Abstract
The early evolution of diapsid reptiles is marked by a deep contrast between our knowledge of the origin and early evolution of archosauromorphs (crocodiles, avian and non-avian dinosaurs) to that of lepidosauromorphs (squamates (lizards, snakes) and sphenodontians (tuataras)). Whereas the former include hundreds of fossil species across various lineages during the Triassic period1, the latter are represented by an extremely patchy early fossil record comprising only a handful of fragmentary fossils, most of which have uncertain phylogenetic affinities and are confined to Europe1,2,3. Here we report the discovery of a three-dimensionally preserved reptile skull, assigned as Taytalura alcoberi gen. et sp. nov., from the Late Triassic epoch of Argentina that is robustly inferred phylogenetically as the earliest evolving lepidosauromorph, using various data types and optimality criteria. Micro-computed tomography scans of this skull reveal details about the origin of the lepidosaurian skull from early diapsids, suggesting that several traits traditionally associated with sphenodontians in fact originated much earlier in lepidosauromorph evolution. Taytalura suggests that the strongly evolutionarily conserved skull architecture of sphenodontians represents the plesiomorphic condition for all lepidosaurs, that stem and crown lepidosaurs were contemporaries for at least ten million years during the Triassic, and that early lepidosauromorphs had a much broader geographical distribution than has previously been thought.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
Data availability
Computed tomography scan data, including surface volume files of the holotype, all morphological and molecular data generated and analysed, along with trees and log files described in the Article are available online as 'Supplementary Data 1' at Harvard Dataverse (https://doi.org/10.7910/DVN/G5EOOC)43.
Code availability
The MrBayes commands for Bayesian analyses are provided as ‘Supplementary Data 2’ and R scripts for reproducing the morphospace analyses and figures are available as the file ‘Supplementary Data 3’, both at Harvard Dataverse (https://doi.org/10.7910/DVN/G5EOOC)43.
References
Sues, H. D. The Rise of Reptiles: 320 Million Years of Evolution (John Hopkins Univ. Press, 2019).
Simões, T. R. & Caldwell, M. W. in Encyclopedia of Geology 2nd ed., vol. 3 (eds Alderton, D. & Elias, S. A.) 165–174 (Academic, 2021).
Simões, T. R. & Pyron, R. A. The squamate tree of life. Bull. Mus. Comp. Zool. 163, 47–95 (2021).
Uetz, P. & Hošek, J. The Reptile Database http://www.reptile-database.org (2021).
Gill, F., Donsker, D. & Rasmussen, F. IOC World Bird List (v.11.1) (2021).
Simões, T. R., Apesteguía, S., Hsiou, A. S. & Daza, J. D. Lepidosaurs from Gondwana: an introduction. J. Herpetol. 51, 297–299 (2017).
Sues, H.-D. & Kligman, B. T. A new lizard-like reptile from the Upper Triassic (Carnian) of Virginia and the Triassic record of Lepidosauromorpha (Diapsida, Sauria). J. Vert. Paleontol. 40, e1879102 (2021).
Schoch, R. R. & Sues, H.-D. A new lepidosauromorph reptile from the Middle Triassic (Ladinian) of Germany and its phylogenetic relationships. J. Vertebr. Paleontol. 38, e1444619 (2018).
Evans, S. E. & Borsuk-Białynicka, M. A small lepidosauromorph reptile from the Early Triassic of Poland. Palaeontol. Pol. 65, 179–202 (2009).
Romo De Vivar, P. R., Martinelli, A. G., Fonseca, P. H. M. & Soares, M. B. To be or not to be: the hidden side of Cargninia enigmatica and other puzzling remains of Lepidosauromorpha from the Upper Triassic of Brazil. J. Vert. Paleontol. 40, e1828438 (2020).
Cavicchini, I., Zaher, M. & Benton, M. J. An enigmatic neodiapsid reptile from the Middle Triassic of England. J. Vertebr. Paleontol. 40, e1781143 (2020).
Sobral, G., Simões, T. R. & Schoch, R. R. A tiny new Middle Triassic stem-lepidosauromorph from Germany: implications for the early evolution of lepidosauromorphs and the Vellberg fauna. Sci. Rep. 10, 2273 (2020).
Simões, T. R. et al. The origin of squamates revealed by a Middle Triassic lizard from the Italian Alps. Nature 557, 706–709 (2018).
Simões, T. R., Vernygora, O., Caldwell, M. W. & Pierce, S. E. Megaevolutionary dynamics and the timing of evolutionary innovation in reptiles. Nat. Commun. 11, 3322 (2020).
Simões, T. R., Caldwell, M. W. & Pierce, S. E. Sphenodontian phylogeny and the impact of model choice in Bayesian morphological clock estimates of divergence times and evolutionary rates. BMC Biol. 18, 191 (2020).
