Abstract
Inflammasomes are important sentinels of innate immune defence that are activated in response to diverse stimuli, including pathogen-associated molecular patterns (PAMPs)1. Activation of the inflammasome provides host defence against aspergillosis2,3, which is a major health concern for patients who are immunocompromised. However, the Aspergillus fumigatus PAMPs that are responsible for inflammasome activation are not known. Here we show that the polysaccharide galactosaminogalactan (GAG) of A. fumigatus is a PAMP that activates the NLRP3 inflammasome. The binding of GAG to ribosomal proteins inhibited cellular translation machinery, and thus activated the NLRP3 inflammasome. The galactosamine moiety bound to ribosomal proteins and blocked cellular translation, which triggered activation of the NLRP3 inflammasome. In mice, a GAG-deficient Aspergillus mutant (Δgt4c) did not elicit protective activation of the inflammasome, and this strain exhibited enhanced virulence. Moreover, administration of GAG protected mice from colitis induced by dextran sulfate sodium in an inflammasome-dependent manner. Thus, ribosomes connect the sensing of this fungal PAMP to the activation of an innate immune response.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
The datasets generated and analysed in this study are contained within the Article, and its Supplementary Information; any other relevant data are available from the corresponding author upon reasonable request. Source data are provided with this paper.
Change history
21 December 2020
A Correction to this paper has been published: https://doi.org/10.1038/s41586-020-03088-5
References
Man, S. M., Karki, R. & Kanneganti, T. D. Molecular mechanisms and functions of pyroptosis, inflammatory caspases and inflammasomes in infectious diseases. Immunol. Rev. 277, 61–75 (2017).
Briard, B. et al. Fungal ligands released by innate immune effectors promote inflammasome activation during Aspergillus fumigatus infection. Nat. Microbiol. 4, 316–327 (2019).
Karki, R. et al. Concerted activation of the AIM2 and NLRP3 inflammasomes orchestrates host protection against Aspergillus infection. Cell Host Microbe 17, 357–368 (2015).
Saïd-Sadier, N., Padilla, E., Langsley, G. & Ojcius, D. M. Aspergillus fumigatus stimulates the NLRP3 inflammasome through a pathway requiring ROS production and the Syk tyrosine kinase. PLoS ONE 5, e10008 (2010).
Latgé, J. P., Beauvais, A. & Chamilos, G. The cell wall of the human fungal pathogen Aspergillus fumigatus: biosynthesis, organization, immune response, and virulence. Annu. Rev. Microbiol. 71, 99–116 (2017).
Fontaine, T. et al. Galactosaminogalactan, a new immunosuppressive polysaccharide of Aspergillus fumigatus. PLoS Pathog. 7, e1002372 (2011).
Gravelat, F. N. et al. Aspergillus galactosaminogalactan mediates adherence to host constituents and conceals hyphal β-glucan from the immune system. PLoS Pathog. 9, e1003575 (2013).
Hua, K. F. et al. Capsular polysaccharide is Involved in NLRP3 inflammasome activation by Klebsiella pneumoniae serotype K1. Infect. Immun. 83, 3396–3409 (2015).
Wolf, A. J. et al. Hexokinase is an innate immune receptor for the detection of bacterial peptidoglycan. Cell 166, 624–636 (2016).
Briard, B., Muszkieta, L., Latgé, J. P. & Fontaine, T. Galactosaminogalactan of Aspergillus fumigatus, a bioactive fungal polymer. Mycologia 108, 572–580 (2016).
Lee, M. J. et al. Deacetylation of fungal exopolysaccharide mediates adhesion and biofilm formation. MBio 7, e00252-16 (2016).
Tzianabos, A. O. Polysaccharide immunomodulators as therapeutic agents: structural aspects and biologic function. Clin. Microbiol. Rev. 13, 523–533 (2000).
Zhou, X., Liao, W. J., Liao, J. M., Liao, P. & Lu, H. Ribosomal proteins: functions beyond the ribosome. J. Mol. Cell Biol. 7, 92–104 (2015).
