Abstract
Disulfide bonds between cysteine residues are important post-translational modifications in proteins that have critical roles for protein structure and stability, as redox-active catalytic groups in enzymes or allosteric redox switches that govern protein function1,2,3,4. In addition to forming disulfide bridges, cysteine residues are susceptible to oxidation by reactive oxygen species, and are thus central not only to the scavenging of these but also to cellular signalling and communication in biological as well as pathological contexts5,6. Oxidized cysteine species are highly reactive and may form covalent conjugates with, for example, tyrosines in the active sites of some redox enzymes7,8. However, to our knowledge, regulatory switches with covalent crosslinks other than disulfides have not previously been demonstrated. Here we report the discovery of a covalent crosslink between a cysteine and a lysine residue with a NOS bridge that serves as an allosteric redox switch in the transaldolase enzyme of Neisseria gonorrhoeae, the pathogen that causes gonorrhoea. X-ray structure analysis of the protein in the oxidized and reduced state reveals a loaded-spring mechanism that involves a structural relaxation upon redox activation, which is propagated from the allosteric redox switch at the protein surface to the active site in the protein interior. This relaxation leads to a reconfiguration of key catalytic residues and elicits an increase in enzymatic activity of several orders of magnitude. The redox switch is highly conserved in related transaldolases from other members of the Neisseriaceae; for example, it is present in the transaldolase of Neisseria meningitides (a pathogen that is the primary cause of meningitis and septicaemia in children). We surveyed the Protein Data Bank and found that the NOS bridge exists in diverse protein families across all domains of life (including Homo sapiens) and that it is often located at catalytic or regulatory hotspots. Our findings will inform strategies for the design of proteins and peptides, as well as the development of new classes of drugs and antibodies that target the lysine–cysteine redox switch9,10.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Data availability
The refined structural protein models and corresponding structure–factor amplitudes have been deposited under PDB accession codes 6XZ4 (NgTAL oxidized citrate 1), 6ZWJ (NgTAL oxidized citrate 2), 6ZWH (NgTAL oxidized acetate), 6ZWF (NgTAL reduced citrate), 7B0L (NgTAL oxidized low-dose), 7BBX (NgTAL Lys8Ala variant) and 7BBW (NgTAL Cys38Ser variant). The structures cited in this publication (1M3Q, 3CLM, 6T3X and 5Y72) are available under their respective PDB accession codes. The data for our protein database are currently private, and can only be accessed with a single reviewer account that has been created. The mass spectrometry proteomics data have been deposited to the ProteomeXchange Consortium via the PRIDE partner repository with the dataset identifier PXD020302 (reviewer account details: reviewer52532@ebi.ac.uk (username), AK3E73R2 (password)). All other data are available from the corresponding author on request.
References
Bardwell, J. C., McGovern, K. & Beckwith, J. Identification of a protein required for disulfide bond formation in vivo. Cell 67, 581–589 (1991).
Wedemeyer, W. J., Welker, E., Narayan, M. & Scheraga, H. A. Disulfide bonds and protein folding. Biochemistry 39, 4207–4216 (2000).
Matsumura, M. & Matthews, B. W. Control of enzyme activity by an engineered disulfide bond. Science 243, 792–794 (1989).
Hogg, P. J. Disulfide bonds as switches for protein function. Trends Biochem. Sci. 28, 210–214 (2003).
Paulsen, C. E. & Carroll, K. S. Cysteine-mediated redox signaling: chemistry, biology, and tools for discovery. Chem. Rev. 113, 4633–4679 (2013).
Depuydt, M. et al. A periplasmic reducing system protects single cysteine residues from oxidation. Science 326, 1109–1111 (2009).
Ito, N. et al. Novel thioether bond revealed by a 1.7 Å crystal structure of galactose oxidase. Nature 350, 87–90 (1991).
Firbank, S. J. et al. Crystal structure of the precursor of galactose oxidase: an unusual self-processing enzyme. Proc. Natl Acad. Sci. USA 98, 12932–12937 (2001).
