Abstract
Antibiotics that target Gram-negative bacteria in new ways are needed to resolve the antimicrobial resistance crisis1,2,3. Gram-negative bacteria are protected by an additional outer membrane, rendering proteins on the cell surface attractive drug targets4,5. The natural compound darobactin targets the bacterial insertase BamA6—the central unit of the essential BAM complex, which facilitates the folding and insertion of outer membrane proteins7,8,9,10,11,12,13. BamA lacks a typical catalytic centre, and it is not obvious how a small molecule such as darobactin might inhibit its function. Here we resolve the mode of action of darobactin at the atomic level using a combination of cryo-electron microscopy, X-ray crystallography, native mass spectrometry, in vivo experiments and molecular dynamics simulations. Two cyclizations pre-organize the darobactin peptide in a rigid β-strand conformation. This creates a mimic of the recognition signal of native substrates with a superior ability to bind to the lateral gate of BamA. Upon binding, darobactin replaces a lipid molecule from the lateral gate to use the membrane environment as an extended binding pocket. Because the interaction between darobactin and BamA is largely mediated by backbone contacts, it is particularly robust against potential resistance mutations. Our results identify the lateral gate as a functional hotspot in BamA and will allow the rational design of antibiotics that target this bacterial Achilles heel.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
The data that support the findings of this study are available from the corresponding author upon request. The atomic coordinates have been deposited in the RCSB Protein Data Bank and are available under the accession codes 7NRE and 7NRF. The cryo-EM map has been deposited in the Protein Data Bank under accession code 7NRI and EMDB accession code 12546. Mass spectrometry data have been deposited in figshare at https://doi.org/10.6084/m9.figshare.12179784. For the study, data were retrieved from the OMPdb (release Dec 1, 2020). Source data are provided with this paper.
Code availability
The codes developed for β-signal analysis have been deposited at https://github.com/hiller-lab/kaur-jakob-2021.
References
Brown, E. D. & Wright, G. D. Antibacterial drug discovery in the resistance era. Nature 529, 336–343 (2016).
Lewis, K. The science of antibiotic discovery. Cell 181, 29–45 (2020).
Tacconelli, E. et al. Discovery, research, and development of new antibiotics: the WHO priority list of antibiotic-resistant bacteria and tuberculosis. Lancet Infect. Dis. 18, 318–327 (2018).
Epand, R. M., Walker, C., Epand, R. F. & Magarvey, N. A. Molecular mechanisms of membrane targeting antibiotics. Biochim. Biophys. Acta 1858, 980–987 (2016).
Srinivas, N. et al. Peptidomimetic antibiotics target outer-membrane biogenesis in Pseudomonas aeruginosa. Science 327, 1010–1013 (2010).
Imai, Y. et al. A new antibiotic selectively kills Gram-negative pathogens. Nature 576, 459–464 (2019).
Bakelar, J., Buchanan, S. K. & Noinaj, N. The structure of the β-barrel assembly machinery complex. Science 351, 180–186 (2016).
Doyle, M. T. & Bernstein, H. D. Bacterial outer membrane proteins assemble via asymmetric interactions with the BamA β-barrel. Nat. Commun. 10, 3358 (2019).
Gu, Y. et al. Structural basis of outer membrane protein insertion by the BAM complex. Nature 531, 64–69 (2016).
Iadanza, M. G. et al. Lateral opening in the intact β-barrel assembly machinery captured by cryo-EM. Nat. Commun. 7, 12865 (2016).
Konovalova, A., Kahne, D. E. & Silhavy, T. J. Outer membrane biogenesis. Annu. Rev. Microbiol. 71, 539–556 (2017).
Lee, J. et al. Formation of a β-barrel membrane protein is catalyzed by the interior surface of the assembly machine protein BamA. eLife 8, e49787 (2019).
Noinaj, N. et al. Structural insight into the biogenesis of β-barrel membrane proteins. Nature 501, 385–390 (2013).
Kaur, H. et al. Identification of conformation-selective nanobodies against the membrane protein insertase BamA by an integrated structural biology approach. J. Biomol. NMR 73, 375–384 (2019).
Hartmann, J. B., Zahn, M., Burmann, I. M., Bibow, S. & Hiller, S. Sequence-specific solution NMR assignments of the β-barrel insertase BamA to monitor its conformational ensemble at the atomic level. J. Am. Chem. Soc. 140, 11252–11260 (2018).
Ni, D. et al. Structural and functional analysis of the β-barrel domain of BamA from Escherichia coli. FASEB J. 28, 2677–2685 (2014).