Vyleta, M. L., Wong, J. & Magun, B. E. Suppression of ribosomal function triggers innate immune signaling through activation of the NLRP3 inflammasome. PLoS One 7, e36044 (2012).
Bronner, D. N. et al. Endoplasmic reticulum stress activates the inflammasome via NLRP3- and caspase-2-driven mitochondrial damage. Immunity 43, 451–462 (2015).
Hetz, C. The unfolded protein response: controlling cell fate decisions under ER stress and beyond. Nat. Rev. Mol. Cell Biol. 13, 89–102 (2012).
Samir, P. et al. DDX3X acts as a live-or-die checkpoint in stressed cells by regulating NLRP3 inflammasome. Nature 573, 590–594 (2019).
Man, S. M. et al. Differential roles of caspase-1 and caspase-11 in infection and inflammation. Sci. Rep. 7, 45126 (2017).
Dupaul-Chicoine, J. et al. Control of intestinal homeostasis, colitis, and colitis-associated colorectal cancer by the inflammatory caspases. Immunity 32, 367–378 (2010).
Zaki, M. H. et al. The NLRP3 inflammasome protects against loss of epithelial integrity and mortality during experimental colitis. Immunity 32, 379–391 (2010).
Chadani, Y. et al. Intrinsic ribosome destabilization underlies translation and provides an organism with a strategy of environmental sensing. Mol. Cell 68, 528–539.e5 (2017).
van der Horst, S., Filipovska, T., Hanson, J. & Smeekens, S. Metabolite control of translation by conserved peptide uORFs: the ribosome as a metabolite multisensor. Plant Physiol. 182, 110–122 (2020).
Menu, P. et al. ER stress activates the NLRP3 inflammasome via an UPR-independent pathway. Cell Death Dis. 3, e261 (2012).
Chen, J. & Chen, Z. J. PtdIns4P on dispersed trans-Golgi network mediates NLRP3 inflammasome activation. Nature 564, 71–76 (2018).
Muñoz-Planillo, R. et al. K+ efflux is the common trigger of NLRP3 inflammasome activation by bacterial toxins and particulate matter. Immunity 38, 1142–1153 (2013).
Topf, U. et al. Quantitative proteomics identifies redox switches for global translation modulation by mitochondrially produced reactive oxygen species. Nat. Commun. 9, 324 (2018).
Willi, J. et al. Oxidative stress damages rRNA inside the ribosome and differentially affects the catalytic center. Nucleic Acids Res. 46, 1945–1957 (2018).
Lamarre, C. et al. Galactofuranose attenuates cellular adhesion of Aspergillus fumigatus. Cell. Microbiol. 11, 1612–1623 (2009).
da Silva Ferreira, M. E. et al. The akuBKU80 mutant deficient for nonhomologous end joining is a powerful tool for analyzing pathogenicity in Aspergillus fumigatus. Eukaryot. Cell 5, 207–211 (2006).
Kayagaki, N. et al. Non-canonical inflammasome activation targets caspase-11. Nature 479, 117–121 (2011).
Kanneganti, T. D. et al. Bacterial RNA and small antiviral compounds activate caspase-1 through cryopyrin/Nalp3. Nature 440, 233–236 (2006).
Jones, J. W. et al. Absent in melanoma 2 is required for innate immune recognition of Francisella tularensis. Proc. Natl Acad. Sci. USA 107, 9771–9776 (2010).
Karki, R. et al. IRF8 regulates transcription of Naips for NLRC4 inflammasome activation. Cell 173, 920–933 (2018).
Hartmann, T. et al. Validation of a self-excising marker in the human pathogen Aspergillus fumigatus by employing the β-rec/six site-specific recombination system. Appl. Environ. Microbiol. 76, 6313–6317 (2010).
Briard, B. et al. Dirhamnolipids secreted from Pseudomonas aeruginosa modify anjpegungal susceptibility of Aspergillus fumigatus by inhibiting β1,3 glucan synthase activity. ISME J. 11, 1578–1591 (2017).
