Abstract
In mammals, cyclic GMP–AMP (cGAMP) synthase (cGAS) produces the cyclic dinucleotide 2′3′-cGAMP in response to cytosolic DNA and this triggers an antiviral immune response. cGAS belongs to a large family of cGAS/DncV-like nucleotidyltransferases that is present in both prokaryotes1 and eukaryotes2,3,4,5. In bacteria, these enzymes synthesize a range of cyclic oligonucleotides and have recently emerged as important regulators of phage infections6,7,8. Here we identify two cGAS-like receptors (cGLRs) in the insect Drosophila melanogaster. We show that cGLR1 and cGLR2 activate Sting- and NF-κB-dependent antiviral immunity in response to infection with RNA or DNA viruses. cGLR1 is activated by double-stranded RNA to produce the cyclic dinucleotide 3′2′-cGAMP, whereas cGLR2 produces a combination of 2′3′-cGAMP and 3′2′-cGAMP in response to an as-yet-unidentified stimulus. Our data establish cGAS as the founding member of a family of receptors that sense different types of nucleic acids and trigger immunity through the production of cyclic dinucleotides beyond 2′3′-cGAMP.
This is a preview of subscription content, access via your institution
Access options



Similar content being viewed by others
Data availability
The authors declare that the data supporting the findings of this study are available within the Article and its Supplementary Information. The sequences and structures used in this study are CG12970 (UniProt: A1ZA55), CG30424 (UniProt: A8DYP7), CG4746 (UniProt: Q9U3W6), CG4766 (UniProt: Q9Y106), CG7194 (UniProt: Q9VSH0), porcine OAS1 (UniProt: Q29599), human OAS1 (UniProt: P00973-1), human cGAS (UniProt: Q8N884-1), porcine cGAS (UniProt: I3LM39), mouse cGAS (UniProt: Q8BSY1) and mouse cGAS in complex with DNA and a cGAMP intermediate analogue (Protein Data Bank code 4K98). Source data are provided with this paper.
References
Whiteley, A. T. et al. Bacterial cGAS-like enzymes synthesize diverse nucleotide signals. Nature 567, 194–199 (2019).
Ablasser, A. et al. cGAS produces a 2′-5′-linked cyclic dinucleotide second messenger that activates STING. Nature 498, 380–384 (2013).
Civril, F. et al. Structural mechanism of cytosolic DNA sensing by cGAS. Nature 498, 332–337 (2013).
Sun, L., Wu, J., Du, F., Chen, X. & Chen, Z. J. Cyclic GMP–AMP synthase is a cytosolic DNA sensor that activates the type I interferon pathway. Science 339, 786–791 (2013).
Wu, J. et al. Cyclic GMP–AMP is an endogenous second messenger in innate immune signaling by cytosolic DNA. Science 339, 826–830 (2013).
Lowey, B. et al. CBASS immunity uses CARF-related effectors to sense 3′-5′- and 2′-5′-linked cyclic oligonucleotide signals and protect bacteria from phage infection. Cell 182, 38–49 (2020).
Morehouse, B. R. et al. STING cyclic dinucleotide sensing originated in bacteria. Nature 586, 429–433 (2020).
Cohen, D. et al. Cyclic GMP–AMP signalling protects bacteria against viral infection. Nature 574, 691–695 (2019).
Guo, Z., Li, Y. & Ding, S. W. Small RNA-based antimicrobial immunity. Nat. Rev. Immunol. 19, 31–44 (2019).
Schneider, J. & Imler, J. L. Sensing and signalling viral infection in Drosophila. Dev. Comp. Immunol. 117, 103985 (2021).
Goto, A. et al. The kinase IKKβ regulates a STING- and NF-κB-dependent antiviral response pathway in Drosophila. Immunity 49, 225–234 (2018).
Liu, Y. et al. Inflammation-induced, STING-dependent autophagy restricts Zika virus infection in the Drosophila brain. Cell Host Microbe 24, 57–68 (2018).
Hua, X. et al. Stimulator of interferon genes (STING) provides insect antiviral immunity by promoting Dredd caspase-mediated NF-κB activation. J. Biol. Chem. 293, 11878–11890 (2018).
Donelick, H. M. et al. In vitro studies provide insight into effects of Dicer-2 helicase mutations in Drosophila melanogaster. RNA 26, 1847–1861 (2020).
