Abstract
Electrical activity in the brain and heart depends on rhythmic generation of action potentials by pacemaker ion channels (HCN) whose activity is regulated by cAMP binding1. Previous work has uncovered evidence for both positive and negative cooperativity in cAMP binding2,3, but such bulk measurements suffer from limited parameter resolution. Efforts to eliminate this ambiguity using single-molecule techniques have been hampered by the inability to directly monitor binding of individual ligand molecules to membrane receptors at physiological concentrations. Here we overcome these challenges using nanophotonic zero-mode waveguides4 to directly resolve binding dynamics of individual ligands to multimeric HCN1 and HCN2 ion channels. We show that cAMP binds independently to all four subunits when the pore is closed, despite a subsequent conformational isomerization to a flip state at each site. The different dynamics in binding and isomerization are likely to underlie physiologically distinct responses of each isoform to cAMP5 and provide direct validation of the ligand-induced flip-state model6,7,8,9. This approach for observing stepwise binding in multimeric proteins at physiologically relevant concentrations can directly probe binding allostery at single-molecule resolution in other intact membrane proteins and receptors.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
All experimental data are available upon reasonable request. Source data are provided with this paper.
Code availability
The DISCO software package is available at https://github.com/ChandaLab/DISC and fully described elsewhere36. All additional MATLAB scripts for single-molecule analysis and image processing are available upon reasonable request.
References
Wahl-Schott, C. & Biel, M. HCN channels: structure, cellular regulation and physiological function. Cell. Mol. Life Sci. 66, 470–494 (2009).
Kusch, J. et al. How subunits cooperate in cAMP-induced activation of homotetrameric HCN2 channels. Nat. Chem. Biol. 8, 162–169 (2011).
Thon, S., Schulz, E., Kusch, J. & Benndorf, K. Conformational flip of nonactivated HCN2 channel subunits evoked by cyclic nucleotides. Biophys. J. 109, 2268–2276 (2015).
Levene, M. J. et al. Zero-mode waveguides for single-molecule analysis at high concentrations. Science 299, 682–686 (2003).
Wainger, B. J., DeGennaro, M., Santoro, B., Siegelbaum, S. A. & Tibbs, G. R. Molecular mechanism of cAMP modulation of HCN pacemaker channels. Nature 411, 805–810 (2001).
Lape, R., Colquhoun, D. & Sivilotti, L. G. On the nature of partial agonism in the nicotinic receptor superfamily. Nature 454, 722–727 (2008).
Jadey, S. & Auerbach, A. An integrated catch-and-hold mechanism activates nicotinic acetylcholine receptors. J. Gen. Physiol. 140, 17–28 (2012).
Abele, R., Keinanen, K. & Madden, D. R. Agonist-induced isomerization in a glutamate receptor ligand-binding domain. A kinetic and mutagenetic analysis. J. Biol. Chem. 275, 21355–21363 (2000).
Cheng, Q., Du, M., Ramanoudjame, G. & Jayaraman, V. Evolution of glutamate interactions during binding to a glutamate receptor. Nat. Chem. Biol. 1, 329–332 (2005).
DiFrancesco, D. The role of the funny current in pacemaker activity. Circ. Res. 106, 434–446 (2010).
DiFrancesco, D. & Tortora, P. Direct activation of cardiac pacemaker channels by intracellular cyclic AMP. Nature 351, 145–147 (1991).
Kusch, J. et al. Interdependence of receptor activation and ligand binding in HCN2 pacemaker channels. Neuron 67, 75–85 (2010).
Chow, S. S., Van Petegem, F. & Accili, E. A. Energetics of cyclic AMP binding to HCN channel C terminus reveal negative cooperativity. J. Biol. Chem. 287, 600–606 (2012).
Hines, K. E., Middendorf, T. R. & Aldrich, R. W. Determination of parameter identifiability in nonlinear biophysical models: a Bayesian approach. J. Gen. Physiol. 143, 401–416 (2014).
Wyman, J. The binding potential, a neglected linkage concept. J. Mol. Biol. 11, 631–644 (1965).
Sigg, D. A linkage analysis toolkit for studying allosteric networks in ion channels. J. Gen. Physiol. 141, 29–60 (2013).
Chowdhury, S. & Chanda, B. Free-energy relationships in ion channels activated by voltage and ligand. J. Gen. Physiol. 141, 11–28 (2013).
Chowdhury, S. & Chanda, B. Estimating the voltage-dependent free energy change of ion channels using the median voltage for activation. J. Gen. Physiol. 139, 3–17 (2012).