Muszkieta, L. et al. Deciphering the role of the chitin synthase families 1 and 2 in the in vivo and in vitro growth of Aspergillus fumigatus by multiple gene targeting deletion. Cell. Microbiol. 16, 1784–1805 (2014).
Dubois, M., Gilles, K., Hamilton, J. K., Rebers, P. A. & Smith, F. A colorimetric method for the determination of sugars. Nature 168, 167 (1951).
Stalhberger, T. et al. Chemical organization of the cell wall polysaccharide core of Malassezia restricta. J. Biol. Chem. 289, 12647–12656 (2014).
Plassard, C. S., Mousain, D. G. & Salsac, L. E. Estimation of mycelial growth of basidiomycetes by means of chitin determination. Phytochemistry 21, 345–348 (1982).
Fontaine, T., Talmont, F., Dutton, G. G. S. & Fournet, B. Analysis of pyruvic acid acetal containing polysaccharides by methanolysis and reductive cleavage methods. Anal. Biochem. 199, 154–161 (1991).
Gressler, M. et al. Definition of the anti-inflammatory oligosaccharides derived from the galactosaminogalactan (GAG) from Aspergillus fumigatus. Front. Cell. Infect. Microbiol. 9, 365 (2019).
Samir, P. et al. Identification of changing ribosome protein compositions using mass spectrometry. Proteomics 18, 1800217 (2018).
Werner, J. L. et al. Requisite role for the dectin-1 β-glucan receptor in pulmonary defense against Aspergillus fumigatus. J. Immunol. 182, 4938–4946 (2009).
Gresnigt, M. S. et al. A polysaccharide virulence factor from Aspergillus fumigatus elicits anti-inflammatory effects through induction of interleukin-1 receptor antagonist. PLoS Pathog. 10, e1003936 (2014).
Schindelin, J. et al. Fiji: an open-source platform for biological-image analysis. Nat. Methods 9, 676–682 (2012).
Mouyna, I. et al. GH16 and GH81 family β-(1,3)-glucanases in Aspergillus fumigatus are essential for conidial cell wall morphogenesis. Cell. Microbiol. 18, 1285–1293 (2016).
Acknowledgements
We thank members of the Kanneganti laboratory for their comments, suggestions and technical assistance; R. Tweedell for scientific editing of the manuscript; the St Jude Children’s Research Hospital Veterinary Pathology Core, SJCRH Center for Proteomics and Metabolomics and SJCRH Cell and Tissue Imaging Center (supported by the NCI P30 CA021765); D. Sheppard for sharing the A. fumigatus deletion mutant ∆agd; and V. M. Dixit and N. Kayagaki for the Casp1−/−Casp11−/− mutant mouse strain. T.-D.K. is supported by NIH grants AI101935, AI124346, AR056296 and CA253095 and by the American Lebanese Syrian Associated Charities. The content is solely the responsibility of the authors and does not necessarily represent the official views of the National Institutes of Health. J.-P.L. is supported by the Aviesan project Aspergillus, the French Government’s Investissement d’Avenir program, Laboratoire d’Excellence ‘Integrative Biology of Emerging Infectious Diseases’ (grant number ANR-10-LABX-62-IBEID) and la Fondation pour la Recherche Médicale (DEQ20150331722 LATGE Equipe FRM 2015).
Author information
Authors and Affiliations
Contributions
B.B. and T.-D.K. conceptualized the study and designed the experiments. B.B., P.S., D.E.P., R.K.S.M., R.K. and S.C. performed the in vitro experiments with BMDMs. B.B., D.E.P., R.K. and S.C. performed the in vivo experiments. T.F., L.M., P.B., R.B., E.M., O.I.-G. and B.H. generated and biochemically characterized the Δgt4c Aspergillus strain. P.V. analysed the in vivo pathology. B.B., P.S. and R.C.K. performed the polysome profiling. P.B. and C.R. performed the electron microscopy. B.B., T.F., J.-P.L. and T.-D.K. analysed the data. B.B. and T.-D.K. wrote the paper. T.-D.K. and J.-P.L. obtained funding. T.-D.K. supervised the study.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature thanks Gordon D. Brown, Osamu Takeuchi and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Identification of A. fumigatus GAG synthase.
