Abstract
The mammalian brain develops through a complex interplay of spatial cues generated by diffusible morphogens, cell–cell interactions and intrinsic genetic programs that result in probably more than a thousand distinct cell types. A complete understanding of this process requires a systematic characterization of cell states over the entire spatiotemporal range of brain development. The ability of single-cell RNA sequencing and spatial transcriptomics to reveal the molecular heterogeneity of complex tissues has therefore been particularly powerful in the nervous system. Previous studies have explored development in specific brain regions1,2,3,4,5,6,7,8, the whole adult brain9 and even entire embryos10. Here we report a comprehensive single-cell transcriptomic atlas of the embryonic mouse brain between gastrulation and birth. We identified almost eight hundred cellular states that describe a developmental program for the functional elements of the brain and its enclosing membranes, including the early neuroepithelium, region-specific secondary organizers, and both neurogenic and gliogenic progenitors. We also used in situ mRNA sequencing to map the spatial expression patterns of key developmental genes. Integrating the in situ data with our single-cell clusters revealed the precise spatial organization of neural progenitors during the patterning of the nervous system.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
RNA-seq data are available at the Sequence Read Archive (https://www.ncbi.nlm.nih.gov/sra) under accession PRJNA637987. HybISS data are available at http://mousebrain.org/downloads. Single-cell gastrulation data were downloaded from https://github.com/MarioniLab/MouseGastrulationData.
Code availability
Source code is available at https://github.com/linnarsson-lab.
References
Camp, J. G. et al. Human cerebral organoids recapitulate gene expression programs of fetal neocortex development. Proc. Natl Acad. Sci. USA 112, 15672–15677 (2015).
Pollen, A. A. et al. Molecular identity of human outer radial glia during cortical development. Cell 163, 55–67 (2015).
Nowakowski, T. J. et al. Spatiotemporal gene expression trajectories reveal developmental hierarchies of the human cortex. Science 358, 1318–1323 (2017).
Zhong, S. et al. A single-cell RNA-seq survey of the developmental landscape of the human prefrontal cortex. Nature 555, 524–528 (2018).
Telley, L. et al. Temporal patterning of apical progenitors and their daughter neurons in the developing neocortex. Science 364, eaav2522 (2019).
La Manno, G. et al. Molecular diversity of midbrain development in mouse, human, and stem cells. Cell 167, 566–580.e19 (2016).
Carter, R. A. et al. A single-cell transcriptional atlas of the developing murine cerebellum. Curr. Biol. 28, 2910–2920.e2 (2018).
Zeisel, A. et al. Molecular architecture of the mouse nervous system. Cell 174, 999–1014.e22 (2018).
Rosenberg, A. B. et al. Single-cell profiling of the developing mouse brain and spinal cord with split-pool barcoding. Science 360, 176–182 (2018).
Cao, J. et al. The single-cell transcriptional landscape of mammalian organogenesis. Nature 566, 496–502 (2019).
Siegenthaler, J. A. & Pleasure, S. J. in Patterning and Cell Type Specification in the Developing CNS and PNS (eds. Rubenstein, J. L. R. & Rakic, P.) 835–849 (Elsevier, 2013).
Niwa, H., Miyazaki, J. & Smith, A. G. Quantitative expression of Oct-3/4 defines differentiation, dedifferentiation or self-renewal of ES cells. Nat. Genet. 24, 372–376 (2000).
Burtscher, I. & Lickert, H. Foxa2 regulates polarity and epithelialization in the endoderm germ layer of the mouse embryo. Development 136, 1029–1038 (2009).
Li, L. et al. Location of transient ectodermal progenitor potential in mouse development. Development 140, 4533–4543 (2013).
Zhang, X. et al. Pax6 is a human neuroectoderm cell fate determinant. Cell Stem Cell 7, 90–100 (2010).
Hatakeyama, J. et al. Hes genes regulate size, shape and histogenesis of the nervous system by control of the timing of neural stem cell differentiation. Development 131, 5539–5550 (2004).
Pijuan-Sala, B. et al. A single-cell molecular map of mouse gastrulation and early organogenesis. Nature 566, 490–495 (2019).
Tamai, H. et al. Pax6 transcription factor is required for the interkinetic nuclear movement of neuroepithelial cells. Genes Cells 12, 983–996 (2007).
Alves dos Santos, M. T. M. & Smidt, M. P. En1 and Wnt signaling in midbrain dopaminergic neuronal development. Neural Dev. 6, 23 (2011).