Vafabakhsh, R., Levitz, J. & Isacoff, E. Y. Conformational dynamics of a class C G-protein-coupled receptor. Nature 524, 497–501 (2015).
Gregorio, G. G. et al. Single-molecule analysis of ligand efficacy in β2AR-G-protein activation. Nature 547, 68–73 (2017).
Liauw, B. W., Afsari, H. S. & Vafabakhsh, R. Conformational rearrangement during activation of a metabotropic glutamate receptor. Nat. Chem. Biol. 17, 291–297 (2021).
Wang, Y. et al. Single molecule FRET reveals pore size and opening mechanism of a mechano-sensitive ion channel. eLife 3, e01834 (2014).
Dolino, D. M. et al. The structure–energy landscape of NMDA receptor gating. Nat. Chem. Biol. 13, 1232–1238 (2017).
Wang, S., Vafabakhsh, R., Borschel, W. F., Ha, T. & Nichols, C. G. Structural dynamics of potassium-channel gating revealed by single-molecule FRET. Nat. Struct. Mol. Biol. 23, 31–36 (2016).
Jiang, Y. et al. Sensing cooperativity in ATP hydrolysis for single multisubunit enzymes in solution. Proc. Natl Acad. Sci. USA 108, 16962–16967 (2011).
Hoskins, A. A. et al. Ordered and dynamic assembly of single spliceosomes. Science 331, 1289–1295 (2011).
Holzmeister, P., Acuna, G. P., Grohmann, D. & Tinnefeld, P. Breaking the concentration limit of optical single-molecule detection. Chem. Soc. Rev. 43, 1014–1028 (2014).
Wu, J. Y., Stone, M. D. & Zhuang, X. A single-molecule assay for telomerase structure-function analysis. Nucleic Acids Res. 38, e16 (2010).
Morse, J. C. et al. Elongation factor–Tu can repetitively engage aminoacyl–tRNA within the ribosome during the proofreading stage of tRNA selection. Proc. Natl Acad. Sci. USA 117, 3610–3620 (2020).
Zheng, J. & Trudeau, M. C. Handbook of Ion Channels (CRC, 2015).
Zhu, P. & Craighead, H. G. in Annual Review of Biophysics Vol. 41 (ed. D. C. Rees) 269–293 (2012).
Goldschen-Ohm, M. P. et al. Structure and dynamics underlying elementary ligand binding events in human pacemaking channels. eLife 5, e20797 (2016).
Uemura, S. et al. Real-time tRNA transit on single translating ribosomes at codon resolution. Nature 464, 1012–1017 (2010).
Eid, J. et al. Real-time DNA sequencing from single polymerase molecules. Science 323, 133–138 (2009).
Goldschen-Ohm, M. P., White, D. S., Klenchin, V. A., Chanda, B. & Goldsmith, R. H. Observing single-molecule dynamics at millimolar concentrations. Angew. Chem. Int. Ed. 56, 2399–2402 (2017).
White, D. S., Goldschen-Ohm, M. P., Goldsmith, R. H. & Chanda, B. Top-down machine learning approach for high-throughput single-molecule analysis. eLife 9, e53357 (2020).
Wang, J., Chen, S. & Siegelbaum, S. A. Regulation of hyperpolarization-activated HCN channel gating and cAMP modulation due to interactions of COOH terminus and core transmembrane regions. J. Gen. Physiol. 118, 237–250 (2001).
Korlach, J. et al. Selective aluminum passivation for targeted immobilization of single DNA polymerase molecules in zero-mode waveguide nanostructures. Proc. Natl Acad. Sci. USA 105, 1176–1181 (2008).
Ding, S. & Sachs, F. Evidence for non-independent gating of P2X2 receptors expressed in Xenopus oocytes. BMC Neurosci. 3, 17 (2002).
Collauto, A. et al. Rates and equilibrium constants of the ligand-induced conformational transition of an HCN ion channel protein domain determined by DEER spectroscopy. Phys. Chem. Chem. Phys. 19, 15324–15334 (2017).
Zagotta, W. N. et al. Structural basis for modulation and agonist specificity of HCN pacemaker channels. Nature 425, 200–205 (2003).
Qin, F., Auerbach, A. & Sachs, F. A direct optimization approach to hidden Markov modeling for single channel kinetics. Biophys. J. 79, 1915–1927 (2000).
Nicolai, C. & Sachs, F. Solving ion channel kinetics with the QuB software. Biophys. Rev. Lett. 8, 191–211 (2013).
Schwarz, G. Estimating dimension of a model. Ann. Stat. 6, 461–464 (1978).
