Abstract
COVID-19 manifests with a wide spectrum of clinical phenotypes that are characterized by exaggerated and misdirected host immune responses1,2,3,4,5,6. Although pathological innate immune activation is well-documented in severe disease1, the effect of autoantibodies on disease progression is less well-defined. Here we use a high-throughput autoantibody discovery technique known as rapid extracellular antigen profiling7 to screen a cohort of 194 individuals infected with SARS-CoV-2, comprising 172 patients with COVID-19 and 22 healthcare workers with mild disease or asymptomatic infection, for autoantibodies against 2,770 extracellular and secreted proteins (members of the exoproteome). We found that patients with COVID-19 exhibit marked increases in autoantibody reactivities as compared to uninfected individuals, and show a high prevalence of autoantibodies against immunomodulatory proteins (including cytokines, chemokines, complement components and cell-surface proteins). We established that these autoantibodies perturb immune function and impair virological control by inhibiting immunoreceptor signalling and by altering peripheral immune cell composition, and found that mouse surrogates of these autoantibodies increase disease severity in a mouse model of SARS-CoV-2 infection. Our analysis of autoantibodies against tissue-associated antigens revealed associations with specific clinical characteristics. Our findings suggest a pathological role for exoproteome-directed autoantibodies in COVID-19, with diverse effects on immune functionality and associations with clinical outcomes.
Similar content being viewed by others
Main
Humoral immunity has dichotomous roles in COVID-19. Although neutralizing antibodies protect against SARS-CoV-2 infection8,9, growing evidence suggests that dysregulated humoral immunity also contributes to the characteristic immunopathology of COVID-1910,11,12,13,14,15,16. Recent reports have identified isolated autoantibody reactivities in patients with COVID-19, including reactivities that are characteristic of systemic autoimmune diseases11,12,13,14,15. Importantly, some autoantibodies—particularly neutralizing autoantibodies against type I interferons (IFNs)—appear to directly contribute to COVID-19 pathophysiology by antagonizing innate antiviral responses11,12. Although notable examples of disease-modifying autoantibody responses have previously been described, the full breadth of autoantibody reactivities in COVID-19 remains undetermined, as does their immunological and clinical effects.
A particularly important class of autoantibodies are those that target the exoproteome. Exoproteome-targeting autoantibodies can exert a wide range of functional effects, such as the perturbation of cell signalling (as with the case of autoantibodies against type I IFNs11,12) and targeted killing of specific cell populations via Fc receptors (FcR) and/or complement. We thus sought to identify functional autoantibody responses in patients with COVID-19 by screening for autoantibody reactivities against the human exoproteome.
Widespread autoantibody increases in COVID-19
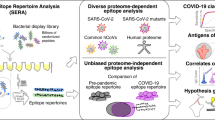
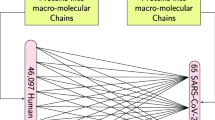
To discover functional autoantibodies that could influence the outcome of COVID-19, we used a high-throughput autoantibody discovery method known as rapid extracellular antigen profiling (REAP)7. REAP enables the highly multiplexed detection of antibody reactivities through the biopanning of IgG from patients against a genetically barcoded library of 2,770 human extracellular proteins displayed on the surface of yeast, converting an antibody–antigen binding event into a quantitative sequencing readout (the REAP score) on the basis of the enrichment of the barcodes of each protein (Extended Data Fig. 1a). To allow for the detection of antibodies against coronavirus proteins, we additionally included the receptor-binding domain (RBD) of SARS-CoV-2 and other common coronaviruses in the library (a full antigen list is provided in Supplementary Table 1).
We used REAP to screen samples from patients infected with SARS-CoV-2 who were prospectively followed as part of the ‘Yale Implementing Medical and Public Health Action Against Coronavirus CT’ (IMPACT) study (Extended Data Fig. 1b). This cohort includes 172 patients who were seen at Yale–New Haven Hospital with a range of clinical severities (as previously reported1) and 22 healthcare workers who had mild illness or asymptomatic infection. We screened longitudinal samples for a subset of the cohort. As a control, we screened 30 healthcare workers who tested negative for SARS-CoV-2 by qPCR with reverse transcription (RT–qPCR) throughout their follow-up period in the IMPACT study. Patient demographics can be found in Extended Data Table 1. To validate the performance of REAP, we compared antibody reactivity to SARS-CoV-2 RBD using REAP to that evaluated using enzyme-linked immunosorbent assays (ELISAs) (Extended Data Fig. 1c), and also compared IL-6R REAP reactivity in patients who received anti-IL-6R therapeutic antibodies with those who did not (Extended Data Fig. 1d). We found a strong concordance between REAP scores for SARS-CoV-2 RBD and ELISA positivity for SARS-CoV-2 RBD, as well as between IL-6R REAP scores and anti-IL-6R treatment.
Next, we examined the total degree of autoreactivity in patients by quantifying the number of autoantibodies at different REAP score thresholds. Irrespective of the REAP score cut off we used, patients with COVID-19 had a greater number of reactivities compared to uninfected individuals, and the highest scoring reactivities were preferentially enriched in patients with severe disease (Fig. 1a, b, Extended Data Fig. 2a). There was not a statistically significant difference in days from symptom onset between patients with severe or moderate COVID-19 (Extended Data Fig. 2b), which suggests that the effects of temporal confounding were minimal. As sex differences in the immune response to SARS-CoV-2 have previously been reported17, we compared the number of autoantibody reactivities between men and women with COVID-19 and found no significant differences in reactivity numbers at any score cut off (Extended Data Fig. 2c). Finally, in a comparison with the REAP profiles of patients with systemic lupus erythematosus (SLE) or autoimmune polyendocrinopathy–candidiasis–ectodermal dystrophy (APECED), patients with COVID-19 had greater numbers of reactivities than did individuals with SLE but fewer numbers of reactivities than did individuals with APECED (Extended Data Fig. 2d). The demographics and clinical characteristics of patients with SLE or APECED have previously been described7. Altogether, these results indicate that autoantibodies that target the exoproteome are increased in COVID-19.
a, Heat map of REAP scores for immune-related proteins and RBDs of the indicated coronaviruses, across all patient samples and stratified by disease severity. CSF, cerebrospinal fluid; gran., granulocyte; mono./lympho., monocytes and lymphocytes; NK, natural killer. b, c, Number of positive (REAP score ≥ 6) total reactivities (b) and immune-targeting reactivities (c) in samples from patients with severe disease (n = 66), moderate disease (n = 160) or mild or asymptomatic disease (n = 36), and uninfected individuals (negative) (n = 54). Dashed lines indicate medians; dotted lines indicate first and third quartiles. NS, not significant. d, Score plot of principal component (PC) analysis performed on immune-targeting reactivities in samples from patients who were hospitalized with COVID-19 (n = 226), coloured by clinical score (CS). e, Loadings for the first (left) and second (right) principal components from d in descending order. In b–d, n values include longitudinal samples from the same patient. Significance in b, c was determined using a linear mixed model with correction for multiple comparisons (Methods).
To investigate the temporal nature of these reactivities relative to the progression of COVID-19, we assessed longitudinal REAP score dynamics. Although definitive assignment was not possible owing to lack of pre-infection samples, we inferred reactivities as ‘likely pre-existing’, ‘newly acquired’ or ‘waning’ on the basis of REAP score trajectories plotted against days from symptom onset and the development of anti-spike S1 IgG. We found that some reactivities were present with high REAP scores within 10 days of symptom onset and before the development of anti-spike S1 IgG responses, which suggests that they were probably pre-existing (Extended Data Fig. 3a, d). Others increased in score and tracked with increasing anti-spike S1 IgG levels, which indicates that they were newly acquired after infection (Extended Data Fig. 3b, e). Finally, some reactivities decreased in REAP score over time while anti-spike S1 IgG increased or remained high, which suggests waning autoantibody titres (Extended Data Fig. 3c, f).
To further explore the potential cellular sources of the increased autoantibody reactivities in patients with COVID-19, we examined B cell phenotypes in peripheral blood mononuclear cells that matched the REAP plasma samples. Similar to previous reports10, we found that extrafollicular double-negative B cells are expanded in patients with moderate or severe COVID-19, as compared to uninfected individuals (Extended Data Fig. 2f).
Autoantibodies target immune-related proteins
Our analysis of the specific reactivities detected by REAP indicated that autoantibodies targeting immune-related proteins were increased in patients with severe COVID-19 (Fig. 1a, c, Extended Data Fig. 2e). These proteins included those involved in lymphocyte function and activation, leukocyte trafficking, the type I and type III IFN responses, type II immunity and the acute phase response. Confirming a previous report11, we identified autoantibodies against type I IFNs in 5.2% of patients who were hospitalized with COVID-19. Using an ELISA, we orthogonally validated a subset of 22 autoantibodies that target cytokines, chemokines, growth factors, complement factors and cell-surface proteins (Extended Data Fig. 4a–d). These results demonstrate that patients with COVID-19 possess autoantibodies that may affect a wide range of immunological functions.
