Abstract
The progression of chronic liver disease to hepatocellular carcinoma is caused by the acquisition of somatic mutations that affect 20–30 cancer genes1,2,3,4,5,6,7,8. Burdens of somatic mutations are higher and clonal expansions larger in chronic liver disease9,10,11,12,13 than in normal liver13,14,15,16, which enables positive selection to shape the genomic landscape9,10,11,12,13. Here we analysed somatic mutations from 1,590 genomes across 34 liver samples, including healthy controls, alcohol-related liver disease and non-alcoholic fatty liver disease. Seven of the 29 patients with liver disease had mutations in FOXO1, the major transcription factor in insulin signalling. These mutations affected a single hotspot within the gene, impairing the insulin-mediated nuclear export of FOXO1. Notably, six of the seven patients with FOXO1S22W hotspot mutations showed convergent evolution, with variants acquired independently by up to nine distinct hepatocyte clones per patient. CIDEB, which regulates lipid droplet metabolism in hepatocytes17,18,19, and GPAM, which produces storage triacylglycerol from free fatty acids20,21, also had a significant excess of mutations. We again observed frequent convergent evolution: up to fourteen independent clones per patient with CIDEB mutations and up to seven clones per patient with GPAM mutations. Mutations in metabolism genes were distributed across multiple anatomical segments of the liver, increased clone size and were seen in both alcohol-related liver disease and non-alcoholic fatty liver disease, but rarely in hepatocellular carcinoma. Master regulators of metabolic pathways are a frequent target of convergent somatic mutation in alcohol-related and non-alcoholic fatty liver disease.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
WGS data in the form of BAM files across samples reported in this study have been deposited in the European Genome-Phenome Archive (accession number EGAD00001006255). RNA-sequencing data have been deposited in the European Nucleotide Archive (https://www.ebi.ac.uk/ena/browser/home) with accession number ERP123192.
Code availability
Detailed methods and custom R scripts for the analysis of clinical features, telomere lengths and metabolomics data are available in the Supplementary Code. Other packages used in the analysis are listed below: R: v.3.5.1, Perl: v.5.3.0, Python: v.3.8.5, MATLAB: v.R2019b, BWA-MEM: v.0.7.17 (https://sourceforge.net/projects/bio-bwa/), cgpCaVEMan: v.1.11.2/1.13.14/1.15.1 (https://github.com/cancerit/CaVEMan), cgpPindel: v.2.2.2/2.2.4/2.2.5/3.2.0/3.3.0 (https://github.com/cancerit/cgpPindel), Brass: v.5.4.1/6.0.5/6.1.2/6.2.0/6.3.4 (https://github.com/cancerit/BRASS), ASCAT NGS: v.4.0.1/ 4.1.2/4.2.1 (https://github.com/cancerit/ascatNgs), JBrowse: v.1.16.1 (https://jbrowse.org/), cgpVAF: v.2.4.0 (https://github.com/cancerit/vafCorrect), alleleCount: v.4.1.0 (https://github.com/cancerit/alleleCount), SigProfiler: v.1.0.0-GRCh37 (https://github.com/AlexandrovLab), HDP: v.0.1.5 (https://github.com/nicolaroberts/hdp), dNdScv: v.0.0.1 (https://github.com/im3sanger/dndscv), Telomerecat: v.3.4.0 (https://github.com/jhrf/telomerecat), STAR: v.2.7.6a (https://github.com/alexdobin/STAR), Picard-tools: v.2.20.7 (https://broadinstitute.github.io/picard/), Samtools: v.1.12 (http://www.htslib.org/), TrimGalore: v.0.6.4 (https://github.com/FelixKrueger/TrimGalore), GATK: v.4.1.4.1 (https://gatk.broadinstitute.org/hc/en-us), GSEA: v.3.0 (https://www.gsea-msigdb.org/gsea/index.jsp), XGBoost: v.0.82.1 (https://xgboost.readthedocs.io/en/latest/), NDP.view2 (https://www.hamamatsu.com/eu/en/product/type/U12388-01/index.html), label.switching: v.1.8 (https://cran.r-project.org/web/packages/label.switching/index.html), philentropy: v.0.3.0 (https://cran.r-project.org/web/packages/philentropy/index.html), MCMCglmm: v.2.29 (https://cran.r-project.org/web/packages/MCMCglmm/index.html), Magick: v.2.0 (https://cran.r-project.org/web/packages/magick/index.html), Pheatmap: v.1.0.12 (https://cran.r-project.org/web/packages/pheatmap/index.html), Thermo Fisher software Tracefinder: v.5.0 (https://www.thermofisher.com/uk/en/home/industrial/mass-spectrometry/liquid-chromatography-mass-spectrometry-lc-ms/lc-ms-software/lc-ms-data-acquisition-software/tracefinder-software.html), CellProfiler: v.4.0.3 (https://cellprofiler.org/), PerkinElmer Harmony: v.4.9 (https://www.perkinelmer.com/category/cellular-imaging-software).