Lolicato, M. et al. Tetramerization dynamics of C-terminal domain underlies isoform-specific cAMP gating in hyperpolarization-activated cyclic nucleotide-gated channels. J. Biol. Chem. 286, 44811–44820 (2011).
Lee, C. H. & MacKinnon, R. Structures of the human HCN1 hyperpolarization-activated channel. Cell 168, 111–120.e11 (2017).
Goehring, A. et al. Screening and large-scale expression of membrane proteins in mammalian cells for structural studies. Nat. Protoc. 9, 2574–2585 (2014).
Idikuda, V. et al. Singlet oxygen modification abolishes voltage-dependent inactivation of the sea urchin spHCN channel. J. Gen. Physiol. 150, 1273–1286 (2018).
Acknowledgements
This research was supported by the NIH grants to B.C. (NS-116850, NS-101723, and NS-081293), D.S.W. (T32 fellowship GM007507) and NSF to R.H.G. (CHE-1856518). We thank K. A. Knapper and C. H. Vollbrecht for assistance with cover glass cleaning at the Wisconsin Centers for Nanoscale Technologies; M. P. Goldschen-Ohm and M. Smith for helpful discussions; C. Lingle and L. Anson for their feedback on the manuscript; and M. Jackson for sparking this collaboration. ZMWs were fabricated at the Center for Nanophase Materials Sciences at Oak Ridge National Laboratory, which is a DOE Office of Science User Facility.
Author information
Authors and Affiliations
Contributions
D.S.W., S.C., R.H.G. and B.C. conceived and designed the studies; V.I. performed electrophysiology experiments; S.C. performed molecular biology and protein purification; D.S.W. and R.Z. performed single-molecule experiments; D.S.W. analysed single-molecule data. D.S.W. fabricated ZMWs under supervision of S.T.R.; D.S.W., S.C., R.H.G. and B.C. wrote the manuscript with input from others.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature thanks the anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Characterization of HCN1SM and HCN2SM.
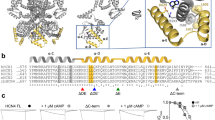
a, Representative electrophysiological recordings (top) of HCN1SM (left) and HCN2SM (right) with voltage protocol. Tail currents (arrow) were collected at −130 mV and were used to generate the activation curves. b, Normalized activation curves of HCN1SM (left) and HCN2SM (right) in the absence or presence of saturating concentrations (500 μM) of internal cAMP with a Boltzmann fit (red). Data points are mean ± s.e.m. (n = 5 patches). V1/2 values for are HCN1SM -71.2 ± 0.4 mV without cAMP and −69.1 ± 0.5 mV with cAMP. V1/2 values for are HCN2SM -105.2 ± 0.6 mV without cAMP and −84.6 ± 0.5 mV with cAMP). c, Size exclusion chromatography (SEC) profiles of HCN1SM (grey) and HCN2SM (orange dashed). Triangles indicate the peak fraction (0.3 ml) used for single-molecule measurements. d, Example fluorescence vs time trajectory of photobleaching eGFP-tagged HCN2SM tetramers via TIRFM. e, Distributions of photobleaching steps overlaid with a maximum likelihood estimate of a zero-truncated binomial distribution (red) for a tetrameric complex with a probability (P) of observing eGFP (HCN1: P = 0.65, 95% CI [0.63, 0.67], n = 752; HCN1: P = 0.67, 95% CI [0.65, 0.69], n = 588).
Extended Data Fig. 2 Non-specific binding in ZMWs.
a, Bright field image of ZMW array on single-molecule imaging set-up featuring a 512 × 512 pixel EMCCD and a 100× objective. Each white dot (about 1,600 per field of view) is a ZMW. b, c, Test of specific binding of eGFP-tagged HCN2SM to ZMWs (b) and of fcAMP (c) to HCN2SM in ZMWs. For b and c, all images shown are averaged over the first 10 frames (1 s) and background subtracted for visualization. Brightness and contrast were adjusted for clarity. d, Representative and randomly selected fluorescence trajectories of empty (no HCN) and passivated ZMWs with 1,000 nM fcAMP fit with DISC (black). The first 50 frames (grey) were removed from analysis.
Extended Data Fig. 3 fcAMP binding to HCN1SM in ZMWs.
Representative fluorescence trajectories of fcAMP (100 nM to 900 nM) binding to HCN1SM in ZMWs with idealized fits (black) imaged at 100-ms resolution. The first 50 frames (grey) were removed from analysis.
Extended Data Fig. 4 fcAMP binding to HCN2SM in ZMWs.
Representative fluorescence trajectories of fcAMP (100 nM to 1,500 nM) binding to HCN2SM in ZMWs with idealized fits (black) imaged at 100-ms resolution. The first 50 frames (grey) were removed from analysis.
