Abstract
The behaviour of an animal is determined by metabolic, emotional and social factors1,2. Depending on its state, an animal will focus on avoiding threats, foraging for food or on social interactions, and will display the appropriate behavioural repertoire3. Moreover, survival and reproduction depend on the ability of an animal to adapt to changes in the environment by prioritizing the appropriate state4. Although these states are thought to be associated with particular functional configurations of large-brain systems5,6, the underlying principles are poorly understood. Here we use deep-brain calcium imaging of mice engaged in spatial or social exploration to investigate how these processes are represented at the neuronal population level in the basolateral amygdala, which is a region of the brain that integrates emotional, social and metabolic information. We demonstrate that the basolateral amygdala encodes engagement in exploratory behaviour by means of two large, functionally anticorrelated ensembles that exhibit slow dynamics. We found that spatial and social exploration were encoded by orthogonal pairs of ensembles with stable and hierarchical allocation of neurons according to the saliency of the stimulus. These findings reveal that the basolateral amygdala acts as a low-dimensional, but context-dependent, hierarchical classifier that encodes state-dependent behavioural repertoires. This computational function may have a fundamental role in the regulation of internal states in health and disease.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Data availability
The data that support the findings of this study are available at https://data.fmi.ch/PublicationSupplementRepo/.
Code availability
Custom-written codes used to analyse data from this study are available from the corresponding author upon request.
References
Darwin, C. The Expression of the Emotions in Man and Animals (John Murray, 1872).
Tinbergen, N. The Study of Instinct (Clarendon and Oxford Univ. Press, 1951).
LeDoux, J. Rethinking the emotional brain. Neuron 73, 653–676 (2012).
Fanselow, M. S. The role of learning in threat imminence and defensive behaviors. Curr. Opin. Behav. Sci. 24, 44–49 (2018).
Salzman, C. D. & Fusi, S. Emotion, cognition, and mental state representation in amygdala and prefrontal cortex. Annu. Rev. Neurosci. 33, 173–202 (2010).
Adolphs, R. & Anderson, D. The Neuroscience of Emotion (Princeton Univ. Press, 2018).
Sah, P., Faber, E. S. L., Lopez De Armentia, M. & Power, J. The amygdaloid complex: anatomy and physiology. Physiol. Rev. 83, 803–834 (2003).
Pape, H.-C. & Pare, D. Plastic synaptic networks of the amygdala for the acquisition, expression, and extinction of conditioned fear. Physiol. Rev. 90, 419–463 (2010).
Tovote, P., Fadok, J. P. & Lüthi, A. Neuronal circuits for fear and anxiety. Nat. Rev. Neurosci. 16, 317–331 (2015).
O’Neill, P.-K., Gore, F. & Salzman, C. D. Basolateral amygdala circuitry in positive and negative valence. Curr. Opin. Neurobiol. 49, 175–183 (2018).
Davis, M. in The Amygdala (ed. Aggleton, J.) 213–288 (Oxford Univ. Press, 2000).
Paton, J. J., Belova, M. A., Morrison, S. E. & Salzman, C. D. The primate amygdala represents the positive and negative value of visual stimuli during learning. Nature 439, 865–870 (2006).
Headley, D. B., Kanta, V., Kyriazi, P. & Paré, D. Embracing complexity in defensive networks. Neuron 103, 189–201 (2019).
Kyriazi, P., Headley, D. B. & Pare, D. Multi-dimensional coding by basolateral amygdala neurons. Neuron 99, 1315–1328.e5 (2018).
Gründemann, J. et al. Amygdala ensembles encode behavioral states. Science 364, eaav8736 (2019).
Remedios, R. et al. Social behaviour shapes hypothalamic neural ensemble representations of conspecific sex. Nature 550, 388–392 (2017).
Li, Y. et al. Neuronal representation of social information in the medial amygdala of awake behaving mice. Cell 171, 1176–1190.e17 (2017).