References
The Cancer Genome Atlas Research Network. Comprehensive and integrative genomic characterization of hepatocellular carcinoma. Cell 169, 1327–1341 (2017).
Schulze, K. et al. Exome sequencing of hepatocellular carcinomas identifies new mutational signatures and potential therapeutic targets. Nat. Genet. 47, 505–511 (2015).
Totoki, Y. et al. Trans-ancestry mutational landscape of hepatocellular carcinoma genomes. Nat. Genet. 46, 1267–1273 (2014).
Fujimoto, A. et al. Whole-genome sequencing of liver cancers identifies etiological influences on mutation patterns and recurrent mutations in chromatin regulators. Nat. Genet. 44, 760–764 (2012).
Letouzé, E. et al. Mutational signatures reveal the dynamic interplay of risk factors and cellular processes during liver tumorigenesis. Nat. Commun. 8, 1315 (2017).
Guichard, C. et al. Integrated analysis of somatic mutations and focal copy-number changes identifies key genes and pathways in hepatocellular carcinoma. Nat. Genet. 44, 694–698 (2012).
Fujimoto, A. et al. Whole-genome mutational landscape and characterization of noncoding and structural mutations in liver cancer. Nat. Genet. 48, 500–509 (2016).
Pinyol, R. et al. Molecular characterization of hepatocellular carcinoma in patients with non-alcoholic steatohepatitis. J. Hepatol. 75, 865–878 (2021).
Nault, J. C. et al. Telomerase reverse transcriptase promoter mutation is an early somatic genetic alteration in the transformation of premalignant nodules in hepatocellular carcinoma on cirrhosis. Hepatology 60, 1983–1992 (2014).
Torrecilla, S. et al. Trunk mutational events present minimal intra- and inter-tumoral heterogeneity in hepatocellular carcinoma. J. Hepatol. 67, 1222–1231 (2017).
Zhu, M. et al. Somatic mutations increase hepatic clonal fitness and regeneration in chronic liver disease. Cell 177, 608–621 (2019).
Kim, S. K. et al. Comprehensive analysis of genetic aberrations linked to tumorigenesis in regenerative nodules of liver cirrhosis. J. Gastroenterol. 54, 628–640 (2019).
Brunner, S. F. et al. Somatic mutations and clonal dynamics in healthy and cirrhotic human liver. Nature 574, 538–542 (2019).
Blokzijl, F. et al. Tissue-specific mutation accumulation in human adult stem cells during life. Nature 538, 260–264 (2016).
Yizhak, K. et al. RNA sequence analysis reveals macroscopic somatic clonal expansion across normal tissues. Science 364, eaaw0726 (2019).
Brazhnik, K. et al. Single-cell analysis reveals different age-related somatic mutation profiles between stem and differentiated cells in human liver. Sci. Adv. 6, eaax2659 (2020).
Barneda, D. et al. The brown adipocyte protein CIDEA promotes lipid droplet fusion via a phosphatidic acid-binding amphipathic helix. Elife 4, e07485 (2015).
Sun, Z. et al. Perilipin1 promotes unilocular lipid droplet formation through the activation of Fsp27 in adipocytes. Nat. Commun. 4, 1594 (2013).
Li, J. Z. et al. Cideb regulates diet-induced obesity, liver steatosis, and insulin sensitivity by controlling lipogenesis and fatty acid oxidation. Diabetes 56, 2523–2532 (2007).
Hammond, L. E. et al. Mitochondrial glycerol-3-phosphate acyltransferase-1 is essential in liver for the metabolism of excess acyl-CoAs. J. Biol. Chem. 280, 25629–25636 (2005).