Dror, R. O. et al. Structural basis for modulation of a G-protein-coupled receptor by allosteric drugs. Nature 503, 295–299 (2013).
Nussinov, R. & Tsai, C. J. The design of covalent allosteric drugs. Annu. Rev. Pharmacol. Toxicol. 55, 249–267 (2015).
Kang, H. J., Coulibaly, F., Clow, F., Proft, T. & Baker, E. N. Stabilizing isopeptide bonds revealed in Gram-positive bacterial pilus structure. Science 318, 1625–1628 (2007).
Uhlin, U. & Eklund, H. Structure of ribonucleotide reductase protein R1. Nature 370, 533–539 (1994).
Lehwess-Litzmann, A. et al. Twisted Schiff base intermediates and substrate locale revise transaldolase mechanism. Nat. Chem. Biol. 7, 678–684 (2011).
Tittmann, K. Sweet siblings with different faces: the mechanisms of FBP and F6P aldolase, transaldolase, transketolase and phosphoketolase revisited in light of recent structural data. Bioorg. Chem. 57, 263–280 (2014).
Wi, T. et al. Antimicrobial resistance in Neisseria gonorrhoeae: global surveillance and a call for international collaborative action. PLoS Med. 14, e1002344 (2017).
Chiu, J. & Hogg, P. J. Allosteric disulfides: sophisticated molecular structures enabling flexible protein regulation. J. Biol. Chem. 294, 2949–2960 (2019).
Neumann, P. & Tittmann, K. Marvels of enzyme catalysis at true atomic resolution: distortions, bond elongations, hidden flips, protonation states and atom identities. Curr. Opin. Struct. Biol. 29, 122–133 (2014).
Ruszkowski, M. & Dauter, Z. On methylene-bridged cysteine and lysine residues in proteins. Protein Sci. 25, 1734–1736 (2016).
Wang, J. Crystallographic identification of spontaneous oxidation intermediates and products of protein sulfhydryl groups. Protein Sci. 28, 472–477 (2019).
Kovacic, P. & Bennett, R. P. Aromatic amination with hydroxylamine-O-sulfonic acid. J. Am. Chem. Soc. 83, 221–224 (1961).
Akter, S. et al. Chemical proteomics reveals new targets of cysteine sulfinic acid reductase. Nat. Chem. Biol. 14, 995–1004 (2018).
Kirby, A. J. et al. Hydroxylamine as an oxygen nucleophile. Structure and reactivity of ammonia oxide. J. Am. Chem. Soc. 128, 12374–12375 (2006).
Lüdtke, S. et al. Sub-Ångström-resolution crystallography reveals physical distortions that enhance reactivity of a covalent enzymatic intermediate. Nat. Chem. 5, 762–767 (2013).
Mesecar, A. D., Stoddard, B. L. & Koshland, D. E. Jr. Orbital steering in the catalytic power of enzymes: small structural changes with large catalytic consequences. Science 277, 202–206 (1997).
Schörken, U. et al. Identification of catalytically important residues in the active site of Escherichia coli transaldolase. Eur. J. Biochem. 268, 2408–2415 (2001).
Dai, S. et al. Low-barrier hydrogen bonds in enzyme cooperativity. Nature 573, 609–613 (2019).
Rabe von Pappenheim, F. et al. Structural basis for antibiotic action of the B1 antivitamin 2′-methoxy-thiamine. Nat. Chem. Biol. 16, 1237–1245 (2020).
Seib, K. L. et al. Defenses against oxidative stress in Neisseria gonorrhoeae: a system tailored for a challenging environment. Microbiol. Mol. Biol. Rev. 70, 344–361 (2006).
Seib, K. L., Tseng, H. J., McEwan, A. G., Apicella, M. A. & Jennings, M. P. Defenses against oxidative stress in Neisseria gonorrhoeae and Neisseria meningitidis: distinctive systems for different lifestyles. J. Infect. Dis. 190, 136–147 (2004).