Schwarz, M. et al. Pax2/5 and Pax6 subdivide the early neural tube into three domains. Mech. Dev. 82, 29–39 (1999).
Sato, S. et al. Regulation of Six1 expression by evolutionarily conserved enhancers in tetrapods. Dev. Biol. 368, 95–108 (2012).
Dasgupta, K. & Jeong, J. Developmental biology of the meninges. Genesis 57, e23288 (2019).
DeSisto, J. et al. Single-cell transcriptomic analyses of the developing meninges reveal meningeal fibroblast diversity and function. Dev. Cell 54, 43–59.e4 (2020).
Hammond, T. R. et al. Single-cell RNA sequencing of microglia throughout the mouse lifespan and in the injured brain reveals complex cell-state changes. Immunity 50, 253–271.e6 (2019).
Lein, E. S. et al. Genome-wide atlas of gene expression in the adult mouse brain. Nature 445, 168–176 (2007).
Murdoch, J. N., Eddleston, J., Leblond-Bourget, N., Stanier, P. & Copp, A. J. Sequence and expression analysis of Nhlh1: a basic helix-loop-helix gene implicated in neurogenesis. Dev. Genet. 24, 165–177 (1999).
Gyllborg, D. et al. Hybridization-based in situ sequencing (HybISS) for spatially resolved transcriptomics in human and mouse brain tissue. Nucleic Acids Res. 48, e112 (2020).
Biancalani, T. et al. Deep learning and alignment of spatially-resolved whole transcriptomes of single cells in the mouse brain with Tangram. Preprint at https://doi.org/10.1101/2020.08.29.272831 (2020).
Kinameri, E. et al. Prdm proto-oncogene transcription factor family expression and interaction with the Notch-Hes pathway in mouse neurogenesis. PLoS ONE 3, e3859 (2008).
Sakamoto, T. & Ishibashi, T. Hyalocytes: essential cells of the vitreous cavity in vitreoretinal pathophysiology? Retina 31, 222–228 (2011).
La Manno, G. et al. RNA velocity of single cells. Nature 560, 494–498 (2018).
Gyllborg, D. & Nilsson, M. HybISS: hybridization-based in situ sequencing. protocols.io., https://doi.org/10.17504/protocols.io.xy4fpyw (2020).
Gopalan, P., Hofman, J. M. & Blei, D. M. Scalable recommendation with Poisson factorization. Preprint at https://arxiv.org/abs/1311.1704 (2013).
Kobak, D. & Berens, P. The art of using t-SNE for single-cell transcriptomics. Nat. Commun. 10, 5416 (2019).
Chalfoun, J. et al. MIST: accurate and scalable microscopy image stitching tool with stage modeling and error minimization. Sci. Rep. 7, 4988 (2017).
Acknowledgements
We thank H. Hochgerner and A. Zeisel for advice; E. Arenas for assistance with anatomical annotation; P. Ernfors for supporting the work of A.F.; and the National Genomics Infrastructure for sequencing services. This work was supported by grants from the Knut and Alice Wallenberg Foundation (2015.0041 and 2018.0220 to S.L., and 2018.0172 to S.L. and M.N.), the Erling Persson Family Foundation (HDCA, to S.L. and M.N.), the Chan Zuckerberg Initiative, Torsten Söderberg Foundation and the Swedish Foundation for Strategic Research (RIF 15-0057 and SB16-0065) to S.L., Hjärnfonden (PS2018-0012) to D.G., the Chan Zuckerberg Initiative DAF, an advised fund of Silicon Valley Community Foundation (2018-191929) to M.N., and the Swiss National Science Foundation (CRSK-3_190495 and PZ00P3_193445) and the School of Life Sciences – EPFL to G.L.M. K.S. was supported by the European Molecular Biology Organization (ALTF 1015-2018).
Author information
Authors and Affiliations
Contributions
S.L. supervised the project. S.L. and G.L.M. conceived the study design. G.L.M., K.S. and S.L. analysed, annotated and interpreted the single-cell data and wrote the manuscript. G.L.M. and A.F. performed the single-cell experiments. A.F. dissociated embryos. A.J. prepared sequencing libraries. P.L. and S.L. built the companion website. E.V., D.G. and C.M.L. performed in situ sequencing experiments. A.M.A. performed and analysed smFISH experiments. E.V. prepared tissue sections and performed anatomical annotation. K.S., I.K., A.R.L., G.L.M. and C.M.L. analysed and interpreted in situ sequencing data. L.M.D. performed clustering sensitivity analysis. M.N. supervised in situ sequencing experiments. All authors critically reviewed the manuscript and approved the final version.
