Abstract
The Toll/interleukin-1 receptor (TIR) domain is a canonical component of animal and plant immune systems1,2. In plants, intracellular pathogen sensing by immune receptors triggers their TIR domains to generate a molecule that is a variant of cyclic ADP-ribose3,4. This molecule is hypothesized to mediate plant cell death through a pathway that has yet to be resolved5. TIR domains have also been shown to be involved in a bacterial anti-phage defence system called Thoeris6, but the mechanism of Thoeris defence remained unknown. Here we show that phage infection triggers Thoeris TIR-domain proteins to produce an isomer of cyclic ADP-ribose. This molecular signal activates a second protein, ThsA, which then depletes the cell of the essential molecule nicotinamide adenine dinucleotide (NAD) and leads to abortive infection and cell death. We also show that, similar to eukaryotic innate immune systems, bacterial TIR-domain proteins determine the immunological specificity to the invading pathogen. Our results describe an antiviral signalling pathway in bacteria, and suggest that the generation of intracellular signalling molecules is an ancient immunological function of TIR domains that is conserved in both plant and bacterial immunity.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
Data that support the findings of this study are available within the Article and its Extended Data. Gene accessions appear in the Methods section of the paper. Plasmid maps of the constructs used for the experiments are attached as Supplementary Files. Additional data are available from the corresponding authors upon reasonable request. Source data are provided with this paper.
References
Fitzgerald, K. A. & Kagan, J. C. Toll-like receptors and the control of immunity. Cell 180, 1044–1066 (2020).
Burch-Smith, T. M. & Dinesh-Kumar, S. P. The functions of plant TIR domains. Sci. STKE 2007, pe46 (2007).
Wan, L. et al. TIR domains of plant immune receptors are NAD+-cleaving enzymes that promote cell death. Science 365, 799–803 (2019).
Horsefield, S. et al. NAD+ cleavage activity by animal and plant TIR domains in cell death pathways. Science 365, 793–799 (2019).
Bayless, A. M. & Nishimura, M. T. Enzymatic functions for Toll/interleukin-1 receptor domain proteins in the plant immune system. Front. Genet. 11, 539 (2020).
Doron, S. et al. Systematic discovery of antiphage defense systems in the microbial pangenome. Science 359, eaar4120 (2018).
Balint-Kurti, P. The plant hypersensitive response: concepts, control and consequences. Mol. Plant Pathol. 20, 1163–1178 (2019).
Duxbury, Z. et al. Induced proximity of a TIR signaling domain on a plant–mammalian NLR chimera activates defense in plants. Proc. Natl Acad. Sci. USA 117, 18832–18839 (2020).
Ka, D., Oh, H., Park, E., Kim, J.-H. & Bae, E. Structural and functional evidence of bacterial antiphage protection by Thoeris defense system via NAD+ degradation. Nat. Commun. 11, 2816 (2020).
Lopatina, A., Tal, N. & Sorek, R. Abortive infection: bacterial suicide as an antiviral immune strategy. Annu. Rev. Virol. 7, 371–384 (2020).
Tzipilevich, E., Pollak-Fiyaksel, O. & Ben-Yehuda, S. Bacteria elicit a phage tolerance response subsequent to infection of their neighbors. Preprint at https://doi.org/10.1101/2021.02.16.428622 (2021).
Morehouse, B. R. et al. STING cyclic dinucleotide sensing originated in bacteria. Nature 586, 429–433 (2020).
Burroughs, A. M. & Aravind, L. Identification of uncharacterized components of prokaryotic immune systems and their diverse eukaryotic reformulations. J. Bacteriol. 202, https://doi.org/10.1128/JB.00365-20 (2020).
Burroughs, A. M., Zhang, D., Schäffer, D. E., Iyer, L. M. & Aravind, L. Comparative genomic analyses reveal a vast, novel network of nucleotide-centric systems in biological conflicts, immunity and signaling. Nucleic Acids Res. 43, 10633–10654 (2015).
Huang, Y., Fliegert, R., Guse, A. H., Lü, W. & Du, J. A structural overview of the ion channels of the TRPM family. Cell Calcium 85, 102111 (2020).
Huang, Y., Roth, B., Lü, W. & Du, J. Ligand recognition and gating mechanism through three ligand-binding sites of human TRPM2 channel. eLife 8, e50175 (2019).
