Abstract
Bile acids are lipid-emulsifying metabolites synthesized in hepatocytes and maintained in vivo through enterohepatic circulation between the liver and small intestine1. As detergents, bile acids can cause toxicity and inflammation in enterohepatic tissues2. Nuclear receptors maintain bile acid homeostasis in hepatocytes and enterocytes3, but it is unclear how mucosal immune cells tolerate high concentrations of bile acids in the small intestine lamina propria (siLP). CD4+ T effector (Teff) cells upregulate expression of the xenobiotic transporter MDR1 (encoded by Abcb1a) in the siLP to prevent bile acid toxicity and suppress Crohn’s disease-like small bowel inflammation4. Here we identify the nuclear xenobiotic receptor CAR (encoded by Nr1i3) as a regulator of MDR1 expression in T cells that can safeguard against bile acid toxicity and inflammation in the mouse small intestine. Activation of CAR induced large-scale transcriptional reprogramming in Teff cells that infiltrated the siLP, but not the colon. CAR induced the expression of not only detoxifying enzymes and transporters in siLP Teff cells, as in hepatocytes, but also the key anti-inflammatory cytokine IL-10. Accordingly, CAR deficiency in T cells exacerbated bile acid-driven ileitis in T cell-reconstituted Rag1−/− or Rag2−/− mice, whereas pharmacological activation of CAR suppressed it. These data suggest that CAR acts locally in T cells that infiltrate the small intestine to detoxify bile acids and resolve inflammation. Activation of this program offers an unexpected strategy to treat small bowel Crohn’s disease and defines lymphocyte sub-specialization in the small intestine.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
RNA-seq data for mouse (GSE149218) and human (GSE149219) CD4+ T cells described here are publicly available at the NCBI Gene Expression Omnibus (GEO) repository (series no. GSE149220). Sequence and other information related to shRNAmir retroviral constructs used in the pooled in vivo RNAi screen (related to Fig. 1, Extended Data Fig. 1), as well as complete gene lists derived from differential gene expression analyses of wild-type and Nr1i3−/− Teff cells (related to Figs. 2a–d, 3a–c) and statistical analysis of differences between in vitro-polarized wild-type and Nr1i3−/− CD4+ T cells (related to Fig. 3h, i), are available in the Supplementary Information. Other relevant data are available from the corresponding authors upon reasonable request. Previously generated ChIP–seq and RNA-seq datasets analysed in this study include GEO accession numbers GSE104734,GSE112199,GSE92940 andGSE21670. Datasets obtained from the Molecular Signature Database (MSigDB; https://www.gsea-msigdb.org/) derive from GEO accession numbers GSE14308 andGSE21379. Source data are provided with this paper.
References
Hofmann, A. F. & Hagey, L. R. Key discoveries in bile acid chemistry and biology and their clinical applications: history of the last eight decades. J. Lipid Res. 55, 1553–1595 (2014).
Poupon, R., Chazouillères, O. & Poupon, R. E. Chronic cholestatic diseases. J. Hepatol. 32 (Suppl. 1), 129–140 (2000).
Arab, J. P., Karpen, S. J., Dawson, P. A., Arrese, M. & Trauner, M. Bile acids and nonalcoholic fatty liver disease: Molecular insights and therapeutic perspectives. Hepatology 65, 350–362 (2017).
Cao, W. et al. The xenobiotic transporter Mdr1 enforces T cell homeostasis in the presence of intestinal bile acids. Immunity 47, 1182–1196 (2017).
Lazar, M. A. Maturing of the nuclear receptor family. J. Clin. Invest. 127, 1123–1125 (2017).
Chen, R. et al. In vivo RNA interference screens identify regulators of antiviral CD4+ and CD8+ T cell differentiation. Immunity 41, 325–338 (2014).
Ludescher, C. et al. Detection of activity of P-glycoprotein in human tumour samples using rhodamine 123. Br. J. Haematol. 82, 161–168 (1992).
Zhang, J., Huang, W., Qatanani, M., Evans, R. M. & Moore, D. D. The constitutive androstane receptor and pregnane X receptor function coordinately to prevent bile acid-induced hepatotoxicity. J. Biol. Chem. 279, 49517–49522 (2004).
Cerveny, L. et al. Valproic acid induces CYP3A4 and MDR1 gene expression by activation of constitutive androstane receptor and pregnane X receptor pathways. Drug Metab. Dispos. 35, 1032–1041 (2007).
Wei, P., Zhang, J., Egan-Hafley, M., Liang, S. & Moore, D. D. The nuclear receptor CAR mediates specific xenobiotic induction of drug metabolism. Nature 407, 920–923 (2000).
Evans, R. M. & Mangelsdorf, D. J. Nuclear receptors, RXR, and the big bang. Cell 157, 255–266 (2014).
Staudinger, J. L. et al. The nuclear receptor PXR is a lithocholic acid sensor that protects against liver toxicity. Proc. Natl Acad. Sci. USA 98, 3369–3374 (2001).