Bruner, S. D., Norman, D. P. & Verdine, G. L. Structural basis for recognition and repair of the endogenous mutagen 8-oxoguanine in DNA. Nature 403, 859–866 (2000).
Chevillard, S. et al. Mutations in OGG1, a gene involved in the repair of oxidative DNA damage, are found in human lung and kidney tumours. Oncogene 16, 3083–3086 (1998).
Huang, P. S., Boyken, S. E. & Baker, D. The coming of age of de novo protein design. Nature 537, 320–327 (2016).
Singh, J., Petter, R. C., Baillie, T. A. & Whitty, A. The resurgence of covalent drugs. Nat. Rev. Drug Discov. 10, 307–317 (2011).
Scott, A. M., Wolchok, J. D. & Old, L. J. Antibody therapy of cancer. Nat. Rev. Cancer 12, 278–287 (2012).
Muyldermans, S. Nanobodies: natural single-domain antibodies. Annu. Rev. Biochem. 82, 775–797 (2013).
Senior, A. W. et al. Improved protein structure prediction using potentials from deep learning. Nature 577, 706–710 (2020).
Callaway, E. ‘It will change everything’: DeepMind’s AI makes gigantic leap in solving protein structures. Nature 588, 203–204 (2020).
Gill, S. C. & von Hippel, P. H. Calculation of protein extinction coefficients from amino acid sequence data. Anal. Biochem. 182, 319–326 (1989).
Inoue, H., Nojima, H. & Okayama, H. High efficiency transformation of Escherichia coli with plasmids. Gene 96, 23–28 (1990).
Bertani, G. Studies on lysogenesis. I. The mode of phage liberation by lysogenic Escherichia coli. J. Bacteriol. 62, 293–300 (1951).
Tsolas, O. & Joris, L. Transaldolase. Methods Enzymol. 42, 290–297 (1975).
Sautner, V., Friedrich, M. M., Lehwess-Litzmann, A. & Tittmann, K. Converting transaldolase into aldolase through swapping of the multifunctional acid–base catalyst: common and divergent catalytic principles in F6P aldolase and transaldolase. Biochemistry 54, 4475–4486 (2015).
Schuck, P. Size-distribution analysis of macromolecules by sedimentation velocity ultracentrifugation and lamm equation modeling. Biophys. J. 78, 1606–1619 (2000).
Laue, M. T., Shah, B. D., Rigdeway, T. M. & Pelletier, S. L. in Analytical Ultracentrifugation in Biochemistry and Polymer Science (eds Harding S. et al.) 90–125 (Royal Society of Chemistry, 1992).
Brautigam, C. A. Calculations and publication-quality illustrations for analytical ultracentrifugation data. Methods Enzymol. 562, 109–133 (2015).
Kabsch, W. Xds. Acta Crystallogr. D 66, 125–132 (2010).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D 66, 213–221 (2010).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D 66, 486–501 (2010).
Williams, C. J. et al. MolProbity: More and better reference data for improved all-atom structure validation. Protein Sci. 27, 293–315 (2018).
Schrödinger. The PyMOL Molecular Graphics System, version 1.8 (2015).
Liebschner, D. et al. Polder maps: improving OMIT maps by excluding bulk solvent. Acta Crystallogr. D 73, 148–157 (2017).
Frisch, M. J. et al. Gaussian 16, revision C.01 (Gaussian, 2016.
Becke, A. D. Density-functional thermochemistry. III. The role of exact exchange. J. Chem. Phys. 98, 5648–5652 (1993).
Grimme, S., Ehrlich, S. & Goerigk, L. Effect of the damping function in dispersion corrected density functional theory. J. Comput. Chem. 32, 1456–1465 (2011).
Rappoport, D. & Furche, F. Property-optimized Gaussian basis sets for molecular response calculations. J. Chem. Phys. 133, 134105 (2010).
Wong, C. P. et al. Two distinct substrate binding modes for the normal and reverse prenylation of hapalindoles by the prenyltransferase AmbP3. Angew. Chem. Int. Ed. 57, 560–563 (2018).