Jin, L. et al. MPYS is required for IFN response factor 3 activation and type I IFN production in the response of cultured phagocytes to bacterial second messengers cyclic-di-AMP and cyclic-di-GMP. J. Immunol. 187, 2595–2601 (2011).
Burdette, D. L. et al. STING is a direct innate immune sensor of cyclic di-GMP. Nature 478, 515–518 (2011).
Diner, E. J. et al. The innate immune DNA sensor cGAS produces a noncanonical cyclic dinucleotide that activates human STING. Cell Rep. 3, 1355–1361 (2013).
Gao, P. et al. Cyclic [G(2′,5′)pA(3′,5′)p] is the metazoan second messenger produced by DNA-activated cyclic GMP–AMP synthase. Cell 153, 1094–1107 (2013).
Kranzusch, P. J., Lee, A. S., Berger, J. M. & Doudna, J. A. Structure of human cGAS reveals a conserved family of second-messenger enzymes in innate immunity. Cell Rep. 3, 1362–1368 (2013).
Zhang, X. et al. Cyclic GMP–AMP containing mixed phosphodiester linkages is an endogenous high-affinity ligand for STING. Mol. Cell 51, 226–235 (2013).
Cai, H. et al. 2′3′-cGAMP triggers a STING- and NF-κB-dependent broad antiviral response in Drosophila. Sci. Signal. 13, eabc4537 (2020).
Wu, X. et al. Molecular evolutionary and structural analysis of the cytosolic DNA sensor cGAS and STING. Nucleic Acids Res. 42, 8243–8257 (2014).
Martin, M., Hiroyasu, A., Guzman, R. M., Roberts, S. A. & Goodman, A. G. Analysis of Drosophila STING reveals an evolutionarily conserved antimicrobial function. Cell Rep. 23, 3537–3550.e6 (2018).
Caygill, E. E. & Brand, A. H. The GAL4 system: a versatile system for the manipulation and analysis of gene expression. Methods Mol. Biol. 1478, 33–52 (2016).
Tanaka, Y. & Chen, Z. J. STING specifies IRF3 phosphorylation by TBK1 in the cytosolic DNA signaling pathway. Sci. Signal. 5, ra20 (2012).
Zhong, B. et al. The adaptor protein MITA links virus-sensing receptors to IRF3 transcription factor activation. Immunity 29, 538–550 (2008).
Palmer, W. H., Medd, N. C., Beard, P. M. & Obbard, D. J. Isolation of a natural DNA virus of Drosophila melanogaster, and characterisation of host resistance and immune responses. PLoS Pathog. 14, e1007050 (2018).
Slavik, K. M. et al. cGAS-like receptors sense RNA and control 3′2′-cGAMP signaling in Drosophila. Nature, https://doi.org/10.1038/s41586-021-03743-5 (2021).
Hartmann, R., Justesen, J., Sarkar, S. N., Sen, G. C. & Yee, V. C. Crystal structure of the 2′-specific and double-stranded RNA-activated interferon-induced antiviral protein 2′-5′-oligoadenylate synthetase. Mol. Cell 12, 1173–1185 (2003).
Gui, X. et al. Autophagy induction via STING trafficking is a primordial function of the cGAS pathway. Nature 567, 262–266 (2019).
McFarland, A. P. et al. Sensing of bacterial cyclic dinucleotides by the oxidoreductase RECON promotes NF-κB activation and shapes a proinflammatory antibacterial state. Immunity 46, 433–445 (2017).
Eaglesham, J. B., McCarty, K. L. & Kranzusch, P. J. Structures of diverse poxin cGAMP nucleases reveal a widespread role for cGAS–STING evasion in host–pathogen conflict. eLife 9, e59753 (2020).
Eaglesham, J. B., Pan, Y., Kupper, T. S. & Kranzusch, P. J. Viral and metazoan poxins are cGAMP-specific nucleases that restrict cGAS–STING signalling. Nature 566, 259–263 (2019).
Hernáez, B. et al. Viral cGAMP nuclease reveals the essential role of DNA sensing in protection against acute lethal virus infection. Sci. Adv. 6, eabb4565 (2020).
Lowey, B. & Kranzusch, P. J. CD-NTases and nucleotide second messenger signaling. Curr. Biol. 30, R1106–R1108 (2020).