Corresponding authors
Ethics declarations
Competing interests
M.N. serves as a scientific advisor for 10x Genomics. All other authors declare no competing financial interests.
Additional information
Peer review information Nature thanks Jennie Close, Mario Suva and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Experimental design and data quality.
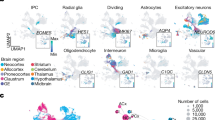
a, Stereo microscope photographs illustrating the tissue dissection strategy. b, Samples composing the dataset. The area of each circle is proportional to the number of cells sampled. Colour indicates the region dissected. c, Histogram showing the distribution of number of UMIs per cell detected across the entire dataset. d, Histogram showing the distribution of number of genes detected per cell across the entire dataset. e, Distribution of UMIs per cell aggregated per age group. f, Distribution of number of genes detected per cell aggregated per age group.
Extended Data Fig. 2 Global properties of gene expression during brain development.
a, Erythrocyte clusters expressing Hbb-bh1 (>500 UMIs per cell) (primitive erythropoiesis) or Hbb-bs (>5,000 UMIs per cell) (definitive erythropoiesis). b, Histogram of cell cycle scores (left) and cell cycle scores indicated on the main t-SNE (right). c, Distribution of total UMIs per cell. d, Distribution of total number of genes detected per cell. e, Distribution of the fraction of unspliced reads detected per cell. f, Average z score for a set of genes with GO term related to the spliceosome. g, Number of clusters observed for each major cell class, as a function of varying hyper-parameters of the clustering algorithm (indicated, left). Top row shows the default clustering. Each subsequent row varies one or more hyper-parameters (number of genes selected, number of HPF components and the stop condition), resulting in different numbers of clusters per class. Deviations from the default settings are indicated by the colours. h, Confusion matrix comparing clustering results for neurons using default settings (vertical axis) and clustering using only 500 genes (horizontal axis). i, Same as h, but for radial glia. The expression of key marker genes is indicated along the axes to reveal major subclasses. j, Distribution of major classes of cells by gestational age.
Extended Data Fig. 3 Computational methods.
a, Illustration of HPF on a small subset of cells using 32 components, showing component loadings across cells for components 4 and 2, illustrating the modularity of components. b, Loadings of all components for two single cells, illustrating the sparseness of components; each cell is approximated by a small number of non-negative components. c, Scatter plot showing observed expression of all genes in all cells, versus predicted expression based on 96 HPF components. Ninety-three per cent of data points were located inside the 95% confidence intervals of the Poisson distribution, demonstrating the accuracy of the HPF representation. d, HPF analysis of all neuronal cells from the full dataset using 32 components, showing that HPF captures essential features of forebrain neurogenesis. Each subplot shows a t-SNE coloured by component loading, a list of genes with high loading in the component, and a bar chart and histogram showing the regional distributions of cells. Nine selected components are shown, related to forebrain based on the regional distribution of cells with high loadings. e, Example of cells (blue) that fall within a Jensen–Shannon distance of 0.36 of a single cell (black circle), corresponding to 0.5 bits of Shannon entropy. f, The relationship between the Kullback–Leibler divergence and the Jensen–Shannon distance, showing that in this case 0.36 JSD corresponds to 0.5 bits. g, Two virtual cells (Poisson samples drawn from a single HPF vector), showing the distribution of pure Poisson expression noise. h, The effect of switching two HPF components, resulting in a change in expression of many genes and a Jensen–Shannon distance of 0.68 bits. i, The ‘curse’ of dimensionality. As the number of HPF components is increased, the difference in Jensen–Shannon distance between nearest-neighbours and far-away cells on the manifold decreases.