Hong, H., Park, S., Jiménez, R. H., Rinehart, D. & Tamm, L. K. Role of aromatic side chains in the folding and thermodynamic stability of integral membrane proteins. J. Am. Chem. Soc. 129, 8320–8327 (2007).
Lee, A. G. Lipid–protein interactions in biological membranes: a structural perspective. Biochim. Biophys. Acta 1612, 1–40 (2003).
Schulz, G. E. The structure of bacterial outer membrane proteins. Biochim. Biophys. Acta 1565, 308–317 (2002).
Gruss, F. et al. The structural basis of autotransporter translocation by TamA. Nat. Struct. Mol. Biol. 20, 1318–1320 (2013).
Chorev, D. S. et al. Protein assemblies ejected directly from native membranes yield complexes for mass spectrometry. Science 362, 829–834 (2018).
Wexler, H. M. Bacteroides: the good, the bad, and the nitty-gritty. Clin. Microbiol. Rev. 20, 593–621 (2007).
Knowles, T. J., Scott-Tucker, A., Overduin, M. & Henderson, I. R. Membrane protein architects: the role of the BAM complex in outer membrane protein assembly. Nat. Rev. Microbiol. 7, 206–214 (2009).
Noinaj, N., Rollauer, S. E. & Buchanan, S. K. The β-barrel membrane protein insertase machinery from Gram-negative bacteria. Curr. Opin. Struct. Biol. 31, 35–42 (2015).
Robert, V. et al. Assembly factor Omp85 recognizes its outer membrane protein substrates by a species-specific C-terminal motif. PLoS Biol. 4, e377 (2006).
Stubenrauch, C. et al. Effective assembly of fimbriae in Escherichia coli depends on the translocation assembly module nanomachine. Nat. Microbiol. 1, 16064 (2016).
Höhr, A. I. C. et al. Membrane protein insertion through a mitochondrial β-barrel gate. Science 359, eaah6834 (2018).
Xiao, L. et al. Structures of the β-barrel assembly machine recognizing outer membrane protein substrates. FASEB J. 35, e21207 (2021).
Tomasek, D. et al. Structure of a nascent membrane protein as it folds on the BAM complex. Nature 583, 473–478 (2020).
Hart, E. M. et al. A small-molecule inhibitor of BamA impervious to efflux and the outer membrane permeability barrier. Proc. Natl Acad. Sci. USA 116, 21748–21757 (2019).
Luther, A. et al. Chimeric peptidomimetic antibiotics against Gram-negative bacteria. Nature 576, 452–458 (2019).
Storek, K. M. et al. Monoclonal antibody targeting the β-barrel assembly machine of Escherichia coli is bactericidal. Proc. Natl Acad. Sci. USA 115, 3692–3697 (2018).
Kaur, H., Grahl, A., Hartmann, J. B. & Hiller, S. Sample preparation and technical setup for NMR spectroscopy with integral membrane proteins. Methods Mol. Biol. 2127, 373–396 (2020).
Kabsch, W. Xds. Acta Crystallogr. D Biol. Crystallogr. 66, 125–132 (2010).
Evans, P. R. & Murshudov, G. N. How good are my data and what is the resolution? Acta Crystallogr. D Biol. Crystallogr. 69, 1204–1214 (2013).
McCoy, A. J. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D Biol. Crystallogr. 60, 2126–2132 (2004).
Schüttelkopf, A. W. & van Aalten, D. M. PRODRG: a tool for high-throughput crystallography of protein-ligand complexes. Acta Crystallogr. D Biol. Crystallogr. 60, 1355–1363 (2004).
Adams, P. D. et al. PHENIX: building new software for automated crystallographic structure determination. Acta Crystallogr. D Biol. Crystallogr. 58, 1948–1954 (2002).
Chen, V. B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr. D Biol. Crystallogr. 66, 12–21 (2010).
DeLano, W. L. Pymol: an open-source molecular graphics tool. CCP4 Newsl. Prot. Crystallogr. 40, 82–92 (2002).
Mastronarde, D. N. Automated electron microscope tomography using robust prediction of specimen movements. J. Struct. Biol. 152, 36–51 (2005).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Pettersen, E. F. et al. UCSF Chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Afonine, P. V. et al. Real-space refinement in PHENIX for cryo-EM and crystallography. Acta Crystallogr. D Struct. Biol. 74, 531–544 (2018).
Vranken, W. F. et al. The CCPN data model for NMR spectroscopy: development of a software pipeline. Proteins 59, 687–696 (2005).