Hung, L. W. et al. Gating of social reward by oxytocin in the ventral tegmental area. Science 357, 1406–1411 (2017).
Kingsbury, L. et al. Correlated neural activity and encoding of behavior across brains of socially interacting animals. Cell 178, 429–446.e16 (2019).
Chen, P. & Hong, W. Neural circuit mechanisms of social behavior. Neuron 98, 16–30 (2018).
Brothers, L. The social brain: a project for integrating primate behavior and neurophysiology in a new domain. Concepts Neurosci. 1, 27–51 (1990).
Adolphs, R. The social brain: neural basis of social knowledge. Annu. Rev. Psychol. 60, 693–716 (2009).
Emery, N. J. et al. The effects of bilateral lesions of the amygdala on dyadic social interactions in rhesus monkeys (Macaca mulatta). Behav. Neurosci. 115, 515–544 (2001).
Hadjikhani, N. et al. The effect of constraining eye-contact during dynamic emotional face perception-an fMRI study. Soc. Cogn. Affect. Neurosci. 12, 1197–1207 (2017).
Kennedy, D. P. & Adolphs, R. The social brain in psychiatric and neurological disorders. Trends Cogn. Sci. 16, 559–572 (2012).
von Heimendahl, M., Rao, R. P. & Brecht, M. Weak and nondiscriminative responses to conspecifics in the rat hippocampus. J. Neurosci. 32, 2129–2141 (2012).
Felix-Ortiz, A. C. & Tye, K. M. Amygdala inputs to the ventral hippocampus bidirectionally modulate social behavior. J. Neurosci. 34, 586–595 (2014).
Liang, B. et al. Distinct and dynamic on and off neural ensembles in the prefrontal cortex code social exploration. Neuron 100, 700–714.e9 (2018).
Kodama, N. X. et al. Anti-correlated cortical networks arise from spontaneous neuronal dynamics at slow timescales. Sci. Rep. 8, 666 (2018).
Levy, D. R. et al. Dynamics of social representation in the mouse prefrontal cortex. Nat. Neurosci. 22, 2013–2022 (2019).
Krabbe, S. et al. Adaptive disinhibitory gating by VIP interneurons permits associative learning. Nat. Neurosci. 22, 1834–1843 (2019).
Froemke, R. C. Plasticity of cortical excitatory–inhibitory balance. Annu. Rev. Neurosci. 38, 195–219 (2015).
Chen, T. W. et al. Ultrasensitive fluorescent proteins for imaging neuronal activity. Nature 499, 295–300 (2013).
Ghosh, K. K. et al. Miniaturized integration of a fluorescence microscope. Nat. Methods 8, 871–878 (2011).
Paxinos, G. & Franklin, K. The Mouse Brain in Stereotaxic Coordinates (Academic, 2001).
Schneider, C. A., Rasband, W. S. & Eliceiri, K. W. NIH Image to ImageJ: 25 years of image analysis. Nat. Methods 9, 671–675 (2012).
Mukamel, E. A., Nimmerjahn, A. & Schnitzer, M. J. Automated analysis of cellular signals from large-scale calcium imaging data. Neuron 63, 747–760 (2009).
Acknowledgements
We thank J. J. Letzkus, A. Ramos-Prats, J. Gründemann, S. Krabbe, P. Tovote, M. S. Esposito, M. B. Pardi, A. Depino and all members of the laboratory of A.L. for comments and discussions; P. Argast, P. Buchmann and C. Müller and all staff of the Friedrich Miescher Institute for Biomedical Research (FMI) Animal Facility for technical assistance; the IT department of the FMI, in particular D. Flanders, S. Grzybek, R. Milani, E. Tagliavini, E. Brondolo, S. van Eden and A. Naylor, for their support with data acquisition and analyses; the Facility for Imaging and Microscopy at the FMI, in particular S. Bourke and J. Eglinger; and M. Stadler for statistical advice. This work was supported by the following grants: the European Research Council under the European Union’s Horizon 2020 research and innovation programme (grant agreement no. 669582) and a Swiss National Science Foundation core grant (310030B_170268) to A.L.; a Novartis Presidential Postdoc fellowship to M.S.F.; as well as by the Novartis Research Foundation.