To uncover the low-dimensional features in immune-targeting reactivities, we performed a principal component analysis (Fig. 1d, e, Extended Data Fig. 4e). We found that the first principal component was primarily composed of autoantibody reactivities against type I IFNs (Fig. 1e) and separated a distinct cluster of samples from individuals with a severe disease phenotype (Fig. 1d), consistent with a previous report11. The second principal component comprised autoantibody reactivities against cytokines, chemokines and the type III IFNs IFNλ2 and IFNλ3 (Fig. 1e). Notably, the severity of COVID-19 disease was a significant predictor of second principal component score (Extended Data Fig. 4f, g). These findings suggest that autoantibodies against cytokines and chemokines may contribute to disease severity in COVID-19.
Virological and immunological autoantibody effects
Because autoantibodies in patients may influence circulating concentrations of their target proteins, we examined the plasma concentrations of cytokines and chemokines in patients with autoantibodies against these proteins. In some cases, autoantibodies were associated with apparent increases in their autoantigen targets (Extended Data Fig. 5b, f, j, m), whereas in other cases they correlated with apparent decreases (Extended Data Fig. 5k, l).
To more directly assess potential immunomodulatory effects of cytokine- and chemokine-targeting autoantibodies in patients with COVID-19, we assessed the in vitro activity of selected autoantibodies. We found that IgG from patients with anti-GM-CSF, anti-CXCL1 or anti-CXCL7 autoantibodies could antagonize the signalling of GM-CSF, CXCL1 and CXCL7, respectively (Fig. 2a, b). In addition, we found that plasma or IgG from patients with anti-CD38 or anti-CD3ε autoantibodies led to increased antibody-dependent cellular phagocytosis by macrophages of Raji B cells or Jurkat T cells, respectively (Fig. 2c, Extended Data Fig. 4h, i). These results demonstrate that immune-targeting autoantibodies in patients with COVID-19 can directly inhibit the activity of cytokines and chemokines, and engage FcR effector functions that could lead to immune-cell depletions in affected patients.
a, GM-CSF signalling assay performed with IgG from a patient with COVID-19 who was positive for anti-GM-CSF autoantibody (pink circles) and two uninfected healthcare workers (grey squares). Results are averages of technical duplicates from one experiment. b, CXCL1 (left) and CXCL7 (right) signalling assays performed with IgG from patients with COVID-19 who were positive for anti-CXCL1 or anti-CXCL7 autoantibody (AAb) and from healthcare workers (HCWs). Results are averages of three technical replicates or duplicates. RLU, relative luminescence unit. c, Macrophage phagocytosis assay performed with Raji (left) or Jurkat (right) cells, using plasma or IgG from patients with COVID-19 who were positive for anti-CD38 or anti-CD3ε autoantibody, respectively, anti-CD38 or anti-CD3ε monoclonal antibodies (positive controls), and plasma or IgG from healthcare workers, respectively (n = 1 for all groups). Technical replicates are shown. Results in b, c are representative of two independent experiments. d, Longitudinal comparisons of SARS-CoV-2 viral load between patients with (positive) and without (negative) autoantibodies against type I IFNs. Linear regressions (solid lines) and 95% confidence bands (shaded areas) for each group are displayed. d, day. n values include longitudinal samples from the same patient. e–g, Average per cent B cells (e), relative proportions of classical, intermediate and nonclassical monocytes (f) and average per cent CD4+ T cells (g) among peripheral leukocytes in healthcare workers and patients with COVID-19 stratified by disease severity and positive REAP reactivity (autoantibody-positive) (REAP score ≥ 2) against B-cell-displayed proteins (CD38, FcμR and FCRL3) (e), proteins preferentially displayed on classical and intermediate monocytes (CCR2, CCRL2, FFAR4, SYND4 and CPAMD8) (f) and CD3ε (g), respectively. Gating strategies for e–g are shown in Extended Data Fig. 8. Data from e, g are presented as box plots with the first quartile, median, third quartile, whiskers (minimum and maximum values within the first or third quartiles ± 1.5× the interquartile range), and individual data points indicated. Significance was determined using a generalized linear mixed model (d) (Methods) or a two-sided Wilcoxon rank-sum test (e). In b, c, e–g, n values indicate samples from unique patients. All error bars represent s.d.
To investigate the potential virological effects of cytokine- and chemokine-targeting autoantibodies, we examined a subset of patients with COVID-19 with autoantibodies against type I IFNs. Consistent with previous reports11, we found that these autoantibodies can neutralize the signalling activity of type I IFNs in vitro (Extended Data Fig. 4j, k). To further assess the functional effects of these autoantibodies, we compared longitudinal composite viral loads in patients who had autoantibodies against type I IFNs to those who did not. After controlling for the contributions of age, sex, days from symptom onset and anti-SARS-CoV-2 IgG responses, patients with autoantibodies against type I IFNs had significantly increased average viral loads relative to patients who lacked these autoantibodies (Fig. 2d, Extended Data Fig. 4l). We also found that patients with autoantibodies against type I IFNs had extended durations of hospital admission relative to those who did not (Extended Data Fig. 4m). These results indicate that autoantibodies against type I IFNs impair virological clearance in patients with COVID-19.
To investigate the in vivo effects of autoantibodies against immune-cell surface proteins in COVID-19, we looked for associations between these autoantibodies and blood leukocyte composition. First, we focused on two groups of antigens: those expressed on B cells (CD38, FcμR and FcRL3) and those preferentially expressed on classical and intermediate monocytes (CCR2, CCRL2, FFAR4, SYND4 and CPAMD8, which we identified from a public RNA-sequencing dataset18). We found that patients with autoantibodies against B cell or classical- and intermediate-monocyte antigens had lower frequencies of B cells (Fig. 2e) and anti-SARS-CoV-2-RBD IgM (Extended Data Fig. 6a) or classical and intermediate monocytes (Fig. 2f, Extended Data Fig. 6b), respectively, as compared to patients who were matched for disease severity but negative for these autoantibodies. Looking at individual reactivities, we found that a patient with anti-CD3ε autoantibodies had intact B and natural killer cell compartments but markedly reduced levels of CD4+, CD8+ and natural killer T cells (Fig. 2g, Extended Data Fig. 6c, d). Similarly, a patient with anti-CD38 autoantibodies exhibited a lower frequency of natural killer cells and activated CD4+ or CD8+ T cells, all of which also express CD38 (Extended Data Fig. 6f). Of note, we found that IgG or plasma from these patients could mediate antibody-dependent cellular phagocytosis against Jurkat or Raji cells, respectively, in vitro (Fig. 2c). In aggregate, these data show that autoantibodies that target immune-cell surface proteins may lead to the depletion of specific immune-cell populations in patients with COVID-19.
Autoantibodies increase disease severity in mice
To directly assess the effect of cytokine-targeting autoantibodies in the pathogenesis of COVID-19 in vivo, we used mice that transgenically express human ACE2 under the human keratin 18 promoter (K18-hACE2 mice) to recapitulate aspects of COVID-19 pathogenesis in humans upon SARS-CoV-2 infection19,20,21. Given the enrichment of autoantibodies targeting type I IFNs in patients with severe COVID-19, we first examined the effect of antibody-mediated blockade of type I IFNs in vivo. We found that mice that were pretreated with neutralizing antibodies against IFNAR (the interferon-α/β receptor) were more susceptible to SARS-CoV-2 infection; they had increased weight loss (Fig. 3a) and reduced survival (Fig. 3b). Additionally, compared to mice infected with SARS-CoV-2 that were treated with phosphate-buffered saline (PBS), infected mice treated with anti-IFNAR exhibited impaired monocyte recruitment, maturation and proinflammatory macrophage differentiation in the lungs (Extended Data Fig. 7b–d), as well as marked decreases in the relative frequency and absolute number of activated (CD44+CD69+) natural killer cells and CD4+, CD8+ and γδ T cells (Extended Data Fig. 7e, f). Collectively, these findings demonstrate that the early blockade of type I IFN signalling by antibodies (which mimics the effects of pre-existing autoantibodies targeting type I IFNs) results in exacerbated disease and interferes with myeloid and lymphoid activation in response to SARS-CoV-2 infection.
a–g, K18-hACE2 mice were intranasally infected with a sublethal dose (b, c) or median lethal dose (d–g) of SARS-CoV-2 (USA-WA1/2020 isolate), and treated with indicated antibodies (administered intraperitoneally at the indicated dose per mouse at the time points indicated in a). b, c, Normalized body weight (b) and survival defined as 10% weight loss or mortality (c) of K18-hACE2 mice treated with PBS or anti-IFNAR from day 1 to 14 after infection. d–g, Survival defined as 20% weight loss or mortality of K18-hACE2 mice treated with anti-IL-18 (d), anti-IL-1β (e), anti-IL-21R (f), anti-GM-CSF (g) or PBS-treated from day 1 to 14 after infection. Significance in c–g was determined using log-rank (Mantel–Cox) test. All error bars are s.e.m. All n values indicate biologically independent mice, examined over two independent experiments.