Ostanin, D. V. et al. T cell transfer model of chronic colitis: concepts, considerations, and tricks of the trade. Am. J. Physiol. Gastrointest. Liver Physiol. 296, G135–G146 (2009).
Arnold, M. A. et al. Colesevelam and colestipol: novel medication resins in the gastrointestinal tract. Am. J. Surg. Pathol. 38, 1530–1537 (2014).
Dawson, P. A., Lan, T. & Rao, A. Bile acid transporters. J. Lipid Res. 50, 2340–2357 (2009).
Cui, J. Y. & Klaassen, C. D. RNA-seq reveals common and unique PXR- and CAR-target gene signatures in the mouse liver transcriptome. Biochim. Biophys. Acta 1859, 1198–1217 (2016).
Niu, B. et al. In vivo genome-wide binding interactions of mouse and human constitutive androstane receptors reveal novel gene targets. Nucleic Acids Res. 46, 8385–8403 (2018).
De Calisto, J. et al. T-cell homing to the gut mucosa: general concepts and methodological considerations. Methods Mol. Biol. 757, 411–434 (2012).
Maglich, J. M. et al. Identification of a novel human constitutive androstane receptor (CAR) agonist and its use in the identification of CAR target genes. J. Biol. Chem. 278, 17277–17283 (2003).
Ramesh, R. et al. Pro-inflammatory human Th17 cells selectively express P-glycoprotein and are refractory to glucocorticoids. J. Exp. Med. 211, 89–104 (2014).
Moore, L. B. et al. Pregnane X receptor (PXR), constitutive androstane receptor (CAR), and benzoate X receptor (BXR) define three pharmacologically distinct classes of nuclear receptors. Mol. Endocrinol. 16, 977–986 (2002).
Karwacz, K. et al. Critical role of IRF1 and BATF in forming chromatin landscape during type 1 regulatory cell differentiation. Nat. Immunol. 18, 412–421 (2017).
Gagliani, N. et al. Coexpression of CD49b and LAG-3 identifies human and mouse T regulatory type 1 cells. Nat. Med. 19, 739–746 (2013).
Korn, T., Bettelli, E., Oukka, M. & Kuchroo, V. K. IL-17 and Th17 cells. Annu. Rev. Immunol. 27, 485–517 (2009).
Maynard, C. L. et al. Regulatory T cells expressing interleukin 10 develop from Foxp3+ and Foxp3− precursor cells in the absence of interleukin 10. Nat. Immunol. 8, 931–941 (2007).
Sano, T. et al. An IL-23R/IL-22 circuit regulates epithelial serum amyloid A to promote local effector Th17 responses. Cell 163, 381–393 (2015).
Wan, Q. et al. Cytokine signals through PI-3 kinase pathway modulate Th17 cytokine production by CCR6+ human memory T cells. J. Exp. Med. 208, 1875–1887 (2011).
Yadava, K. et al. Natural Tr1-like cells do not confer long-term tolerogenic memory. eLife 8, e44821 (2019).
Barrat, F. J. et al. In vitro generation of interleukin 10-producing regulatory CD4+ T cells is induced by immunosuppressive drugs and inhibited by T helper type 1 (Th1)- and Th2-inducing cytokines. J. Exp. Med. 195, 603–616 (2002).
Song, X. et al. Microbial bile acid metabolites modulate gut RORγ+ regulatory T cell homeostasis. Nature 577, 410–415 (2020).
Knott, S. R. V. et al. A computational algorithm to predict shRNA potency. Mol. Cell 56, 796–807 (2014).
Fellmann, C. et al. An optimized microRNA backbone for effective single-copy RNAi. Cell Rep. 5, 1704–1713 (2013).
Kim, J. J., Shajib, M. S., Manocha, M. M. & Khan, W. I. Investigating intestinal inflammation in DSS-induced model of IBD. J. Vis. Exp. 60, 3678 (2012).
Berg, D. J. et al. Enterocolitis and colon cancer in interleukin-10-deficient mice are associated with aberrant cytokine production and CD4+ TH1-like responses. J. Clin. Invest. 98, 1010–1020 (1996).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Zhang, Y. et al. Model-based analysis of ChIP-Seq (MACS). Genome Biol. 9, R137 (2008).
Robinson, J. T. et al. Integrative genomics viewer. Nat. Biotechnol. 29, 24–26 (2011).
Patro, R., Duggal, G., Love, M. I., Irizarry, R. A. & Kingsford, C. Salmon provides fast and bias-aware quantification of transcript expression. Nat. Methods 14, 417–419 (2017).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102, 15545–15550 (2005).
Mootha, V. K. et al. PGC-1α-responsive genes involved in oxidative phosphorylation are coordinately downregulated in human diabetes. Nat. Genet. 34, 267–273 (2003).