Author information
Authors and Affiliations
Contributions
M.S.F., T.B. and A.L. designed the experiments. M.S.F. performed the experiments, with help from T.E. Y.B. and M.S.F. analysed behaviour and deep-brain imaging data. M.S.F., Y.B. and A.L. wrote the paper, and all authors contributed to the interpretation of the data and commented on the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature thanks Ekaterina Likhtik, Hee-Sup Shin and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Combined automatic description of social behaviour and deep-brain calcium imaging.
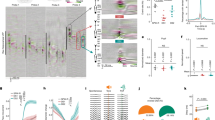
a, Social exploration session between two freely behaving mice. The arrows and letters represent the quantities used by the behavioural detection algorithm. a and b denote the x–y coordinates of the head positions of each of the two mice; Va and Vb are velocity vectors for each mouse; d is the distance vector between the mice; αa,b is the angle between the velocity vector V and the distance vector d. b, Criteria used to detect the different behaviours. c, Agreement of the algorithm with the scoring of the experimenter (algorithm) and the agreement between manual scoring performed independently by two experimenters (manual). Numbers indicate the percentage of matched frames. Wilcoxon matched-pairs signed-rank test, P = 0.12, n = 4 mice. d, Representation of the behaviour descriptors assigned by the algorithm and the behavioural scoring of the experimenter for a representative mouse. e, Location of the GRIN lens for mice in the study. A, anterior; BA, basal amygdala; CeA, central nucleus of the amygdala; D, dorsal; L, lateral; LA, lateral amygdala; M, medial; P, posterior; V, ventral. f, Top left, schematic of virus injection and implantation of the GRIN lens. Bottom left, GCaMP6f expression in an example coronal section. Scale bar, 500 μm. Bottom right, close-up image of the BLA region in fixed sections. Scale bar, 200 μm. Top right, maximum projection over the entire session of the miniature microscope image from the GRIN lens. Scale bar, 100 μm. g, Behavioural comparison between microscope (n = 10) and no-microscope (n = 9) mice. Multiple unpaired t-test between microscope and no-microscope mice for each behaviour. Holm–Sidak multiple comparisons test; neutral, P = 0.91; interaction, P = 0.91; approach, P = 0.75; reciprocated, P = 0.86; avoidance, P = 0.99; freezing, P = 0.13; and aggression, P = 0.86. h, Example calcium traces of six recorded BLA cells during a free social exploration session. Background colours denote the behaviours of the mice (as in b). zs, z-score. Example images and traces in f, h are representative of 10 mice. Data in c, g are mean ± s.e.m. n.s., P ≥ 0.05. Statistical tests are two-sided.
Extended Data Fig. 2 Social exploration encoded by two BLA ensembles.