Beyond type I IFNs, we identified patients with COVID-19 with autoantibodies that target components of the interleukin-18 (IL-18) pathway (in particular, IL-18Rβ) (Extended Data Fig. 4d), which has a critical role in antiviral responses of natural killer and CD8+ T cells22,23. To examine the effect of disruptions to the IL-18 pathway in SARS-CoV-2 infection, we administered neutralizing anti-IL-18 antibodies to K18-hACE2 mice immediately before infection. We found that IL-18 blockade greatly enhanced susceptibility of these mice to SARS-CoV-2 infection (Fig. 3d, Extended Data Fig. 7k), resulted in a significantly higher viral burden (Extended Data Fig. 7g, h) and led to a decreased frequency and number of effector natural killer cells with enhanced cytotoxic properties (CD11b+ or KLRG1+) (Extended Data Fig. 7i, j). These results highlight the disruptive role that autoantibody-mediated blockade of IL-18 can have in the immune response to SARS-CoV-2.
Furthermore, we identified IL-1β, IL-21 and GM-CSF as cytokine autoantibody targets in patients with COVID-19. IL-1β and IL-21 both participate directly in host antiviral defence24,25, and IL-21 is a major contributor to the second principal component in our principal component analysis of immune-targeting autoantibodies in COVID-19 (Fig. 1e). GM-CSF has a critical role in augmenting innate antiviral defence mediated by alveolar macrophages26. Consistent with the antiviral properties of these three cytokines, we found that mice that received anti-IL-1β, anti-IL-21R or anti-GM-CSF antibodies became substantially more susceptible to SARS-CoV-2 infection, as they had significantly decreased survival (Fig. 3e–g) and lost more weight (Extended Data Fig. 7l–n) upon SARS-CoV-2 infection that did mice treated with PBS.
Tissue-targeting autoantibody correlations
In addition to immune-targeting autoantibodies, we also observed a high prevalence of tissue-associated autoantibodies in patients with COVID-19 (Fig. 4a). These autoantibodies were directed against vascular cells, coagulation factors and platelets, connective tissue, extracellular matrix components and various organ systems, including lung, the central nervous system compartment, skin, gastrointestinal tract and other tissues. To assess whether any of these putative autoantigens were associated with significant perturbations in clinical phenotype, we performed exploratory data analysis using a common, generalized linear mixed-effects model (Extended Data Fig. 9). We found that some of the autoantigens (such as NXPH1, PCSK1, SLC2A10 and DCD) significantly correlated with clinical markers that are known to be associated with an increased severity of COVID-19 disease (such as d-dimer, ferritin, C-reactive protein and lactate)27,28. Given the extent of autoantigens specific to the central nervous system that we identified in our REAP screen, and previous reports on the potential for SARS-CoV-2 neuroinvasion29, we further examined whether any of the autoantibodies correlated with the Glasgow Coma Scale scores of any of the patients. We found that ten patients with COVID-19 developed autoantibodies against HCRTR2, which is an orexin receptor that is enriched in the hypothalamus. We noted a marked negative correlation between levels of HCRTR2 autoantibodies in these patients and exceptionally low Glasgow coma scale scores, encompassing the time of sample collection (Fig. 4b). Furthermore, we validated the presence of these autoantibodies in two patients using an ELISA (Fig. 4c) and, using an in vitro orexin signalling assay, found that HCRTR2 autoantibodies in one patient antagonized HCRTR2 activity (Fig. 4d).
a, Heat map of tissue-associated REAP scores stratified by disease severity. CNS, central nervous system; coag., coagulation factors; GI, gastrointestinal; pleio., pleiotropic. b, Correlation of HCRTR2 REAP scores with Glasgow Coma Scale (GCS) scores in patients with COVID-19 (n = 89, Spearman’s ρ = −0.20, P = 0.052). Linear regression (solid lines) and 95% confidence bands (shaded areas) are displayed. Patients are coloured by clinical score. Significance of Spearman’s correlation was determined via asymptotic t-approximation with a two-sided test. c, Single-point pan-IgG HCRTR2 autoantibody ELISA conducted with 1:50 plasma dilution. Dotted line represents average of uninfected individuals (healthcare workers) + 3 s.d. Results are averages of technical duplicates from one experiment. OD, optical density. d, Orexin signalling assay performed with IgG from patients with COVID-19 who were positive for anti-HCRTR2 autoantibody and from healthcare workers. Results are averages of three technical replicates from one experiment. All n values indicate samples from unique patients. All error bars represent s.d.
Discussion
The extent of autoantibody reactivities seen in patients with COVID-19 suggests that humoral immunopathology is an intrinsic aspect of the pathogenesis of COVID-19. Screening patient samples with the REAP platform, we have identified and validated numerous protein targets across a wide range of tissues and immunological and physiological functions. These autoantibodies had potent functional activities and could be directly correlated with various virological, immunological and clinical parameters in vivo within samples from patients with COVID-19. Our analysis of REAP score trajectories and comparisons to SARS-CoV-2 humoral responses suggest that some of these autoantibodies probably predated infection, whereas others were induced after infection. Furthermore, mouse surrogates of these autoantibodies led to increased disease severity in a mouse model of SARS-CoV-2 infection. These results provide evidence that autoantibodies are capable of altering the course of COVID-19 by perturbing the immune response to SARS-CoV-2 and tissue homeostasis.
The diversity of autoantibody responses in patients with COVID-19 also underscores the importance of high-throughput and unbiased proteome-scale surveys for autoantibody targets. Beyond validating the biologically compelling example of autoantibodies targeting type I IFNs in COVID-19, our studies implicated numerous other immune pathways that are targeted by autoantibodies in COVID-19. We also detected autoantibodies against various tissue-associated antigens and identified correlations between these autoantibodies and inflammatory clinical markers such as d-dimer, ferritin, C-reactive protein and lactate in patients with COVID-19. Many of the tissue autoantibodies that we identified were also present across diverse physiological compartments that have frequently been implicated in post-COVID-19 syndrome30. For example, we identified autoantibodies against the orexin receptor HCRTR2 that—ex vivo—could inhibit orexin signalling, which has an important role in regulating wakefulness and appetite31. Ultimately, whether the specific autoantibodies identified here have a role in the establishment of post-COVID-19 syndrome, and whether they persist beyond the acute phase of COVID-19, warrants further investigation.
In summary, our analyses have revealed an expansive autoantibody landscape in patients with COVID-19 and identified distinct autoantibodies that exerted notable immunological and clinical outcomes. These results implicate previously underappreciated immunological pathways in the aetiology of COVID-19 and suggest therapeutic paradigms that are centred around modulating these pathways—as well as attenuating the autoantibodies themselves. Finally, our findings provide a strong rationale for the wider investigation of autoantibodies in the pathogenesis of infectious diseases.
Methods
No statistical methods were used to predetermine sample size. For research performed on human samples, randomization was not performed as these studies were observational in nature. For experiments involving mice, age- and sex-matched mice were randomly assigned to experimental groups. For the REAP screen, human peripheral blood mononuclear cell flow cytometry measurements and plasma cytokine measurements, investigators were blinded to patients’ clinical information and clinical scores until after data collection and analysis. For all other experiments, investigators were not blinded to patients’ clinical information and clinical scores during data collection and analysis.
Ethics statement
This study was approved by Yale Human Research Protection Program Institutional Review Boards (FWA00002571, protocol ID 2000027690). Informed consent was obtained from all enrolled patients and healthcare workers.
Patients
As previously described1 and reproduced here for accessibility, 197 patients admitted to Yale New Haven Hospital with COVID-19 between 18 March and 5 May 2020 were included in this study. No statistical methods were used to predetermine sample size. Nasopharyngeal and saliva samples were collected as previously described32, approximately every four days, for SARS-CoV-2 reverse-transcription qPCR analysis when clinically feasible. Paired whole blood for flow cytometry analysis was collected simultaneously in sodium-heparin-coated vacutainers and kept on gentle agitation until processing. All blood was processed on the day of collection. Patients were scored for COVID-19 disease severity through review of electronic medical records at each longitudinal time point. Scores were assigned by a clinical infectious disease physician according to a custom-developed disease severity scale. Moderate disease status (clinical score 1–3) was defined as: SARS-CoV-2 infection requiring hospitalization without supplementary oxygen (1); infection requiring noninvasive supplementary oxygen (<3 l min−1 to maintain SpO2 > 92%) (2); and infection requiring noninvasive supplementary oxygen (>3 l min−1 to maintain SpO2 > 92%, or >2 l min−1 to maintain SpO2 > 92% and had a high-sensitivity C-reactive protein (CRP) >70) and received tocilizumab) (3). Severe disease status (clinical score 4 or 5) was defined as infection meeting all criteria for clinical score 3, and also requiring admission to the intensive care unit and >6 l min−1 supplementary oxygen to maintain SpO2 > 92% (4); or infection requiring invasive mechanical ventilation or extracorporeal membrane oxygenation in addition to glucocorticoid or vasopressor administration (5). Clinical score 6 was assigned for deceased patients. For all patients, days from symptom onset were estimated as follows: (1) highest priority was given to explicit onset dates provided by patients; (2) next highest priority was given to the earliest reported symptom by a patient; and (3) in the absence of direct information regarding symptom onset, we estimated a date through manual assessment of the electronic medical records by an independent clinician. Demographic information was aggregated through a systematic and retrospective review of patient electronic medical records and was used to construct Extended Data Table 1. The clinical data were collected using EPIC EHR and REDCap 9.3.6 software. At the time of sample acquisition and processing, investigators were unaware of the conditions of the patients. Blood acquisition was performed and recorded by a separate team. Information about the conditions of the patients was not available until after processing and analysis of raw data by flow cytometry and ELISA. A clinical team, separate from the experimental team, performed chart reviews to determine relevant statistics. Cytokines and fluorescence-activated cell sorting analyses were performed blinded. Patients’ clinical information and clinical score coding were revealed only after data collection.
