN-LAT (n = 8 mice, paired t-test, P = 0.223). (f) Experimental timeline: 24-h food deprivation followed by measurement of high fat high sugar (HFHS) diet intake over 12 h (top). Cumulative kcal of HFHS diet intake in food-deprived mice with hM3D(Gq) expression in the aDCN-LAT following vehicle or CNO treatment (bottom; n = 9 hM3D(Gq) mice; two-way repeated measures ANOVA interaction P < 0.001, main effect P<0.001; Holm-Sidak’s, P < 0.001 (30-min), P < 0.001 (1-h), P<0.001 (2-h), P < 0.001 (4-h), P<0.001 (6-h), P < 0.001 (8-h), P<0.001 (10-h), P < 0.001 (12-h)). (g) 12-h HFHS diet intake in food-deprived mice expressing hM3D(Gq) in the aDCN-LAT (n = 9 mice, paired t-test, P < 0.001). (h) 12-h HFHS diet intake in food-deprived mice expressing mCherry in the aDCN-LAT (n = 8 mice, paired t-test, P = 0.527). (i) Calorie intake during the first 2-h of HFHS diet refeeding in food-deprived mice expressing hM3D(Gq) in the aDCN-LAT (n = 9 mice, paired t-test, P < 0.001). (j) Calorie intake during the first 2-h of HFHS diet refeeding in food-deprived mice expressing mCherry in the aDCN-LAT (n=9 mice, paired t-test, P = 0.686). Data are expressed as mean ± SEM, two-sided P values, t-tests and post-hoc comparisons: **P<0.01, ***P < 0.001, ANOVA interaction: ∞∞∞P<0.001; ANOVA main effect of group: ¤¤¤P < 0.001. Statistical analysis in Supplementary Table 1
Extended Data Fig. 6 Glutamatergic neurons in the DCN are activated by food intake.
(a-h) Fluorescent In Situ Hybridization (FISH) histochemistry in the lateral nucleus of the DCN in food-deprived (a, b, e, f) and chow-refed mice (c, d, g, h) (red, Homer1a; green, vGluT2; blue, vGAT in b and d, GlyT2 in f and h). Scale bars, 20 µm. (i) Homer1a expression in excitatory (vGluT2+) and inhibitory (vGAT+ or GlyT2+) DCN neurons following food deprivation or refeeding (n = 3 mice per group, two-way ANOVA main effect P < 0.001; Holm-Sidak’s, P = 0.034). (j) Number of excitatory (vGluT2+) and inhibitory (vGAT+ or GlyT2+) DCN neurons that express Homer1a following food deprivation or refeeding (n = 3 mice per group, two-way ANOVA main effect P = 0.009; Holm-Sidak’s, P = 0.023). (k, l) Expression level (k) and number (l) of Homer1a+ vGluT2+ neurons within the 3 major cerebellar nuclei following food deprivation or refeeding (n = 3 mice per group, two-way ANOVA, expression level main effect P = 0.013, number main effect P = 0.010; Holm-Sidak’s, expression level, P = 0.006, number, P = 0.025). Data are expressed as mean ± SEM, two-sided P values, t-tests and post-hoc comparisons: *P < 0.05, **P<0.01, ANOVA interaction: ∞∞∞P < 0.001; ANOVA main effect of group: ¤P < 0.05, ¤¤¤P < 0.001. Statistical analysis in Supplementary Table 1
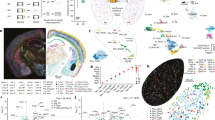
Extended Data Fig. 7 Gene expression gradient along the anterior-posterior axis of the DCN.