Scheyer, T. M. et al. Colobops: a juvenile rhynchocephalian reptile (Lepidosauromorpha), not a diminutive archosauromorph with an unusually strong bite. R. Soc. Open Sci. 7, 192179 (2020).
Hsiou, A. S., De França, M. A. G. & Ferigolo, J. New data on the Clevosaurus (Sphenodontia: Clevosauridae) from the Upper Triassic of Southern Brazil. PLoS ONE 10, e0137523 (2015).
Fraser, N. C. The osteology and relationships of Clevosaurus (Reptilia: Sphenodontida). Phil. Trans. R. Soc. Lond. B 321, 125–178 (1988).
Martinez, R. N. et al. A basal dinosaur from the dawn of the dinosaur era in southwestern Pangaea. Science 331, 206–210 (2011).
Garberoglio, F. F. et al. New skulls and skeletons of the Cretaceous legged snake Najash, and the evolution of the modern snake body plan. Sci. Adv. 5, eaax5833 (2019).
Bittencourt, J. S., Simões, T. R., Caldwell, M. W. & Langer, M. C. Discovery of the oldest South American fossil lizard illustrates the cosmopolitanism of early South American squamates. Commun. Biol. 3, 201 (2020).
Bertin, T. J. C., Thivichon-Prince, B., LeBlanc, A. R. H., Caldwell, M. W. & Viriot, L. Current perspectives on tooth implantation, attachment, and replacement in Amniota. Front. Physiol. 9, 1630 (2018).
Fraser, N. C. A new rhynchocephalian from the British Upper Trias. Palaeontology 25, 709–725 (1982).
Evans, S. E. The skull of a new eosuchian reptile from the Lower Jurassic of South Wales. Zool. J. Linn. Soc. 70, 203–264 (1980).
Whiteside, D. I. The head skeleton of the Rhaetian sphenodontid Diphydontosaurus avonis gen. et sp. nov. and the modernizing of a living fossil. Phil. Trans. R. Soc. Lond. B 312, 379–430 (1986).
Herrera‐Flores, J. A., Stubbs, T. L. & Benton, M. J. Macroevolutionary patterns in Rhynchocephalia: is the tuatara (Sphenodon punctatus) a living fossil? Palaeontology 60, 319–328 (2017).
Gemmell, N. J. et al. The tuatara genome reveals ancient features of amniote evolution. Nature 584, 403–409 (2020).
Jones, M. E. H. et al. Integration of molecules and new fossils supports a Triassic origin for Lepidosauria (lizards, snakes, and tuatara). BMC Evol. Biol. 13, 208 (2013).
Hsiou, A. S. et al. A new clevosaurid from the Triassic (Carnian) of Brazil and the rise of sphenodontians in Gondwana. Sci. Rep. 9, 11821 (2019).
Vernygora, O. V., Simões, T. R. & Campbell, E. O. Evaluating the performance of probabilistic algorithms for phylogenetic analysis of big morphological datasets: a simulation study. Syst. Biol. 69, 1088–1105 (2020).
Maddison, W. P. & Maddison, D. R. Mesquite: a modular system for evolutionary analysis, version 3.04, http://mesquiteproject.org (2015).
Goloboff, P. A., Farris, J. S. & Nixon, K. C. TNT, a free program for phylogenetic analysis. Cladistics 24, 774–786 (2008).
Ronquist, F. et al. MrBayes 3.2: efficient Bayesian phylogenetic inference and model choice across a large model space. Syst. Biol. 61, 539–542 (2012).
Miller, M. A., Pfeiffer, W. & Schwartz, T. Creating the CIPRES Science Gateway for inference of large phylogenetic trees in Gateway Computing Environments Workshop (GCE) 1–8 (IEEE, 2010).
Lanfear, R., Frandsen, P. B., Wright, A. M., Senfeld, T. & Calcott, B. PartitionFinder 2: new methods for selecting partitioned models of evolution for molecular and morphological phylogenetic analyses. Mol. Biol. Evol. 34, 772–773 (2017).
Lewis, P. O. A likelihood approach to estimating phylogeny from discrete morphological character data. Syst. Biol. 50, 913–925 (2001).
Hughes, M., Gerber, S. & Wills, M. A. Clades reach highest morphological disparity early in their evolution. Proc. Natl Acad. Sci. USA 110, 13875–13879 (2013).
Sutherland, J., Flannery, T., Moon, B. C., Stubbs, T. L. & Benton, M. J. Does exceptional preservation distort our view of disparity in the fossil record? Proc. R. Soc. Lond. Biol. Sci. 286, 20190091 (2019).
Lloyd, G. T. Estimating morphological diversity and tempo with discrete character-taxon matrices: implementation, challenges, progress, and future directions. Biol. J. Linn. Soc. 118, 131–151 (2016).