Clinical data acquisition
Clinical data for patients and healthcare workers were extracted from the Yale–New Haven Health computational health platform33,34 in the Observational Medical Outcomes Partnership data model. For each research specimen, summary statistics including minimum, mean, median and maximum values were obtained for relevant clinical measurements, including the Glasgow coma scale, within ±1 day of biospecimen collection. Disease severity end points, including admission, supplementary oxygen use and invasive ventilation, were validated as previously described35.
Viral RNA measurements from human nasopharyngeal samples
RNA concentrations were measured from human nasopharyngeal samples by RT–qPCR as previously described32. In brief, total nucleic acid was extracted from 300 μl of viral transport medium (nasopharyngeal swab) using the MagMAX Viral/Pathogen Nucleic Acid Isolation kit (ThermoFisher) and eluted into 75 μl elution buffer. For SARS-CoV-2 RNA detection, 5 μl of RNA template was tested as previously described36, using the US Centers for Disease Control and Prevention real-time RT–PCR primer/probe sets 2019-nCoV_N1 and 2019-nCoV_N2, as well as the human RNase P as an extraction control. Virus RNA copies were quantified using a tenfold dilution standard curve of RNA transcripts that was previously generated. If the RNA concentration was lower than the limit of detection (ND) that was previously determined, the value was set to 0 and used for the analyses.
Yeast induction
All yeast were induced as previously described7. In brief, one day before induction, yeast were expanded in synthetic dextrose medium lacking uracil (SDO −Ura) at 30 °C. The following day, yeast were induced by resuspension at an optical density of 1 in synthetic galactose medium lacking uracil (SGO −Ura) supplemented with 10% SDO −Ura and culturing at 30 °C for approximately 18 h.
REAP
IgG antibody isolation for REAP was performed as previously described7. In brief, Triton X-100 and RNase A were added to plasma samples at final concentrations of 0.5% and 0.5 mg ml−1, respectively, and incubated at room temperature for 30 min before use to reduce risk from any potential virus in plasma. Twenty μl protein G magnetic resin (Lytic Solutions) was washed with sterile PBS, resuspended in 75 μl sterile PBS, and added to 25 μl plasma. Plasma–resin mixture was incubated overnight at 4 °C with shaking. Resin was washed with sterile PBS, resuspended in 90 μl 100 mM glycine pH 2.7, and incubated for five min at room temperature. Supernatant was extracted and added to 10 μl sterile 1M Tris pH 8.0. At this point, IgG concentration was measured using a NanoDrop 8000 Spectrophotometer (Thermo Fisher Scientific). To generate yeast-depleted IgG for use in REAP, purified IgG was added to 108 induced empty vector (pDD003) yeast and incubated for 3 h at 4 °C with shaking. Yeast–IgG mixtures were placed into 96-well 0.45-μm filter plates (Thomas Scientific) and yeast-depleted IgG was eluted into sterile 96-well plates by centrifugation at 3,000g for 3 min.
Yeast library selection for REAP was performed as previously described7. In brief, 400 μl of the induced yeast library was set aside to allow for comparison to post-selection libraries. Then, 108 induced yeast were added to wells of a sterile 96-well v-bottom microtitre plate, resuspended in 100 μl PBE (PBS with 0.5% BSA and 0.5 mM EDTA) containing 10 μg patient-derived antibody, and incubated with shaking for 1 h at 4 °C. Yeast were washed twice with PBE, resuspended in 100 μl PBE with a 1:100 dilution of biotin anti-human IgG Fc antibody (clone HP6017, BioLegend), and incubated with shaking for 1 h at 4 °C. Yeast were washed twice with PBE, resuspended in 100 μl PBE with a 1:20 dilution of Streptavidin MicroBeads (Miltenyi Biotec), and incubated with shaking for 30 min at 4 °C. All following steps were carried out at room temperature. Multi-96 Columns (Miltenyi Biotec) were placed into a MultiMACS M96 Separator (Miltenyi Biotec) in positive selection mode and the columns were equilibrated with 70% ethanol and degassed PBE. Yeast were resuspended in 200 μl degassed PBE and placed into the columns. The columns were washed three times with degassed PBE. To elute the selected yeast, columns were removed from the separator and placed over 96-well deep well plates. Then, 700 μl degassed PBE was added to each well of the column and the column and deep-well plate were centrifuged briefly. This process was repeated three times. Yeast were recovered in 1 ml SDO −Ura at 30 °C.
DNA was extracted from yeast libraries using Zymoprep-96 Yeast Plasmid Miniprep kits or Zymoprep Yeast Plasmid Miniprep II kits (Zymo Research) according to standard manufacturer protocols. A first round of PCR was used to amplify a DNA sequence containing the protein display barcode on the yeast plasmid. PCR reactions were conducted using 1 μl plasmid DNA, 159_DIF2 and 159_DIR2 primers, and the following PCR settings: 98 °C denaturation, 58 °C annealing, 72 °C extension, 25 rounds of amplification. A second round of PCR was conducted using 1 μl first round PCR product, Nextera i5 and i7 dual-index library primers (Illumina) along with dual-index primers containing custom indices, and the following PCR settings: 98 °C denaturation, 58 °C annealing, 72 °C extension, 25 rounds of amplification. PCR products were pooled and run on a 1% agarose gel. The band corresponding to 257 base pairs was cut out and DNA (NGS library) was extracted using a QIAquick Gel Extraction Kit (Qiagen) according to standard manufacturer protocols. NGS library was sequenced using an Illumina NextSeq 500 and NextSeq 500/550 75 cycle High Output Kit v2.5 with 75 base pair single-end sequencing according to standard manufacturer protocols. A minimum of 50,000 reads per sample was collected and the preselection library was sampled at ten times greater depth than other samples.
REAP data analysis
REAP score was calculated as previously described7. First, barcode counts were extracted from raw NGS data using custom codes. Next, aggregate and clonal enrichment was calculated using edgeR37 and custom codes. For aggregate enrichment, barcode counts across all unique barcodes associated with a given protein were summed, library sizes across samples were normalized using default edgeR parameters, common and tagwise dispersion were estimated using default edgeR parameters, and exact tests comparing each sample to the preselection library were performed using default edgeR parameters. Aggregate enrichment is thus the log2-transformed fold change values from these exact tests with zeroes in the place of negative fold changes. log2-transformed fold change values for clonal enrichment were calculated in an identical manner, but barcode counts across all unique barcodes associated with a given protein were not summed. Clonal enrichment for a given reactivity was defined as the fraction of clones out of total clones that were enriched (log2-transformed fold change ≥ 2). Thus, the clonal enrichment metric progressively penalizes proteins with lower fractions of clones enriched. This metric was implemented because a true reactivity should theoretically enrich all yeast clones displaying a given protein.
Aggregate (Ea) and clonal enrichment (Ec) for a given protein, a scaling factor (βu) on the basis of the number of unique yeast clones (yeast that have a unique DNA barcode) displaying a given protein, and a scaling factor (βf) on the basis of the overall frequency of yeast in the library displaying a given protein were used as inputs to calculate the REAP score, which is defined as REAP score = Ea × (Ec)2 × βu × βf. βu and βf are logarithmic scaling factors that progressively penalize the REAP score of proteins with low numbers of unique barcodes or low frequencies in the library. βu is applied to proteins with ≤5 unique yeast clones in the library and βf is applied to proteins with a frequency of ≤ 0.0001 in the library. βf was implemented to mitigate spurious enrichment signals from low-frequency proteins, which could occur owing to sequencing errors or stochasticity in the selection process. βu was implemented because the clonal enrichment metric is less valid for proteins with low numbers of unique yeast clones, decreasing confidence in the validity of the reactivity. βu is defined as βu = ln(xu + 0.5)/1.705 and βf is defined as βf = ln(xf + 7.1)/1.16, in which xu is the number of unique yeast clones for a given protein and xf is the log10-transformed frequency of a given protein in the library. Antigens (excluding coronavirus RBDs and IL-6R) with an average REAP score greater than 0.5 across all samples were defined as ‘sticky’ and excluded from further analysis. Antigens defined as immune-targeting and tissue-associated were manually identified.