Cohen, D. et al. Cyclic GMP–AMP signalling protects bacteria against viral infection. Nature 574, 691–695 (2019).
Ye, Q. et al. HORMA domain proteins and a Trip13-like ATPase regulate bacterial cGAS-like enzymes to mediate bacteriophage immunity. Mol. Cell 77, 709–722 (2020).
Essuman, K. et al. The SARM1 Toll/interleukin-1 receptor domain possesses intrinsic NAD+ cleavage activity that promotes pathological axonal degeneration. Neuron 93, 1334–1343 (2017).
Essuman, K. et al. TIR domain proteins are an ancient family of NAD+-consuming enzymes. Curr. Biol. 28, 421–430 (2018).
Coronas-Serna, J. M. et al. The TIR-domain containing effectors BtpA and BtpB from Brucella abortus impact NAD metabolism. PLoS Pathog. 16, e1007979 (2020).
Watanabe, S., Shiwa, Y., Itaya, M. & Yoshikawa, H. Complete sequence of the first chimera genome constructed by cloning the whole genome of Synechocystis strain PCC6803 into the Bacillus subtilis 168 genome. J. Bacteriol. 194, 7007 (2012).
Wilson, G. A. & Bott, K. F. Nutritional factors influencing the development of competence in the Bacillus subtilis transformation system. J. Bacteriol. 95, 1439–1449 (1968).
Mazzocco, A., Waddell, T. E., Lingohr, E. & Johnson, R. P. Enumeration of bacteriophages using the small drop plaque assay system. Methods Mol. Biol. 501, 81–85 (2009).
Zheng, L. et al. Fumarate induces redox-dependent senescence by modifying glutathione metabolism. Nat. Commun. 6, 6001 (2015).
Pluskal, T., Castillo, S., Villar-Briones, A. & Orešič, M. MZmine 2: modular framework for processing, visualizing, and analyzing mass spectrometry-based molecular profile data. BMC Bioinformatics 11, 395 (2010).
Myers, O. D., Sumner, S. J., Li, S., Barnes, S. & Du, X. One step forward for reducing false positive and false negative compound identifications from mass spectrometry metabolomics data: new algorithms for constructing extracted ion chromatograms and detecting chromatographic peaks. Anal. Chem. 89, 8696–8703 (2017).
Bernheim, A. et al. Prokaryotic viperins produce diverse antiviral molecules. Nature 589, 120–124 (2020).
Berman, H. M. et al. The Protein Data Bank. Nucleic Acids Res. 28, 235–242 (2000).
Sonn-Segev, A. et al. Quantifying the heterogeneity of macromolecular machines by mass photometry. Nat. Commun. 11, 1772 (2020).
Acknowledgements
We thank the Sorek laboratory members, M. Voichek, A. Levy, D. Dar, V. Šikšnys and M. Zaremba for comments on earlier versions of this manuscript; Y. M. Bar-On for useful discussion during the project; C. Avraham and T. Fedorenko for their assistance with the experiments; A. Bernheim for her continuous advice and support throughout this project; A. Sonn-Segev for her assistance with mass photometry experiments; and Y. Peleg and S. Albeck for their help in purification of the ThsA protein. R.S. was supported, in part, by the European Research Council (grant ERC-CoG 681203), the Ernest and Bonnie Beutler Research Program of Excellence in Genomic Medicine, the Minerva Foundation with funding from the Federal German Ministry for Education and Research, the Knell Family Center for Microbiology, and the Yotam project and the Weizmann Institute Sustainability And Energy Research (SAERI) initiative. G.O. was supported by the SAERI doctoral fellowship. A.M. was supported by a fellowship from the Ariane de Rothschild Women Doctoral Program and, in part, by the Israeli Council for Higher Education via the Weizmann Data Science Research Center, and by a research grant from O. Klein-Astrachan.