a, Empirical distribution of the absolute pairwise Pearson’s correlations between single-neuron activations for all mice (663 neurons and 10 mice). b, Mean cellular pairwise distance within- and across-ensemble of all individual neurons (663 neurons and 10 mice) (left), and the cumulative probability distributions of the mean cellular pairwise distance (PWD) of neurons in each ensemble and in the total neuronal population (right) reveal that the two ensembles are not spatially clustered. Two-sample Kolmogorov–Smirnov test (K–S), ensemble 1 versus total, K–S = 0.01, P = 0.82; ensemble 2 versus total, K–S = 0.02, P = 0.62. c, Mean empirical probability distributions of the instantaneous speed of the experimental mouse during activation of the social exploration (blue) and no social exploration (red) ensembles. Two-sample Kolmogorov–Smirnov test, K–S = 0.11, P = 0.55, n = 2 distributions from n = 10 mice. d, Instantaneous speed of the mice plotted against ensemble activation for the social exploration (blue) and no social exploration (red) ensembles. Data binned in 1-s bins. Only two mice showed a weak correlation. Pearson’s correlation values indicated in the respective panels: mouse 7: ensemble 1, P = 0.003; ensemble 2, P = 0.002; mouse 8: ensemble 1, P < 0.0001; ensemble 2, P < 0.0001. zs, z-score. e, Average entropy of the spatial occupancy maps of the experimental mouse, calculated during epochs of activation of the social exploration (ensemble 1) (blue) and no social exploration (ensemble 2) (red) ensembles. Paired t-test, t9 = 1.65 P = 0.13, n = 10 mice. f, Mean activity of three ensembles for one mouse. k-means clustering into three groups with correlation as a distance measure. Colours as in Fig. 1b. g, Mean probability of different behaviours during epochs of high (left columns) and low (right columns) ensemble activation for the three ensembles, calculated as in f. n = 10 mice. h, Probability of social exploration behaviours during epochs of high (+) and low (−) ensemble activation (defined as in f) for the different ensembles. Paired t-test, activated versus inactivated; ensemble 1, t9 = 7.6, P = 3.3 × 10−5; ensemble 2, t9 = 6.3, P = 0.0001; ensemble 3, t9 = 4.2, P = 0.002. n = 10 mice. Ensembles are colour-coded as in f. Data in b (right), c, e, h are mean ± s.e.m. **P < 0.01, ***P < 0.001, n.s., P ≥ 0.05. Statistical tests are two-sided.
Extended Data Fig. 3 Distributed representation of social exploration.
a, Correlation between neurons and ensemble activity for an example mouse. Colours denote the ensemble membership of the neurons. Circles mark the 20% most-correlated neurons (10% most correlated with the mean of each ensemble activity). b, Correlation values between the mean activity of the two ensembles calculated for: the full ensembles; the 20% highly correlated neurons; the remaining 80% least-correlated neurons; and the shift control in an example mouse (left) and the population (right). One-way repeated-measures analysis of variance, main group effect F(1.9,17.1) = 38.67, P < 0.0001, Dunnett’s multiple comparisons test, shifted versus full ensembles, P < 0.0001; shifted versus 20%, P = 0.002; shifted versus 80%, P < 0.0001. n = 10 mice. c, Mean activity calculated using the full ensembles (top), 20% most-correlated (middle) and 80% least-correlated neurons (bottom). Colours as in Fig. 1b. d, Probability of social exploration behaviours during ensemble activation for the 20% most-correlated (paired t-test, t9 = 3.8, P = 0.004, n = 10 mice) and the 80% least-correlated neurons (paired t-test, t9 = 5.35, P = 0.0005, n = 10 mice). e, Average change in decoder F1 score between ensemble data and circular temporal shift of neuronal activity. One-sample t-test, t9 = 5.18, P = 0.0006. n = 10 mice. Box-and-whisker plot indicating median, interquartile range and the minimum to maximum values of the data distribution. Data in b, d, e are mean ± s.e.m. **P < 0.01, ***P < 0.001, n.s., P ≥ 0.05. Statistical tests are two-sided.
Extended Data Fig. 4 Ensemble stability and single-neuron correlates of social behaviour.