Gerber, S. Use and misuse of discrete character data for morphospace and disparity analyses. Palaeontology 62, 305–319 (2019).
Cisneros, J. C. & Ruta, M. Morphological diversity and biogeography of procolophonids (Amniota: Parareptilia). J. Syst. Palaeontology 8, 607–625 (2010).
Ciampaglio, C. N., Kemp, M. & McShea, D. W. Detecting changes in morphospace occupation patterns in the fossil record: characterization and analysis of measures of disparity. Paleobiology 27, 695–715 (2001).
Martinez, R., Simões, T. R., Sobral, G. & Apesteguía, S. Supplementary Data for “A Triassic stem lepidosaur illuminates the origin of lizard-like reptiles” Harvard Dataverse (2021).
Acknowledgements
R.N.M. thanks the Secretaría de Ciencia, Técnica e Innovación of San Juan (SECITI) and the field crew and EarthWatch volunteers of 2001 fieldwork. T.R.S. thanks the Natural Sciences and Engineering Research Council of Canada (NSERC) for providing a postdoctoral fellowship. We thank J. Antonio González for his line reconstruction of Taytalura skull and jaw and J. Blanco for his artwork of Taytalura.
Author information
Authors and Affiliations
Contributions
R.N.M. led the project, and conducted fieldwork and specimen preparation. T.R.S. conducted phylogenetic and morphospace analyses. G.S. performed CT scan data segmentation. T.R.S., G.S. and S.A. produced the figures. All authors contributed to interpretation of the results, discussions and manuscript writing.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature thanks Hans-Dieter Sues and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Location and geology of the Ischigualasto–Villa Unión Basin.
a, Geologic map of the southern outcrops of the Ischigualasto–Villa Unión Basin. b, Stratigraphic section of the Ischigualasto Formation. Black star indicates the type locality of PVSJ 698. Black line indicates the location of the stratigraphic section.
Extended Data Fig. 2 Cranial anatomy of T. alcoberi on the basis of segmented micro-computed tomography scan data.
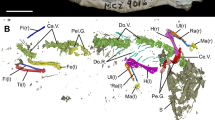
a, b, Three-dimensional model of Taytalura with segmented bones in right lateral and ventral views. c, Segmented right squamosal and quadrate in lateral view (left) and corresponding line drawing (right). d, Segmented left quadrate in posterior view. e, f, Line drawings of the left jaw in medial and dorsal views. af, adductor fossa; ap, anterior process; ar, articular; bc, braincase; co, coronoid; dp, dorsal process, fr, frontal; gf, glenoid fossa; hy, hyoid arch; ju, jugal; lc, lateral condyle, lj, lower jaw; mc, medial condyle, mf, medial flange; mx, maxilla; na, nasal; oc, otic conch; pa, parietal; pm, pre-maxilla; pl, palate; po, post-orbital; pof, post-frontal; pr, pre-artciular; prf, pre-frontal; qd, quadrate; qj, quadrato-jugal; sp., splenial; sq, squamosal; vp, ventral process. Scale bars, 3.5 mm.
Extended Data Fig. 3 Further details of cranial anatomy of T. alcoberi on the basis of segmented micro-computed tomography scan data.
a, b, Segmented palate in right lateral and left dorsolateral views. c, Segmented right posterior half of the braincase with stapes in red in lateral view. d, Segmented left posterior half of the braincase in lateral view. e, f, Segmented anterior region of the braincase in left lateral and anterior views. ap, ascending process; bc, braincase; bp, basipterygoid process; CN, cranial nerve; cp, cultriform process; cr, raised crest formed by the pterygoids meeting medially; clp, clinoid process; crp, crista prootica; ds, dorsum sella; ec, ectopterygoid; ep, epipterygoid; fo, fenestra ovalis; hf, hypophyseal fossa; lf, lateral flange; oc, otic conch; os, orbitosphenoid; pa, palatine; pp, paroccipital process; qw, quadrate wing; rc, recess; st, stapes. Roman numerals indicate corresponding cranial nerves. Scale bars, 3.5 mm (a, b), 1.5 mm (c–f).