Author information
Authors and Affiliations
Contributions
G.O. designed all the experiments, performed all of the experiments unless otherwise noted, analysed the data and wrote the manuscript. E.H. performed the experimental design and data analysis for the mass spectrometry experiments presented in Figs. 1d, 3a, b, Extended Data Figs. 1h, i, 3a. M.B. and D.C. performed the cloning and plaque assay experiments presented in Fig. 4b, c, Extended Data Fig. 4. A.M. and S.D. performed bioinformatic analysis leading to the identification of the Thoeris systems studied. N.T. participated in writing the manuscript. D.B.A.M. performed the mass spectrometry experiments and data analysis presented in Fig. 3a, b. S.M. performed the mass spectrometry experiments presented in Fig. 1d, Extended Data Fig. 1h, i. G.A. performed, together with G.O., the enzymatic assay experiments presented in Figs. 2b, c, 4d; the structural analysis presented in Fig. 3c; and the mass photometry experiments presented in Fig. 3d, e, Extended Data Fig. 3c–f. R.S. supervised the study and wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
R.S. is a scientific cofounder and advisor of BiomX, Pantheon Bioscience and Ecophage. The other authors declare no competing interests.
Additional information
Peer review information Nature thanks the anonymous reviewers for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Effects of mutations in Thoeris genes on defence against phage SPO1.
a, Plaques of phage SPO1 on control cells (black), cells expressing both WT Thoeris proteins (green), cells expressing mutant ThsA(N112A) and WT ThsB (magenta), or cells expressing WT ThsA and mutant ThsB(E85Q) (cyan). Ten-fold serial dilution of the phage lysate were dropped on the plates. b, Efficiency of plating (EOP) of phage SPO1 on control and Thoeris-containing strains, representing plaque-forming units per millilitre (PFU/ml). Asterisk marks statistically significant reduction in EOP (One-way ANOVA, followed by pairwise multiple comparison analysis according to Tukey’s honest significant difference criterion p=2*10−8). c, Replication of phage SPO1 in the presence of Thoeris-containing and Thoeris-lacking (control) cells, or without cells (no bacteria). Lysates were collected 2.5 h following infection of liquid cultures at an initial MOI of 5, and phage titer was quantified by plating serial dilution of the lysates on the control strain. Asterisk marks statistically significant reduction in EOP compared to the control strain (One-way ANOVA, followed by pairwise multiple comparison analysis according to Tukey’s honest significant difference criterion p=0.016). d, Remaining phage titre after culture recovery. Samples were collected 12 h following infection of liquid cultures at an initial MOI of 5, and phage titre was quantified by plating serial dilution of the samples on the control strain. Asterisk marks statistically significant reduction in EOP compared to the control strain (One-way ANOVA, followed by pairwise multiple comparison analysis according to Tukey’s honest significant difference criterion p = 0.0004). e, Growth curves of infections with phage SPO1 at MOI of 5 of control cells (black), Thoeris-expressing cells (dark green), and 11 colonies isolated from recovered Thoeris-expressing cells 12 h post infection (light green). 3–4 Colonies were isolated from each of three independent infections. f, g, Growth curves of uninfected Thoeris mutants (f) and during infection by phage SPO1 at MOI of 5 or 0.05 (g). Three independent experiments are presented as individual curves. h, Adenosine diphosphate ribose (ADPR) levels in control culture (black) and cells expressing wild type Thoeris (green). Time 0 represents uninfected cells. Cells were infected by phage SPO1 at an MOI of 5. Each line represents the mean of three independent experiments, with individual data points shown. ADPR levels were measured by LC–MS and calculated from the area under the curve of the identified ADPR peak. i, NAD+ levels in cell expressing WT ThsB + ThsA(N112A) (magenta) and cells expressing WT ThsA + ThsB(E85Q) (cyan). Time 0 represents uninfected cells. Each line represents the mean of three independent experiments, with individual data points shown. Cells were infected by phage SPO1 at an MOI of 5. NAD+ levels were measured by LC–MS and calculated from the area under the curve of the identified NAD+ peak.
Extended Data Fig. 2 NADase activity of ThsA after addition of standard cADPR.
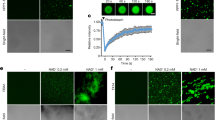
NADase activity of ThsA, when incubated with filtered cell lysates derived from ThsB-expressing cells 70-min post infection by phage SPO1, or with buffer containing 0 µM – 1000 µM of synthetic cyclic ADP-ribose (cADPR). NADase activity was calculated as the rate of change in εNAD fluorescence during the linear phase of the reaction. Bars represent mean of three experiments, with individual data points overlaid. NADase activity between the cADPR-containing samples and the blank sample (0 µM cADPR) are not statistically significant (One-way ANOVA, followed by pairwise multiple comparison analysis according to Tukey’s honest significant difference).