a, Probability of social exploration behaviours in a second social exploration session during ensemble activation epochs. k-means clustering performed on this session; paired t-test, t9 = 6.33, P = 0.0001, n = 10 mice. b, Mean within- and across-ensemble distance of individual neurons (663 neurons and 10 mice) (left) and cumulative probability distribution of the mean cellular pairwise distance of neurons in each ensemble and in the total neuronal population (right) reveal that the two ensembles are not spatially clustered. Two-sample Kolmogorov–Smirnov test (K–S), ensemble 1 versus total, K–S = 0.02, P = 0.35; ensemble 2 versus total, K–S = 0.01, P = 0.62. n = 2 distributions from n = 10 mice. c, Correlation between single neurons and no social ensemble mean activities during the two social exploration sessions. Ensembles were defined independently for each session. Pearson’s correlation, R = 0.59, P < 0.0001, n = 663 neurons. d, Calcium traces of single neurons activated by social exploration (left) and by no social exploration (right) obtained by single-neuron analysis for an example mouse (Methods). Colours are as in Fig. 1b. e, Average percentage of neurons activated during social exploration (blue) or no social exploration (red) in the first social exploration session. n = 10 mice. f, Percentage of social-exploration-activated neurons included in each ensemble. Wilcoxon matched-pairs signed-rank test, P = 0.003, n = 10 mice. Activity of neurons included in the no social exploration ensemble was very sparse. g, Percentage of neurons significantly activated by social exploration or no social exploration for each social exploration session and the corresponding overlaps. Wilcoxon matched-pairs signed-rank test; activated by social exploration, P = 0.002; activated by no social exploration, P = 0.009. n = 10 mice. None of the neurons switched its activity correspondence with the behaviour across the two social sessions. h, Left, mean activity of the social-exploration-activated neurons (n = 106 neurons), aligned to initiation of interaction, approach and avoidance. Neurons divided into five groups by k-means clustering using combined response to the three behaviours and cosine as a distance measure. Dashed lines separate clusters. Right, mean responses of the four clusters that show locked responses to the initiation of the behaviours. Data in a, b (right), f, h (right) are mean ± s.e.m. **P < 0.01, ***P < 0.001, n.s., P ≥ 0.05. Statistical tests are two-sided.
Extended Data Fig. 5 Spatial exploration ensembles predict exploratory and self-centred behaviours.
a, Distribution of the magnitude of the response for each of the ensembles in the two sessions. Two-sample Kolmogorov–Smirnov test (K–S); social versus no social, K–S = 0.14, P = 0.26; social versus spatial, K–S = 0.14, P = 0.26; social versus no spatial, K–S = 0.17, P = 0.099; spatial versus no spatial, K–S = 0.02, P = 0.99; spatial versus no social, K–S = 0.06, P = 0.99; no social versus no spatial, K–S = 0.12, P = 0.44. n = 2 distributions from n = 10 mice. Comparison of correlation of the ensembles: social versus spatial, paired t-test, t9 = 0.02, P = 0.98. n = 10 mice. b, Two spatial exploration sessions. c, Correlation between single-neuron and spatial-exploration-ensemble mean activities in the first spatial exploration session versus the second spatial exploration session. Ensembles were defined independently for each session. The strong correlation indicates stability across the two spatial sessions. Pearson’s correlation, R = 0.45, P < 0.0001, n = 663 neurons. Grey and white quadrants as in Fig. 2k. Grey quadrant: 31.1 ± 3.1%; white quadrant: 68.3 ± 3.1%; paired t-test, t9 = 5.9, P = 0.0002, n = 10 mice. Mean absolute Pearson correlation with the spatial exploration ensemble. Grey quadrant, 0.18 ± 0.01%; white quadrant 0.22 ± 0.01; paired t-test, t9 = 3.0, P = 0.01, n = 10 mice. d, Average entropy calculated from the occupancy maps of each mouse during the second spatial exploration session, collected as in Fig. 3c. Paired t-test, t9 = 6.89, P < 0.0001, n = 10 mice. Ensembles were as defined in the first spatial exploration session. e, Probability of object exploration detected using the behavioural detection algorithm during ensemble activation. Left, object exploration divided by the different behaviours for one mouse. Colours denote the behaviour of the mouse with the object: (1) neutral; (2) interaction; (3) approach; (5) avoidance; and (6) freezing. Right, object exploration averaged across mice. f, Rearing (left), grooming (middle) and freezing (right) probability during epochs of high relative ensemble activation. In e, f, Wilcoxon matched-pairs signed-rank test; object exploration, P = 0.002; rearing, P = 0.04; grooming, P = 0.002; freezing, P = 0.002. n = 10 mice. g, Trajectories of an example mouse during activation epochs of the ensemble in the spatial exploration session. Colours denote the behaviour of the mouse as in e, along with rearing (8) and grooming (9). Different behaviours can occur in the same spatial location and activate different ensembles. Data in a, e, f are mean ± s.e.m. *P < 0.05, **P < 0.01, ***P < 0.001, n.s., ≥ 0.05. Statistical tests are two-sided.