Wendel, A. A., Cooper, D. E., Ilkayeva, O. R., Muoio, D. M. & Coleman, R. A. Glycerol-3-phosphate acyltransferase (GPAT)−1, but not GPAT4, incorporates newly synthesized fatty acids into triacylglycerol and diminishes fatty acid oxidation. J. Biol. Chem. 288, 27299–27306 (2013).
Jeon, S. & Carr, R. Alcohol effects on hepatic lipid metabolism. J. Lipid Res. 61, 470–479 (2020).
Friedman, S. L., Neuschwander-Tetri, B. A., Rinella, M. & Sanyal, A. J. Mechanisms of NAFLD development and therapeutic strategies. Nat. Med. 24, 908–922 (2018).
Clugston, R. D. et al. Altered hepatic lipid metabolism in C57BL/6 mice fed alcohol: a targeted lipidomic and gene expression study. J. Lipid Res. 52, 2021–2031 (2011).
Puri, P. et al. A lipidomic analysis of nonalcoholic fatty liver disease. Hepatology 46, 1081–1090 (2007).
Meister, G. et al. Identification of novel argonaute-associated proteins. Curr. Biol. 15, 2149–2155 (2005).
Rheinbay, E. et al. Analyses of non-coding somatic drivers in 2,658 cancer whole genomes. Nature 578, 102–111 (2020).
Yaffe, M. B. et al. The structural basis for 14-3-3:phosphopeptide binding specificity. Cell 91, 961–971 (1997).
Saline, M. et al. AMPK and AKT protein kinases hierarchically phosphorylate the N-terminus of the FOXO1 transcription factor, modulating interactions with 14-3-3 proteins. J. Biol. Chem. 294, 13106–13116 (2019).
Kleiner, D. E. et al. Design and validation of a histological scoring system for nonalcoholic fatty liver disease. Hepatology 41, 1313–1321 (2005).
Ishak, K. et al. Histological grading and staging of chronic hepatitis. J. Hepatol. 22, 696–699 (1995).
Ellis, P. et al. Reliable detection of somatic mutations in solid tissues by laser-capture microdissection and low-input DNA sequencing. Nat. Protoc. 16, 841–871 (2021).
Jones, D. et al. cgpCaVEManWrapper: simple execution of CaVEMan in order to detect somatic single nucleotide variants in NGS data. Curr. Protoc. Bioinformatics 56, 15.10.1–15.10.18 (2016).
Yoshida, K. et al. Tobacco smoking and somatic mutations in human bronchial epithelium. Nature 578, 266–272 (2020).
Papastamoulis, P. label.switching: an R package for dealing with the label switching problem in MCMC outputs. J. Stat. Softw. 69, Code Snippet 1 (2015).
Nik-Zainal, S. et al. The life history of 21 breast cancers. Cell 149, 994–1007 (2012).
Martincorena, I. et al. Universal patterns of selection in cancer and somatic tissues. Cell 171, 1029–1041 (2017).
Raine, K. M. et al. cgpPindel: identifying somatically acquired insertion and deletion events from paired end sequencing. Curr. Protoc. Bioinformatics 52, 15.7.1–15.7.12 (2015).
Campbell, P. J. et al. Identification of somatically acquired rearrangements in cancer using genome-wide massively parallel paired-end sequencing. Nat. Genet. 40, 722–729 (2008).
Stephens, P. J. et al. Massive genomic rearrangement acquired in a single catastrophic event during cancer development. Cell 144, 27–40 (2011).
Sohlenius-Sternbeck, A. K. Determination of the hepatocellularity number for human, dog, rabbit, rat and mouse livers from protein concentration measurements. Toxicol. Vitr. 20, 1582–1586 (2006).
Lipscomb, J. C., Fisher, J. W., Confer, P. D. & Byczkowski, J. Z. In vitro to in vivo extrapolation for trichloroethylene metabolism in humans. Toxicol. Appl. Pharmacol. 152, 376–387 (1998).
Bergstrom, E. N. et al. SigProfilerMatrixGenerator: a tool for visualizing and exploring patterns of small mutational events. BMC Genomics 20, 685 (2019).
Alexandrov, L. B. et al. The repertoire of mutational signatures in human cancer. Nature 578, 94–101 (2020).
Drost, H.-G. Philentropy: information theory and distance quantification with R. J. Open Source Softw. 3, 765 (2018).
Qiao, W. et al. PERT: a method for expression deconvolution of human blood samples from varied microenvironmental and developmental conditions. PLoS Comput. Biol. 8, (2012).