Durant, L. et al. Diverse targets of the transcription factor STAT3 contribute to T cell pathogenicity and homeostasis. Immunity 32, 605–615 (2010).
Ritchie, M. E. et al. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 43, e47 (2015).
Wei, G. et al. Global mapping of H3K4me3 and H3K27me3 reveals specificity and plasticity in lineage fate determination of differentiating CD4+ T cells. Immunity 30, 155–167 (2009).
Yusuf, I. et al. Germinal center T follicular helper cell IL-4 production is dependent on signaling lymphocytic activation molecule receptor (CD150). J. Immunol. 185, 190–202 (2010).
Suino, K. et al. The nuclear xenobiotic receptor CAR: structural determinants of constitutive activation and heterodimerization. Mol. Cell 16, 893–905 (2004).
Acknowledgements
We thank Core Facility staff at Scripps Florida and Baylor College of Medicine for technical support, and P. Dawson and A. Rao for discussions. This work was supported by Scripps Florida via the State of Florida (M.S.S.), the R. P. Doherty Jr.–Welch Chair in Science Q‐0022 at Baylor College of Medicine (D.D.M.), National Institute of Health grants R21AI119728 (M.S.S.), R01AI118931 (M.S.S.), U19AI109976 (M.E.P.), P01AI145815 (M.E.P.), R01DK113789 (C.T.W.) and F30DK127865 (B.F.), and Crohn’s and Colitis Foundation (CCF) grants (Senior Research Award no. 422515 (M.S.S.) and Litwin IBD pioneer award no. 572171 (H.W.)).
Author information
Authors and Affiliations
Contributions
Study design: M.L.C., X.H., H.W., J.S., H.P., B.F., L.A.S., D.J.K., A.R.-P., D.A.S., C.T.W., M.E.P., D.D.M. and M.S.S. Data generation: M.L.C., X.H., H.W., C.H., Y.L., J.S., A.E. and S.A.M. Bioinformatics: M.L.C., H.D., G.W., A.R.-P. and M.E.P. Manuscript: M.L.C., X.H., C.H., A.E., D.J.K., A.R.-P., C.T.W., M.E.P., D.D.M. and M.S.S. Principal investigators: L.A.S., D.J.K., A.R.-P., D.A.S., C.T.W., M.E.P., D.D.M. and M.S.S.
Corresponding authors
Ethics declarations
Competing interests
M.S.S. is a consultant to Sigilon Therapeutics and Sage Therapeutics.
Additional information
Peer review information Nature thanks Richard Blumberg and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Nuclear receptor-dependent regulation of effector T cell persistence and MDR1 expression in vivo.
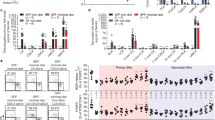
a, Left, abundance of shRNAmirs in ex vivo-isolated spleen and in vitro-transduced (input) Teff cells. shRNAmirs with one normalized read or fewer in both ex vivo spleen and input Teff cell pools were considered ‘poorly represented’ (highlighted green). Well-represented shRNAmirs with ≤10-fold change between ex vivo spleen and input Teff cell pools (between blue lines) were considered for downstream analysis. Right, abundance of shRNAmirs, filtered for minimal effects on in vivo Teff cell persistence, in ex vivo-isolated MDR1hi (Rh123lo) and MDR1lo (Rh123hi) siLP Teff cells. b, Mean ± s.e.m. log2[fold change in abundance] of shRNAmirs against Cd19 (n = 3), Abcb1a (n = 2), Nr1i3 (n = 5), Thra (n = 6) and Esrra (n = 3) in FVB/N wild-type Rh123lo (MDR1hi) versus Rh123hi (MDR1lo) Teff cells (sorted as in Fig. 1a) recovered from spleens or siLP of transferred FVB/N.Rag1−/− mice. a, b, Data incorporate shRNAmir abundance, determined by DNA-seq, in two independent screens using pooled spleens and siLP from 10 transferred FVB/N.Rag1−/− mice per screen. c, Top, schematic of the mouse Nr1i3 locus. Positions of seed sequences for Nr1i3-specific shRNAmirs are shown. Untranslated regions (UTRs) are indicated by open boxes. Bottom, ex vivo Rh123 efflux, determined by flow cytometry, in FVB/N wild-type Teff cells transduced with ametrine-expressing retroviruses containing control (shCD8a) or one of five Nr1i3-specific shRNAmirs re-isolated from spleens of FVB/N.Rag1−/− mice six weeks after T cell transfer. Rh123 efflux in transduced (ametrine+; red) cells is overlaid with that in bystander untransduced (ametrine−; blue) Teff cells from the same mouse. Background Rh123 efflux in untransduced Teff cells treated with the MDR1 inhibitor elacridar is shown in grey. Representative of at least ten mice per group analysed in three independent experiments. d, Mean ± s.e.m. normalized ex vivo Rh123 efflux in FVB/N wild-type spleen Teff cells expressing