Andersen, L. L. et al. Frequently used bioinformatics tools overestimate the damaging effect of allelic variants. Genes Immun. 20, 10–22 (2019).
Andersen, L. L. et al. Functional IRF3 deficiency in a patient with herpes simplex encephalitis. J. Exp. Med. 212, 1371–1379 (2015).
Schulz, A., Jankowski, V., Zidek, W. & Jankowski, J. Highly sensitive, selective and rapid LC-MS method for simultaneous quantification of diadenosine polyphosphates in human plasma. J. Chromatogr. B Analyt. Technol. Biomed. Life Sci. 961, 91–96 (2014).
Wang, Y., Holleufer, A., Gad, H. H. & Hartmann, R. Length dependent activation of OAS proteins by dsRNA. Cytokine 126, 154867 (2020).
Holleufer, A. & Hartmann, R. A highly sensitive anion exchange chromatography method for measuring cGAS activity in vitro. Bio Protoc. 8, e3055 (2018).
Dalskov, L. et al. SARS-CoV-2 evades immune detection in alveolar macrophages. EMBO Rep. 21, e51252 (2020).
Acknowledgements
We thank K. Slavik and P. Kranzusch for discussing and sharing information before publication; P. E. Andersen for help with generating Sting-knockout S2 cells; D. Obbard for providing Kallithea virus; and C. Meignin, J. Marques, J. Schneider and G. Haas for helpful discussions. R.H was supported by grants from the Novo Nordisk Foundation (NNF17OC0028184) and the Danish Council for Independent Research (4183-0032B and 0135-00338B). J.-L.I. was supported by Agence Nationale de la Recherche (ANR-17-CE15-0014), Investissement d’Avenir Programs (ANR-10-LABX-0036, ANR-11-EQPX-0022), Institut Universitaire de France and the Chinese National Overseas Expertise Introduction Center for Discipline Innovation (Project ‘111’ (D18010)). H.C was supported by the Natural Science Foundation (32000662) and the Foreign Experts Program (2020A1414010306). A.P. was supported by an ERC consolidator grant (ERC-CoG ProDAP, 817798) and grants from the German Research Foundation (PI 1084/5, TRR179 and TRR237).
Author information
Authors and Affiliations
Contributions
A.H., K.G.W., H.H.G., H.C., J.-L.I. and R.H. conceived and designed experiments for this study. A.H., K.G.W., H.H.G., X.A., Y.C., L.L., Z.W., H.D., J.L., N.A.F., B.S., L.L.A., K.K., L.D. and H.C. performed experiments. A.H., H.H.G. and H.C. created the figures. H.C., A.P., J.-L.I. and R.H. supervised the study. A.H., J.-L.I. and R.H. wrote the original draft of the manuscript and all authors participated in reviewing and editing it.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature thanks Osamu Nureki and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Alignment of mammalian cGAS and OAS1 proteins and cGAS-like proteins from D. melanogaster.
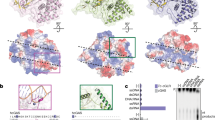
a, Extended view of the alignment shown in Fig. 1a with the positions of the GS/GG duplet, the metal ion coordinating acidic residues and the Zn finger indicated above the sequences with stars, arrows and bar, respectively. The annotated start methionine in the NCBI reference sequence for isoform D of CG30424 (cGLR2), which would delete the entire spine helix as well as part of the active site, is highlighted in yellow. Deletion of the spine helix is incompatible with a folded enzyme. Furthermore, the GS-containing loop, which coordinates the γ-phosphate of the donor nucleotide, is a universally conserved feature of nucleotidyltransferases. b, c, Structure (Protein Data Bank code 4K98) of mouse cGAS in complex with DNA and a cGAMP intermediate analogue (represented in sticks) and two magnesium ions (yellow spheres). The side chains of the acidic active site residues and the serine in the GS motif are also represented in sticks and their oxygen atoms are coloured red. Grey colouring represents the proportion of the protein upstream of the annotated start methionine site in CG30424 (cGLR2). b, Full view of the structure. c, Enhanced view of the active site.