a, Schematic of GAG synthase cluster. b, RNA sequencing analysis during A. fumigatus growth (0, 4 and 8 h) from ref. 46; gene expression is represented by mean of normalized read count per gene for the GT4C cluster. n = 3 biologically independent samples. Data are mean ± s.e.m. c, Heat map showing differential gene expression of A. fumigatus at 4 h (swollen conidia) and 8 h (germinated conidia) compared to 0 h (resting conidia). d, Schematic of the GT4C protein with transmembrane regions (black), α-glycosyltransferase domains (green) and major facilitator superfamily domain (MFS) predicted from amino acid sequence with InterProscan 5. e, Schematic of wild-type (WT) and Δgt4c locus with NcoI restriction sites and Southern blot probe used to control the GT4C gene deletion. f, Southern blot using GT4C probe with WT and Δgt4c purified DNA from one experiment. g, RT–PCR analysis of GT4C, AGD3, EGA3, SPH3 and UGE3 A. fumigatus genes in WT and Δgt4c strains (8 h in LB medium, 37 °C and 250 rpm) presented relative to that of TEF1. n.d., not detected. n = 3 biologically independent samples. Data are mean ± s.e.m.
Extended Data Fig. 2 Absence of GAG does not affect release of non-inflammasome dependent cytokines.
a, b, Release of IL-6 (a) and TNF (b) from unprimed BMDMs left uninfected (med.) or assessed 20 h after infection with A. fumigatus wild type (WT) or Δgt4c strain (MOI of 10). n = 3 independent biological samples. Data are mean ± s.e.m.
Extended Data Fig. 3 UGE3 potentiates A.-fumigatus-induced inflammasome activation.
a, Assessment of biofilm formation on an abiotic surface with A. fumigatus (A. f) wild type (WT) and deletion mutant strain Δuge3. b, Immunoblot analysis of pro-caspase 1 (pro-Casp1; p45) and the active caspase 1 subunit (p20) from unprimed BMDMs left untreated (medium alone (med.)) or measured 20 h after infection with the indicated A. fumigatus live resting conidia (MOI of 10). Representative images. n ≥ 3 independent experiments. c, Immunoblot analysis of phospho-IκBα and total IκBα (t-IκBα) or phospho-ERK1/2 (p-ERK) and total ERK1/2 (t-ERK) from unprimed WT BMDMs 0–8 h after infection with WT or Δuge3 mutant A. fumigatus live resting conidia (MOI of 10). Representative images. n ≥ 3 independent experiments. d, Immunoblot analysis of pro-IL-1β from unprimed BMDMs 0–8 h after infection with WT or Δuge3 mutant A. fumigatus live resting conidia (MOI of 10). Representative images. n ≥ 3 independent experiments. e–g, RT–PCR analysis of Nlrp3, Il1b and Tnf genes from WT BMDMs 0–8 h after infection with WT or Δuge3 mutant A. fumigatus live resting conidia presented relative to that of the gene encoding β-actin. n = 4 biologically independent samples. Data are mean ± s.e.m.
Extended Data Fig. 4 Over-synthesis of GAG induces hyper-inflammasome activation.
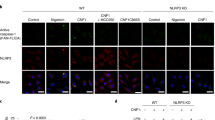
a, Immunofluorescence staining of A. fumigatus GAG (green) and BMDM nuclei (blue) in unprimed BMDMs 4 h after infection with A. fumigatus wild-type (WT) or Δugm1 resting conidia (MOI of 10). Scale bars, 10 μm. Representative images. n ≥ 3 independent experiments. b, Immunoblot analysis of pro-caspase 1 (pro-Casp1; p45) and the active caspase 1 subunit (p20) of unprimed BMDMs left untreated (medium alone (med.)) or assessed 20 h after infection with the indicated live A. fumigatus resting conidia genotype (WT or A. fumigatus deletion mutant Δugm1) (MOI of 10). Representative image. n ≥ 3 independent experiments. c, Release of IL-1β from unprimed BMDMs left uninfected (med.) or assessed 20 h after infection with A. fumigatus (MOI of 10). **P = 0.0046. Unpaired two-tailed t-test. n = 4 biologically independent samples. Data are mean ± s.e.m.