control (shCd8a; n = 11) or Nr1i3-specific shRNAmirs (shNr1i3.1 (n = 10), shNr1i3.2 (n = 10), shNr1i3.3 (n = 12), shNr1i3.4 (n = 10) and shNr1i3.5 (n = 10)), determined by flow cytometry as in c. Rh123 efflux was normalized to control shCd8a-expressing Teff cells after calculating the change (Δ) in Rh123 mean fluorescence intensity (MFI) between bystander transduced (ametrine+) and untransduced (ametrine−) Teff cells. *P = 0.049, **P = 0.003, ***P = 0.0006, one-way ANOVA with Dunnett’s correction for multiple comparisons. e, Mean ± s.e.m. relative Abcb1a, Nr1i3, and Cyp2b10 expression determined by qPCR in FVB/N spleen Teff cells FACS-sorted from FVB/N.Rag1−/− recipient mice expressing either a negative control shRNAmir against CD8 (shCd8a; n = 8), or the indicated shRNAmirs against CAR (shNr1i3.1 (n = 8), shNr1i3.2 (n = 8), shNr1i3.3 (n = 8), shNr1i3.4 (n = 8) and shNr1i3.5 (n = 8). P < 0.05 versus shCd8a-expressing cells (one-way ANOVA with Tukey’s correction for multiple comparisons) are shown. f, Median log2[fold change] in shRNAmir abundance between FVB/N wild-type ex vivo-isolated spleen and in vitro-transduced (input) Teff cells. a, f, shRNAmir abundance reflects the mean number of normalized reads, by DNA-seq, in the indicated Teff subsets obtained in two independent screens, each using cells transferred into 10 FVB/N.Rag1−/− mice. g, Ex vivo Rh123 efflux, determined by flow cytometry, in CD45.1 wild-type (B6; blue) or CD45.2 (red) CAR-deficient (B6.Nr1i3−/−), PXR-deficient (B6.Nr1i2−/−) or CAR/PXR double-deficient (B6.Nr1i3−/−Nr1i2−/−) Teff cells (gated as in Extended Data Fig. 6a) recovered from spleens of B6.Rag1−/− mice six weeks after co-transfer of mixed naive T cells. Background Rh123 efflux in CD45.1 B6 Teff cells treated with the MDR1 inhibitor elacridar is shown in grey. Representative of a total of seven mice per group analysed in two independent experiments. h, Mean ± s.e.m. normalized Rh123 efflux in congenically transferred CD45.1 wild-type (B6; n = 7) or CD45.2 CAR-deficient (B6.Nr1i3−/−; n = 7), PXR-deficient (B6.Nr1i2−/−; n = 7) or CAR/PXR double-deficient (B6.Nr1i3−/−Nr1i2−/−; n = 7) spleen Teff cells, determined by flow cytometry as in g. *P < 0.019, **P < 0.002, one-way ANOVA with Tukey’s correction for multiple comparisons. i, Mean ± s.e.m. relative Abcb1a expression determined by ex vivo qPCR in CD45.1 wild-type (B6; n = 5) or CD45.2 CAR-deficient (B6.Nr1i3−/−; n = 5), PXR-deficient (B6.Nr1i2−/−; n = 4) or CAR/PXR double-deficient (B6.Nr1i3−/−Nr1i2−/−; n = 5) spleen Teff cells as in g. P < 0.05 versus wild-type (B6) Teff cells (one-way ANOVA with Tukey’s correction for multiple comparisons) are shown.
Extended Data Fig. 2 Inhibition of bile acid reabsorption rescues ileitis induced by CAR-deficient T cells in reconstituted Rag−/− mice.
a, Mean ± s.e.m. weight loss of co-housed B6.Rag2−/− mice transplanted with wild-type (B6; blue; n = 15) or CAR-deficient (B6.Nr1i3−/−; red; n = 13) naive CD4+ T cells and treated with 2% (w:w) cholestyramine (CME) beginning three weeks after T cell transfer. NS, not significant by two-way ANOVA. b, Top, H&E-stained sections of colons or terminal ilea from B6.Rag2−/− mice reconstituted with wild-type or CAR-deficient T cells and treated with or without CME as in a. Representative of 12 mice per group analysed in 3 independent experiments. Magnification, 10×; scale bars, 100 μm. Bottom, mean ± s.e.m. histology scores (n = 12) for colons or terminal ilea as in a. NS, not significant by unpaired two-tailed Student’s t-test. c, Mean ± s.e.m. weight loss (n = 16 per group) of co-housed B6.Rag1−/− mice with or without ASBT (encoded by Slc10a2) after reconstitution with wild-type (B6) or CAR-deficient (B6.Nr1i3−/−) naive CD4+ T cells. **P = 0.005 (CAR-deficient T cell transfer into control versus Slc10a2−/−Rag1−/− recipients), two-way ANOVA. d, Top, H&E-stained colon or terminal ileum sections from control or ASBT-deficient B6.Rag1−/− mice reconstituted with wild-type or CAR-deficient T cells as in c. Representative of five mice per group. Bottom, mean ± s.e.m. histology scores for colons or terminal ilea (n = 5) as above. P values (one-way ANOVA with Tukey’s correction for multiple comparisons) are listed. NS, not significant.
