Abstract
Mitochondrial DNA double-strand breaks (mtDSBs) are toxic lesions that compromise the integrity of mitochondrial DNA (mtDNA) and alter mitochondrial function1. Communication between mitochondria and the nucleus is essential to maintain cellular homeostasis; however, the nuclear response to mtDSBs remains unknown2. Here, using mitochondrial-targeted transcription activator-like effector nucleases (TALENs)1,3,4, we show that mtDSBs activate a type-I interferon response that involves the phosphorylation of STAT1 and activation of interferon-stimulated genes. After the formation of breaks in the mtDNA, herniation5 mediated by BAX and BAK releases mitochondrial RNA into the cytoplasm and triggers a RIG-I–MAVS-dependent immune response. We further investigated the effect of mtDSBs on interferon signalling after treatment with ionizing radiation and found a reduction in the activation of interferon-stimulated genes when cells that lack mtDNA are exposed to gamma irradiation. We also show that mtDNA breaks synergize with nuclear DNA damage to mount a robust cellular immune response. Taken together, we conclude that cytoplasmic accumulation of mitochondrial RNA is an intrinsic immune surveillance mechanism for cells to cope with mtDSBs, including breaks produced by genotoxic agents.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
All figures and extended data figures are associated with raw data used for statistical derivation, immunofluorescence images, raw RNA-sequencing data and DE gene analysis, uncropped western blots and flow cytometry files. These data have been uploaded in Mendeley Data (https://doi.org/10.17632/jszd6bwvtw.1). RNA-sequencing .fastq files are available through the NCBI GEO repository under accession codes GSE164979 and GSE164980.
References
Phillips, A. F. et al. Single-molecule analysis of mtDNA replication uncovers the basis of the common deletion. Mol. Cell 65, 527–538.e6 (2017).
Fu, Y., Tigano, M. & Sfeir, A. Safeguarding mitochondrial genomes in higher eukaryotes. Nat. Struct. Mol. Biol. 27, 687–695 (2020).
Bacman, S. R., Williams, S. L., Pinto, M., Peralta, S. & Moraes, C. T. Specific elimination of mutant mitochondrial genomes in patient-derived cells by mitoTALENs. Nat. Med. 19, 1111–1113 (2013).
Renaud, J. B. et al. improved genome editing efficiency and flexibility using modified oligonucleotides with TALEN and CRISPR–Cas9 nucleases. Cell Rep. 14, 2263–2272 (2016).
McArthur, K. et al. BAK/BAX macropores facilitate mitochondrial herniation and mtDNA efflux during apoptosis. Science 359, eaao6047 (2018).
Schon, E. A. et al. A direct repeat is a hotspot for large-scale deletion of human mitochondrial DNA. Science 244, 346–349 (1989).
Peeva, V. et al. Linear mitochondrial DNA is rapidly degraded by components of the replication machinery. Nat. Commun. 9, 1727 (2018).
Nissanka, N., Bacman, S. R., Plastini, M. J. & Moraes, C. T. The mitochondrial DNA polymerase gamma degrades linear DNA fragments precluding the formation of deletions. Nat. Commun. 9, 2491 (2018).
Moretton, A. et al. Selective mitochondrial DNA degradation following double-strand breaks. PLoS ONE 12, e0176795 (2017).
Liao, X. & Butow, R. A. RTG1 and RTG2: two yeast genes required for a novel path of communication from mitochondria to the nucleus. Cell 72, 61–71 (1993).
Nargund, A. M., Pellegrino, M. W., Fiorese, C. J., Baker, B. M. & Haynes, C. M. Mitochondrial import efficiency of ATFS-1 regulates mitochondrial UPR activation. Science 337, 587–590 (2012).
Guo, X. et al. Mitochondrial stress is relayed to the cytosol by an OMA1–DELE1–HRI pathway. Nature 579, 427–432 (2020).
West, A. P. et al. Mitochondrial DNA stress primes the antiviral innate immune response. Nature 520, 553–557 (2015).
Dhir, A. et al. Mitochondrial double-stranded RNA triggers antiviral signalling in humans. Nature 560, 238–242 (2018).
Waugh, D. S. & Sauer, R. T. Single amino acid substitutions uncouple the DNA binding and strand scission activities of Fok I endonuclease. Proc. Natl Acad. Sci. USA 90, 9596–9600 (1993).