Extended Data Fig. 5 GAG induces caspase 1 cleavage in a dose- and charge–charge interaction-dependent manner and interacts with ribosomes.
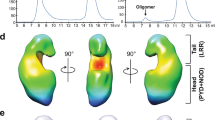
a, Representative images of BMDMs in medium (med.) or during treatment with DOTAP alone (green fluorescence corresponds to Sytox green nuclei, and Sytox green-positive nuclei are marked with a red circle). Scale bars, 10 μm. Representative images. n ≥ 3 independent experiments. b, Immunoblot analysis of pro-caspase 1 (pro-Casp1; p45) and the active caspase 1 subunit (p20) of BMDMs left untreated (medium alone (med.)) or assessed 3 h after transfection with increasing concentrations of GAG or vehicle alone (DOTAP). Representative image. n ≥ 3 independent experiments. c, Immunoblot analysis of caspase 1 during transfection with GAG in wild-type (WT) and Gsdmd−/− BMDMs. Representative image. n ≥ 3 independent experiments. d, Volcano plot of the polysaccharide pull-down mass spectrometry analysis of the β-glucan interactome with BMDM cytosolic proteins versus control. Proteins with P < 0.005 are highlighted in orange and proteins with P < 0.005 and log2-transformed fold change > 7 compared to control are highlighted in red (none identified); P value was determined by the G test; exact P values are presented in Supplementary Table 1. e, Immunoblot analysis of ribosomal proteins interacting with GAG, Ac-GAG, d-GAG or β-glucan or vehicle with or without NaCl. Representative images. n ≥ 3 independent experiments. f, Immunoblot analysis of caspase 1 from BMDMs assessed after 3 h incubation with GAG, Ac-GAG or d-GAG, with or without NaCl. Representative image. n ≥ 3 independent experiments. g, h, Measurement of cell death by Sytox green staining during GAG and d-GAG treatment, with or without NaCl. n = 3 biologically independent samples. Data are mean ± s.e.m. i, Electron microscopy pictures of ribosomes, GAG + ribosomes and chitin + ribosomes with negative staining; data from one experiment. Scale bars, 100 nm.
Extended Data Fig. 6 GAG inhibits translation and induces endoplasmic stress.
a–c Immunoblot analysis of translation rate in BMDMs by puromycin integration into proteins during vehicle (DOTAP) or PBS incubation (a), poly(dA:dT) transfection (b) or A. fumigatus infection and pro-caspase 1 (pro-Casp1; p45) and the active caspase 1 subunit (p20) during A. fumigatus infection (c). Representative images. n ≥ 2 independent experiments. d, Polysome profiling during DOTAP or DOTAP + GAG treatments. e, Immunoblot analysis of the cell pellet after polysome profiling. Representative images. n ≥ 2 independent experiments. f, Polysome:monosome ratio during DOTAP or DOTAP + GAG treatments. Data are mean ± s.e.m. *P = 0.0366. Paired two-tailed t-test. n = 3 biologically independent samples. g−i, Immunoblot analysis of pro-caspase 1 and the active caspase 1 subunit of wild-type (WT) or Nlrp3−/− BMDMs assessed after 16 h incubation with 25 μg ml−1 anisomycin (aniso) (g), 50 μg ml−1 puromycin (puro) (h) or 50 μg ml−1 cycloheximide (CHX) (i). Representative images. n ≥ 2 independent experiments. j, k, Immunoblot analysis of PERK activation (p-PERK) and IRE1α induction during GAG transfection (j) or PERK activation during treatment with translation inhibitors (k). Representative images. n ≥ 2 independent experiments. l, Immunoblot analysis of proteins ubiquitinated during GAG transfection. Representative images. n ≥ 2 independent experiments. m, Immunoblot analysis of caspase 1 of BMDMs left untreated (medium alone (med.)) or assessed 3 h after transfection with DOTAP alone, GAG or GAG with MG132 treatment. Representative image. n ≥ 2 independent experiments.