Extended Data Fig. 4 Emergence of cell-type heterogeneity from gastrulation to neurulation.
a, Scheme of tissue sampling between E7 and E8.5. b, t-SNE from Fig. 1 overlaid with cells sampled E7–E8.5 and belonging to dendrogram clades 1–4. c, t-SNE overlaid with expression of lineage-specific marker genes. d, Uniform manifold approximation and projection (UMAP) of the mouse gastrulation single-cell atlas from ref. 17. e, Cells from b projected onto the embedding in d17. f, Pairwise correlation between neurulation-stage clusters of this Article and those of ref. 17. g–i, UMAP embedding of cells of the first (g), second (h) and third (i) cluster clade of the dendrogram. Cells are coloured and labelled by cluster identity. J–l, Heat map showing genes enriched in of the first (j), second (k) and third (l) cluster clade. m, UMAP embedding of subclustered neuroepithelial cells obtained from time points E7–E8.5. Cells are coloured by cluster. n, Heat map displaying expression of genes enriched in each of the neuroepithelial populations. o, Expression of broad neuroepithelial markers overlaid on the UMAP in m. p, Bar plot reporting the number of E7–E8.5 cells that clustered with the indicated neuroblast clusters. Clusters are sorted by number of cells. Bars are coloured as in l. q, Bar plots showing the dissection origins of the cells belonging to the clusters in p. r, UMAP representation of the E8.5 cells belonging to clusters no. 377, no. 366 and no. 338 (top left), and overlaid with expression of enriched transcription factors.
Extended Data Fig. 5 Characterization of molecular identity and signal-receptor repertoire of neural tube organizers.
a, Cell types sorted on the basis of the average Z score of morphogens genes grouped in categories on the basis of gene family and neurulation literature. Hedgehog, Shh and Ptch1; R-spondins, Rspo1, Rspo2, Rspo3 and Rspo4; Wnts, Wnt1, Wnt3a, Wnt5a and Wnt8b; neuregulins, Nrg1, Nrg3 and Fgf7; Fgfs, Fgf8, Fgf15, Fgf17 and Fgf18; Bmps, Bmp6, Bmp7 and Gdf7. Cell types displayed in red were localized and analysed in Fig. 2 and in the panels below. b, Summary of the plots in a showing the maximum average z score achieved. c, Beeswarm plots showing the expression of a selection of transcription factors that were found to be enriched in the different organizer cell populations. d, Beeswarm plots showing the gene expression of enriched receptors and ligands with a distribution in relation to organizers that, to our knowledge, had not previously been described. Genes in c, d were selected for HybISS profiling (Fig. 4).
Extended Data Fig. 6 Glial diversity.
a, Cells from clusters 172 to 320 were pulled from the complete dataset, and a t-SNE embedding was calculated. The dendrogram was cut into 15 metaclusters. Cells are coloured by their original cluster (left) and metacluster cut from the dendrogram (right). b, Cells are coloured by their labels for differential gene-expression testing: brown, glioblast; coral, neurogenic; grey, not included in testing. c, Select genes are annotated on a volcano plot illustrating differential expression between neurogenic and gliogenic radial glia. The dotted line denotes the significance threshold (q = 10−5). The range of the x axis was chosen to capture statistically significant genes. Cell-cycle genes are coloured red. d, For each metacluster, the size of the bar indicates the percentage of cells from each tissue (top), each embryonic stage (middle) or that are cycling (>1% cell-cycle UMIs) (bottom). e, Expression dot plots are shown for select genes. The size of the dot represents the fraction of cells expressing a gene. The colour represents normalized mean expression, in which the mean expression for each gene is normalized by the maximum mean expression of the gene across the metaclusters. f, Cells selected for region classification are coloured on the t-SNE. Grey cells were not selected. g, For a range of parameter values (max_features), a gradient boosting classifier was fit to each training set and tested on the five remaining training sets, coloured as in f. Average precision, recall and F1 scores are plotted for each parameter value and training set. h, F1 scores for each gradient boosting classifier (max_features = 50) are shown for forebrain, midbrain and hindbrain cells. The colour of each line indicates which cells were used to train the classifier (coloured as in g). i, Feature importance for each gradient boosting classifier was used to rank genes, and the intersection of the 100 most important genes for all 6 classifiers is shown on the x axis. Expression dot plots are shown for each of these genes in radial glia, glioblasts and OPCs from the forebrain, midbrain and hindbrain . j, Cells are coloured by log-transformed gene expression for the indicated genes. Grey indicates no expression. k, Cells that are positive for both Egfr and Dll1 are coloured blue.
Extended Data Fig. 7 Fibroblasts.
Cells from clusters 109 to 125 were pulled from the dataset, and a t-SNE embedding was calculated. a, Cells are coloured by their clusters. Putative doublets are grey. b, Cells are coloured by their tissues of origin. c, Cycling cells (>1% cell-cycle UMIs) are coloured blue. d, Expression dot plots are shown for select genes. e, Cells from the indicated embryonic age are coloured blue on each t-SNE. f, Cells are coloured if they express the indicated gene. g, Multiplexed single-molecule RNA fluorescence in situ hybridization for six marker genes (columns) at three different ages (rows). Cartoons on the left indicate the approximate location of each image. Cartoons on the right show an interpretation of the observed gene expression patterns. This experiment was performed once.