Extended Data Fig. 3 Shared features of CAR-dependent gene expression in mucosal T cells and hepatocytes.
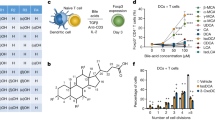
a, Overlap between genes induced in B6 wild-type mouse hepatocytes by in vivo treatment with TC or PCN, relative to vehicle (CO, corn oil). b, Enrichment of genes induced by TC, but not PCN, in mouse hepatocytes (as in a), within those reduced in CAR-deficient (B6.Nr1i3−/−) versus wild-type (B6) siLP Teff cells from week-3 congenically co-transferred Rag1−/− mice (as in Fig. 2a–c). Nominal P values compare observed NESs to the null distribution for 1,000 permutations. c, Differential gene expression, determined by DESeq2 and shown as a volcano plot, between CAR-deficient (B6.Nr1i3−/−) and wild-type (B6) siLP Teff cells re-isolated from transferred B6.Rag1−/− mice, as in Fig. 2a. Genes induced by TC, but not PCN, treatment in mouse hepatocytes (as in a; purple), bound by CAR in ChIP–seq analysis of hepatocytes from TC-treated mice (blue), or both (red) are highlighted. Chi-square P values are indicated. d, CAR occupancy, determined by ChIP–seq, at representative loci whose expression is regulated by CAR in both mucosal T cells and hepatocytes within mouse hepatocytes ectopically expressing epitope-tagged mouse (m) or human (h) CAR proteins and re-isolated from mice after treatment with TC or CITCO. *False discovery rate (FDR) q < 0.05 determined using MACS2 base settings.
Extended Data Fig. 4 CAR promotes effector T cell persistence in the presence of small intestinal bile acids.
a, Percentages of live CD44hi wild-type (B6; CD45.1+; blue) or CAR-deficient (B6.Nr1i3−/−; CD45.1−; red) Teff cells, determined by flow cytometry and gated as in Extended Data Fig. 6a, in tissues from reconstituted B6.Rag1−/− mice over time. Numbers indicate percentages; representative of five mice per tissue and time point. b, Fitness, defined as mean ± s.e.m. log2[fold change (F.C.)] of CAR-deficient (B6.Nr1i3−/−) versus wild-type (B6) Teff cell percentages (n = 5) in tissues from congenically co-transferred Rag1−/− mice over time, determined by flow cytometry as in a. c, Percentage of wild-type (B6, CD45.1+; blue) and CAR-deficient (B6.Nr1i3−/−, CD45.1−; red) naive (CD62Lhi) CD4+ T cells after sorting and mixing, and before in vivo transfer into Rag1−/− mice (input Tnaive); representative of three mixtures used for analysing resulting Teff cells at 2, 4 or 6 weeks after transfer. d, Equal numbers of CD45.1 wild-type (B6; blue) and CD45.2 CAR-deficient (B6.Nr1i3−/−; red) naive CD4+ T cells were transferred together into co-housed Rag1−/− mice with or without the ileal bile acid reuptake transporter ASBT (gene symbol Slc10a2). The resulting Teff cells from siLP were analysed two weeks after T cell transfer via flow cytometry. e, Percentages of live CD44hi wild-type (B6; CD45.1+; blue) or CAR-deficient (B6.Nr1i3−/−; CD45.1−; red) Teff cells, determined by flow cytometry and gated as in Extended Data Fig. 6a, in siLP from week-2 reconstituted B6.Rag1−/− mice. Numbers indicate percentages; representative of 8–10 mice analysed over two independent experiments. f, Mean ± s.e.m. absolute numbers of live CD45.1 wild-type (B6; left) or CD45.2 CAR-deficient (B6.Nr1i3−/−; right) Teff cells, determined by ex vivo flow cytometry as in e, from siLP two weeks after mixed T cell transfer into control (Slc10a2+/+; blue; n = 8) or ASBT-deficient (Slc10a2−/−; red; n = 10) Rag1−/− recipients. Fold changes in cell numbers recovered from ASBT-deficient versus control recipients, as well as P values (two-tailed unpaired Student’s t-test), are indicated.