Extended Data Fig. 2 Transient expression of candidate cGLRs in S2 cells.
a, Owing to issues with the anti-Flag M2 antibody giving rise to nonspecific bands when performing immunoblots on S2 cell lysates, we replaced the Flag tag we initially used with a V5 tag and reproduced the experiment from Fig. 1b. Cells were transfected with cGAS, cGLR1, cGLR2 and CG7194 or mutants thereof, as well as plasmids encoding firefly or Renilla luciferase under transcriptional control of the Sting or Actin5C promoter, respectively. At 24 h after transfection, luciferase activity was measured. Data are from one experiment performed in biological triplicates and are shown with mean ± s.d. (n = 3). b, Immunoblot showing the expression of the V5-tagged proteins from a. cGLR2 appears to be rapidly degraded in S2 cells, whereas it could easily be detected in HEK293T cells following transfection (Extended Data Fig. 10a). For gel source data, see Supplementary Fig. 1.
Extended Data Fig. 3 Ectopic expression of cGLR1 or cGLR2 in transgenic flies induces expression of Srg2 and Srg3.
a–d, Expression of cGLR1 (a), cGLR2 (b), Srg2 (c) and Srg3 (d) was monitored by RT–qPCR in transgenic flies ectopically expressing wild-type or mutant cGLRs. Expression was normalized to the housekeeping gene RpL32. Data are from three independent experiments (red, blue and green, each performed in biological triplicate) and shown with mean (n = 9). e, Flies of the indicated genotypes were injected with Tris and survival was monitored daily. Data are from 3 independent experiments, each with 30 flies (n = 90). In a–d, data were analysed using two-way ANOVA and a two-tailed Holm–Šídák post hoc test and compared to relevant AFA mutants. In e, log-rank test was used to compare cGLR1 and cGLR1 AFA (P = 0.0030) and cGLR2 and cGLR2 AFA (P = 0.0109)
Extended Data Fig. 4 cGLR signalling depends on Sting and Relish.
a, Sanger sequencing showing the 2-bp deletion in the Sting gene in Sting-knockout S2 cells. b, Immunoblot showing the lack of expression of Sting in Sting-knockout S2 cells. The arrow indicates the position of the Sting band. The presented immunoblot is representative out of two independent immunoblots. For gel source data, see Supplementary Fig. 1. c, d, S2 cells transfected with expression vectors for cGAS, cGLR1 or cGLR2 and plasmids encoding firefly or Renilla luciferase under transcriptional control of the Sting or Actin5C promoter, respectively, were used to monitor activation of the Sting pathway. c, S2 cells with or without co-transfection with an expression vector for Sting. d, Sting-knockout S2 cells with or without co-transfection with an expression vector for Sting. c, d, Data are from three independent experiments (blue, red, and green, each performed in biological triplicates) and are shown with mean (n = 9). Data were analysed using two-way ANOVA and a two-tailed Holm–Šídák post hoc test. e, S2 cells transfected with expression vectors for cGAS, cGLR1 or cGLR2 and plasmid encoding firefly luciferase under transcriptional control of the Sting promoter or a mutated version containing two point mutations in the Relish binding site. Data are from three independent experiments (blue, red and green, each performed in biological triplicates) and are shown with mean (n = 9). Data were analysed using two-way ANOVA and a two-tailed Holm–Šídák post hoc test, in which the groups were pairwise compared to mock.
Extended Data Fig. 5 Generation of cGLR1- and cGLR2-knockout flies.
The cGLR1 and cGLR2 genes, both located on the right arm of the second chromosome, are shown together with their annotated transcripts. Open reading frames are indicated in green. For cGLR1, an 8-bp deletion was introduced in the first exon using CRISPR–Cas9 technology. The deletion creates a frameshift after the asparagine residue at position 31, leading to termination of translation after insertion of single glycine residue. For cGLR2, a 5-bp deletion was created in exon 3, which is shared by all isoforms. The deletion results in a frameshift after the glutamate residue at position 338, leading to termination of translation after insertion of a 32 amino acid extension (HDRRIDPGSSLGNVPVRAKDSKRPEGRRDQPE).