Farmery, J. H. R. et al. Telomerecat: a ploidy-agnostic method for estimating telomere length from whole genome sequencing data. Sci. Rep. 8, 1300 (2018).
Hadfield, J. D. MCMC methods for multi-response generalized linear mixed models: the MCMCglmm R package. J. Stat. Softw. 33, v033i02 (2010).
Hoare, M. et al. NOTCH1 mediates a switch between two distinct secretomes during senescence. Nat. Cell Biol. 18, 979–992 (2016).
Liao, Y., Smyth, G. K. & Shi, W. FeatureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30, 923–930 (2014).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140 (2009).
Acknowledgements
This work was supported by a Cancer Research UK Grand Challenge Award (C98/A24032) and the Wellcome Trust. S.W.K.N. holds an EMBO Long Term Fellowship (ALTF 721-2019). S.F.B. was supported by the Swiss National Science Foundation (P2SKP3-171753 and P400PB-180790). M.A.S. is supported by a Rubicon fellowship from NWO (019.153LW.038). The Cambridge Human Research Tissue Bank is supported by the NIHR Cambridge Biomedical Research Centre. M.H. is supported by a CRUK Clinician Scientist Fellowship (C52489/A19924) and a CRUK Accelerator Award (C18873/A26813). P.J.C. was supported by a Wellcome Senior Clinical Fellowship until 2020 (WT088340MA).
Author information
Authors and Affiliations
Contributions
P.J.C., M.H. and S.W.K.N. designed the experiments. S.W.K.N. performed mutation calling and computational analyses including visualization of results for mutation calling; identification of SNV clusters and the inference of phylogenetic relationships between them; assignment of indels and FOXO1 hotspot mutations to SNV clusters; clone size estimation and comparisons; mutational signature extraction; identification of protein-coding and non-coding drivers; telomere length estimation; processing and normalization of RNA-sequencing data; gene set enrichment analysis; and estimation of the liver-wide mass of driver-mutation-bearing hepatocytes. S.W.K.N. developed software for the refinement of indel calling, phylogenetic inference and visualization of clonal structure, and clone size estimation, visualization and mapping to histological images. P.J.C. assisted with the filtering of structural variants, performed statistical inference of factors that affect telomere length using mixed effects models and supervised all statistical analyses. N. Brzozowska performed telomere length estimation. F.A. and I.M. provided support for running variants of dNdScv. M.R.S. advised on mutational signature extraction. T.H.H.C. provided support for running beta-binomial-based variant filtering. M.A.S. provided support and advice for performing LCM-specific variant-filtering algorithms for SNV and structural variant calls. D.L. and T.B. provided insights into indel filtering associated with homopolymers and problematic genomic loci. F.J.R., S.F.B., Y.H., B.W. and N. Birtchnell performed tissue sectioning, fixing, staining and histology image generation. S.F.B. also performed LCM and submission for WGS, and was responsible for the initial development of source code for producing diagnostic plots to facilitate the manual determination of clonal relationships, and the visualization of phylogenetic tree structures. P.R., A.I. and T.B. provided wet laboratory support. N.W., J.T., K.R. and A.P.B. provided technical support for computational analyses. M.H. and F.J.R. provided biological samples used in this study, and the associated clinical annotations were curated with assistance from S.J.A. and S.E.D. S.J.A. and S.E.D. analysed histology sections of background liver and HCC from all patients in the study, and L.M. supervised microdissection of tissue samples for sequencing. M.H. coordinated all validation experiments relating to FOXO1 hotspot mutations using HCC cell lines, with additional support from H.N. M.Y., E.N. and C.F. performed analysis of metabolites from HCC cell lines. L.C. and R.R. performed processing and quality control of RNA-sequencing samples and data. P.J.C., S.W.K.N. and M.H. drafted the manuscript with input and guidance from M.R.S. and I.M., and updated the paper after contributions from all authors.
Corresponding authors
Ethics declarations
Competing interests
A patent has been filed by CRUK’s technology transfer office, with support from that of Wellcome Sanger Institute (named inventors: S.W.K.N., M.H. and P.J.C.), covering the use of somatic mutations in liver tissue for stratifying diagnosis and treatment of patients with metabolic diseases.