Extended Data Fig. 5 Preferential CAR expression and function in human Teff cells expressing small bowel homing receptors.
a, FACS-based identification of human CD4+ T cell subsets in PBMCs from healthy adult human donors. Right, expression of integrin α4 (α4 int.) in gated naive (grey), Treg (blue), and Teff (red) T cells. b, Expression of integrin β7 (β7 int.) and CCR9 in total naive CD4+ T cells, or in α4 int. with or without Treg or Teff subsets (gated as in a). Representative of seven independent experiments using PBMCs from different donors. c, Percentages of α4+β7+CCR9+ naive T, Treg, and Teff cells, determined by flow cytometry as in a, b. Individual data points for seven independent experiments are shown and connected by lines. P values (one-way ANOVA with Tukey’s correction for multiple comparisons) are shown. d, Ex vivo Rh123 efflux in CD4+ T cell subsets (gated as in a, b) in the presence (grey) or absence (red) of the selective MDR1 inhibitor elacridar. Representative of seven experiments. e, Mean ± s.e.m. percentages (n = 7) of Rh123lo (MDR1+) Teff subsets, determined by flow cytometry as in d. P values (one-way ANOVA with Tukey’s correction for multiple comparisons) are shown. f, Mean ± s.e.m. ex vivo expression, determined by qPCR, of NR1I3 (n = 12), ABCB1 (n = 12) and CYP2B6 (n = 10) in α4−β7−CCR9− and α4+β7+CCR9+ naive T, Treg and Teff cells, FACS-sorted as in a, b. P values (one-way ANOVA with Tukey’s correction for multiple comparisons) are shown. g, Mean ± s.e.m. relative CYP2B6 expression (n = 5), determined by qPCR, in CD4+ T cell subsets (as in f) stimulated ex vivo in the presence of titrating concentrations of CITCO. Gene expression analysed 24 h after activation. ***P < 0.0001 by two-way ANOVA. h, Mean ± s.e.m. normalized ABCB1 or CYP2B6 expression, determined by RNA-seq and presented as TPM, in FACS-sorted α4+β7+CCR9+ Teff cells stimulated in vitro with or without CITCO. Data from four independent RNA-seq experiments using cells sorted from different healthy adult donors; P values (paired two-tailed Student’s t-test) are indicated. i, Identification of CD4+ naive (Tnaive; CD25−CD45RO−; grey) or Teff (CD25−CD45RO+; red) cells, by flow cytometry, from healthy adult human PBMCs. For improved purity of TH1, TH2, TH17 and TH17.1 cells, CCR10-expressing TH22 cells were excluded. CCR6 expression in naive T (grey) or non-TH22 Teff cells (red) is shown on the right; CCR6+ or CCR6− Teff cells were gated to enrich for TH17 or non-TH17 lineages, respectively. j, Expression of CCR4 and CXCR3 in CCR6− (non-TH17; left) or CCR6+ (TH17; right) Teff cells identifies enriched CCR6−CCR4loCXCR3hi (TH1; orange), CCR6−CCR4hiCXCR3lo (TH2; blue), CCR6+CCR4hiCXCR3lo (TH17; green), and CCR6+CCR4loCXCR3hi (TH17.1; red) subsets. k, Expression of integrin α4 (α4 int.; top) in TH2, TH1, TH17 and TH17.1 human Teff cells gated as in a, b. Expression of integrin β7 (β7 int.) and CCR9 within α4 int− (middle) or α4 int+ (bottom) TH2, TH1, TH17 or TH17.1 cells gated as above. i–k, Representative of nine independent experiments using PBMCs from different healthy adult donors. l, Percentages (n = 9) of α4+β7+CCR9+ cells (gated as in a, b) amongst ex vivo TH1, TH2, TH17 and TH17.11 Teff cells (gated as in i–k). Data from independent donors are connected by lines; P values (one-way ANOVA with Tukey’s correction for multiple comparisons) are shown. m, MDR1-dependent Rh123 efflux in the indicated TH1, TH2, TH17 and TH17.1 Teff subsets gated by expression of α4 int., β7 int., and/or CCR9 in the presence (grey) or absence (red) of elacridar. Representative of eight independent experiments using PBMCs from different donors. n, Mean ± s.e.m. percentages (n = 8) of Rh123lo (MDR1+) cells within TH1, TH2, TH17 and TH17.11 Teff subsets gated according to expression of α4 int., β7 int., and/or CCR9 as in e. P values (one-way ANOVA with Tukey’s correction for multiple comparisons) are indicated; ****P < 0.0001. N.D., not detected; NS, not significant.
Extended Data Fig. 6 TCPOBOP promotes CAR-dependent gene expression in ex vivo-isolated effector T cells.