Nelson, I., Hanna, M. G., Wood, N. W. & Harding, A. E. Depletion of mitochondrial DNA by ddC in untransformed human cell lines. Somat. Cell Mol. Genet. 23, 287–290 (1997).
Niu, X. et al. A small-molecule inhibitor of Bax and Bak oligomerization prevents genotoxic cell death and promotes neuroprotection. Cell Chem. Biol. 24, 493–506 (2017).
Ichim, G. et al. Limited mitochondrial permeabilization causes DNA damage and genomic instability in the absence of cell death. Mol. Cell 57, 860–872 (2015).
Seth, R. B., Sun, L., Ea, C. K. & Chen, Z. J. Identification and characterization of MAVS, a mitochondrial antiviral signaling protein that activates NF-κB and IRF3. Cell 122, 669–682 (2005).
Shokolenko, I. N., Wilson, G. L. & Alexeyev, M. F. Persistent damage induces mitochondrial DNA degradation. DNA Repair 12, 488–499 (2013).
Reynders, K., Illidge, T., Siva, S., Chang, J. Y. & De Ruysscher, D. The abscopal effect of local radiotherapy: using immunotherapy to make a rare event clinically relevant. Cancer Treat. Rev. 41, 503–510 (2015).
Apetoh, L. et al. Toll-like receptor 4-dependent contribution of the immune system to anticancer chemotherapy and radiotherapy. Nat. Med. 13, 1050–1059 (2007).
Mackenzie, K. J. et al. cGAS surveillance of micronuclei links genome instability to innate immunity. Nature 548, 461–465 (2017).
Harding, S. M. et al. Mitotic progression following DNA damage enables pattern recognition within micronuclei. Nature 548, 466–470 (2017).
Jourdain, A. A., Boehm, E., Maundrell, K. & Martinou, J. C. Mitochondrial RNA granules: compartmentalizing mitochondrial gene expression. J. Cell Biol. 212, 611–614 (2016).
Fazal, F. M. et al. Atlas of subcellular RNA localization revealed by APEX-seq. Cell 178, 473–490 (2019).
Iacovoni, J. S. et al. High-resolution profiling of γH2AX around DNA double strand breaks in the mammalian genome. EMBO J. 29, 1446–1457 (2010).
Raab, M. et al. ESCRT III repairs nuclear envelope ruptures during cell migration to limit DNA damage and cell death. Science 352, 359–362 (2016).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Thomas, P. D. et al. PANTHER: a library of protein families and subfamilies indexed by function. Genome Res. 13, 2129–2141 (2003).
Mi, H. et al. PANTHER version 7: improved phylogenetic trees, orthologs and collaboration with the Gene Ontology Consortium. Nucleic Acids Res. 38, D204–D210 (2010).
Draghici, S. et al. A systems biology approach for pathway level analysis. Genome Res. 17, 1537–1545 (2007).
Stewart, S. A. et al. Lentivirus-delivered stable gene silencing by RNAi in primary cells. RNA 9, 493–501 (2003).
Sarbassov, D. D., Guertin, D. A., Ali, S. M. & Sabatini, D. M. Phosphorylation and regulation of Akt/PKB by the rictor-mTOR complex. Science 307, 1098–1101 (2005).
Bernardini, J. P. et al. Parkin inhibits BAK and BAX apoptotic function by distinct mechanisms during mitophagy. EMBO J. 38, e99916 (2019).
Liu, G. et al. Nuclear-resident RIG-I senses viral replication inducing antiviral immunity. Nat. Commun. 9, 3199 (2018).
Schuster, S., Tholen, L. E., Overheul, G. J., van Kuppeveld, F. J. M. & van Rij, R. P. Deletion of cytoplasmic double-stranded RNA sensors does not uncover viral small interfering RNA production in human cells. mSphere 2, e00333-17 (2017).
Kim, E. et al. Precision genome engineering with programmable DNA-nicking enzymes. Genome Res. 22, 1327–1333 (2012).
Schindelin, J. et al. Fiji: an open-source platform for biological-image analysis. Nat. Methods 9, 676–682 (2012).