Autoantibody ELISA measurement
In brief, 200 ng of purchased or independently produced recombinant protein in 100 μl of PBS pH 7.0 was added to 96-well flat-bottom Immulon 2HB plates (Thermo Fisher Scientific) and placed at 4 °C overnight. Plates were washed once with 225 μl ELISA wash buffer (PBS + 0.05% Tween 20) and 150 μl ELISA blocking buffer (PBS + 2% human serum albumin) was added to the well. Plates were incubated for 2 h at room temperature. ELISA blocking buffer was removed from the wells and appropriate dilutions of sample plasma in 100 μl ELISA blocking buffer were added to each well. Plates were incubated for 2 h at room temperature. Plates were washed 6 times with 225 μl ELISA wash buffer and 1:5,000 goat anti-human IgG HRP (Millipore Sigma) or anti-human IgG isotype-specific HRP (Southern Biotech; IgG1: clone HP6001, IgG2: clone 31-7-4, IgG3: clone HP6050, IgG4: clone HP6025) in 100 μl ELISA blocking buffer was added to the wells. Plates were incubated for 1 h at room temperature. Plates were washed 6 times with 225 μl ELISA wash buffer. Then, 50 μl TMB substrate (BD Biosciences) was added to the wells and plates were incubated for 20–30 min in the dark at room temperature. Next, 50 μl 1 M sulfuric acid was added to the wells and absorbance at 450 nm was measured in a Synergy HTX Multi-Mode Microplate Reader (BioTek). Proteins used are as follows: ACKR1–mIgG2a-Fc (produced in-house), BAMBI (Sino Biological, 10890-H08H-20), C1qB (Sino Biological, 10941-H08B-20), CCL15 (PeproTech, 300-43), CCL16 (PeproTech, 300-44), CNPY3 (produced in-house), CNPY4 (produced in-house), CST5 (produced in-house), CD38 (R&D Systems, 2404-AC-010), GM-CSF (produced in-house), CXCL1 (PeproTech, 300-11), CXCL3 (PeproTech, 300-40), CXCL7 (PeproTech, 300-14), FcμR (R&D Systems, 9494-MU-050), HCRTR2–mIgG2a-Fc (produced in-house), IFNω (PeproTech, 300-02J), IL-13 (PeproTech, 200-13), IL-1α (RayBiotech, 228-10846-1), IL-6 (produced in-house), leptin (R&D Systems, 398-LP-01M), SLC2A12–mIgG2a-Fc (produced in-house), TSLP (PeproTech, 300-62) and IL-18Rβ (produced in-house).
SARS-CoV-2-specific antibody ELISA measurement
SARS-CoV-2-specific antibodies were measured as previously described38. In brief, plasma samples were first treated with 0.5% Triton X-100 and 0.5 mg ml−1 RNase A at room temperature for 30 min to inactivate potentially infectious viruses. Meanwhile, recombinant SARS-CoV-2 S1 protein (ACRO Biosystems, S1N-C52H3) or recombinant SARS-CoV-2 RBD protein (ACRO Biosystems, SPD-C82E9) was used to coat 96-well MaxiSorp plates (Thermo Scientific) at a concentration of 2 μg ml−1 in PBS in 50 μl per well, followed by overnight incubation at 4 °C. The coating buffer was removed, and plates were incubated for 1 h at room temperature with 200 μl of blocking solution (PBS with 0.1% Tween-20 and 3% milk powder). Plasma was diluted 1:50 in dilution solution (PBS with 0.1% Tween-20 and 1% milk powder) and 100 μl of diluted serum was added for 2 h at room temperature. Plates were washed three times with PBS-T (PBS with 0.1% Tween-20) and 50 μl of HRP anti-human IgG antibody at 1:5,000 dilution (GenScript) or anti-human IgM–peroxidase antibody at 1:5,000 dilution (Sigma-Aldrich) in dilution solution were added to each well. After 1 h of incubation at room temperature, plates were washed six times with PBS-T. Plates were developed with 100 μl of TMB Substrate Reagent Set (BD Biosciences 555214) and the reaction was stopped after 12 minutes by the addition of 100 μl of 2 N sulfuric acid. Plates were then read at a wavelength of 450 nm and 570 nm. The cut-off values for seropositivity were determined as 0.392, 0.436 and 0.341 for anti-S1-IgG, anti-S1 IgM and anti-RBD IgG, respectively. Eighty and sixty-nine pre-pandemic plasma samples were assayed to establish the negative baselines for the S1 and RBD antigens, respectively. These values were statistically determined with a confidence level of 99%.
Functional validation of anti-GM-CSF autoantibody and autoantibodies targeting type I IFNs
TF-1 cells (ATCC, CRL-2003) were cultured in RPMI (+ 10% heat inactivated FBS, 10 U ml−1 penicillin, 100 mg ml−1 streptomycin, 1 mM sodium pyruvate, 2 ng ml−1 GM-CSF (PeproTech, 300-03)) and incubated at 37 °C, 5% CO2. THP-1 cells (ATCC, TIB-202) were cultured in RPMI (+ 10% heat inactivated FBS, 10 U ml−1 penicillin, 100 mg ml−1 streptomycin) and incubated at 37 °C, 5% CO2. For validation of GM-CSF autoantibodies, TF-1 cells were starved of recombinant GM-CSF 18 h before experiments. GM-CSF at 200 pg ml−1 was incubated with dilutions of purified IgG for 15 min at room temperature and then used to stimulate TF-1 cells in a 96-well plate (2 × 105 cells per well) in a final volume of 100 μl (final concentration of 100 pg ml−1). For validation of IFN autoantibodies, IFNα2 (R&D Systems, 11100-1) and IFNω (Peprotech, 300-02J) at 1,500 pg ml−1 and 2,000 pg ml−1, respectively, were incubated with dilutions of purified IgG for 15 min at room temperature and then used to stimulate THP-1 cells in a 96-well plate (3.5 × 105 cells per well) in a final volume of 100 μl (final concentrations of 750 and 1,000 pg ml−1, respectively). IgG was purified from plasma using protein G magnetic beads (Lytic Solutions) as previously described7. After 15 min of stimulation, cells were fixed in 4% paraformaldehyde for 30 min, washed with PBS and permeabilized in 100% methanol on ice for 45 min. Cells were then washed twice with PBE and stained with PE-conjugated anti-STAT5 pY694 (1:50) (BD Biosciences, 562077) or anti-STAT1 pY701 (1:50) (BD Biosciences, 612564) and human TruStain FcX (1:100) (Biolegend, 422302) for 1 h at room temperature. Cells were washed with PBE and acquired on a SONY SA3800 flow cytometer. Data were analysed using FlowJo software version 10.6 software (Tree Star). pSTAT signal was measured as a function of mean fluorescence intensity (MFI). Per cent maximum signal was calculated by subtracting background MFI and calculating values as a percentage of cytokine-induced pSTAT MFI in the absence of IgG. Curves were fit using a sigmoidal four-parameter logistic curve. TF-1 and THP-1 cells were purchased commercially and were not authenticated by us. TF-1 and THP-1 cells were not tested for mycoplasma contamination.
Functional validation of anti-CXCL1, anti-CXCL7 and anti-HCRTR2 autoantibodies
CXCL1, CXCL7 and orexin signalling was assayed using the PRESTO-TANGO system39. HTLA cells, a HEK293-derived cell line that stably expresses β-arrestin-TEV and tTA-luciferase, were seeded in wells of a sterile tissue-culture-grade flat-bottom 96-well plate (35,000 cells per well) in 100 μl DMEM (+ 10% FBS, 1% penicillin–streptomycin) and incubated at 37 °C, 5% CO2. At 18–24 h after seeding (approximately 80–90% cell confluence), 200 ng CXCR2-Tango or HCTR2-Tango plasmid in 20 μl DMEM and 600 ng polyethylenimine-max (Polysciences, 24765-1) in 20 μl DMEM were mixed, incubated at room temperature for 20 min and added to each well. At 18–24 h after transfection, medium was replaced with 100 μl DMEM (+ 1% penicillin–streptomycin, 10 mM HEPES) containing 10 ng CXCL7 (Peprotech, 300-14) or CXCL1 (PeproTech, 300-46), or 100 nM orexin A (Millipore Sigma, O6012) and 5 μg isolated IgG. IgG was purified from plasma using protein G magnetic beads (Lytic Solutions) as previously described7. At 18–24 h after stimulation, supernatant was replaced with 50 μl Bright-Glo solution (Promega) diluted 20-fold with PBS with 20 mM HEPES. The plate was incubated at room temperature for 20 min in the dark and luminescence was quantified using a Synergy HTX Multi-Mode Microplate Reader (BioTek). HTLA cells were a gift from N. Palm and were derived as previously described39. HTLA cells were authenticated based on their performance in PRESTO-TANGO and were not tested for mycoplasma contamination. Tango plasmids were a gift from B. Roth (Addgene plasmid no. 66260)
Functional validation of anti-CD38 and anti-CD3ε autoantibodies
Bone marrow stem cells were isolated from the femur and tibia of 8-week-old C57BL/6 mice. Cells were plated in RPMI (+ 10% heat inactivated FBS, 10 U ml−1 penicillin, 100 mg ml−1 streptomycin (cRPMI)) with 30% (v/v) L929 fibroblast conditioned medium as a source of M-CSF and incubated at 37 °C, 5% CO2. On day 3 after isolation, 10 ml L929 fibroblast conditioned medium was added to plates. Two hundred thousand bone-marrow-derived macrophages (7 days after isolation) were plated on non-TC treated 6-well plates in cRPMI + 10% L929 medium. The following day, Raji (ATCC, CCL-86) or Jurkat (ATCC, TIB-152) cells were labelled using CellTrace Far Red (Thermo Fisher) according to standard manufacturer protocols. For the T cell antibody-dependent cellular phagocytosis assay, labelled Jurkat cells were incubated with 100 μg ml−1 healthy control IgG or CD3ε autoantibody+ patient IgG for 30 min on ice. Mouse IgG2a anti-human CD3ε (clone OKT3, Biolegend) was used at 5 μg ml−1 as a positive control. Jurkat cells were washed with 10 ml PBS. One million Jurkat cells were added to each well and incubated for 3 h. For the B cell antibody-dependent cellular phagocytosis assay, Raji cells were incubated with complement inactivated patient plasma at 1:50 dilution in PBS for 30 min on ice. Mouse IgG2a anti-human CD38 (clone MAB2404, R&D systems) was used at 5 μg ml−1 as a positive control. Raji cells were washed with 10 ml PBS once. Five hundred thousand Raji cells were added to each well and incubated for 3 h at 37 °C, 5% CO2. Bone marrow-derived macrophages were detached from the plate after a 3-h incubation using 10 mM EDTA PBS and stained with anti-mouse CD45–Pacific blue (clone 30-F11, Biolegend) for 30 min at 4 °C. Samples were acquired on a CytoFLEX flow cytometer (Beckman Coulter). Raji and Jurkat cells were purchased commercially and were not authenticated by us. Raji and Jurkat cells were not tested for mycoplasma contamination.