(a) Experimental design of single nucleus RNA sequencing of DCN neurons. (b) Uniform Manifold Approximation and Projection for Dimension Reduction (UMAP) plot of cerebellar cell types derived from microdissection of the DCN and surrounding tissues. (c) Principal component (PC) 1 loadings of select class I and class II defining genes expressed by vGluT2+ DCN neurons. (d) Celf4 (red) and Spp1 (blue) expression level in vGluT2+ neurons in the DCN, 492 neurons (two-way ANOVA interaction P < 0.001; Holm-Sidak’s, P<0.001, P < 0.001, P < 0.001, P < 0.001). (e, f) vGluT2 (red, e, f), Spp1 (green, e) and Celf4 (green, f) expression in the aDCN. Scale bar, 25 μm. (g) PC embedding of Miat expression, fluorescent in situ hybridization (FISH) and quantification of Miat levels in vGluT2+ neurons in the three major cerebellar nuclei (n = 1,434 neurons, one-way ANOVA P < 0.001; Holm-Sidak’s, P = 0.165, P < 0.001, P < 0.001). (h) PC embedding of Crhr1 expression, FISH, and quantification of Crhr1 levels in Miat+ neurons in the three major cerebellar nuclei (n = 1,434 neurons, one-way ANOVA P < 0.001; Holm-Sidak’s, P = 0.003, P=0.006, P < 0.001). (i) PC embedding of Dpp10 expression, FISH, and quantification of Dpp10 levels in Celf4+ neurons in the three major cerebellar nuclei (n = 2,261 neurons, one-way ANOVA P<0.001; Holm-Sidak’s, P < 0.001, P<0.001, P < 0.001). (j) PC embedding of Unc5d expression, FISH, and quantification of Unc5d levels in Celf4+ neurons in the three major cerebellar nuclei (n = 2,261 neurons, one-way ANOVA P < 0.001; Holm-Sidak’s, P<0.001, P < 0.001, P < 0.001). Scale bar, 100 µm. (k) FISH of Spp1 and Celf4 expression in vGluT2+ neurons of the interposed nucleus (left image: red, vGluT2; green, Spp1; right image: red, vGluT2; green, Celf4; n = 3, unpaired t-test, P < 0.001). Scale bar,100 µm. (l) FISH of Spp1 and Celf4 expression in vGluT2+ neurons in the medial nucleus (left image: red, vGluT2; green, Spp1; right image: red, vGluT2; green, Celf4; n = 3, unpaired t-test, P < 0.001). Scale bar, 100 µm. (m) Spp1 expression levels in vGluT2+ neurons across the three major cerebellar nuclei (n = 3, one-way ANOVA P < 0.001; Holm-Sidak’s, P = 0.946, P < 0.001, P < 0.001). (n) Celf4 expression levels in vGluT2+ neurons across the three major cerebellar nuclei (n=3, one-way ANOVA P < 0.001; Holm-Sidak’s, P = 0.026, P<0.001, P < 0.001). (o-r) Quantification of Spp1+ (o), Celf4+ (p), Spp1+Celf4+ (q) and Spp1–Celf4– (r) vGluT2+ neurons across the three major cerebellar nuclei (n = 3, one-way ANOVA (o) P = 0.002, (p) P<0.001; Holm-Sidak’s, (o) P = 0.190, P = 0.005, P = 0.002, (p) P = 0.982, P < 0.001, P<0.001, lateral versus interposed, lateral versus medial, and interposed versus medial, respectively). Data are expressed as mean ± SEM, two-sided P values, post-hoc comparisons: *P < 0.05, **P<0.01, ***P < 0.001; ANOVA interaction: ∞∞∞P < 0.001. Statistical analysis in Supplementary Table 1
Extended Data Fig. 8 Molecular and topographical distinctions of DCN neurons that respond to food intake.