a, Top left, equal numbers of CD45.1 wild-type (B6; blue) and CD45.2 CAR-deficient (B6.Nr1i3−/−; red) naive CD4+ T cells were transferred together into B6.Rag1−/− mice. The resulting Teff cells were FACS-purified from spleen after 3 weeks. Right, sequential gating strategy for re-isolating wild-type and CD45.2 CAR-deficient spleen Teff cells. Bottom left, mean ± s.e.m. relative expression of Abcb1a (n = 4), Cyp2b10 (n = 4), and Il10 (n = 3), determined by qPCR, in ex vivo-isolated wild-type (B6) or CAR-deficient (B6.Nr1i3−/−) spleen Teff cells. These cells were used for ex vivo stimulation experiments with or without small-molecule ligands (b, c). P values (paired two-tailed Student’s t-test) are indicated. b, Mean ± s.e.m. relative expression of Abcb1a (n = 4), Cyp2b10 (n = 4), and Il10 (n = 3), by qPCR, in wild-type (B6) or CAR-deficient Teff cells isolated from transferred Rag1−/− mice (as in a), and stimulated ex vivo with anti-CD3 and anti-CD28 antibodies in the presence or absence of TC (10 μM), the mCAR inverse agonist And (10 μM), or both. P values (one-way ANOVA with Tukey’s correction for multiple comparisons) are indicated; ****P < 0.0001. c, Mean ± s.e.m. relative expression of Abcb1a, Cyp2b10, and Il10 (n = 5), determined by qPCR, in wild-type (B6) or CAR-deficient (B6.Nr1i3−/−) Teff cells isolated and stimulated as in a, b in the presence or absence of TC (10 μM) or PCN (10 μM). Data are presented as fold change in mRNA abundance relative to vehicle-treated cells (DMSO for TC; ethanol for PCN). ****P < 0.0001, one-way ANOVA with Dunnett’s correction for multiple comparisons. NS, not significant.
Extended Data Fig. 7 Characteristics of endogenous intestinal metabolites that activate the CAR LBD.
a, Mean ± s.e.m. activation (triplicate samples) of recombinant hRXRα LBD homodimers, determined by TR-FRET in the presence of TC (blue) or 9-cis retinoic acid (RA; red). Median effective concentration (EC50) of 9-cis RA-dependent hRXRα LBD homodimer activation is indicated. Representative of more than five independent experiments. b, Mean ± s.e.m. activation (n = 3) of hRXRα LBD homodimers, determined by TR-FRET as in a, in the presence of titrating concentrations of siLC, bile, cLC or serum from wild-type B6 mice. NS, not significant by one-way ANOVA with Dunnett’s correction for multiple comparisons. c, Mean ± s.e.m. activation (n = 3) of CAR–RXR LBD heterodimers, determined by TR-FRET, in the presence of titrating concentrations of siLC isolated from wild-type B6 mice housed under specific pathogen-free (SPF) or germ-free (GF) conditions and pre-treated with or without cholestyramine (CME) to deplete free bile acids. ****P < 0.0001, one-way ANOVA with Dunnett’s correction for multiple comparisons. a–c, The bars for each tissue extract indicate dilution series (left to right): diluent (PBS) alone; 0.01%, 0.1%, 1%. Data are shown from three independent experiments using extracts from different wild-type mice, with each concentration from each individual mouse run in triplicate. d, Mean ± s.e.m. TR-FRET signals (n = 3) of CAR–RXR LBD heterodimers in the presence of titrating concentrations of individual bile acid (BA) species. NS, not significant by one-way ANOVA with Dunnett’s correction for multiple comparisons. The bars for bile acids indicate concentrations (left to right): vehicle (DMSO), 10 μM, 100 μM, 1,000 μM. Data are shown from three independent experiments, in which each bile acid concentration was run in triplicate.
Extended Data Fig. 8 CAR promotes IL-10 expression in mucosal Teff cells and regulates TR1 and TH17 cell development in the small intestine.
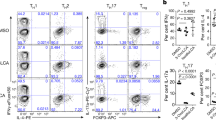
a, Equal numbers of CD45.1 wild-type (B6; blue) and CD45.2 CAR-deficient (B6.Nr1i3−/−; red) naive CD4+ T cells were transferred together into Rag1−/− mice. The resulting Teff cells were analysed using surface and intracellular flow cytometry after ex vivo stimulation with PMA and ionomycin at 2, 4, and 6 weeks from spleen, MLN, siLP, or cLP. Gating hierarchy is shown from a representative sample of MLN mononuclear cells at two weeks after T cell transfer. b, Intracellular IL-10 and IFNγ expression, determined by flow cytometry, in wild-type (B6, blue; left) and CAR-deficient (B6.Nr1i3−/−, red; right) non-TH17 Teff cells, gated as in a, from tissues of T cell-reconstituted B6.Rag1−/− mice over time. Numbers indicate percentages; representative of five mice per tissue and time point. c, d, Mean ± s.e.m. percentages (c) or numbers (d) (n = 5) of IL-10-expressing wild-type (B6, left) or CAR-deficient (B6.Nr1i3−/−, right) Teff cells, determined by ex vivo flow cytometry as in a, b from tissues of transferred B6.Rag1−/− mice over time. e, Specificity of IL-10 intracellular staining, as validated by analysis of IL-10 production by CD45.1 wild-type (B6; blue) or CD45.2 Il10−/− (red) Teff cells isolated from spleen or siLP of congenically co-transferred Rag1−/− mice. Representative of six mice analysed over two independent experiments. f, Expression of RORγt and IL-17A, determined by intracellular FACS analysis as in Extended Data Fig. 8a, in wild-type (B6) or CAR-deficient (B6.Nr1i3−/−) CD4+ Teff cells from tissues of reconstituted Rag1−/− mice two weeks after mixed T cell transfer. Numbers indicate percentages; representative of five mice per tissue and time point. g, Mean ± s.e.m. percentages of (n = 5) wild-type (B6; blue) or CAR-deficient (B6.Nr1i3−/−; red) RORγt+IL-17A− Teff cells, determined by intracellular flow cytometry as in a. *P = 0.029, paired two-tailed Student’s t-test. h, Expression of RORγt and IL-17A, determined by intracellular FACS analysis, in wild-type (B6) or Il10 −/− Teff cells from tissues of reconstituted Rag1−/− mice two weeks after mixed T cell transfer. Numbers indicate percentages; representative of five mice per tissue and time point. i, Mean ± s.e.m. percentages of (n = 7) wild-type (B6; blue) or Il10 −/− (red) RORγt+IL-17A− Teff cells, determined by intracellular flow cytometry as in c. *P = 0.033, **P = 0.0099, paired two-tailed Student’s t-test.