Muller, Y. A. et al. High-resolution crystal structures of two prototypical β- and γ-herpesviral nuclear egress complexes unravel the determinants of subfamily specificity. J. Biol. Chem. 295, 3189–3201 (2020).
Acknowledgements
This study was supported by the Max-Planck Society and the DFG-funded Göttingen Graduate Center for Neurosciences, Biophysics and Molecular Biosciences GGNB. We acknowledge access to beamline P14 at DESY/EMBL (Hamburg), and thank G. Bourenkov and T. Schneider for local support. We thank G. Bourenkov, A. Pearson, H. Stark, R. Kluger and M. Alcarazo for discussion; and A. Linden for help with the mass spectrometry experiments.
Author information
Authors and Affiliations
Contributions
K.T. designed and coordinated research; M.W. expressed, purified, biochemically characterized and crystallized NgTAL proteins with initial support from P.K.; L.-M.F. expressed, purified and biochemically characterized NmTAL; F.R.v.P. and V.S. conducted crystallographic data collection, processing, model building and refinement with support from K.T.; F.R.v.P., V.S. and K.T. interpreted crystallographic data; F.R.v.P. did phylogenetic analysis; U.C. conducted and analysed analytical ultracentrifugation experiments; P.F. and K.-T.P. performed and analysed mass spectrometry experiments under the supervision of H.U.; R.A.M. supervised and planned theoretical investigations; J.U. performed electronic structure calculations including potentials, pKa value and structures of reaction intermediates; J.Y. performed electronic structure calculations of reaction intermediates and pathways; U.D., J.U., R.A.M. and K.T. discussed reaction mechanisms; K.T. wrote the manuscript with input from all authors.
Corresponding author
Ethics declarations
Competing interests
M.W., F.R.v.P., L.-M.F., U.D., J.U., R.A.M., V.S. and K.T. have filed a European patent application (application number EP21164101.4) for regulating protein activities by targeting the NOS redox switches.
Additional information
Peer review information Nature thanks Deborah Fass, Martin Högbom and Judith P. Klinman for their contribution to the peer review of this work. Peer reviewer reports are available.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
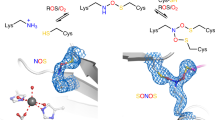
Extended Data Fig. 1 Redox chemistry of cysteine residues in proteins and electron density maps, showing the presence of the lysine–cysteine NOS bridge in independent crystal structures of NgTAL.
a, Redox reactions of cysteine with key species involved, including cysteine oxidation by reactive oxygen species (ROS) and reduction by thiol antioxidants (RSH) or enzymes (Srx). b, Mechanism of allosteric redox switch containing an intramolecular allosteric disulfide bridge. Reduction of the disulfide to the corresponding dithiol results in a structural reorganization of the protein that is propagated to the active-site altering activity. c, NgTAL crystal 1. Top left, 2mFo − DFc electron density map after final refinement contoured at 3σ. No additional positive or negative electron density was observed at ±3σ. Top right, mFo − DFc omit electron density map contoured at 5σ. Bottom, alternative refinement with a methylene bridge yields unexplained positive difference electron density at the bridging atom. The 2mFo − DFc map is shown in grey at 3σ, positive (green) and negative (red) electron density maps are shown at 3σ and −3σ, respectively. d, NgTAL crystal 2. Left, 2mFo − DFc electron density map after final refinement, contoured at 1.5σ. Right, mFo − DFc omit electron density map contoured at 5σ. e, NgTAL crystal 3. Left, 2mFo − DFc electron density map after final refinement, contoured at 1.5σ. Right, mFo − DFc omit electron density map contoured at 5σ. f, NgTAL crystal 4. Low-dose dataset of an independent crystal measured in-house at a rotating anode. The NOS bridge is also present in the corresponding NgTAL structure (left, 2mFo − DFc electron density map contoured at 1σ; right, mFo − DFc omit electron density map contoured at 3σ), thus ruling out that formation of the covalent linkage seen in crystals 1–3 results from radiation damage at the high-energy synchrotron beamline. g, Structure of the Lys8–Cys38 redox switch in the oxidized (left) and reduced (right) state, showing electron density for neighbouring waters (W1 and W2) and a presumed dioxygen molecule (O2) that is exclusively observed in the reduced state. The corresponding 2mFo − DFc electron density map is shown in blue at a contour level of 1.5σ.