Mice
B6.Cg-Tg(K18-ACE2)2Prlmn/J (K18-hACE2) mice (stock no. 034860) and C57BL/6 mice (stock no. 000664) were purchased from the Jackson Laboratories and were subsequently bred and housed at Yale University. Six- to ten-week-old mixed-sex mice were used throughout the study. All mice were housed as groups of 5 to 6 mice per cage and maintained on a 12-h light/dark cycle (lights on at 07:00) at 22–25 °C and 30–70% relative humidity under specific-pathogen-free conditions. All procedures used in this study (sex-matched and age-matched) complied with federal guidelines and the institutional policies of the Yale School of Medicine Animal Care and Use Committee. All infection studies were performed in animal biosafety level 3 facilities at Yale University in accordance with approved institutional protocols.
SARS-CoV-2 mouse infections and antibody treatments
Before infection, mice were anaesthetized using 30% (v/v) isoflurane diluted in propylene glycol. Then, 50 μl of SARS-CoV-2 isolate USA-WA1/2020 (NR-52281; BEI Resources) at 2 × 104 or 6 × 104 plaque-forming units (PFU) per ml was delivered intranasally to mice, equivalent of 1,000 (sublethal dose) or 3,000 (median lethal dose) PFU per mouse, respectively. Following infection, weight loss and survival were monitored daily. Mice reported as ‘dead’ in this study were found dead, moribund or euthanized at 90% or 80% of their starting body weight. For IFNAR blockade, mice were treated intraperitoneally once with 2 mg of blocking antibodies one day before infection (clone MAR1-5A3). For IL-18 blockade, mice were treated intraperitoneally three times each with 0.5 mg of blocking antibodies at 0, 2 and 4 days after infection (clone YIGIF74-1G7). For blockade of IL-1β, GM-CSF or IL-21R, mice were treated intraperitoneally three times each with 0.2 mg of blocking antibodies at 0, 2 and 4 days after infection (anti-IL-1β: clone B122; anti-GM-CSF: clone MP1-22E9; and anti-IL-21R: clone 4A9). The first injection of anti-IL18, anti-IL-1β, anti-GM-CSF or anti-IL-21R antibodies was given at least 8 to 10 h before infection. All blocking antibodies were purchased from BioXCell.
Statistical analysis
Details of linear models and principal component analysis can be found in the Supplementary Methods. Specific details of other statistical analysis are found in the figure legends. Data analysis was performed using MATLAB, GraphPad Prism, R, and the following R packages: ggplot2, edgeR, tidyverse, tidyr, dplyr, stringr, forcats, lme4, emmeans and ggpubr.
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this paper.
Data availability
The previously published RNA-sequencing dataset is publicly available in the Gene Expression Omnibus under accession number GSE107011. All data analysed in this study are available in the Article and Supplementary Information. Data that are not available within the manuscript are available from the corresponding authors upon reasonable request. Source data are provided with this paper.
Code availability
Custom codes used for analysis in this study are available publicly at https://github.com/ring-lab/COVID-19_REAP_nature_2021.
References
Lucas, C. et al. Longitudinal analyses reveal immunological misfiring in severe COVID-19. Nature 584, 463–469 (2020).
Maucourant, C. et al. Natural killer cell immunotypes related to COVID-19 disease severity. Sci. Immunol. 5, eabd6832 (2020).
Mann, E. R. et al. Longitudinal immune profiling reveals key myeloid signatures associated with COVID-19. Sci. Immunol. 5, eabd6197 (2020).
Lee, J. S. et al. Immunophenotyping of COVID-19 and influenza highlights the role of type I interferons in development of severe COVID-19. Sci. Immunol. 5, eabd1554 (2020).
Rodrigues, T. S. et al. Inflammasomes are activated in response to SARS-CoV-2 infection and are associated with COVID-19 severity in patients. J. Exp. Med. 218, e20201707 (2021).
Vabret, N. et al. Immunology of COVID-19: current state of the science. Immunity 52, 910–941 (2020).
Wang, E. Y. et al. REAP: a platform to identify autoantibodies that target the human exoproteome. Preprint at https://doi.org/10.1101/2021.02.11.430703 (2021).
Hassan, A. O. et al. A SARS-CoV-2 infection model in mice demonstrates protection by neutralizing antibodies. Cell 182, 744–753.e4 (2020).
Garcia-Beltran, W. F. et al. COVID-19 neutralizing antibodies predict disease severity and survival. Cell 184, 476–488 (2021).
Woodruff, M. C. et al. Extrafollicular B cell responses correlate with neutralizing antibodies and morbidity in COVID-19. Nat. Immunol. 21, 1506–1516 (2020).
Bastard, P. et al. Autoantibodies against type I IFNs in patients with life-threatening COVID-19. Science 370, eabd4585 (2020).
Combes, A. J. et al. Global absence and targeting of protective immune states in severe COVID-19. Nature 591, 124–130 (2021).
Zuo, Y. et al. Prothrombotic autoantibodies in serum from patients hospitalized with COVID-19. Sci. Transl. Med. 12, eabd3876 (2020).
Woodruff, M. C., Ramonell, R. P., Lee, F. E.-H. & Sanz, I. Clinically identifiable autoreactivity is common in severe SARS-CoV-2 infection. Preprint at https://doi.org/10.1101/2020.10.21.20216192 (2020).
Zhou, Y. et al. Clinical and autoimmune characteristics of severe and critical cases of COVID-19. Clin. Transl. Sci. 13, 1077–1086 (2020).
Chang, S. E. et al. New-onset IgG autoantibodies in hospitalized patients with COVID-19. Preprint at https://doi.org/10.1101/2021.01.27.21250559 (2021).
Takahashi, T. et al. Sex differences in immune responses that underlie COVID-19 disease outcomes. Nature 588, 315–320 (2020).
Monaco, G. et al. RNA-seq signatures normalized by mRNA abundance allow absolute deconvolution of human immune cell types. Cell Rep. 26, 1627–1640.e7 (2019).
McCray, P. B., Jr et al. Lethal infection of K18-hACE2 mice infected with severe acute respiratory syndrome coronavirus. J. Virol. 81, 813–821 (2007).
Zheng, J. et al. COVID-19 treatments and pathogenesis including anosmia in K18-hACE2 mice. Nature 589, 603–607 (2021).
Winkler, E. S. et al. SARS-CoV-2 infection of human ACE2-transgenic mice causes severe lung inflammation and impaired function. Nat. Immunol. 21, 1327–1335 (2020).
Mantovani, A., Dinarello, C. A., Molgora, M. & Garlanda, C. Interleukin-1 and related cytokines in the regulation of inflammation and immunity. Immunity 50, 778–795 (2019).
Zhou, T. et al. IL-18BP is a secreted immune checkpoint and barrier to IL-18 immunotherapy. Nature 583, 609–614 (2020).
Orzalli, M. H. et al. An antiviral branch of the IL-1 signaling pathway restricts immune-evasive virus replication. Mol. Cell 71, 825–840.e6 (2018).
Spolski, R. & Leonard, W. J. Interleukin-21: a double-edged sword with therapeutic potential. Nat. Rev. Drug Discov. 13, 379–395 (2014).
Trapnell, B. C. & Whitsett, J. A. GM-CSF regulates pulmonary surfactant homeostasis and alveolar macrophage-mediated innate host defense. Annu. Rev. Physiol. 64, 775–802 (2002).