Piñeiro, Á. et al. AFFINImeter: A software to analyze molecular recognition processes from experimental data. Anal. Biochem. 577, 117–134 (2019).
Kemmer, G. & Keller, S. Nonlinear least-squares data fitting in Excel spreadsheets. Nat. Protocols 5, 267–281 (2010).
Zhao, H., Piszczek, G. & Schuck, P. SEDPHAT—a platform for global ITC analysis and global multi-method analysis of molecular interactions. Methods 76, 137–148 (2015).
Sievers, F. et al. Fast, scalable generation of high-quality protein multiple sequence alignments using Clustal Omega. Mol. Syst. Biol. 7, 539 (2011).
Pei, J. & Grishin, N. V. AL2CO: calculation of positional conservation in a protein sequence alignment. Bioinformatics 17, 700–712 (2001).
Datsenko, K. A. & Wanner, B. L. One-step inactivation of chromosomal genes in Escherichia coli K-12 using PCR products. Proc. Natl Acad. Sci. USA 97, 6640–6645 (2000).
Bennion, D., Charlson, E. S., Coon, E. & Misra, R. Dissection of β-barrel outer membrane protein assembly pathways through characterizing BamA POTRA 1 mutants of Escherichia coli. Mol. Microbiol. 77, 1153–1171 (2010).
Ruiz, N., Wu, T., Kahne, D. & Silhavy, T. J. Probing the barrier function of the outer membrane with chemical conditionality. ACS Chem. Biol. 1, 385–395 (2006).
Aoki, S. K. et al. Contact-dependent growth inhibition requires the essential outer membrane protein BamA (YaeT) as the receptor and the inner membrane transport protein AcrB. Mol. Microbiol. 70, 323–340 (2008).
Marty, M. T. et al. Bayesian deconvolution of mass and ion mobility spectra: from binary interactions to polydisperse ensembles. Anal. Chem. 87, 4370–4376 (2015).
Huang, J. et al. CHARMM36m: an improved force field for folded and intrinsically disordered proteins. Nat. Methods 14, 71–73 (2017).
Vanommeslaeghe, K. et al. CHARMM general force field: a force field for drug-like molecules compatible with the CHARMM all-atom additive biological force fields. J. Comput. Chem. 31, 671–690 (2010).
Jo, S., Kim, T., Iyer, V. G. & Im, W. CHARMM-GUI: a web-based graphical user interface for CHARMM. J. Comput. Chem. 29, 1859–1865 (2008).
Lomize, M. A., Pogozheva, I. D., Joo, H., Mosberg, H. I. & Lomize, A. L. OPM database and PPM web server: resources for positioning of proteins in membranes. Nucleic Acids Res. 40, D370–D376 (2012).
Van Der Spoel, D. et al. GROMACS: fast, flexible, and free. J. Comput. Chem. 26, 1701–1718 (2005).
Jorgensen, W. L., Chandrasekhar, J., Madura, J. D., Impey, R. W. & Klein, M. L. Comparison of simple potential functions for simulating liquid water. J. Chem. Phys. 79, 926–935 (1983).
Essmann, U. et al. A smooth particle mesh Ewald method. J. Chem. Phys. 103, 8577–8593 (1995).
Hoover, W. G. Canonical dynamics: equilibrium phase–space distributions. Phys. Rev. A 31, 1695–1697 (1985).
Nose, S. A unified formulation of the constant temperature molecular-dynamics methods. J. Chem. Phys. 81, 511–519 (1984).
Parrinello, M. & Rahman, A. Polymorphic transitions in single crystals—a new molecular-dynamics method. J. Appl. Phys. 52, 7182–7190 (1981).
Castillo, N., Monticelli, L., Barnoud, J. & Tieleman, D. P. Free energy of WALP23 dimer association in DMPC, DPPC, and DOPC bilayers. Chem. Phys. Lipids 169, 95–105 (2013).
Paramasivam, N., Habeck, M. & Linke, D. Is the C-terminal insertional signal in Gram-negative bacterial outer membrane proteins species-specific or not? BMC Genomics 13, 510 (2012).
Tsirigos, K. D., Bagos, P. G. & Hamodrakas, S. J. OMPdb: a database of β-barrel outer membrane proteins from Gram-negative bacteria. Nucleic Acids Res. 39, D324–D331 (2011).
Hunter, J. D. Matplotlib: a 2D graphics environment. Comput. Sci. Eng. 9, 90–95 (2007).