Extended Data Fig. 9 CAR expression and function in mucosal Teff cells is increased in response to intestinal inflammation.
a, Percentages of CD3+CD4+ T cells in tissues from Rag1−/− mice transplanted with congenic mixtures of wild-type and CAR-deficient naive CD4+ T cells over time, determined by flow cytometry. Representative of five mice per tissue and time point. b, Mean ± s.e.m. absolute numbers of CD3+CD4+ TH cells (n = 5) in tissues from transferred B6.Rag1−/− mice over time, determined by flow cytometry as in a. c, Mean ± s.e.m. relative ex vivo Nr1i3, Abcb1a, Cyp2b10, and Il10 gene expression (n = 3), determined by qPCR, in wild-type (B6) CD4+ Teff cells (sorted as in Extended Data Fig. 8a) from spleens of transferred B6.Rag1−/− mice over time. d, Top, expression of FOXP3 and RORγt, determined by intracellular staining after ex vivo (PMA + ionomycin) stimulation, in CD4+CD44hi cells from spleen (left) or siLP (right) of wild-type (B6, blue) or CAR-deficient (B6.Nr1i3−/−, red) mice injected with isotype control (IgG) or soluble anti-CD3. Bottom, expression of IL-10 and IL-17A in wild-type or CAR-deficient spleen or siLP T cell subsets from mice treated with isotype control (IgG) or anti-CD3 antibodies. Cells were gated and analysed by flow cytometry as above. Numbers indicate percentages; representative of three mice per group and genotype analysed over two independent experiments. e, f, Mean ± s.e.m. percentages of IL-10-expressing T cell subsets (n = 3), gated and analysed by ex vivo flow cytometry as in a, in spleen (e) or siLP (f) TH cell subsets from wild-type (B6, blue) or CAR-deficient (B6.Nr1i3−/−, red) mice injected with isotype control (IgG) or anti-CD3 antibody. P values (unpaired Student’s t-test) are indicated; NS, not significant.
Extended Data Fig. 10 TCPOBOP protection against bile acid-induced ileitis requires CAR expression in T cells.
a, Mean ± s.e.m. weight loss (n = 5 per group) of co-housed B6.Rag2−/− mice transplanted with CAR-deficient (B6.Nr1i3−/−) CD4+ naive T cells and maintained on a cholic acid-supplemented diet with (CA/TC) or without (CA/Veh) TC treatment. Weights are shown relative to three weeks after transfer, when TC treatments were initiated. NS, not significant; two-way ANOVA. b, H&E-stained sections of terminal ilea or colons from B6.Rag2−/− mice reconstituted with CAR-deficient T cells and treated as above and as indicated. Representative of five mice per group analysed in two independent experiments. Magnification, 10×; scale bars, 100 μm. c, Mean ± s.e.m. histology scores for colons and terminal ilea as in b. NS, not significant by unpaired two-tailed Student’s t-test.
Supplementary information
Supplementary Tables
This file contains Supplementary Tables 1–3.
Source data
Rights and permissions
About this article
Cite this article
Chen, M.L., Huang, X., Wang, H. et al. CAR directs T cell adaptation to bile acids in the small intestine. Nature 593, 147–151 (2021). https://doi.org/10.1038/s41586-021-03421-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-021-03421-6
This article is cited by
-
Diindoles produced from commensal microbiota metabolites function as endogenous CAR/Nr1i3 ligands
Nature Communications (2024)
-
Gut microbiome modulates tacrolimus pharmacokinetics through the transcriptional regulation of ABCB1
Microbiome (2023)
-
TH17 cell heterogeneity and its role in tissue inflammation
Nature Immunology (2023)
-
Lipid metabolism in T cell signaling and function
Nature Chemical Biology (2022)
-
Regulation of tissue-resident memory T cells by the Microbiota
Mucosal Immunology (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.