Extended Data Fig. 2 Thermal unfolding, analytical ultracentrifugation and X-ray structures of NgTAL Lys8Ala and Cys38Ser variants.
a, Thermal unfolding of NgTAL wild type and the Lys8Ala and Cys38Ser variants under oxidizing and reducing conditions, as monitored by far-UV CD spectroscopy at 222 nm. Different unfolding temperatures are seen for the oxidized and reduced states in case of the wild-type enzyme and the Lys8Ala variant, whereas the Cys38Ser variant does not exhibit this feature. This suggests an oxidation of Cys38 in the Lys8Ala variant, despite the absence of Lys8. b, Analytical ultracentrifugation analyses of NgTAL wild-type and variants in the oxidized and reduced state shows the predominant formation of the monomeric form in all of the cases we tested. Under oxidizing conditions and high protein concentrations, a small fraction of higher oligomers is observed (presumably resulting from incorrectly linked monomers). c, X-ray crystallographic structure of the NgTAL Lys8Ala variant, showing the allosteric redox switch site with residues Ala8 (mutation site), Cys38, Glu93 and Thr101. For residues Ala8 and Cys38, the corresponding 2mFo − DFc electron density maps are shown in blue at a contour level of 2σ. Inset, peaks in the mFo − DFc difference electron density map (in green, contour level 3σ) around the sulfur atom of residue Cys38 suggest that this atom is oxidized. Owing to the structural flexibility of Cys38, the discrete oxidation state (mono-oxidized and/or dioxidized) cannot be unambiguously assigned. Notwithstanding this ambiguity, this observation supports our proposed mechanism of an initial cysteine oxidation as part of the formation of the NOS bridge. d, X-ray crystallographic structure of the NgTAL Cys38Ser variant, showing the allosteric redox switch site with residues Lys8, Ser38 (mutation site), Glu93 and Thr101. For residues Lys8 and Ser38, the corresponding 2mFo − DFc electron density maps are shown in blue at a contour level of 1.5σ. Lys8 is chemically unmodified, thus ruling out that the covalent linkage between Lys8 and Cys38 seen in the wild-type enzyme results from the addition of CO2 or formaldehyde potentially establishing an NCS linkage18.
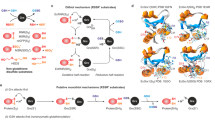
Extended Data Fig. 3 Putative reaction mechanisms of lysine–cysteine NOS bridge formation and associated computational calculations.
a, Initial reaction of the cysteine thiolate with dioxygen and subsequent attack of the lysine amine onto the α-oxygen atom of the thio-(hydro)peroxy intermediate concomitantly with proton transfers and water release. b, Oxidation of both cysteine and lysine in either concerted fashion (top path) or independently (bottom path), followed by nucleophilic attack of the oxidized lysine as a O-nucleophile onto the cysteine sulfenic acid with concomitant water release. c, Initial attack of the lysine amine onto the sulfur atom of sulfenic acid or sulfinic acid to afford a sulfinamide species followed by [1,2] rearrangement driven by orbital steering. d, Depiction of selected reaction intermediates and relative free Gibbs energies (T = 298.15 K) computed at the B3LYP-D3(BJ)/def2-TZVPD//B3LYP-D3(BJ)/def2-SVPD level of theory (as described in ‘Computational details’ in Methods). The reference is given by the sulfinic acid state (the thermodynamically most stable intermediate in our investigations). Top, starting thio-(hydro)peroxy species, which bears a barrier of 12.2 kcal mol−1 for the heterolytic cleavage of the O–O bond and concurrent oxidation of the lysine residue. We compare two different pathways (using the same nomenclature as in a–c) through the sulfenic and sulfinic acids. The mechanism in a is not directly depicted, as we have not observed any concerted O–O cleavage with amino nucleophilic attack and NOS bridge formation. Only the mechanisms in b, c are depicted. A path that leads through the sulfinic acid or sulfinamide species is not viable, because the two species are far too stable. Instead, we suggest that the formal oxidation state of sulfur [0] is kept, with concurrent oxidation of the Lys8 (pathway from b). This would represent a reversible mechanism. For each intermediate, three different protonation states were investigated with a total cluster charge of −1, 0 and 1. Only the most stable species are shown.