Kitay, B. M., McCormack, R., Wang, Y., Tsoulfas, P. & Zhai, R. G. Mislocalization of neuronal mitochondria reveals regulation of Wallerian degeneration and NMNAT/WLD(S)-mediated axon protection independent of axonal mitochondria. Hum. Mol. Genet. 22, 1601–1614 (2013).
Valente, A. J., Maddalena, L. A., Robb, E. L., Moradi, F. & Stuart, J. A. A simple ImageJ macro tool for analyzing mitochondrial network morphology in mammalian cell culture. Acta Histochem. 119, 315–326 (2017).
Faccenda, D., Tan, C. H., Seraphim, A., Duchen, M. R. & Campanella, M. IF1 limits the apoptotic-signalling cascade by preventing mitochondrial remodelling. Cell Death Differ. 20, 686–697 (2013).
Jourdain, A. A. et al. A mitochondria-specific isoform of FASTK is present in mitochondrial RNA granules and regulates gene expression and function. Cell Rep. 10, 1110–1121 (2015).
Acknowledgements
We thank R. Greenberg, E. Brunet and D. E. Levy for providing reagents; S. Zaganelli, J.-C. Martinou (Université de Genève) and D. Moreau (ACCESS Geneva) for help with assessing the effect of irradiation on mtRNA granules; the Genome Technology Center (GTC) and the Microscopy Laboratory at NYU School of Medicine; and E. Lazzerini-Denchi, L. Walton Masters, A. Barrientos, F. Fontanesi and members of the Sfeir laboratory for reading the manuscript. This work was supported by the David and Lucile Packard Foundation (A.S.), Mallinckrodt Scholars Program (A.S.), Pew-Innovator fund (A.S.) and Pershing Square award (A.S.).
Author information
Authors and Affiliations
Contributions
A.S. and M.T. conceived the experimental design. M.T. performed all experiments with help from Y.F. (Extended Data Fig 2), S.T.-B. (Figs. 3a, f, 4e) and D.C.V. (Fig. 1b, e and Extended Data Figs. 2, 5, 9). A.S. and M.T. wrote the manuscript. All authors discussed the results and commented on the manuscript.
Corresponding author
Ethics declarations
Competing interests
A.S. is a co-founder, consultant and shareholder in Repare Therapeutics. The other authors declare no competing interests.
Additional information
Peer review information Nature thanks the anonymous reviewers for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Mitochondria-targeted TALENs as a tool to trigger mtDSBs.
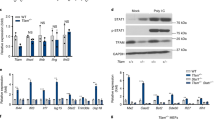
a, Schematic of a circular mtDNA molecule (16.6 kb) depicting the cleavage sites of mTLND-loop, mTLNND5 and mTLNATP8. Both origins of replication, OriH and OriL, are highlighted. b, Sanger sequencing denoting the C>A base substitution in FokI that generates the corresponding Asp>Ala missense mutation that blocks the catalytic activity of the enzyme. c, Representative immunofluorescence images to visualize the mitochondrial localization of mTLNATP8 and dmTLNATP8. mTLNs are identified with a HA antibody (green); mitochondria are counterstained with mito-dsRed (red) and nuclei with DAPI (blue). d, Western blot analysis using the anti-HA antibody to mark the expression of the indicated mTLN, 20 h after transfection (RNA from the same samples were analysed using RNA sequencing and are reported in Fig. 1). e, qPCR for mtDNA copy number in the same samples as shown in d. Cells were analysed 48 h after transfection with the indicated mTLN and controls. Negative (-ve) control represents cells transfected with no plasmid and positive (+ve) control represent cells transfected with calf thymus DNA. Normalized expression data as mean ± s.d. from 1 biological and 3 technical replicates and the normalized error are shown. f, Representative immunofluorescence image showing the absence of nuclear DNA damage marked by the formation of 53BP1 foci (green) in cells treated with mTLN. g, Quantification of 53BP1 foci from f. Mock transfection with no DNA serves as a negative control. As a positive control, cells were exposed to irradiation (20 Gy) followed by 1-h recovery. Data are the mean percentage ± s.d. of n = 3 biological replicates; two-tailed unpaired Student’s t-test with CI = 95%. ns, not significant. h, Western blot analysis of phosphorylated Chk2 (T68), 48 h after mTLN expression. Irradiated cells served as a positive control. i, RT–qPCR of RSAD2 in A549 (n = 6 biological replicates), U2-OS (n = 3 biological replicates) and RPE-1 (n = 4 biological replicates) cells treated with mTLN and analysed 48 h after transfection. Expression levels in mTLND-loop-treated cells were normalized to control cells treated with dmTLND-loop and ratios are plotted as mean ± s.d.; one-tailed ratio paired Student’s t-test with CI = 95%, P values are shown at the top of the graph.