Wang, G. et al. C-reactive protein level may predict the risk of COVID-19 aggravation. Open Forum Infect. Dis. 7, ofaa153 (2020).
Cheng, L. et al. Ferritin in the coronavirus disease 2019 (COVID-19): a systematic review and meta-analysis. J. Clin. Lab. Anal. 34, e23618 (2020).
Song, E. et al. Neuroinvasion of SARS-CoV-2 in human and mouse brain. J. Exp. Med. 218, e20202135 (2021).
Nalbandian, A. et al. Post-acute COVID-19 syndrome. Nat. Med. 27, 601–615 (2021).
Nixon, J. P. et al. Sleep disorders, obesity, and aging: the role of orexin. Ageing Res. Rev. 20, 63–73 (2015).
Wyllie, A. L. et al. Saliva or nasopharyngeal swab specimens for detection of SARS-CoV-2. N. Engl. J. Med. 383, 1283–1286 (2020).
McPadden, J. et al. Health care and precision medicine research: analysis of a scalable data science platform. J. Med. Internet Res. 21, e13043 (2019).
Schulz, W. L., Durant, T. J. S., Torre, C. J., Jr, Hsiao, A. L. & Krumholz, H. M. Agile health care analytics: enabling real-time disease surveillance with a computational health platform. J. Med. Internet Res. 22, e18707 (2020).
McPadden, J. et al. Clinical characteristics and outcomes for 7,995 patients with SARS-CoV-2 infection. Preprint at https://doi.org/10.1101/2020.07.19.20157305 (2020).
Vogels, C. B. F. et al. Analytical sensitivity and efficiency comparisons of SARS-CoV-2 RT-qPCR primer-probe sets. Nat. Microbiol. 5, 1299–1305 (2020).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140 (2010).
Amanat, F. et al. A serological assay to detect SARS-CoV-2 seroconversion in humans. Nat. Med. 26, 1033–1036 (2020).
Kroeze, W. K. et al. PRESTO-Tango as an open-source resource for interrogation of the druggable human GPCRome. Nat. Struct. Mol. Biol. 22, 362–369 (2015).
Acknowledgements
We thank all members of the laboratories of A.M.R. and A.I., as well as E. Meffre, J. Craft and K. O’Connor, for helpful conversation and technical assistance; M. Linehan and H. Dong for logistical assistance; H. P. Young for technical assistance with data management; N. K. Savalia and T. J. Shelby for discussion regarding statistical techniques; and L. Steinman for helpful discussions about autoantibodies to HCRTR2. This work was supported by the Mathers Family Foundation (to A.M.R. and A.I.), the Ludwig Family Foundation (to A.M.R. and A.I.), a supplement to the Yale Cancer Center Support Grant 3P30CA016359-40S4 (to A.M.R.), the Beatrice Neuwirth Foundation, Yale Schools of Medicine and Public Health and NIAID grant U19 AI08992. IMPACT received support from the Yale COVID-19 Research Resource Fund. A.M.R. is additionally supported by an NIH Director’s Early Independence Award (DP5OD023088), a Pew-Stewart Award, and the Robert T. McCluskey Foundation. T.M. and Y.D. are supported by the Yale Interdisciplinary Immunology Training Program T32AI007019. J.K. and J.R.J. are supported by the Yale Medical Scientist Training Program T32GM007205. Schematics in Fig. 3a, Extended Data Figs. 1a, 7a were created with Biorender.com.
Author information
Authors and Affiliations
Consortia
Contributions
E.Y.W., T.M., J.K., Y.D., J.D.H., J.R.J., A.I. and A.M.R. designed experiments. E.Y.W. and Y.D. performed the REAP assay. E.Y.W., Y.D., J.D.H., J.R.J., F.L., E.S.P. and S.F. performed biochemical and functional validations. T.M., B.I. and E. Song performed mouse experiments. C.L., P.W., J.K., J.S., T.M. and J.E.O. defined parameters for flow cytometry experiments, and collected and processed peripheral blood mononuclear cell samples from patients. A.L.W., C.B.F.V., I.M.O., C.C.K., M.E.P. and A.E.W. performed the virus RNA concentration assays. N.D.G. supervised the virus RNA concentration assays. B.I. and J.K. collected epidemiological and clinical data. C.D.C., S.F., A.I.K., M.C. and J.B.F. assisted in designing, recruiting and following in-patient and healthcare worker cohorts. A.C. performed clinical data aggregation and management. W.L.S. supervised clinical data aggregation and management. E.Y.W., T.M., J.K., N.S.Z., Y.D., J.D.H., J.R.J., A.I. and A.M.R. analysed data. S.M. supervised the statistical analysis. E.Y.W., T.M., J.K., A.I. and A.M.R. wrote the paper. A.M.R. and A.I. supervised the research. Authors from the Yale IMPACT Research Team contributed to collection and storage of patient samples, as well as the collection of the patients’ epidemiological and clinical data.
Corresponding authors
Ethics declarations
Competing interests
A.M.R., E.Y.W. and Y.D. are inventors of a pending US patent application (PCT/US21/23521) filed by Yale University describing the REAP technology used in this Article. A.M.R. is the founder of Seranova Bio, the commercial licensee of the REAP technology. A.I. serves as a consultant for Spring Discovery, Boehringer Ingelheim and Adaptive Biotechnologies.
Additional information
Peer review information Nature thanks Petter Brodin, Karl Dane Wittrup and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 REAP screen of patients with COVID-19.
a, A simplified schematic of REAP. Antibodies are incubated with a barcoded yeast library displaying members of the exoproteome. Antibody-bound yeast are enriched by magnetic-column-based sorting and enrichment is quantified by next-generation sequencing. b, Heat map of all profiled reactivities across all patient samples stratified by disease severity and using the same colour scale as in Fig. 1a. Sticky antigens (as defined in Methods) were removed from the heat map. c, SARS-CoV-2 RBD REAP scores for samples from patients with COVID-19 stratified by positive or negative ELISA RBD reactivity. d, IL-6R REAP scores for samples from patients with COVID-19 stratified by treatment with an anti-IL-6R biologic therapy (tocilizumab or sarilumab). Samples collected at least one day after infusion were considered treated. Samples collected on the day of infusion were excluded from analysis owing to uncertainty in the timing of sample collection. Significance in c, d was determined using a linear mixed model (Methods). In c, d, n values include longitudinal samples from the same patient.
Extended Data Fig. 2 Comparisons of reactivities and clinical or immunological parameters between patient groups.
a, Number of positive reactivities per sample at various REAP score cut offs, stratified by disease severity. b, Days from symptom onset (DFSO) in samples from patients with severe or moderate COVID-19. Days from symptom onset data were not available for a limited number of samples from each group and were not available for any samples from individuals with mild COVID-19 or no symptoms. The median (solid line) and first and third quartile (dashed lines) are shown. c, Number of positive reactivities per sample at various REAP score cut offs, stratified by patient sex. d, Number of positive reactivities in samples from patients with COVID-19, SLE or APECED at various score cut offs. Patients with SLE or APECED were screened as previously described7. Owing to the smaller size of the yeast exoproteome library used to screen the samples from patients with SLE or APECED, reactivities in samples from patients with COVID-19 against proteins that were not in the previously described yeast exoproteome library were removed from these analyses. e, Number of positive immune-targeting reactivities per sample at various REAP score cut offs, stratified by disease severity. f, Average percentages of double-negative (IgD−CD27−) B cells among peripheral leukocytes in patients with COVID-19 stratified by disease severity and uninfected controls (neg.). In a–e, n values include longitudinal samples from the same patient. In f, n values indicate samples from unique patients. Significance was determined using linear mixed models (a–e) (Methods) or a Kruskal–Wallis test followed by a two-sided Dunn’s test (f). Medians are represented by a dashed line and first and third quartiles are represented by dotted lines for all plots in this figure.
Extended Data Fig. 3 Autoantibodies exhibit varied developmental kinetics in patients with COVID-19.
a, Percentage of reactivities (REAP score greater than score cut off) in patients with COVID-19 present within 10 days of symptom onset at various score cut offs. b, Percentage of reactivities in patients with COVID-19 that had a REAP score less than the score cut off (using various score cut offs) at the first time point sampled and an increase in REAP score of at least 1 at the last time point. c, Percentage of reactivities in patients with COVID-19 that had a REAP score greater than the score cut off (using various score cut offs) at the first time point sampled and a decrease in REAP score of at least 1 at the last time point. For a–c, all patients with COVID-19 with longitudinal samples available (n = 77 patients) were included in calculations. d–f, Plots of longitudinal changes in REAP score for ‘likely pre-existing’ (d), ‘newly acquired’ (e) and ‘waning’ (f) REAP reactivities in individual patients alongside scaled anti-spike S1 ELISA values in the same patients. Scaled ELISA values are defined as anti-spike S1 ELISA optical density (450–570 nm) values multiplied by four. In each plot, unique patients are represented by uniquely coloured lines. For a given patient, solid lines connect REAP scores of reactivities against respective antigens at various time points and dashed lines connect scaled anti-spike S1 ELISA values at those same time points. The dotted grey line indicates the scaled ELISA positivity cut off value (Methods).