Extended Data Fig. 6 Stable representation of spatial exploration across behavioural contexts.
a, Neuronal subpopulations defined by their activity in both the spatial (left) and social (right) exploration sessions: neurons that stably correlate with exploration (group 1 (blue)) or no exploration (group 3 (purple)), and neurons that correlate with exploration in one behavioural context and no exploration in the other (group 2 (yellow) and group 4 (red)). Lines connect the same neurons. b, Proportion of different behaviours during high (+) and low (−) activation of different subpopulation defined as in a, during the spatial (left) and the social (right) exploration sessions. n = 10 mice. Bar colours denote the behaviour of the experimental mouse, as in Fig. 3h. c, Magnitude of the difference between the mean activity of the two subpopulations in the no social exploration ensemble (top left) is quantified by the interquartile range of this quantity over long epochs (bottom left). Interquartile range during the spatial exploration session (green) and social exploration session, split between no social exploration (grey) and social exploration epochs (blue). The subpopulations of the no social exploration ensemble exhibit a stronger difference signal during no social exploration epochs than during epochs of social exploration. One-way repeated-measures analysis of variance, main effect between groups F(1.7,15.2) = 88.9, P < 0.0001; Tukey’s multiple comparisons test, spatial versus no social, P < 0.0001; spatial versus social, P < 0.0001; no social versus social, P = 0.008. n = 10 mice. d, Probability of rearing (left), grooming (middle) and entropy of the spatial occupancy maps (right) during epochs of high (+) and low (−) ensemble activation for the subpopulations defined in a during the entire social exploration session for rearing and grooming, and during only the no social exploration epochs for entropy. Entropy results are consistent with those obtained for the two spatial exploration ensembles in Fig. 3i left. Wilcoxon matched-pairs signed rank test; activated versus inactivated. Rearing: group 1, P = 0.32; group 2, P = 0.03; group 3, P = 0.55; and group 4, P = 0.06. Grooming: group 1, P = 0.13; group 2, P = 0.91; group 3, P = 0.01; and group 4, P = 0.84. Entropy: group 1, P = 0.23; group 2, P = 0.49; group 3, P = 0.92; and group 4, P = 0.27. n = 10 mice. Data in c, e are mean ± s.e.m. *P < 0.05, **P < 0.01, ***P < 0.001, n.s., P ≥ 0.05. Statistical tests are two-sided.
Supplementary information
Rights and permissions
About this article
Cite this article
Fustiñana, M.S., Eichlisberger, T., Bouwmeester, T. et al. State-dependent encoding of exploratory behaviour in the amygdala. Nature 592, 267–271 (2021). https://doi.org/10.1038/s41586-021-03301-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-021-03301-z
This article is cited by
-
Integrated cardio-behavioral responses to threat define defensive states
Nature Neuroscience (2023)
-
Functional specialization and interaction in the amygdala-hippocampus circuit during working memory processing
Nature Communications (2023)
-
Adolescent social isolation induces distinct changes in the medial and lateral OFC-BLA synapse and social and emotional alterations in adult mice
Neuropsychopharmacology (2022)
-
Distinct serotonergic pathways to the amygdala underlie separate behavioral features of anxiety
Nature Neuroscience (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.