Extended Data Fig. 2 mtDSBs activate ISGs.
a, Extended statistics for Fig. 1b. Cells were transfected with mTLND-loop or dmTLND-loop and collected 48 h later. RT–qPCR analysis of mitochondrial factors was performed and data were normalized to dmTLND-loop expression. Data are mean ± s.d. of n = 5 independent biological experiments; one-tailed ratio paired Student’s t-test with CI = 95%, P values are shown at the top of the graph. b, Additional validation of mitochondrial genes shown in Fig. 1b, using an independent mTLN. For each gene, expression levels in mTLNND5-expressing cells were normalized to cells treated with dmTLNND5 collected 48 h after transfection. Normalized data as mean ± s.d. from RT–qPCR for two independent biological replicates, with additional calf thymus DNA and no DNA controls. c, Extended statistics for additional independent experiments as in b. Data are mean ± s.d. of n = 5 independent biological experiments; one-tailed ratio paired Student’s t-test with CI = 95%. d, Extended statistics for Fig. 1c, e. Cells were transfected with mTLNATP8 or dmTLNATP8 and collected 48 h later. RT–qPCR analysis of cytoplasmic ISGs was performed and fold expression was calculated by normalizing to dmTLNATP8 expression. For all genes, n = 4 biological replicates were analysed, except ISG15 (n = 5). Data are mean ± s.d.; one-tailed ratio paired Student’s t-test with CI = 95%. e, Experiment as in Fig. 1e. Cells transfected with mTLNATP8 or dmTLNATP8 were collected 24 h or 48 h later. Normalized data as mean ± s.d. from RT–qPCR for two independent biological replicates. f, Extended statistics for additional RT–qPCR validation of nuclear-DNA-encoded genes (as per Fig. 1c, e) after transfection with mTLND-loop and normalized to the dead counterpart (dmTLND-loop). Data are mean ± s.d. of n = 7 independent biological replicates for ISG15, n = 5 for all other genes; one-tailed ratio paired Student’s t-test with CI = 95%. g, Two independent experiments for mTLND-loop are shown as described in e and Fig. 1e. Normalized data as mean ± s.d.from RT–qPCR. h, Extended statistics for additional validation experiments of the activation of cytoplasmic ISGs after transfection with a third independent TLN, which cuts in ND5. Cells were collected 48 h after transfection with mTLNND5 and fold expression ratios were calculated by normalizing to dmTLNND5. Data are mean ± s.d. of n = 6 independent experiments for ISG15, n = 3 for all other genes; one-tailed ratio paired Student’s t-test with CI = 95%.
Extended Data Fig. 3 Paracrine signalling in response to mtDSBs.
a, RT–qPCR analysis of IFNB1, ISG15 and RSAD2 in cells with the indicated treatment that were collected 48 h after transfection. Bar graphs, expression levels were normalized to the indicated samples. Mean ± s.d.for n = 1 biological, 3 technical replicates. Scatter plot, data are mean ± s.d. of n = 3 biological replicates; one-tailed unpaired Student’s t-test with CI = 95%. b, Western blot analysis for pSTAT1(Y701) in naive cells cultured with medium transferred from cells with the indicated treatment as in a. c, qPCR analysis to measure mtDNA copy number in ARPE-19 cells treated with ethidium bromide (EtBr) (50 ng ml−1 for 6 weeks) or ddC (5 μM for 15 days). d, Representative image displaying the mitochondrial network in cells treated with ddC, analysed in e. e, Analysis of mitochondrial length and branching in the indicated ARPE-19 cells. Data are mean ± s.d.; individual cells are shown as circles; two-tailed Welch’s t-test with CI = 95%.