Extended Data Fig. 8 Immune cells.
Cells from clusters 157 to 171 were pulled from the dataset, and a t-SNE embedding was calculated. a, Cells coloured by their clusters. Putative doublets are grey. Infiltrating immune includes six clusters, starred in d. ATM, axon tract-associated microglia; MGL, microglia; PVM, perivascular macrophage. b, Cells coloured by their tissues of origin. c, Cycling cells (>1% cell-cycle UMIs) coloured blue. d, Expression dot plots shown for select genes. e, Cells from the indicated gestational age coloured blue on each t-SNE.
Extended Data Fig. 9 Pseudolineage analysis and neurogenesis.
a, Pseudolineage tree algorithm, computing shortest paths to the root cell passing only through neighbours inside the information radius (that is, neighbours within a fixed maximum Jensen–Shannon distance). b, Isolating the pseudolineages terminating in a selected cluster, and projecting to the pseudoage axis. c, Ten selected pseudolineages coloured by major class, on the t-SNE of Fig. 1 with grey scale showing the geodesic distance to the root from every cell. d, The same ten selected pseudolineages coloured by tissue. e–i, Pseudolineages of astrocytes (e), OPCs (f), fibroblasts (g), cortical excitatory neurons (h), forebrain GABAergic neurons (i), midbrain dopaminergic neurons (j) and hindbrain serotonergic neurons (k). Each plot shows a randomly selected subset of pseduolineages terminating in the indicated clusters, as well as expression of selected genes in pseudoage bins along the lineage (calculated for all cells in the lineage). The region and class of each cell is indicated at the bottom. l, Expression of Nhlh1 and Nhlh2. m, In situ hybridization for Nhlh1 and Nhlh2 on sagittal sections of the E115 mouse embryo (Allen Brain Atlas). n, RNA in situ hybridization of mouse embryonic brain at the indicated time points, showing genes relevant to the cortical lineage (image credit: Allen Institute). Each subpanel shows a strip from ventricular zone to pia at four different ages.
Extended Data Fig. 10 Spatial mapping of single-cell clusters.
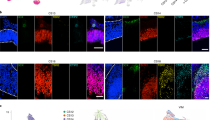
a, Expression of Shh detected by in situ sequencing on all sections, with close up images showing the detection of individual mRNA molecules. Inset shows the expression of Shh by in situ hybridization (Allen Brain Atlas) for reference. b, Manually curated anatomical annotation of all tissue sections (coloured as in the Allen Brain Atlas). c, Illustration of Tangram (mapping single cells to spatial voxels) and our generalized Tangram (mapping clusters to voxels). d, Inferred spatial distribution of roof plate organizers (clusters no. 213, no. 214 and no. 39). e, Inferred spatial distribution of cluster no. 206 on all tissue sections. f, Imputed expression of Prdm12, shown for four informative sections (left) and Allen Brain Atlas ground truth (right). g, Spatial gene expression of Pax6 and imputed gene expression of dorsal (Mitf and Vsx2) and ventral (Vax2 and Pax2) optic vesicle markers in the left and right eye.
Supplementary information
Supplementary Table 1
Sample metadata.
Supplementary Table 2
Cluster annotations and metadata.
Supplementary Table 3
Oligonucleotides used for HybISS.
Rights and permissions
About this article
Cite this article
La Manno, G., Siletti, K., Furlan, A. et al. Molecular architecture of the developing mouse brain. Nature 596, 92–96 (2021). https://doi.org/10.1038/s41586-021-03775-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-021-03775-x
This article is cited by
-
Characterizing neuroinflammation and identifying prenatal diagnostic markers for neural tube defects through integrated multi-omics analysis
Journal of Translational Medicine (2024)
-
Intestinal cell diversity and treatment responses in a parasitic nematode at single cell resolution
BMC Genomics (2024)
-
Cellular development and evolution of the mammalian cerebellum
Nature (2024)
-
Drug targeting in psychiatric disorders — how to overcome the loss in translation?
Nature Reviews Drug Discovery (2024)
-
Gatad2b, associated with the neurodevelopmental syndrome GAND, plays a critical role in neurodevelopment and cortical patterning
Translational Psychiatry (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.