(a-f) Expression of activity-regulated transcript Homer1a63 (red) in the three major cerebellar nuclei following food deprivation (a, c, e) or refeeding (b, d, f). (g-l) Spp1 expression (green) in vGluT2+ neurons (blue) (g, i, k), and colocalised Spp1 (cyan, Spp1+vGluT2+ neurons) (h, j, l) in the three major cerebellar nuclei. (m-r) Celf4 expression (blue) in vGluT2+ neurons (red) (m, o, q), and colocalised Celf4 (magenta, Celf4+vGluT2+ neurons) (n, p, r) in the three major cerebellar nuclei. Scale bar, 100 µm. (s) Summary of the expression of Spp1 and Celf4 and the distribution of Homer1a+ DCN neurons. (t) Schematic of fibre photometry system. (u-w) Fibre targeting aDCN-LAT glutamatergic neurons in vGluT2::Cre mouse (u), expression of GCaMP6s (green) and vGluT2 (red) (v, w). Scale bar, 20 µm. (x-z) Heatmaps depicting ∆F/F of GCaMP6 signals in the aDCN-LAT glutamatergic neurons of ad libitum fed (x) and food-deprived (y) mice response to chow, and ad libitum fed mice response to non-food object (z, marble). Signals are aligned to the introduction of chow or non-food object (red line) (n = 7 mice). (aa) Average ∆F/F of GCaMP6 signals in the aDCN-LAT glutamatergic neurons (490 nm, green, and control 405 nm, magenta). Signals are aligned to the introduction of non-food object (red line). Dark line represents the mean and lighter shaded area represents SEMs (n = 7). (bb-cc) Mean (bb) and max (cc) ∆F/F GCaMP6s signals of aDCN-LAT glutamatergic neurons in response to chow, in ad libitum fed (grey) and food-deprived (red) mice, and response to non-food object in ad libitum fed mice (n=7, one-way ANOVA (bb) P < 0.001, (cc) P < 0.001; Holm-Sidak’s, (bb) P < 0.006, P = 0.475, P < 0.003, (cc) P<0.009, P=0.651, P = 0.011, ad libitum fed chow versus food deprivation chow, ad libitum fed chow versus non-food, food deprivation chow versus non-food, respectively). Data are expressed as mean ± SEM, two-sided P values, post-hoc comparisons: *P < 0.05, **P < 0.01. Statistical analysis in Supplementary Table 1
Extended Data Fig. 9 Activation of arcuate AgRP neurons does not fully restore food intake suppression mediated by aDCN-LAT activation.
(a) Schematic depicting hM3D(Gq) expression in the aDCN-LAT, ChR2 expression and fibre implant in the arcuate nucleus (ARC) of a AgRP::Cre; Ai32 mouse for either individual or simultaneous activation.(b, c) ChR2-eYFP expression in AgRP ARC neurons (b) and hM3d(Gq) expression in the aDCN (c). Scale bar, 500 µm in b, 1000 µm in c. (d) Chow intake following AgRP neuron activation in ad libitum state (blue), aDCN neuron activation in food-deprived state (red), or AgRP and aDCN neuron activation in food-deprived state (pink) (n = 11, repeated measures one-way ANOVA, P = 0.001; Holm-Sidak’s, P < 0.001, P < 0.001, P = 0.024). Data are expressed as mean ± SEM, two-sided P values, post-hoc comparisons: *P < 0.05, **P < 0.01, ***P < 0.001. Statistical analysis in Supplementary Table 1
Extended Data Fig. 10 Activation of aDCN neurons robustly increases striatal dopamine signalling that correlates with reduced food intake.