Extended Data Fig. 4 Biochemical and functional validation of autoantibodies in patients with COVID-19.
a, Single-point pan-IgG autoantibody ELISAs conducted with 1:25 or 1:50 plasma dilution (indicated in graph titles). Dotted line represents the uninfected individual (healthcare worker) average plus 3 s.d. For controls, results (averages of technical duplicates) from biologically independent samples are displayed in the same column (n indicated below each column). For patients with COVID-19, results from one patient are displayed in each column and technical duplicates are depicted as distinct points. b–d, GM-CSF (b), CD38 (c) and IL-18Rβ (d) pan-IgG autoantibody ELISAs conducted with serial dilutions of plasma from patients with COVID-19 or uninfected individuals. Results are averages of two technical replicates. Curves were fit using a sigmoidal four-parameter logistic curve. Experiments in a–d were performed once. e, Per cent of variance explained for principal components from the principal component analysis in Fig. 1d. f, Second principal component scores of samples from patients with COVID-19 stratified by clinical score. Solid black lines depict group means. g, Fixed-effects model fits from a generalized linear mixed effects model with second principal component score as the dependent variable (Methods). h, i, Flow cytometry gating for the Raji (h) and Jurkat (i) macrophage phagocytosis assay in Fig. 2c. j, k, IFNα2 (j) and IFNω (k) signalling assay performed with IgG from patients with COVID-19 who were positive for anti-IFNα2 or anti-IFNω autoantibody or from uninfected individuals. Results are averages of two technical replicates from one experiment. l, Fixed-effects model fits for the generalized linear mixed-effects model in Fig. 2d (Methods). m, Hospital stay length in patients with and without autoantibodies targeting type I IFNs, stratified by disease severity. Significance in m was determined using a two-sided Wilcoxon rank-sum test. In f, n values include longitudinal samples from the same patient. All other n values in this figure indicate samples from unique patients.
Extended Data Fig. 5 Effects of anti-cytokine autoantibodies on corresponding cytokine plasma concentrations.
a–t, Average concentration of plasma CCL11 (a), CCL15 (b), CCL2 (c), CCL26 (d), CCL8 (e), CXCL1 (f), CXCL12 (g), CXCL13 (h), FLT3LG (i), IFNα2 (j), IL-1A (k), IL-1B (l), IL-13 (m), IL-16 (n), IL-21 (o), IL-22 (p), IL6 (q), PDGFA (r), TGFα (s) and TSLP (t) measured by a Luminex assay in patients stratified by COVID-19 disease severity and REAP reactivity (autoantibody positive) (REAP score ≥ 2 at any time point) against the corresponding cytokine. Data are presented as box plots with the first quartile, median, third quartile, whiskers (minimum and maximum values within the first and third quartiles ± 1.5× the interquartile range), and individual data points indicated. Significance was determined using two-sided, Wilcoxon rank-sum tests. All n values in this figure indicate samples from unique patients.
Extended Data Fig. 6 Additional correlations between autoantibodies targeting immune-cell surface proteins.
a, Average anti-SARS-CoV-2 RBD IgM reactivity as measured by ELISA in patients stratified by COVID-19 disease severity and REAP reactivity against B-cell-displayed proteins (as defined in Fig. 2e). b, Average percentage among total monocytes of classical monocytes, intermediate monocytes and nonclassical monocytes in patients stratified by COVID-19 disease severity and REAP reactivity against proteins preferentially displayed on classical and intermediate monocytes (as defined in Fig. 2f). c, Average per cent CD8+ T cells, natural killer T (NKT) cells and natural killer cells among peripheral leukocytes in patients stratified by COVID-19 disease severity and REAP reactivity against CD3ε. d, Average CD4+ T cell-to-natural killer cell ratio, CD8+ T cell-to-natural killer cell ratio and natural killer T cell-to-natural killer cell ratio among peripheral leukocytes in patients stratified by COVID-19 disease severity and REAP reactivity against CD3ε. e, Representative flow plot of T cells (CD3+), natural killer cells (CD56+) and natural killer T cells (CD3+CD56+) for c, d. f, Average per cent B cells, natural killer cells, activated CD4+ T cells and activated CD8+ T cells among peripheral leukocytes in patients stratified by COVID-19 disease severity and REAP reactivity against CD38. g, Representative flow plot of B cells (CD19+HLA-DR+) for f. h, Representative flow plot of activated CD8+ T cells (CD38+HLA-DR+) for f. Significance in a was determined using a two-sided Wilcoxon rank-sum test. All n values in this figure indicate samples from unique patients. All box plots are presented with the median, first and third quartile, whiskers (minimum and maximum values within the first and third quartiles ± 1.5× the interquartile range), and individual data points indicated.
Extended Data Fig. 7 Additional immunological and clinical characterization of autoantibody effects in a mouse model of COVID-19.
a–n, Schematic of experiment (a) in which K18-hACE2 mice were intranasally infected with a sublethal (b–f) or median lethal (g–n) dose of SARS-CoV-2 (USA-WA1/2020 isolate) and treated with indicated antibodies. b, c, Relative frequency (b) and absolute number (c) of lung Ly6C+CD11b+CD64+ macrophages from mock-infected, SARS-CoV-2-infected and PBS-treated, and SARS-CoV-2-infected and anti-IFNAR-treated K18-hACE2 mice. d, Expression of CD64 on lung-infiltrating CD11b+Ly6Chigh monocytes. e, f, Relative frequency (e) and absolute number (f) of CD44+CD69+ lymphocytes (CD4+ T cells, CD8+ T cells, NK1.1+ cells and γδ T cells). g, h, Viral RNA loads (g) and infectious titres (h) from lung tissue homogenates of mock-infected, SARS-CoV-2-infected and PBS-treated, and SARS-CoV-2-infected and anti-IL-18-treated mice measured by reverse-transcription qPCR and plaque assay, respectively. i, j, Relative frequency (i) or absolute number (j) of CD11b+ and KLRG1+NK1.1+ cells in lung tissues of PBS- and anti-IL-18-treated mice. k–n, Normalized body weight of anti-IL-18- (k), anti-IL-1β- (l), anti-IL-21R- (m), anti-GM-CSF- (n) and PBS-treated, SARS-CoV-2-infected K18-hACE2 mice from day 1 to 14 after infection. Significance was determined using one-way analysis of variance followed by Tukey correction (b–f, g), and unpaired two-tailed t-tests (i, j). All n values in this figure represent biologically independent mice examined over two independent experiments.
Extended Data Fig. 8 Flow cytometry gating strategies for immunophenotyping experiments.
a, Gating strategy to identify the B cells described in Fig. 2e, monocytes described in Fig. 2f, Extended Data Fig. 6b, and T cells, natural killer T cells and natural killer cells described in Fig. 2g, Extended Data Fig. 6c, d, f, in human peripheral blood mononuclear cells. b, Gating strategy to identify CD11b+Ly6Chigh monocytes and Ly6C+CD11b+CD64+ macrophages in mouse lung tissues described in Extended Data Fig. 7b–d. c, Gating strategy to identify CD44+CD69+ lymphocytes in mouse lung tissues described in Extended Data Fig. 7e, f. d, Gating strategy to identify KLRG1+ and CD11b+ natural killer cells described in Extended Data Fig. 7i, j.
Extended Data Fig. 9 Tissue-associated autoantibodies are correlated with clinical parameters in patients with COVID-19.
Heat map of pairwise, −log10-transformed REAP protein P values from generalized linear model fits accounting for age, sex and maximum REAP score in estimating average or minimum clinical values from hospital admission. A total of 135 samples from unique patients were included in this analysis (n = 135). Intensity represents P values and colour indicates directionality of gene-parameter pairs (red indicates positive change, blue indicates negative change. Grey columns indicate genes excluded from analysis due to insufficient patient number (n < 2). Asterisks denote pairwise relationships that were significant after correction for multiple comparisons (false-discovery rate). Specific details of the generalized linear models can be found in the Methods. Proteins were classified as in Fig. 4a.
Supplementary information
Supplementary Information
This file contains the Supplementary Methods.
Supplementary Table 1
The full antigen list.
Source data
Rights and permissions
About this article
Cite this article
Wang, E.Y., Mao, T., Klein, J. et al. Diverse functional autoantibodies in patients with COVID-19. Nature 595, 283–288 (2021). https://doi.org/10.1038/s41586-021-03631-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-021-03631-y
This article is cited by
-
Acute neutrophilic vasculitis (leukocytoclasia) in 36 COVID-19 autopsy brains
Diagnostic Pathology (2024)
-
Characterisation, symptom pattern and symptom clusters from a retrospective cohort of Long COVID patients in primary care in Catalonia
BMC Infectious Diseases (2024)
-
Muscle abnormalities worsen after post-exertional malaise in long COVID
Nature Communications (2024)
-
The neurobiology of SARS-CoV-2 infection
Nature Reviews Neuroscience (2024)
-
Autoantibodies to ACE2 and immune molecules are associated with COVID-19 disease severity
Communications Medicine (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.