Extended Data Fig. 3 MS/MS spectra of the cADPR isomer produced by ThsB following phage infection and the activity of the SLOG domain.
a, MS/MS fragmentation spectra of standard cADPR (top) and the Thoeris-derived cADPR isomer (bottom). Hypothesized structures of MS/MS fragments of cADPR are presented. b, EOP of phages on bacteria expressing WT ThsB and ThsA(R371A) compared to WT Thoeris and other mutants. Results for the control, WT Thoeris and ThsB(E85Q) and ThsA(N112A) are those presented in Extended Data Fig. 1. Asterisk marks statistically significant reduction in EOP (One-way ANOVA, followed by pairwise multiple comparison analysis according to Tukey’s honest significant difference criterion p = 2*10−8). EOP of ThsA(R371A) is not statistically significant compared to the control strain (p = 0.7). c–f, Two additional replicates of the mass photometry measurement presented in Figs 3d, e. ThsA purified protein was incubated with lysates derived from infected cells expressing ThsB(E85Q) (c, e) or ThsB (d, f). Dashed lines represent masses of ThsA monomer, dimer and tetramer.
Extended Data Fig. 4 Efficiency of plating on cells expressing Thoeris systems of B. cereus, B. dafuensis and hybrid systems.
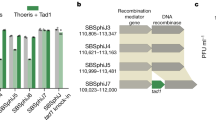
a, EOP of phages on control cells (black) and cells expressing combinations of ThsA from either B. cereus (green) or B. dafuensis (blue), TIR protein from B. cereus (ThsB), TIR proteins from B. dafuensis (TIR1 and TIR2) or both TIRs from B. dafuensis (TIR1+TIR2). Bars represent mean of 3 replicates, with individual data points overlaid. Asterisk marks statistically significant reduction in EOP (One-way ANOVA, followed by pairwise multiple comparison analysis according to Tukey’s honest significant difference criterion p<0.0015). b, EOP of phages on control cells (black) and cells expressing mutant ThsA(N112A) from B. cereus together with combinations of TIR proteins from B. cereus and B. dafuensis.
Supplementary information
Supplementary Information
This file contains Supplementary Tables 1 and 2.
Supplementary File 1
Plasmid map of pGO1_thrC_Pxyl_cereus_ThsA, shuttle vector for B. cereus MSX-D12 thsA gene.
Supplementary File 2
Plasmid map of pGO2_thrC_Pxyl_dafuensis_ThsA, shuttle vector for B. dafuensis FJAT-25496 thsA
Supplementary File 3
Plasmid map of pGO3_amyE_hspank_cereus_ThsB, shuttle vector for B. cereus MSX-D12 thsB gene.
Supplementary File 4
Plasmid map of pGO4_amyE_hspank_dafuensis_TIR1, shuttle vector for B. dafuensis FJAT-25496 tir1 gene.
Supplementary File 5
Plasmid map of pGO5_amyE_hspank_dafuensis_TIR2, shuttle vector for B. dafuensis FJAT-25496 tir2 gene.
Supplementary File 6
Plasmid map of pGO6_amyE_hspank_GFP, shuttle vector for GFP control construct.
Supplementary File 7
Plasmid map of pGO7_amyE_hspank_dafuensis_TIR1+TIR2, shuttle vector for B. dafuensis FJAT-25496 tir1+tir2 genes.
Supplementary File 8
Plasmid map of pGO8_pET30a_strep_BdTIR_His, protein expression plasmid for Brachypodium distachyon BdTIR gene.
Supplementary File 9
Plasmid map of pGO9_pAB151_cereus_ThsA_strep, protein expression plasmid for B. cereus ThsA purification.
Rights and permissions
About this article
Cite this article
Ofir, G., Herbst, E., Baroz, M. et al. Antiviral activity of bacterial TIR domains via immune signalling molecules. Nature 600, 116–120 (2021). https://doi.org/10.1038/s41586-021-04098-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-021-04098-7
This article is cited by
-
Phages overcome bacterial immunity via diverse anti-defence proteins
Nature (2024)
-
Activation of Thoeris antiviral system via SIR2 effector filament assembly
Nature (2024)
-
Anti-phage defence through inhibition of virion assembly
Nature Communications (2024)
-
Structural basis for phage-mediated activation and repression of bacterial DSR2 anti-phage defense system
Nature Communications (2024)
-
Insights into the modulation of bacterial NADase activity by phage proteins
Nature Communications (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.