Tareen, A. & Kinney, J. B. Logomaker: beautiful sequence logos in Python. Bioinformatics 36, 2272–2274 (2020).
Acknowledgements
We thank H. Bernstein and C. Bieniossek for expression plasmids, and T. Sharpe, T. Müntener and the Biozentrum Bio-EM lab for scientific support and discussions. The Scientific Computing Center at the University of Basel (sciCORE) and the National Supercomputing Centre Singapore (NSCC) are acknowledged for providing computational resources. This work was supported by the Swiss National Science Foundation (grants 167125, 185388 and 187170 to S.H. and R’Equip 177084 to T.M.), AntiResist: new approaches to combat antibiotic-resistant bacteria (51AU40_180541), the Medical Research Council (program grant MR/N020413/1 awarded to C.V.R.), the Bioinformatics Institute (BII) A*STAR and NRF (NRF2017NRF-CRP001-027 to P.J.B. and J.K.M.), and the NIH (grant P01 AI118687 to K.L.).
Author information
Authors and Affiliations
Contributions
C.V.R., K.L., T.M. and S.H. designed the study and supervised experiments. R.G. and Y.I. performed the microbiology experiments. J.R.B. performed the mass spectrometry experiments. H.K. and R.P.J. performed all other experiments. J.K.M. and P.J.B. ran simulations. E.A. performed sequence analysis. All authors analysed data and discussed the findings. H.K., K.L., T.M. and S.H. wrote the manuscript. All authors edited and approved the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature thanks Harris Bernstein, Susan Buchanan, John Rubinstein and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Cryo-electron microscopy structure of the BAM–darobactin complex.
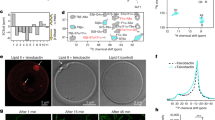
a, Flow chart of data processing to generate the structure (see Methods). b, Purified BAM–darobactin sample used for cryo-EM structure determination analysed on SDS–PAGE. This experiment was repeated at least three times independently with similar results. c, Representative electron micrograph of BAM–darobactin. This experiment was repeated at least three times independently with similar results. d, Selected examples of 2D classes from cryoSPARC. e, Viewing direction distribution plot for the final 3D reconstruction. f, Fourier shell correlation (FSC) curves for unmasked, spherical, loose and tight masks, and corrected FSC curve for the final reconstruction, yielding a gold standard FSC resolution of 3.03 Å. g, Local resolution variations of the EM reconstruction. POTRA domains 1 and 2 are at a local resolution below 4.5 Å and are visualized only at a lower contour level where micelle density obscures the view onto the BamA barrel. h, Plot of directional FSC (red; mean ± s.d.) and histogram of per angle FSC (blue). FSC curve indicates a resolution of 3.15 Å. i, Overview of the cryo-EM reconstruction of the BAM complex. BAM is shown in ribbon representation and the coulomb potential map as blue mesh. Note that the density of POTRA domains P1 and P2 is below the display threshold chosen here because of motional averaging. j–n, Expanded local views, showing the map around selected atoms in stick representation from the directions and viewpoints indicated by arrows and letters in i.
Extended Data Fig. 2 Structural details of the cryo-EM and crystal structures of BAM.
a, Superposition of the BAM–darobactin cryo-EM structure (salmon) with the ligand-free BAM crystal structure (green, PDB 5D0O). b, Superposition of POTRA domains P1–P5 and the individual components BamB–BamE, as indicated. The dashed horizontal line indicates the pivot between P2 and P3 around which P1 and P2 are rotated by rigid-body movement. c, Superposition of the BamA β-barrel–darobactin crystal structure (salmon) with a closed-gate BamA β-barrel crystal structure (cyan, PDB 4N75). d, Crystallographic omit map for darobactin bound at the lateral gate region of the BamA β-barrel after refinement of the model without darobactin. The 2mFo − DFc map is shown at 1σ in slate and the mFo − DFc difference map at ±3σ level in green and red. Top, overview of an entire BamA barrel. Bottom left and right, expanded views without and with overlay of the refined model coordinates, respectively. The cyclizations of darobactin can clearly be observed at 2.3 Å resolution. e, Omit map for strands β1 (top) and β16 (bottom) of the BamA β-barrel visualized as in d. f, Superposition of the cryo-EM (Bordeaux and white for BamA and darobactin, respectively) and X-ray structures (salmon and blue). g, As in f for the ligand darobactin only.