Additional information
Peer review information Nature thanks the anonymous reviewers for their contribution to the peer review of this work. Peer reviewer reports are available.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Mutations in ACVR2A.
a, Distribution of somatic mutations in ACVR2A according to genomic location. Pie charts show fraction of sequencing reads reporting the mutant allele in each microdissection. b, Two microdissections in different patients showing structural variants generating copy loss of ACVR2A. Black points represent corrected read depth along the chromosome. Lines and arcs represent structural variants, coloured by the orientation of the joined ends (purple, deletion-type orientation; brown, tandem-duplication-type orientation; turquoise, head-to-head inverted; green, tail-to-tail inverted).
Extended Data Fig. 2 Mutations in TNRC6B and NEAT1.
a, Distribution of somatic mutations in CLCN5 according to genomic location. Pie charts show fraction of sequencing reads reporting the mutant allele in each microdissection. b, Distribution of somatic mutations in the long non-coding RNA, NEAT1, according to genomic location. Pie charts show fraction of sequencing reads reporting the mutant allele in each microdissection.
Extended Data Fig. 3 Structural variants affecting FOXO1 and GPAM.
a, A chromothripsis event affecting chromosome 13 in one of the microdissections from PD37907, a patient with NAFLD. Black points represent corrected read depth along the chromosome. Lines and arcs represent structural variants, coloured by the orientation of the joined ends (purple, deletion-type orientation; brown, tandem-duplication-type orientation; turquoise, head-to-head inverted; green, tail-to-tail inverted). The structural variant that breaks FOXO1 is highlighted, and would be predicted to break the gene within the first intron, preserving the first coding exon but deleting the remaining coding exons. b, A tandem duplication upstream of GPAM in a microdissection from PD37110, a patient with ARLD. GPAM is left intact, but the tandem duplication starts 20kb upstream of the gene.
Extended Data Fig. 4 Multiple independent acquisitions of FOXO1 mutations in PD37239.
The clone map from Fig. 1b is shown, laid onto an H&E-stained section. On the left of the figure, raw sequencing data from representative samples with and without FOXO1 mutations are shown, with their physical locations on the H&E section shown by the arrows. In the sequencing data, reads mapping to the forward strand of the reference genome are in pink; the reverse strand in blue. Base calls that do not match the reference genome are shown as coloured squares. The locations of the S22W and R21L mutations are marked with arrows. The scatterplots arranged around the H&E section represent VAF plots of mutations in pairs of samples. The colours of the x and y axis titles match the clone map colours of the H&E section. Individual mutations called in either sample are shown in orange, according to their VAF, with the FOXO1 S22W mutation shown in dark green. In clonally related pairs of samples, most of the mutations are shared by both samples, evident as a cloud of mutations with non-zero VAF. In clonally unrelated samples, the mutations line the x and y axes, with the one exception being the FOXO1 mutation, indicating that it is independently acquired in the two clones.
Extended Data Fig. 5 Further examples of FOXO1 mutations in patients with chronic liver disease.
a–c, Phylogenetic trees and clone maps are shown for PD37234 (a), PD37105 (b) and PD37245 (c). The left panel shows the phylogenetic tree, with coloured branches showing independently acquired mutations. Solid lines indicate that nesting is in accordance with the pigeonhole principle; dashed lines indicate that nesting is in accordance with the pigeonhole principle, assuming that hepatocytes represent < 100% of cells. The right panel shows the clones from the phylogenetic tree mapped onto an H&E-stained photomicrograph of the liver, with FOXO1-mutant clones coloured to match the tree.
Extended Data Fig. 6 Somatic mutations of FOXO1 impair its phosphorylation and nuclear export.
a, HepG2 cells were transfected with the indicated wild-type or mutant constructs of FOXO1 fused with a C-terminal GFP. Cells were counterstained with DAPI to highlight the nucleus, and imaged after overnight serum starvation conditions (left) and after 15 min of exposure to 100 nM insulin (right). Studies were performed in triplicate. b, HepG2 cells, expressing ectopic eGFP-tagged wild-type or mutant FOXO1 constructs as indicated and treated for 15 min with vehicle or insulin (100nM), were analysed for the indicated proteins by immunoblotting. Molecular weight markers (kDa) indicated. Studies were performed in triplicate. Uncropped versions of the blots are shown in Supplementary Fig. 4.