Extended Data Fig. 4 Investigating the effect of BAX–BAK macropore formation and cGAS sensing during immune activation in response to mtDSBs.
a, Western blot analysis of pSTAT1(Y701) in ARPE-19 cells with the indicated treatment. b, Western blot analysis of BAX and BAK in BAX−/−BAK−/− ARPE-19 cells generated with CRISPR–Cas9 targeting (see Methods for details on double-knockout generation). c, RT–qPCR analysis of RSAD2, IFI44 and ISG15 in ARPE-19 cells treated with 400 μM BAX V5 inhibitor during the indicated mTLN treatments. For each gene, expression levels in mTLND-loop-treated cells were normalized to control cells treated with dmTLND-loop and collected 48 h after transfection. Mean ± s.d. for n = 1 biological and 3 technical replicates are shown. d, Similar to c but using BAX V5 inhibitor as well as 20 μM of the pan-caspase inhibitor Q-Vd-Ph for 48 h during mTLN treatment. Mean ± s.d. for n = 1 biological and 3 technical replicates are shown. Scatter plot shows the mean ± s.d. of n = 2 and 3 biological replicates for c and d, respectively; two-tailed unpaired Student’s t-test with CI = 95%. e, Western blot analysis to monitor caspase cleavage in cells treated with the indicated mTLN or dmTLN. ABT-737 serves as a positive control f, Assessment of mRNA levels of different DNA and RNA cytosolic sensors in ARPE-19 cells based on read abundance from RNA-seq (from Fig. 1). g, Western Blot surveying the expression of DNA/RNA sensing proteins in ARPE-19 and MCF10A cells with the indicated treatment. h, Bar graphs, RT–qPCR analysis of RSAD2 and ISG15 in ARPE-19 cells with exogenously expressed cGAS, collected 48 h after transfection with mTLND-loop or dead counterpart. Mean ± s.d. for n = 1 biological replicate and 3 technical replicates are shown. Scatter plot, ratio of activation of ISG15 and RSAD2 with or without exogenously expressed cGAS; n = 2 biological replicates. Although expression of cGAS increases ISG expression in response to transfection, the ratio of ISG induction in mTLND-loop compared with dmTLND-loop did not increase.
Extended Data Fig. 5 Investigating the role of cytoplasmic sensors in innate immune signalling in response to mtDSB.
a, b, Western blot analyses of STING in cells with the indicated genotype after CRISPR targeting and isolation of clonally derived lines (as described in the Methods). c, Western blot analysis of RIG-I in ARPE-19 cells with the indicated genotype. d, Detection of RIG-I protein levels upon complementation of DDX58−/− ARPE-19 cells with wild-type and catalytically inactive RIG-I(K270A). e, Western blot analysis of MDA5 in cells with the indicated genotype after CRISPR targeting and isolation of clonally derived lines. f, RT–qPCR analysis of ISG15 and RSAD2 in DDX58−/− ARPE-19 cells complemented with wild-type or RIG-I(K270A) with the indicated treatment. Expression levels of ISGs in mTLND-loop-treated cells were normalized to cells treated with dmTLND-loop collected 48 h after transfection. Bar graphs, mean ± s.d. of n = 1 biological and 3 technical replicates are shown. Scatter plot, data are represents mean ± s.d. of n = 2 biological replicates. g, Western blot analysis of MAVS and STING in cells with the indicated genotype after CRISPR targeting and isolation of clonally derived lines. e, g, Analysed clones are numbered and highlighted in a red box.
Extended Data Fig. 6 Characterization of cellular responses to irradiation in wild-type and Rho0 MCF10A cells.