Extended Data Fig. 4 Phylogenetic analyses and sequence conservation of the lysine–cysteine redox switch in the TAL protein family using NgTAL as reference.
Two related consensus motifs were identified that contain the lysine and cysteine residues of the redox switch, the active site serine and asparagine residues required for catalytic activation of the Schiff-base-forming lysine, and the linker region that connects the redox switch with the active site. The identified motifs are highly conserved in Betaproteobacteria (in particular, Neisseriales (motif 1), and in Cyanophycea (motif 2)). NmTAL is highly similar (95% identity) to NgTAL, and also contains motif 1. Structural and functional analyses of NmTAL could confirm the existence of a redox switch (Extended Data Fig. 5).
Extended Data Fig. 5 Structural and functional analyses of NmTAL indicate the presence of a redox switch.
a, Steady-state kinetic analysis of enzymatic activity of NmTAL in the oxidized (black) and reduced (red) state. There is a multi-fold increase of kcat and concomitant decrease of substrate KM upon reduction. The catalytic constant of the oxidized form represents an upper limit, as oxidized and reduced species cannot be quantitatively separated by chromatographic methods as in the case of NgTAL. All measurements were carried out in triplicate and are shown as mean ± s.d. Kinetic and thermodynamic constants are provided in Extended Data Table 1. Experiments were repeated twice with similar results. b, Far-UV CD spectra of NmTAL in the oxidized (black) and reduced (red) state, showing both the natively folded states (solid lines) and states after thermal unfolding (dashed lines). The reduced enzyme completely unfolds and does not contain residual secondary structure. By contrast, the oxidized enzyme exhibits only partial unfolding of mostly helical elements and contains thermally stable β-sheet structures. c, Thermal unfolding of NmTAL in the oxidized (black) and reduced (red) state, monitored by far-UV CD spectroscopy at 222 nm. Although the oxidized enzyme displays a monophasic unfolding with a melting temperature of 50.6 °C corresponding to unfolding of the α-helices (as shown in b), the reduced enzyme shows a biphasic unfolding with melting temperatures of 51.9 °C (first transition) and 71.9 °C (second transition). This observation suggests that the putative NOS bridge in the oxidized state of NmTAL specifically stabilizes the interior β-sheet structure of the TIM barrel, in agreement with the position of switch residue Cys38.
Extended Data Fig. 6 Structure of the human DNA repair enzyme 8-oxoguanine glycosylase 1 with a putative NOS bridge at the active site.
Structure is from PDB 1M3Q, and is at 1.90 Å resolution. a, Overall structure of human 8-oxoguanine glucosylase 1 in complex with DNA and product analogue 8-aminoguanine. b, Close-up view of the active site, showing the product and residues Cys253 and Lys249. Residues Cys253 and Lys249 are superposed with the calculated 2mF − DFc electron density map (in blue, contour level 1.5σ). A strong, unexplained positive peak in the mFo − DFc difference electron density map (in green, contour level 3σ) is observed between the cysteine sulfur atom and the lysine nitrogen atom. The S–N interatomic distance (2.80 Å in the previously deposited structure) is too short for a hydrogen-bond interaction and too long for a direct S–N linkage. c, Refinement of the previously deposited structure with a geometrically parametrized NOS bridge linking Cys253 and Lys249 resulted in a structural model with no remaining unexplained electron density (2mFo − DFc electron density map in blue, contour level 1.5σ; mFo − DFc difference electron density map in green and red, contour level ± 3σ), suggesting the presence of an NOS bridge.