Extended Data Fig. 4 Cranial anatomy of T. alcoberi on the basis of 2D cross-sectional slices from micro-computed tomography scan data.
a–c, Transverse cross sections of the skull showing the paired nasals (a), frontals (b) and parietals (c). d, Longitudinal cross-section of the paired frontals. e, Transverse cross section of the left lower and upper jaws in anterior view, showing labial and lingual walls of the tooth grooves of the maxilla and dentary. f, Transverse cross section of the posterior region of the skull through the braincase and the left lower jaw in anterior view. g, Transverse cross section of the left maxilla, showing the replacement tooth going into position. h, Longitudinal cross-section of the left dentary in occlusal view. i, j, Longitudinal cross section of the anterior (i) and posterior (j) sectors of the right maxilla in occlusal view. In h–i, the absence of the interdental ridges and less dense attachment tissue (alveolar bone and/or cementum) in the interdental space of the anterior dentition is shown, whereas j shows the direct contact between the posterior teeth and the absence of interdental space. Planes of the sections in the corresponding details. ac, adductor crest; alv.t., alveolar tissue; de, dentary; mx, maxilla; na, nasals; f-f, inter-frontal suture; fr, frontals; la, labial wall; li, lingual wall; ot, otolith; pa, parietals; pof, post-frontal; prf, pre-frontal; rp, resorption pit; rt, replacement tooth; to, tooth. Scale bars, 1.5 mm (a), 1 mm (b–e, h–j), 2.5 mm (f), 2 mm (g).
Extended Data Fig. 5 Skull and jaw of T. alcoberi.
a–d, Photographs of PVSJ 698 in dorsal (a), ventral (b), left lateral (c) and right lateral (d) views. e–h, Reconstruction of the skull of T. alcoberi in dorsal (e), ventral (f), anterior (g) and left lateral (h) views (reconstructed on the basis of information from both sides). i–k, Reconstruction of the lower jaw of T. alcoberi in lateral (i), medial (j) and occlusal (k) views. d, dentary; ec, ectopterygoid; fr, frontal; ju, jugal; mx, maxilla; na, nasal; pa, parietal; pm, pre-maxilla; pl, palatine; po, post-orbital; pof, post-frontal; prf, pre-frontal; pt, pterygoid; qd, quadrate; qj, quadrato-jugal; sq, squamosal; Scale bar, 10 mm.
Extended Data Fig. 6 Phylogenetic analyses using morphological data only, including data from all species.
a, Equal weights maximum parsimony analysis. Strict consensus of 602 most parsimonious trees (2,481 steps each). b, Bayesian inference analysis. Majority rule consensus tree. Numbers at nodes indicate posterior probabilities.
Extended Data Fig. 7 Bayesian inference phylogenetic analysis of combined morphological and molecular data, including data from all species.
Maximum compatibility tree (illustrating all nodes, including those with very low support). Numbers at nodes indicate posterior probabilities.
Extended Data Fig. 8 Phylogenetic analyses using morphological data only, with data from the rogue taxon (Vellbergia) removed.
a, Equal weights maximum parsimony analysis. Strict consensus of 572 most parsimonious trees (2,480 steps each). b, Bayesian inference analysis. Majority rule consensus tree. Numbers at nodes indicate posterior probabilities.
Extended Data Fig. 9 Bayesian inference phylogenetic analysis of combined morphological and molecular data, with data from the rogue taxon (Vellbergia) removed.
Majority rule consensus tree. Numbers at nodes indicate posterior probabilities.
Extended Data Fig. 10 Morphospace occupation by early diapsid reptiles and lepidosauromorphs, with the first 10 pairwise principal coordinate comparisons illustrated.
PC1 distinguishes squamates from all other groups when contrasted against all other principal coordinates (red), whereas PC3 distinguishes sphenodontians from all other groups when contrasted against all other principal coordinates (cyan). Major clades of interest are highlighted within convex hulls: squamatans (yellow), sphenodontians (light green), early lepidosaurs of uncertain placement (blue), all other diapsids (purple) and Taytalura (turquoise). For individual taxon names for each data point and relative contribution of each principal coordinate, see supplementary data files at Harvard Dataverse (‘Data availability’ in Methods).
Supplementary information
Supplementary Information
This file contains Geological and paleontological settings, Description of Taytahura alcoberi, Historical overview of lepidosauromorph classification, Additional taxa added to phylogenetic dataset, Synapomorphies Supplementary References.
Rights and permissions
About this article
Cite this article
Martínez, R.N., Simões, T.R., Sobral, G. et al. A Triassic stem lepidosaur illuminates the origin of lizard-like reptiles. Nature 597, 235–238 (2021). https://doi.org/10.1038/s41586-021-03834-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-021-03834-3
This article is cited by
-
A new sphenodontian (Diapsida: Lepidosauria) from the Upper Triassic (Norian) of Germany and its implications for the mode of sphenodontian evolution
BMC Ecology and Evolution (2024)
-
Ontogenetic variation in the cranium of Mixosaurus cornalianus, with implications for the evolution of ichthyosaurian cranial development
Swiss Journal of Palaeontology (2023)
-
Extended embryo retention and viviparity in the first amniotes
Nature Ecology & Evolution (2023)
-
Africa’s oldest dinosaurs reveal early suppression of dinosaur distribution
Nature (2022)
-
Synchrotron tomography of a stem lizard elucidates early squamate anatomy
Nature (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.