a, qPCR analysis to evaluate mtDNA copy numbers in MCF10A cells with the indicated treatments. Wild-type and ddC-treated cells were exposed to 20 Gy irradiation and collected 6 days later. Left, bar graph, normalized mtDNA copy number data are mean ± s.d. of n = 1 biological and 3 technical replicates. Blue, cells treated with ddC for 10 + 6 days; purple, cells in which ddC treatment was discontinued after 10 days, after which the cells were exposed to irradiation and collected 6 days after irradiation and ddC withdrawal. Right, scatter plot, mean ± s.d. of n = 3 biological replicates; one-way analysis of variance (ANOVA) with CI = 95%. b, Analysis of mtDNA copy number in cells treated with 20 Gy irradiation at indicated times. Data are mean ± s.d. of n = 2 biological replicates. c, Western blot analysis of pSTAT1(Y701) in MCF10A wild-type and ddC-treated cells after 20 Gy irradiation and 6 days of incubation. Data are shown for two independent experiments. d, RT–qPCR showing expression of ISG15 and RSAD2 in MCF10A cells subjected to increasing levels of irradiation and collected 6 days after. Data are mean ± s.d. of the indicated number of independent biological replicates; one-tailed unpaired Student’s t-test with CI = 95%. e, Western blot analysis for pChk2 and pSTAT1(Y701) in wild-type and Rho0 MCF10A cells collected 24 h after treatment with the indicated DNA damaging agents (zeocin (zeo) and etoposide (eto)) or bacterial lipopolysaccharide (LPS). f, Cell cycle analysis of MCF10A cells with the indicated treatment. Cells were analysed 6 days after irradiation (20 Gy). Data are mean ± s.d. of n = 3 independent biological replicates. g, Western blot analysis of pChk2(T68) in wild-type and Rho0 (ddC-treated) MCF10A cells collected at the indicated time points after irradiation (20 Gy). h, RT–qPCR for ISG15 and RSAD2 in wild-type and Rho0 (ddC-treated) MCF10A cells after the indicated treatments. Data are mean ± s.d. of n = 3 biological replicates. i, Western blot analysis of pSTAT1(Y701) in Rho0 (ddC-treated) MCF10A cells treated with IFNβ1 for 24 h. j, Western blot analysis of pSTAT1(Y701) in wild-type and Rho0 (ddC-treated) MCF10A cells transfected with calf thymus DNA. k, Quantification of the two biological replicates described in j. Mean ± s.e.m.
Extended Data Fig. 7 BAX–BAK macropore formation in response to irradiation.
a, Additional representative images for Fig. 4e. Active BAX was recognized with the specific 6A7 monoclonal antibody. ARPE-19 cells were treated with 20 Gy irradiation and incubated for 72 h before staining. Shown are images of untreated cells and cells incubated for 72 h after 20 Gy irradiation. As a positive control, ARPE-19 cells were treated with staurosporine 5 μM for 24 h to induce apoptosis. b, Western blot analysis of cleaved caspase-3 showing no detectable apoptosis 0–6 days after delivery of 20 Gy irradiation. As a positive control, cells were incubated with ABT-737 and S63845 for 3 h. c, RT–qPCR showing RSAD2 expression in wild-type and BAX−/−BAK−/− cells after 20 Gy irradiation. Data are mean ± s.d. of n = 3 biological replicates; one-tailed unpaired Student’s t-test with CI = 95%.
Extended Data Fig. 8 Irradiation and mTLN expression trigger similar transcriptional responses.
a, Volcano plot representation of expression profiles for deregulated genes after irradiation of Rho0 MCF10A cells versus mtDNA-proficient cells. The log2 ratio (fold change) against the negative log10[P value] (calculated by DeSeq2 using a Wald test) is shown. Red dots highlight factors in the RNA-sensing pathway. b, Negative log[fold change] values of genes that were not upregulated in irradiated Rho0 MCF10A cells compared with cells proficient for mitochondrial genomes. The differential expression was calculated through Rosalind and overlaid onto components of the cytosolic DNA-sensing pathway (GO term) using Advaita iPathwayGuide software (P = 0.027, false discovery rate correction, enrichment calculated using Advaita’s iPathwayGuide; Methods, ‘Genome-wide RNA-sequencing and bioinformatics analysis’). c, log[fold change] values of genes deregulated after mTLN treatment (compared to dmTLN) as in Fig. 1c and those deregulated upon irradiation of mtDNA-proficient versus -deficient cells (Fig. 4a) were analysed for co-regulation through Rosalind (meta-analysis) and overlaid on the cytosolic DNA-sensing pathway GO term using Advaita iPathwayGuide (P = 0.004, false discovery rate correction, enrichment calculated using Advaita’s iPathwayGuide; Methods, ‘Genome-wide RNA-sequencing and bioinformatics analysis’).