(a) Schematic depicting hM3D(Gq) expression in the DCN combined with GRABDA expression41 and fibre implant in the ventral striatum which receives projections from the ventral tegmental area (VTA) dopamine (DA) neurons64,65. (b) GRABDA expression and fibre placement in the ventral striatum. Scale bar, 1000 µm. (c, d) Average ∆F/F of GRABDA signals in the ventral striatum of food-deprived mCherry control (c) or aDCN-LAT hM3D(Gq) (d) mice treated with vehicle or CNO. Signals are aligned to the vehicle or CNO injection (red line). Dark line represents the mean and lighter shaded area represents SEMs. Corresponding heatmaps (right) depict ∆F/F of GRABDA signals in each mouse (n = 6 control mice, grey; n = 6 hM3D(Gq) aDCN-LAT mice, green). (e) Average ∆F/F of GRABDA signals in 3-min bins (n = 6 control mice, grey; n = 6 hM3D(Gq) aDCN-LAT mice, repeated measures two-way ANOVA interaction P < 0.001, main effect P < 0.001; Holm-Sidak’s, P = 0.027/ = 0.397/=0.625 (12-15 min), P < 0.001/<0.001/ < 0.001 (15-18 min), P < 0.001/<0.001/ < 0.001 (18-21 min), P<0.001/ < 0.001/ < 0.001 (21-24 min), P < 0.001/ < 0.001/ < 0.001 (24-27 min), P < 0.001/ < 0.001/ < 0.001 (27-30 min), hM3D(Gq) CNO to hM3D(Gq) vehicle/control CNO/control vehicle respectively). (f) Maximum ∆F/F of GRABDA signals in the ventral striatum of vehicle or CNO treated food-deprived mice with aDCN-LAT hM3D(Gq) (n = 6, paired t-test, P = 0.011). (g) Mean ∆F/F of GRABDA signals in the ventral striatum (n = 6, paired t-test, P = 0.002). (h) Maximum ∆F/F of GRABDA signals in the ventral striatum of vehicle or CNO treated food-deprived mice with aDCN mCherry control mice (n = 6, paired t-test, P = 0.242). (i) Mean ∆F/F of GRABDA signals in the ventral striatum of vehicle or CNO treated food-deprived mice with aDCN mCherry control mice (n = 6, paired t-test, P = 0.418). (j) Scatter plot comparing changes in GRABDA signals to amount of chow consumed in 1 h following activation of the aDCN in hM3D(Gq)-expressing mice treated with CNO (n = 13, Pearson correlation). (k) Average ∆F/F of GRABDA signals in the ventral striatum of food-deprived aDCN-INT hM3D(Gq) mice treated with vehicle or CNO. Signals are aligned to the vehicle or CNO injection (red line). Dark line represents the mean and lighter shaded area represents SEMs. Corresponding heatmaps (right) depict ∆F/F of GRABDA signals in each mouse (n = 7). (l) Average ∆F/F of GRABDA signals in 3-min bins following vehicle or CNO treatment of the aDCN-INT with hM3D(Gq) (n = 7, repeated measures two-way ANOVA interaction P = 0.301). (m) Maximum ∆F/F of GRABDA signals in the ventral striatum of vehicle or CNO treated food-deprived mice with aDCN-INT hM3D(Gq) mice (n = 7, paired t-test, P = 0.410). (n) Mean ∆F/F of GRABDA signals in the ventral striatum of vehicle or CNO treated food-deprived mice with aDCN-INT hM3D(Gq) mice (n = 7, paired t-test, P = 0.367). Data are expressed as mean ± SEM, two-sided P values, t-tests and post-hoc comparisons: *P < 0.05, **P < 0.01, ***P < 0.001; ANOVA interaction: ∞∞∞P < 0.001; ANOVA main effect of group: ¤¤¤P < 0.001. Statistical analysis in Supplementary Table 1
Extended Data Fig. 11 Selective activation of glutamatergic aDCN neurons is sufficient to induce striatal dopamine surge and suppression of food intake.