Extended Data Fig. 3 Comparison of BamA β-barrel conformations in aqueous solution.
a, Comparison of 2D [15N,1H]-TROSY fingerprint spectra of different BamA preparations in LDAO. Left, overlay of BamA-β fingerprint spectra in the absence and presence of darobactin. Middle, overlay of fingerprint spectra of BamA-β–nanobodyF7 and BamA-β–darobactin. Right, overlay of fingerprint spectra of BamA+9 (BamA-β + C-terminal extension MENVALDFS) and BamA-β–darobactin complex. Bottom panels show expanded views of boxed areas in main spectra. b, Backbone amide chemical shift perturbations between the fingerprint spectra of BamA-β with and without darobactin (left, black), Bam-β–nanobodyF7 in comparison with BamA–darobactin (middle, blue) and BamA+9 in comparison with BamA–darobactin (right, purple). The dotted lines indicate the average chemical shift perturbation (CSP), which can be interpreted as a measure of dissimilarity between two spectra. c, Top, structures of the BamA β-barrel in various conformations of the gate region. Bottom, expanded views of part of the backbone showing hydrogen bonds between β1 and β16 or darobactin. From left to right: open gate (6QGW, red); closed gate (6QGX, blue); BamA+9 (6FSU, purple); and BamA–darobactin complex. d, In vivo functional assay of BamA barrel mutants and C-terminal extensions using JCM166 cells in the absence and presence of arabinose. fl-BamAMENVALDFS and fl-BamA serve as a negative and positive controls, respectively.
Extended Data Fig. 4 ITC of BamA β-barrel in detergent micelles and its variants titrated with darobactin.
Experiments were repeated independently twice with similar results.
Extended Data Fig. 5 MD simulations of BamA β-barrel.
a, Representative snapshots of lipid PE molecule (left) and PG molecule (right) anchored by Ile430 and Leu780 in the gate region. b, The most dominant conformation of BamA–darobactin showing contacts consistently observed between the BamA β-barrel and darobactin throughout the simulation sampling. c, Partial densities of all lipids (top-down view of the membrane); the white arrow highlights the darobactin-binding region. d, Partial densities of lipid phosphate groups (side view of the membrane). e–g, Structural drift and fluctuations of key β-strands around the darobactin-binding site. Time-dependent r.m.s.d. measured with respect to the initial structure for backbone atoms of β-strands, after performing a least-squares fit. The resulting r.m.s.d. is shown for β16 in ligand-free BamA (e), β16 in BamA–darobactin complex (f), and hairpin β1/β2 in BamA–darobactin complex (g).
Extended Data Fig. 6 Interaction of BAM complex with lipids in the absence and presence of bound darobactin.
a, Mass spectra of the BAM complex with different lipids (top, CL; middle, PG; bottom, PE). b, Deconvolution of the mass spectra in a indicates that all the subcomplexes have lipids bound. c, Relative intensities of lipid binding peaks from a suggest that the negatively charged PG and CL have higher affinity than PE. d, Mass spectra of BAM complex with lipids and darobactin. Bottom, PE and darobactin; top, PG and darobactin. Below, expanded view of a section of the 23+ charge state highlights the bound peaks; bar charts show relative ratio of darobactin binding. No significant increase in darobactin binding is observed in these two cases, suggesting that PE and PG lipids do not affect darobactin binding. e, Mass spectra of BAM complex with lipid mixtures (bottom, PE and CL; top, PG and CL) and darobactin. Below, expanded view of a section of the 23+ charge state with lipids, darobactin and their various combination binding peaks highlighted. Bar charts of relative peak intensities indicate that darobactin bound with CL is observed to a greater extent than bound alone or bound with PE or PG. This increase is even higher for 2 × CL bound species and is slightly lower for PE or PG bound to 1 × CL species. However, no change in darobactin binding is observed for PE and PG. Bars (c–e) represent mean ± s.d., points show data from three independent experiments. *P < 0.05; **P < 0.01; ***P < 0.001; NS, not significant; two-tailed unpaired Student’s t-test. Exact P values are indicated in the figures.
Extended Data Fig. 7 MD simulations of the effect of darobactin-resistance mutations on the BamA-β–darobactin interaction.
a, Left, representative snapshots with expanded views from a simulation with strain 1 bound to darobactin. Mutations G429V and G807V are shown as yellow spheres at the α-carbon position. Right, time-dependent r.m.s.d. of β16 backbone atoms relative to the initial structure. b, As in a for strain 2 (mutations E435K and G443D, cyan) and r.m.s.d. of hairpin β1/β2. c, As in a for strain 3 (mutations T434A, Q445P and A705T, green) and r.m.s.d. of hairpin β1/β2. In each panel, protein is shown in cartoon representation, darobactin as sticks (blue, carbon; red, oxygen; white, proton; navy, nitrogen). Hydrogen bonds are shown as black dotted lines.