Extended Data Fig. 7 Representative examples of protein structures deposited in the PDB that probably contain an NOS bridge akin to that of NgTAL.
a, Prenyltransferase AmbP3. b, Human cytomegalovirus pUL50–pUL53 complex. For both examples, the overall structure is shown in cartoon representation in the top panel, highlighting the Lys–Cys linkage and providing the PDB code56,57. In the corresponding middle panels, the structure of the lysine–cysteine pair as deposited in the PDB is shown enlarged, including the calculated 2mFo − DFc (in blue, contour level 1σ) and mFo − DFc difference (in green, contour level 3σ) electron density maps. There is a pronounced positive difference peak in the electron density maps in between the lysine nitrogen atoms and the cysteine sulfur atoms, indicating the presence of a covalent bridge. In the bottom panels, the refined structural models that include the covalent lysine–cysteine NOS bridges are shown with the corresponding 2mF − DFc electron density maps. The mFo − DFc difference electron density maps are shown in green and are contoured at 3σ. The calculated occupancies of the NOS bridges amount to 62% (a) and 76% (b). The NOS bridge is prominently located at either the substrate binding site or the protein–protein binding interface.
Extended Data Fig. 8 Superposition of NgTAL in the oxidized and reduced state.
Both of these structures are from this Article. a, Structure of the active site of NgTAL in the oxidized state, showing catalytic residues Lys138, Asp17 and the ligand citrate. Citrate is partially disordered. The corresponding 2mFo − DFc (blue) and mFo − DFc omit (green) electron density maps are shown at contour levels of 1σ and 5σ, respectively. b, Structure of the active site of NgTAL in the reduced state, showing catalytic residues Lys138, Asp17 and the ligand citrate. Citrate is structurally well-defined in this state. A covalent conjugate between Lys138 and a two-carbon fragment formed during crystallization. The corresponding 2mFo − DFc (blue) and mFo − DFc omit (green) electron density maps are shown at contour levels of 1σ and 5σ, respectively. c, Superposition of the oxidized (yellow) and reduced (grey) NgTAL, showing selected active site residues and the ligand citrate. The structural change of ligand citrate is accompanied by a redistribution of the two conformers of Arg204 (oxidized 70:30% occupancy, reduced 30:70% occupancy). There is also a subtle repositioning of the active-site residues (for example, Asp17 and water molecules).
Supplementary information
Supplementary Information
This file contains Supplementary Results: Mass spectrometric analyses of NgTAL in the reduced and oxidized state; Supplementary Fig. 1: Chromatographic analysis of NgTAL in the reduced and oxidized state; and Supplementary Fig. 2: Mass spectrometric analysis of post-translational modifications of NgTAL under oxidizing and reducing conditions.
Rights and permissions
About this article
Cite this article
Wensien, M., von Pappenheim, F.R., Funk, LM. et al. A lysine–cysteine redox switch with an NOS bridge regulates enzyme function. Nature 593, 460–464 (2021). https://doi.org/10.1038/s41586-021-03513-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-021-03513-3
This article is cited by
-
Simulation of gap junction formation reveals critical role of Cys disulfide redox state in connexin hemichannel docking
Cell Communication and Signaling (2024)
-
An allosteric redox switch involved in oxygen protection in a CO2 reductase
Nature Chemical Biology (2024)
-
Multiple redox switches of the SARS-CoV-2 main protease in vitro provide opportunities for drug design
Nature Communications (2024)
-
The H163A mutation unravels an oxidized conformation of the SARS-CoV-2 main protease
Nature Communications (2023)
-
Structural and functional analysis of hyper-thermostable ancestral L-amino acid oxidase that can convert Trp derivatives to D-forms by chemoenzymatic reaction
Communications Chemistry (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.