Extended Data Fig. 9 The RIG-I RNA-sensing machinery is a major pathway that is activated when cells are exposed to irradiation.
a, b, Western blot analysis of RIG-I (a) and MAVS (b) in cells with the indicated genotype after CRISPR targeting and isolation of clonal lines (as described in the Methods). c, The ratio of pSTAT1/GAPDH is shown. Quantification of western blot signals of pSTAT1 and GAPDH in cells with the indicated genotype, 6 days after 20 Gy irradiation is shown (as in Fig. 4f). Data are mean of n = 3 or >3 biological replicates; ordinary one-way ANOVA with CI = 95%. d, Western blot analysis to confirm depletion of RIG-I and TBK1 in cells transduced with the indicated shRNA vectors. e, Western blot analysis of pSTAT1(Y701) in MCF10A cells depleted of DDX58 and TBK1 with shRNA. Blots from three out of four independent experiments quantified in f are shown. f, The ratio of pSTAT1/GAPDH is shown. Quantification of western blot signals of pSTAT1 and GAPDH in cells from e. Data are mean ± s.d. of n = 4 biological replicates; ordinary one-way ANOVA with CI = 95%.
Extended Data Fig. 10 Detection of mtRNA in the cytoplasm after irradiation.
a, Representative images of cells at the indicated time points after irradiation treatment and stained with anti-FASTKD2 to mark mtRNA granules in green, mitochondria in red (cytochrome c) and DAPI to highlight nuclei. Images were acquired with high-throughput ImageXpress Micro Confocal platform (Molecular Devices). b, Quantification of the average area and average integrated intensity of mitochondrial FASTKD2 foci during a 6-day time course after delivery of 20 Gy. Data analysis was performed with MetaExpress software (Molecular Devices). DAPI-, mitochondria- and FASTKD2-specific masks were generated and FASTKD2 signal parameters were computed in the areas overlaying mitochondria. Data were further analysed using Microsoft Excel and normalized to the total number of nuclei or number of granules. Fields with a minimum of 3 and a maximum of 60 nuclei were considered for analysis. Data are mean and individual fields of view; ordinary one-way ANOVA with Dunnett’s multiple comparison correction with CI = 95%. Results were reproduced in two biological replicates. c, Subcellular fractionation was performed as described in the Methods section ‘Subcellular fractionation for cytoplasmic mtRNA quantification’. MCF10A cells were irradiated with 15 Gy and incubated for 3 and 6 days; equal volumes of the indicated fractions were resolved by SDS–PAGE. Immunoblotting was performed to ensure the quality of the fractions using antibodies against VDAC (mitochondria), lamin B1 (nuclear membrane) and GAPDH (cytosol). For further details, see Methods. d, Cytosolic fractions from c were used to purify RNAs accumulated in the cytosol 3 days after irradiation and quantify those of mitochondrial origin using RT–qPCR. The amount of each mtRNA tested is expressed as a fold ratio between irradiated and unirradiated cells. Data are mean fold change ± s.d. of n = 3 biological replicates; one-tailed ratio paired Student’s t-test with CI = 95%. e, DDX58−/− ARPE-19 cells were complemented by lentiviral delivery with a fusion of RIG-I and APEX2. This construct was placed under the control of a SFFV promoter and translated as a fusion with a GFP moiety that is self-processed by T2A cleavage. After infection, GFP-expressing cells were sorted and used for proximity biotinylation experiments. RIG-I–APEX2 expression was confirmed by western blot.
Extended Data Fig. 11 Investigating the interplay between nuclear and mitochondrial DNA damage.