Extended Data Fig. 6 Expression of Sting, Srg2 and Srg3 in flies upon infection with DCV.
a, Corresponding control to Fig. 2a. w1118, cGLR1-knockout, cGLR2-knockout or cGLR1/2-knockout flies were injected with Tris and survival was monitored daily. Data are from 3 independent experiments, each with 3 groups of around 10 flies, and shown with s.e.m. b–d, w1118, cGLR1-knockout, cGLR2-knockout or cGLR1/2-knockout flies were injected with DCV or Tris and expression of Sting (b), Srg2 (c) and Srg3 (d) was monitored by RT–qPCR at 2 and 3 days after injection (days post-infection (dpi)). Expression was normalized to the housekeeping gene RpL32. Data are from three independent experiments, each performed in biological triplicates (n = 9) shown with mean ± s.e.m. In a, log-rank test was used to test whether the survival curves differed (P = 0.0569). In b–d, data were analysed using two-way ANOVA and a two-tailed Holm–Šídák post hoc test
Extended Data Fig. 7 Expression of Sting, Srg2 and Srg3 in flies upon infection with Kallithea virus.
a, Corresponding control to Fig. 2b. w1118, cGLR1-knockout or cGLR2-knockout flies were injected with Tris and survival was monitored daily. Data are from 3 independent experiments, each with 3 groups of around 10 flies, and shown with s.e.m. b–d, w1118, cGLR1-knockout or cGLR2-knockout flies were injected with Kallithea virus or Tris and expression of Sting (b), Srg2 (c) and Srg3 (d) was monitored by RT–qPCR at 5 and 10 days after injection. Expression was normalized to the housekeeping gene RpL32. Data are from three independent experiments, each performed in biological triplicates (n = 9) shown with mean ± s.e.m. e, f, w1118, cGLR1-knockout, cGLR2-knockout or cGLR1/2-knockout flies were injected with Kallithea virus (e) or Tris (f) and survival was monitored daily. Data are from 3 independent experiments, each with 3 groups of around 10 flies, and shown with s.e.m. g, w1118, cGLR1-knockout, cGLR2 or cGLR1/2-knockout flies were injected with Kallithea virus and viral load was monitored by RT–qPCR at 5 days after infection. Expression was normalized to the housekeeping gene RpL32. Data are from three independent experiments, each performed in triplicates (n = 9) shown with mean ± s.e.m. In a, e, f, log-rank test was used to compare the survival curves pairwise followed by a Holm–Šídák multiple comparison correction (a, e) or all and once (f) (P = 0.1452). In b–d, data were analysed using two-way ANOVA and a two-tailed Holm–Šídák post hoc test. In g, log-transformed data were analysed using one-way ANOVA and a two-tailed Dunnett T3 post hoc test and compared to w1118 flies
Extended Data Fig. 8 Loss of cGLR1 or cGLR2 has a limited effect on VSV infection in flies.
a, b, w1118, cGLR1-knockout or cGLR2-knockout flies were injected with VSV (a) or Tris (b) and survival was monitored daily. Data are from 3 independent experiments, each with 3 groups of around 10 flies, and shown with s.e.m. c–g, w1118, cGLR1-knockout, cGLR2-knockout or cGLR1/2-knockout flies were injected with VSV or Tris and viral load (c) as well as expression of Sting (d), Srg1 (e), Srg2 (f) and Srg3 (g) were monitored by RT–qPCR at 4 and 5 days after infection. Expression was normalized to the housekeeping gene RpL32. Data are from three independent experiments, each performed in biological triplicates (n = 9) shown with mean ± s.e.m.In a, b, log-rank test was used to test whether the survival curves differed (a, P = 0.0772; b, P = 0.1942). In c, log-transformed data were analysed using two-way ANOVA and a two-tailed Holm–Šídák post hoc test and compared to w1118 flies. In d–g, data were analysed using two-way ANOVA and a two-tailed Holm–Šídák post hoc test
Extended Data Fig. 9 Loss of cGLR1 or cGLR2 has a limited effect on IIV6 infection in flies.
a, b, w1118, cGLR1-knockout or cGLR2-knockout flies were injected with IIV6 (a) or Tris (b) and survival was monitored daily. Data are from 3 independent experiments, each with 3 groups of around 10 flies, and shown with s.e.m. c–g, w1118, cGLR1-knockout or cGLR2-knockout flies were injected with IIV6 or Tris and viral load (c) as well as expression of Sting (d), Srg1 (e), Srg2 (f) and Srg3 (g) were monitored by RT–qPCR at 5 and 10 days after infection. Expression was normalized to the housekeeping gene RpL32. Data are from three independent experiments, each performed in biological triplicates (n = 9) shown with mean ± s.e.m. In a, b, log-rank test was used to test whether the survival curves differed (a, P = 0.3536; b, P = 0.7337). In c, log-transformed data were analysed using two-way ANOVA and a two-tailed Holm–Šídák post hoc test and compared to w1118 flies. In d–g, data were analysed using two-way ANOVA and a two-tailed Holm–Šídák post hoc test
Extended Data Fig. 10 cGLR2 produce 2′3′-cGAMP and 3′2′-cGAMP, which can activate human and Drosophila STING.