(a) Schematic depicting hM3D(Gq) expression in the DCN combined with GRABDA expression and fibre implant in the striatum of a vGluT2::Cre mouse. (b-d) IHC analysis of Cre dependent hM3D(Gq) expression in the DCN of vGluT2::Cre mouse (green, vGluT2, red, hM3D(Gq), blue, DAPI). Scale bar, 25 µm in b-d. (e) Average ∆F/F of GRABDA signals in the ventral striatum of food-deprived mice expressing hM3D(Gq) in glutamatergic neurons of the aDCN-LAT following vehicle or CNO injection. Signals are aligned to the vehicle or CNO injection (red line). Dark line represents the mean and lighter shaded area represents SEMs. Corresponding heatmaps (right) depict ∆F/F of GRABDA signals in each mouse (n = 7). (f) Average ∆F/F of GRABDA signals in the ventral striatum of food-deprived mice expressing hM3D(Gq) in glutamatergic neurons of the aDCN-INT following vehicle or CNO injection. Signals are aligned to the vehicle or CNO injection (red line). Dark line represents the mean and lighter shaded area represents SEMs. Corresponding heatmaps (right) depict ∆F/F of GRABDA signals in each mouse (n = 6). (g) Average ∆F/F of GRABDA signals in 3-min bins of food-deprived mice expressing hM3D(Gq) in glutamatergic neurons of the aDCN-INT or aDCN-LAT (n = 7 vGluT2 aDCN-LAT mice, green, n = 6 vGluT2 aDCN-INT mice, grey, two-way ANOVA, interaction P < 0.001, main effect P < 0.001; Holm-Sidak’s, P = 0.001/ = 0.001/ < 0.001 (9-12 min), P < 0.001/ < 0.001/ < 0.001 (12-15 min), P < 0.001/ < 0.001/ < 0.001 (15-18 min), P < 0.001/ < 0.001/ < 0.001 (18-21 min), P < 0.001/ < 0.001/ < 0.001 (21-24 min), P<0.001/ < 0.001/ < 0.001 (24-27 min), P < 0.001/ < 0.001/ <0.001 (27-30 min), aDCN-LAT CNO to vehicle/aDCN-INT CNO/aDCN-INT vehicle respectively). (h) Maximum ∆F/F GRABDA signals in the ventral striatum of food-deprived mice expressing hM3D(Gq) in glutamatergic neurons of the aDCN-LAT following vehicle or CNO treatment (n = 7, paired t-test, P < 0.001). (i) Mean ∆F/F GRABDA signals in the ventral striatum of food-deprived mice expressing hM3D(Gq) in glutamatergic neurons of the aDCN-LAT following vehicle or CNO treatment (n = 7, paired t-test, P = 0.001). (j) Maximum ∆F/F GRABDA signals in the ventral striatum of food-deprived mice expressing hM3D(Gq) in glutamatergic neurons of the aDCN-INT following vehicle or CNO treatment (n = 6, paired t-test, P = 0.644). (k) Mean ∆F/F GRABDA signals in the ventral striatum of food-deprived mice expressing hM3D(Gq) in glutamatergic neurons of the aDCN-INT following vehicle or CNO treatment (n = 6, paired t-test, P = 0.367). (l) Maximum ∆F/F GRABDA signals in the striatum following non-specific aDCN-LAT activation or vGluT2+ aDCN-LAT neuron activation (aDCN-LAT: n = 6 mice, vGluT2 aDCN-LAT: n = 7, unpaired t-test, P = 0.250). (m) Mean ∆F/F GRABDA signals in the striatum of following non-specific aDCN-LAT activation or vGluT2+ aDCN-LAT neuron activation (aDCN-LAT: n = 6 mice, vGluT2 aDCN-LAT: n = 7, unpaired t-test, P = 0.323). (n) Plot of GRABDA signals and corresponding food intake in food-deprived mice treated following glutamatergic aDCN activation (n = 13, Pearson correlation). Solid line indicates the linear trend line fit to the data. Data are expressed as mean ± SEM, two-sided P values, t-tests and post-hoc comparisons: **P < 0.01, ***P < 0.001; ANOVA interaction: ∞∞∞P < 0.001; ANOVA main effect of group: ¤¤¤P < 0.001. Statistical analysis in Supplementary Table 1.
Extended Data Fig. 12 Increased striatal dopamine suppresses food intake.