Extended Data Fig. 7 Nuclear–cytoplasmic ratios for wild-type and mutant FOXO1-GFP constructs in HCC cell lines.
a, b, Wide-field view of Hep3B (a) and PLC/PRF5 (b) cells pseudocoloured on a blue-to-red scale by the nuclear-cytoplasmic ratio of FOXO1-GFP. Cells were imaged under conditions of serum starvation (left), after exposure to insulin 100nM for 15 min (middle) or foetal calf serum (FCS) for 15 min (right).
Extended Data Fig. 8 RNA sequencing from cell lines transduced with either wild-type or mutant FOXO1-GFP constructs.
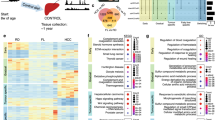
a, Heat map showing gene expression levels for genes in the ‘Canonical Glycolysis’ gene set from GO (GO:0061621). The order of genes on the x axis is determined by the level of significance (and direction of change) and the order of samples on the y axis is by condition (FOXO1 status and insulin status). b, Heat map showing gene expression levels for genes in the ‘Cell cycle, mitotic’ gene set from Reactome (R-HSA-69278). The order of genes on the x axis is determined by the level of significance (and direction of change) and the order of samples on the y axis is by condition (FOXO1 status and insulin status). c–e, Enrichment plots for the ‘FOXO-mediated transcription of oxidative stress, metabolic and neuronal genes’ gene set of Reactome (9615017) (c); ‘Lipid catabolic process’ gene set of GO (0016042) (d); and ‘Apoptotic process’ gene set of GO (0006915) (e). In each, the top panel reflects the cumulative enrichment score as the gene set is traversed from most up-regulated to most down-regulated in the presence of FOXO1-mutant constructs. The bottom panel in each shows the ranking of each gene in the gene set across all genes measured.
Extended Data Fig. 9 CIDEB mutations in patients with chronic liver disease.
a, Distribution of somatic mutations in CIDEB. Amino acid residues are coloured by type, with observed mutations in chronic liver disease shown above the wild-type protein sequence. b, Phylogenetic trees and clone maps are shown for one of the Couinaud segments of PD48367 with CIDEB mutations. The left panel shows the phylogenetic tree, with coloured branches showing independently acquired driver mutations. Solid lines indicate that nesting is in accordance with the pigeonhole principle; dashed lines indicate that nesting is in accordance with the pigeonhole principle, assuming that hepatocytes represent < 100% of cells. The right panel shows the clones from the phylogenetic tree mapped onto an H&E-stained photomicrograph of the liver, with mutant clones coloured to match the tree.
Extended Data Fig. 10 GPAM mutations in patients with chronic liver disease.
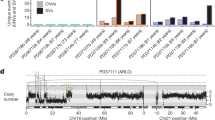
a, Distribution of somatic mutations in GPAM according to genomic location. Pie charts show fraction of sequencing reads reporting the mutant allele in each microdissection. b, Phylogenetic trees and clone maps are shown for a biopsy from PD37111 with GPAM mutations. The left panel shows the phylogenetic tree, with coloured branches showing independently acquired driver mutations. Solid lines indicate that nesting is in accordance with the pigeonhole principle; dashed lines indicate that nesting is in accordance with the pigeonhole principle, assuming that hepatocytes represent < 100% of cells. The right panel shows the clones from the phylogenetic tree mapped onto an H&E-stained photomicrograph of the liver, with mutant clones coloured to match the tree.
Extended Data Fig. 11 Properties of clones and patients with driver mutations.
a, Stacked bar chart showing the estimated cumulative liver mass carrying driver mutations, extrapolated from samples analysed in each patient. The calculations assume a total liver mass of 1500g for each patient. Bars are coloured for each of the 6 recurrently mutated genes identified in the study, and patient codes on the x axis are coloured for disease status. b, Estimated clone size for the 4 most frequently mutated genes compared to wild-type clones. The points are overlaid on box-and-whisker plots where the median is marked with a heavy black line and the interquartile range in a thin black box. The whiskers denote mark the full range of the data or 25th/75th centile plus 1.5x the interquartile range (whichever is smaller). The p values are two-sided, derived from Wilcoxon rank-sum tests and have not been corrected for multiple hypothesis testing. Sample sizes are n = 25 mutant clones for FOXO1; n = 17 mutant clones for CIDEB; n = 15 mutant clones for GPAM; and n = 32 mutant clones for ACVR2A. c, Scatter plot showing the distribution of ages of patients in the cohort by whether they carried clones with mutations in the specified genes or not. The p values are two-sided, derived from Wilcoxon rank-sum tests and have not been corrected for multiple hypothesis testing. Sample sizes were n = 7 FOXO1 mutant versus n = 22 FOXO1 wild-type; n = 6 CIDEB mutant versus n = 23 CIDEB wild-type; and n = 7 GPAM mutant versus n = 22 GPAM wild-type. d, Stacked bar charts showing the proportion of patients with or without type 2 diabetes by whether they carried driver mutations in each gene. The p values are two-sided, derived from Fisher’s exact tests and have not been corrected for multiple hypothesis testing. Sample sizes were as for c. e, Stacked bar charts showing the distribution of the NAFLD Activity Score (NAS) by whether they carried driver mutations in each gene, with low scores denoting a low degree of histological abnormality. The p values are two-sided, derived from chi-squared tests for trend and have not been corrected for multiple hypothesis testing. Sample sizes were as for c.