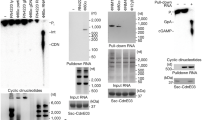
a, Immunoblot showing the expression of STING and Flag-tagged nucleotidyltransferases from Fig. 3a. b, HEK293T cells transfected with cGAS, cGAS AFA, cGLR2, cGLR2 AFA and STING as indicated and plasmids encoding firefly or Renilla luciferase under transcriptional control of the IFNB1 or a constitutive promoter, respectively. Data are from three independent experiments (blue, red and green, each performed in biological triplicates) and shown with mean (n = 9). c, Immunoblot showing the expression of STING and Flag-tagged nucleotidyltransferases from b. For gel source data, see Supplementary Fig. 1. d, Representative chromatograms from mass spectrometry analysis of 2′3′-cGAMP or 3′2′-cGAMP spiked lysates from GFP-transfected cells. e, Representative chromatogram from mass spectrometry analysis of cGAS-transfected cells. f, Representative chromatograms from mass spectrometry analysis of GFP- or cGLR2-transfected cells. g–i, HT-1080 (g) or S2 cells (h, i) permeabilized using digitonin and treated with the indicated cGAMPs. Expression of IFNB1 (g), Sting (h) or Srg3 (i) were monitored by RT–qPCR at 8 and 6 h after treatment for HT-1080 and S2 cells, respectively. Expression was normalized to the housekeeping genes GADPH (g) and RpL32 (h, i). Data are shown with mean ± s.e.m. (n = 4 for g; n = 3 for h, i) and are biological replicates from one representative experiment out of two independent experiments. In b, data were analysed using two-way ANOVA and a two-tailed Holm–Šídák post hoc test and compared to mock. In g–i, data were analysed using one-way ANOVA and a two-tailed Dunnett’s T3 post hoc test and compared to mock.
Extended Data Fig. 11 cGLR1 produces 3′2′-cGAMP in response to dsRNA.
a, Representative chromatograms from anion exchange chromatography analysis of different nucleotides. b, Representative chromatograms from anion exchange chromatography analysis of reaction products from activity assays with recombinant cGLR1 in the presence of different nucleic acids. c, Immunoblot showing the expression of Flag-tagged cGLR2 and mutants thereof from Fig. 3f. For gel source data, see Supplementary Fig. 1. d, S2 cells transfected with cGLR2 or mutants thereof as well as plasmids encoding firefly or Renilla luciferase under transcriptional control of the Sting or Actin5C promoter, respectively. Data are from three independent experiments (blue, red and green, each performed in biological triplicates) and shown with mean (n = 9). In d, data were analysed using two-way ANOVA and a two-tailed Holm–Šídák post hoc test.
Supplementary information
Supplementary Figure
This file contains the uncropped gel source data for Extended Data Figs 2, 4, 10 and 11.
Rights and permissions
About this article
Cite this article
Holleufer, A., Winther, K.G., Gad, H.H. et al. Two cGAS-like receptors induce antiviral immunity in Drosophila. Nature 597, 114–118 (2021). https://doi.org/10.1038/s41586-021-03800-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-021-03800-z
This article is cited by
-
Cytosolic DNA sensors in neurodegenerative diseases: from physiological defenders to pathological culprits
EMBO Molecular Medicine (2024)
-
Condensin-mediated restriction of retrotransposable elements facilitates brain development in Drosophila melanogaster
Nature Communications (2024)
-
Second messenger 2'3'-cyclic GMP-AMP (2'3'-cGAMP): the cell autonomous and non-autonomous roles in cancer progression
Acta Pharmacologica Sinica (2024)
-
Conservation and similarity of bacterial and eukaryotic innate immunity
Nature Reviews Microbiology (2024)
-
Evolution of the Major Components of Innate Immunity in Animals
Journal of Molecular Evolution (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.