(a) Schematic depicting hM3D(Gq) expression in the VTA of a DAT::Cre mouse, and GRABDA expression and fibre implant in the ventral striatum. (b-c) Representative images of hM3D(Gq) expression in the VTA (b), and GRABDA expression and fibre track in the ventral striatum (c) of a DAT::Cre mouse. Scale bar, 500 µm in B, 200 µm in C. (d) Average ∆F/F of GRABDA signals in 3-min bins following VTA neuron activation with vehicle and varying concentrations of CNO (0.025 mg/Kg, 0.25 mg/Kg, 1 mg/Kg and 2.5 mg/Kg; n = 8 per group, repeated measures two-way ANOVA interaction P < 0.001, main effect P < 0.001; Holm-Sidak’s). (e) Net area under curve ∆F/F of GRABDA signals following VTA neuron activation with vehicle and varying concentrations of CNO (0.025 mg/Kg, 0.25 mg/Kg, 1 mg/Kg and 2.5 mg/Kg; n = 8 per group, repeated measures one-way ANOVA P < 0.001; Holm-Sidak’s, P = 0.012 (vehicle versus 0.025 mg/Kg), P=0.010 (vehicle versus 0.25 mg/Kg), P = 0.023 (vehicle versus 1.0 mg/Kg), P = 0.010 (vehicle versus 2.5 mg/Kg)). (f) Food intake of food-deprived mice following VTA neuron activation with vehicle and varying concentrations of CNO (0.025 mg/Kg, 0.25 mg/Kg, 1 mg/Kg and 2.5 mg/Kg; n = 8 per group, repeated measures one-way ANOVA P<0.001; Holm-Sidak’s, P < 0.001 (vehicle versus 0.025 mg/Kg), P < 0.001 (vehicle versus 0.25 mg/Kg), P < 0.001 (vehicle versus 1.0 mg/Kg), P < 0.001 (vehicle versus 2.5 mg/Kg)). (g) Plot of GRABDA signals and corresponding food intake in food-deprived mice treated following VTA neuron activation (n = 8 per group, Pearson correlation). Solid line indicates the linear trend line fit to the data. (h) Average ∆F/F of GRABDA signals from 0 to 30 min following treatment with vehicle and varying concentrations of CNO (n = 8 per group, repeated measures one-way ANOVA P < 0.001; Holm-Sidak’s, P = 0.004 (vehicle versus 0.025 mg/Kg), P = 0.004 (vehicle versus 0.25 mg/Kg), P = 0.004 (vehicle versus 1.0 mg/Kg), P = 0.004 (vehicle versus 2.5 mg/Kg)). (i) Maximum ∆F/F of GRABDA signals following treatment with vehicle and varying concentrations of CNO (n = 8 per group, repeated measures one-way ANOVA P=0.023; Holm-Sidak’s, P = 0.034 (vehicle versus 2.5 mg/Kg)). (j) Average ∆F/F of GRABDA signals during presentation of food in fasted mice following treatment with vehicle and varying concentrations of CNO (0.025, 0.25, 1.0 and 2.5 mg/Kg). Signals are aligned to food presentation. Dark lines represent mean values and lighter shaded areas represent SEM (n = 8). (k) Heatmaps reporting ∆F/F of GRABDA signals in individual mice in (j) (n = 8). (l) Maximum ∆F/F of GRABDA signals during food presentation in mice following treatment with vehicle and varying concentrations of CNO (n = 8 per group, one-way ANOVA P = 0.0117; Holm-Sidak’s, P = 0.0389 (vehicle versus 0.25 mg/Kg), P = 0.0056 (vehicle versus 1.0 mg/Kg), P=0.0121 (vehicle versus 2.5 mg/Kg)). (m) Scatter plot depicting the maximal ∆F/F GRABDA response to food following pre-stimulation of VTA DA neurons and the associated amount of food intake following pre-stimulation of VTA DA neurons (n = 8 per group, Pearson correlation, P < 0.01). Solid line shows the linear trend line fit to the data. (n-p) Images of hM4D(Gi) expression (red) in TH+, VTA neurons (green) of a DAT::Cre mouse (n). Higher magnification of white box (o-p). Scale bar, 500 µm (n), 50 µm (p). (q) Neurons transduced with hM4D(Gi) in the VTA and SNC (n = 3, 1047, 2745, 2710 neurons each mouse, unpaired t-test, P = 0.02). (r) Average ∆F/F of DA signals in aDCN-LAT hM3D(Gq) mice and aDCN-LAT hM3D(Gq); VTA hM4D(Gi) mice (n = 6 per group, unpaired t-test, P = 0.003). (g) Distance travelled by aDCN-LAT hM3D(Gq) mice and aDCN-LAT hM3D(Gq); VTA hM4D(Gi) mice during a 10-min open field session (n = 6 and 7, respectively, unpaired t-test, P = 0.382). Data are expressed as mean ± SEM, two-sided P values, t-tests and post-hoc comparisons: *P < 0.05, **P<0.01, ***P < 0.001; ANOVA interaction: ∞∞∞P < 0.001; ANOVA main effect of group: ¤¤¤P < 0.001. SNC, substantia nigra pars compacta; TH, tyrosine hydroxylase; VTA, ventral tegmental area. Statistical analysis in Supplementary Table 1.