Extended Data Fig. 12 Analysis of telomere lengths.
a, Scatter plot showing the distribution of telomere lengths for samples grouped by disease status, and ranked from lowest to highest age within each disease category. b, Posterior distributions of the effect size of clone size (per log10(μm2)), age (per decade of life) and disease state (NAFLD and ARLD versus normal) on telomere lengths. Density plots are shown from the MCMC sampler, coloured by decile. Posterior ‘p values’ are calculated from the posterior samples of the MCMC chain and are two-sided and not corrected for multiple hypothesis testing. c, Telomere lengths layered onto two representative phylogenetic trees from patients with ARLD. Branches are coloured on a yellow-to-blue scale according to telomere lengths of the sample with the highest VAF assigned to that branch. The internal nodes are estimated using maximum likelihood and colours are interpolated along each branch.
Extended Data Fig. 13 Distribution of mutational signatures across the phylogenetic trees within the cohort.
Estimated proportional contributions of each mutational signature to each phylogenetically defined cluster of somatic substitutions. Stacked bar plots show proportional contributions of signatures in normal controls (top), patients with ARLD (middle), and patients with NAFLD (bottom).
Extended Data Fig. 14 Distribution of the new T>A signature across three samples.
a, Signatures for a sample with high rates of the novel signature (PD37240). The left panel shows phylogenetic trees with each branch coloured by the proportion of mutations in that branch assigned to the different mutational signatures. The contribution from the new signature is coloured purple. The middle panel shows the overlay of clones onto an H&E-stained liver section. Clones are coloured on a grey-to-purple scale according to the proportion of mutations attributed to the novel signature. The right panel shows observed mutation spectra for representative clones with low (top) or high (bottom) burden of the novel signature, laid out as for Fig. 4b. Purple arrows indicate parts of the mutation spectrum that are characteristic of the new mutational signature. b, c, In one patient with NAFLD, we had three samples from 2008 (not shown as the signature was absent), 2011 (b) and 2013 (c), with the relative contribution of the signature increasing over time. The photomicrograph of the H&E section in c was captured after the microdissections were excised, hence the white gaps in the tissue.
Supplementary information
Supplementary Information
This file contains Supplementary Notes 1 and 2, Supplementary Methods including further details on indel calling and mutational signature extraction not included in the main Methods section, Supplementary References and Supplementary Figures 1–6.
Supplementary Tables
This file contains Supplementary Tables 1–9 and a Supplementary Table Guide.
Supplementary Data
This file contains the Supplementary Code: HTMLs of Jupyter notebooks outlining key statistical analyses presented in the manuscript, including analysis of clinical variables, telomere lengths and metabolomics data.
Rights and permissions
About this article
Cite this article
Ng, S.W.K., Rouhani, F.J., Brunner, S.F. et al. Convergent somatic mutations in metabolism genes in chronic liver disease. Nature 598, 473–478 (2021). https://doi.org/10.1038/s41586-021-03974-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-021-03974-6
This article is cited by
-
FOXO transcription factors as mediators of stress adaptation
Nature Reviews Molecular Cell Biology (2024)
-
Deep whole-genome analysis of 494 hepatocellular carcinomas
Nature (2024)
-
Tissue mosaicism following stem cell aging: blood as an exemplar
Nature Aging (2024)
-
Genetic variation across and within individuals
Nature Reviews Genetics (2024)
-
Reconstructing phylogenetic trees from genome-wide somatic mutations in clonal samples
Nature Protocols (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.