Extended Data Fig. 5 All state occupancy distributions.
a, b, Normalized state occupancy distributions for HCN1SM (a) and HCN2SM (b) across all recorded fcAMP concentrations. Each plot indicates the total number of molecules (n) and data points (that is, frames, m) included in the analysis. P is the success rate of the optimized binomial distribution considering four binding sites. All obtained and expected state occupancies values are in Supplementary Table 2.
Extended Data Fig. 6 Isolated-B1 Events of HCN1SM.
a, Dwell time distributions of isolated-B1 events for HCN1SM at various fcAMP concentrations overlaid with maximum likelihood estimates for monoexponential (blue dashed) and biexponential (red) distributions (Supplementary Table 3). For inset, error bars are the error of a binomial distribution (Methods). b, c, Coordinates of identified single-molecules in the 512 × 512 pixel field of view superimposed across all ZMW arrays. The colour bars denote the average dwell time (b) and fluorescence (c) of the isolated-B1 state for each molecule (n). d, Correlation of fluorescence intensity and dwell times for each isolated-B1 event (m), where r is the Pearson correlation coefficient. Data are binned for visualization.
Extended Data Fig. 7 Isolated-B1 Events of HCN2SM.
a, Dwell time distributions of isolated-B1 events for HCN2SM at various fcAMP concentrations overlaid with maximum likelihood estimates for monoexponential (blue dashed) and biexponential (red) distributions (Supplementary Table 3). For inset, error bars are the error of a binomial distribution (Methods). b, c, Coordinates of identified single-molecules in the 512 × 512 pixel field of view superimposed across all ZMW arrays. The colour bars denote the average dwell time (b) and fluorescence (c) of the isolated-B1 state for each molecule (n). d, Correlation of fluorescence intensity and dwell times for each isolated-B1 event (m), where r is the Pearson correlation coefficient. Data are binned for visualization.
Extended Data Fig. 8 Isolated-B1 events do no exhibit static heterogeneity.
a, e, Average isolated-B1 dwell time of HCN1SM (a) and HCN2SM (e) for each molecule at 100 nM fcAMP. Outliers (diamonds) were identified by three scaled median absolute deviations. Data plotted as mean ± s.d. of exponential distribution. The blue dashed line indicates the average B1 dwell time across all molecules (HCN1SM: n = 176; HCN2SM: n = 77). b, f, Histograms of average isolated-B1 dwell times for each HCN1SM (b) and HCN2SM (f) molecule. c, g, Parameters for a monoexponential fit (τ) to isolated-B1 dwell times of HCN1SM (c) and HCN2SM (g). d, h, Parameters for a biexponential fit (τ1, τ2, A1) to isolated-B1 dwell times of HCN1SM (d) and HCN2SM (h). For c, g, d, h, the ordinate corresponds to the obtained parameter (τ, τ1, τ2, A1) and error bars are 95% confidence intervals. All parameters were obtained using maximum likelihood estimates across all isolated-B1 events in either all data (HCN1SM: n = 8,229; HCN2SM: n = 2,676) or inlier (HCN1SM: n = 7,816; HCN2SM: n = 2,575) groups, as indicated on the abscissa.
Extended Data Fig. 9 HCN1SM dwell time distributions.
Dwell time distributions of all liganded states of HCN1SM across all fcAMP concentrations overlaid with expectations from the optimized rates in Fig. 4b.
Extended Data Fig. 10 HCN2SM dwell time distributions.
Dwell time distributions of all liganded states of HCN2SM across all fcAMP concentrations overlaid with expectations from the optimized rates from in Fig. 4b.
Supplementary information
Suppkementary Information
This file contains Supplementary Note 1, Supplementary Methods, Supplementary Tables 1-7, and Supplementary Figures 1-2.
Source data
Rights and permissions
About this article
Cite this article
White, D.S., Chowdhury, S., Idikuda, V. et al. cAMP binding to closed pacemaker ion channels is non-cooperative. Nature 595, 606–610 (2021). https://doi.org/10.1038/s41586-021-03686-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-021-03686-x
This article is cited by
-
Plant adenylate cyclases have come full circle
Nature Plants (2023)
-
Discrimination between cyclic nucleotides in a cyclic nucleotide-gated ion channel
Nature Structural & Molecular Biology (2023)
-
Enlightening activation gating in P2X receptors
Purinergic Signalling (2022)
-
Single-molecule imaging with cell-derived nanovesicles reveals early binding dynamics at a cyclic nucleotide-gated ion channel
Nature Communications (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.