Extended Data Fig. 8 Relatedness of predicted Gram-negative BamA proteins.
a, Sequence alignment of BamA sequence from Gram-negative bacteria. The region highlighted in yellow is the predicted interaction site of darobactin in sheet β1. Right, these six-amino acid sequences from A. baumannii and B. fragilis were substituted into E. coli BamA for in vivo assays. b, Phylogenetic tree of full-length BamA sequences from various species of Gram-negative bacteria. Colours indicate branches belonging to the species specified next to the branch. Multiple alignments for the tree were carried out using CLUSTAL-W and the phylogenetic tree was derived using SEAVIEW software. c, Topology plot of BamA from E. coli with bound darobactin (blue). For the chimeric mutants, the red amino acids were exchanged with the local sequence from either A. baumannii or B. fragilis.
Extended Data Fig. 9 Comparison of BamA structures involved in molecular interactions and analysis of β-signals.
a, Overlay of the BamA subunit from the BAM–darobactin complex (salmon–blue; this work) with a BamA engaged with a substrate in a late-stage insertion intermediate state (green; PDB 6V05). The substrate has been omitted in this panel and the structures have been globally aligned to the protein backbone. b, Expanded view of the BamA β-barrel from a with strand β16subs of the substrate shown in purple. It is paired to strand β1mem of the catalytic BamA. Bold green and red arrows depict the directions of strand β1, forming an ~90° angle. c, Expanded view of the gate region indicating the spatial proximity of the substrate and darobactin interaction sites and their relative rotation of ~90°. d–f, Comparison of the register of β16 complementation to b1. d, In BamA–darobactin (salmon), residue Ile806 pairs with Tyr432. e, In the late-stage intermediate, Ile806 of the substrate BamAsubs (purple) pairs with Phe428 in catalyst BamAmem (green), corresponding to a register shift of 4. f, Hypothetical position of the four C-terminal residues of substrate BamA, which are not resolved in the available electron density. When paired to β1mem in canonical antiparallel β-strand conformation, they locate exactly at the darobactin-binding site, with the C-terminal Trp810 at the position of Phe7 of darobactin. g, Frequency logo of known and putative β-signals from bacterial OMPs, coloured by amino acid type. Numbering refers to distance from the C terminus. h, Distribution of log-likelihood scores in three sets of sequences, as indicated. The score obtained by the darobactin sequence is indicated by a blue line. The percentile rank of darobactin within each of the three sets is given in parentheses. i, j, As in g, h, but based on amino acid chemistry. H, hydrophobic and non-polar residue; A, aromatic; N, neutral; C, charged; P, polar non-charged.
Extended Data Fig. 10 Interaction of β-signal peptides with BamA β-barrel in detergent micelles by ITC.
The first four panels show direct titration of each of the ten-amino acid β-signal peptides of BamA, BtuB, FhuA, and OmpF to the BamA β-barrel. The next five panels show a competition experiment with darobactin titrated to the BamA β-barrel in the presence of ten-amino acid β-signal peptides of OmpT (0.7 mM), BtuB (2.6 mM), OmpF (1.4 mM) and FhuA (1.1 mM) and a β-consensus-peptide (1.2 mM). The results from fitting of the data to the competition model are given in Supplementary Table 7.
Supplementary information
Supplementary Information
This file contains Supplementary Tables 1-8 and Supplementary Fig. 1.
Rights and permissions
About this article
Cite this article
Kaur, H., Jakob, R.P., Marzinek, J.K. et al. The antibiotic darobactin mimics a β-strand to inhibit outer membrane insertase. Nature 593, 125–129 (2021). https://doi.org/10.1038/s41586-021-03455-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-021-03455-w
This article is cited by
-
High-throughput screening of BAM inhibitors in native membrane environment
Nature Communications (2023)
-
Structural basis of BAM-mediated outer membrane β-barrel protein assembly
Nature (2023)
-
Systematic mining of the human microbiome identifies antimicrobial peptides with diverse activity spectra
Nature Microbiology (2023)
-
Hydroxytryptophan biosynthesis by a family of heme-dependent enzymes in bacteria
Nature Chemical Biology (2023)
-
Modular synthesis of clickable peptides via late-stage maleimidation on C(7)-H tryptophan
Nature Communications (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.