Extended Data Fig. 13 Proposed role of the cerebellum in feeding control.
The cerebellum is well-positioned to integrate homeostatic satiation signals and is capable of orchestrating adaptive feeding responses by modulating motor, cognitive, affective and endocrine functions20,66,67,68,69,70,71,72,73,74,75. Visual, gustatory and olfactory inputs are all known to activate the cerebellum76,77,78 which could provide salience update to control appetitive drive. It functions as a comparator of physiological nutrient state (interoception) and post-ingestion nutritional outcome (nutrient feedback) to fine-tune predictive reward signals (reward network)79 and ultimately influence meal size (feeding network). While cerebellar output has been shown to influence VTA neuron activity12,80, our observed changes in DA signalling are tightly associated with decreases in food intake, suggesting a dedicated role of the cerebellum in regulating DA circuits that influence feeding that is distinct from motor80 or social12 behaviours. Based on our mechanistic studies into the changes in the reward system mediated by the cerebellum, it is possible that previously discovered differences between PWS and control subjects arise because of cerebellar alterations8,81,82. In response to a predicted meal size (predicted nutritional reward outcome) by either food cues or food, cerebellar activity increases dopamine efflux that blunts dopamine transients. Consequently, the reward value of consuming food reduces and meals are terminated. In PWS patients8,81,82, food-dependent cerebellar activity is absent and thus, dopamine transients remain regardless of amount of food consumed, leading to excessive eating. Conversely, in dopamine-deficient animals, there is a complete absence of drive to eat14. A better understanding of the mechanisms and circuits underlying cerebellar-mediated behaviours can guide brain stimulation strategies to control food intake recently shown to have the capability of ameliorating symptoms for disorders associated with the cerebellum83,84,85,86.
Supplementary information
Supplementary Table 1
Details of statistics used and statistical results. Related to Figs. 1–4, and Extended Data Figs. 2–12.
Source data
Rights and permissions
About this article
Cite this article
Low, A.Y.T., Goldstein, N., Gaunt, J.R. et al. Reverse-translational identification of a cerebellar satiation network. Nature 600, 269–273 (2021). https://doi.org/10.1038/s41586-021-04143-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-021-04143-5
This article is cited by
-
Cytokine enrichment in deep cerebellar nuclei is contributed by multiple glial populations and linked to reduced amyloid plaque pathology
Journal of Neuroinflammation (2023)
-
Modeling tissue co-regulation estimates tissue-specific contributions to disease
Nature Genetics (2023)
-
Purkinje cell dopaminergic inputs to astrocytes regulate cerebellar-dependent behavior
Nature Communications (2023)
-
Ghrelin signaling in the cerebellar cortex enhances GABAergic transmission onto Purkinje cells
Scientific Reports (2023)
-
Linking the cerebellum to Parkinson disease: an update
Nature Reviews Neurology (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.