a, Representative immunofluorescence images monitoring 53BP1 formation in cells after 6 h of AsiSI induction with Shield-1 and 4-OHT or 1 h after irradiation (20 Gy). 53BP1 is shown in green and DAPI in blue. The total percentage of counted cells with more than 5 foci per nucleus is shown under each image. b, Western blot analysis of pChk2(T68) in MCF10A cells expressing an inducible AsiSI (AsiSIInd). Cells were collected 1 h after irradiation (20 Gy) or after 6 h of treatment with 4-OHT and Shield-1, which induces AsiSI expression and translocation to the nucleus. c, Representative immunofluorescence images of cGAS+ micronuclei analysed in MCF10A expressing AsiSIInd 3 days after AsiSI induction (treatment with Shield-1 and 4-OHT for 6 h) or irradiation (20 Gy). cGAS is labelled in red and DAPI in blue. d, Quantification of c. The total percentage of cells with at least 1 cGAS+ micronucleus is shown. Data are mean ± s.d. of n = 5 biological replicates; ordinary one-way ANOVA with CI = 95%. e, Western blot analysis of pSTAT1 in MCF10A cells expressing AsiSIInd. Cells were collected 6 days after 20 Gy irradiation and after withdrawal of 4-OHT and Shield-1 (24 h treatment). f, Western blot analysis of pSTAT1 in ARPE-19 cells expressing AsiSIInd (transduced with cGAS–GFP) treated and analysed as in e. g, The ratio of pSTAT1/GAPDH is shown. Quantification of western blot signals of pSTAT1 and GAPDH obtained from independent experiments as in e and f. Data are mean ± s.d. of n = 3 or 4 independent biological replicates (shown as dots); ordinary one-way ANOVA with CI = 95%. h, RT–qPCR analysis of RSAD2 and ISG15 in AsiSIInd MCF10A and AsiSIInd ARPE-19 cells after 20 Gy irradiation or 24 h of AsiSI induction (for a total of 6 days). The fold increase in ISG expression using untreated cells for normalization is shown. Data are mean ± s.d. of n = 4 or 6 independent biological replicates (shown as dots), ordinary one-way ANOVA with CI = 95%. i, AsiSIInd ARPE-19 cells were induced with 4-OHT and Shield-1 for a total of 24 h, incubated for an additional 24 h, treated with the indicated mTLNs and collected 48 h after transfection (4 days after the initial AsiSI induction). In each experiment, ISG15 and RSAD2 expression in mTLND-loop-treated AsiSI-induced cells was normalized to control cells treated with dmTLND-loop in the absence of AsiSI induction. Data are mean fold expression ± s.d. of n = 6 biological replicates; one-tailed ratio paired Student’s t-test with CI = 95%. j, Cells treated as in i were collected and analysed for cell cycle stage. Data are mean ± s.d. of n = 3 biological replicates.
Supplementary information
Supplementary Data
Source Data Figure 1: Uncropped Western Blot. Provided are uncropped Western blot images for all figures. Pseudo-colors are introduced during multichannel imaging on ChemiDocMP Biorad running on ImageLab 5.2. Exposures might be different from main figures as MW marker imaging was performed as last step after the longest exposure. MW bands are in blue, size is indicated.
Supplementary Data
Source Data Figure 2: FACS Gating Strategies. The manuscript did not require complex gating strategies. Reported are three examples of in three different flow cytometry analysis. In general, live cells were first gated through SSC-A vs FSC-A plot, single cells were then gated through FSC-A vs FSC-H plot. Final gating was experiment specific and based on positive or negative controls and performed by plotting fluorescence intensity vs. number of events. Cell cycle analysis was performed by correlating the PE-bound DNA signal to cell size, and phase was attributed by FlowJo 10.0 Cell Cycle Biological analysis package.
Supplementary Data
Source Data Figure 3: GO Analysis. Due space limitation on main figures, the GO analysis presented in Figure 1d was modified by removing certain redundant GO terms. Here, we provide original version, including official GO code.
Supplementary Tables
This file contains Supplementary Tables 1-3 and a Guide.
Rights and permissions
About this article
Cite this article
Tigano, M., Vargas, D.C., Tremblay-Belzile, S. et al. Nuclear sensing of breaks in mitochondrial DNA enhances immune surveillance. Nature 591, 477–481 (2021). https://doi.org/10.1038/s41586-021-03269-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-021-03269-w
This article is cited by
-
Construction of an ER stress-related prognostic signature for predicting prognosis and screening the effective anti-tumor drug in osteosarcoma
Journal of Translational Medicine (2024)
-
Ancestral allele of DNA polymerase gamma modifies antiviral tolerance
Nature (2024)
-
When DNA-damage responses meet innate and adaptive immunity
Cellular and Molecular Life Sciences (2024)
-
Neuronal STING activation in amyotrophic lateral sclerosis and frontotemporal dementia
Acta Neuropathologica (2024)
-
Unscheduled excessive R-loops in immune response
Functional & Integrative Genomics (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.