Extended Data Fig. 7 Stress granules are not induced by GAG.
a, Immunofluorescence staining of G3BP1 (green), DDX3X (red) and BMDM nuclei (blue) in unprimed BMDMs 40 min after transfection with GAG or incubation with arsenite (Ars). Representative images. n ≥ 2 independent experiments. b, Quantification of the percentage of stress-granule-positive cells after transfection with GAG, vehicle (DOTAP) alone or Ars. n > 10 biologically independent fields of cells. Data are mean ± s.e.m. c, Immunofluorescence staining of G3BP1 (green), DDX3X (red) and BMDM nuclei (blue) in unprimed BMDMs 15 h after infection with A. fumigatus. Representative images. n ≥ 2 independent experiments. d, e, Immunoblot analysis of pro-caspase 1 (pro-Casp1; p45) and the active caspase 1 subunit (p20) of BMDMs assessed 3 h after transfection with vehicle (DOTAP), GAG (d) or d-GAG (e). Representative images. n ≥ 2 independent experiments. f, Immunoblot analysis of caspase 1 from BMDMs left untreated (medium alone (med.)) or infected with A. fumigatus wild-type (WT) or deletion mutant ∆gt4c (MOI of 10). Representative image. n ≥ 2 independent experiments. Scale bars, 10 μm (a, c).
Extended Data Fig. 8 GAG-induced pro-inflammatory cytokine secretion during aspergillosis and DSS-induced colitis.
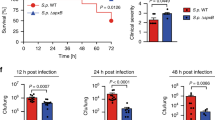
a, b, Level of IL-1β (a) and IL-18 (b) in bronchioalveolar lavage 2 days after infection with wild-type (WT) or Δugm1 strains of A. fumigatus. a, *P = 0.036. Unpaired two-tailed t-test. n = 6 independent samples. Data are mean ± s.e.m. c, Survival of 7–8-week-old immunocompetent WT mice infected intravenously with 1 × 106 A. fumigatus resting conidia (WT or Δugm1). **P = 0.0014. Log-rank (Mantel–Cox) test. d, e, Levels of IL-1β (d) and IL-18 (e) in liver homogenates after infection with WT or Δgt4c strains. d, *P = 0.0209; e, **P = 0.0041. Unpaired two-tailed t-test. n = 5 independent samples. Data are mean ± s.e.m. f–k, Concentration of cytokines in colon homogenates after DSS water supplementation and treatment with GAG or vehicle (vehicle and GAG, n = 10 mice each). f, *P = 0.0336; g, *P = 0.0181; h, *P = 0.0188; i, *P = 0.0115; j, **P = 0.0066; k, *P = 0.0154. Unpaired two-tailed t-test. Data are mean ± s.e.m.
Supplementary information
Supplementary Table
Supplementary Table 1: Spectral count comparison of proteins identified from the Control and GAG-treated samples. P values were determined by the G-test.
Supplementary Figure
Supplementary Figure 1: Uncropped blots with molecular weight and size markers and an indication of how the images were cropped.
Source data
Rights and permissions
About this article
Cite this article
Briard, B., Fontaine, T., Samir, P. et al. Galactosaminogalactan activates the inflammasome to provide host protection. Nature 588, 688–692 (2020). https://doi.org/10.1038/s41586-020-2996-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-020-2996-z
This article is cited by
-
Targeting NLRP3 inflammasome for neurodegenerative disorders
Molecular Psychiatry (2023)
-
Cryo-electron tomography of NLRP3-activated ASC complexes reveals organelle co-localization
Nature Communications (2023)
-
Structural adaptation of fungal cell wall in hypersaline environment
Nature Communications (2023)
-
Organellar homeostasis and innate immune sensing
Nature Reviews Immunology (2022)
-
Filamentous fungal biofilms: Conserved and unique aspects of extracellular matrix composition, mechanisms of drug resistance and regulatory networks in Aspergillus fumigatus
npj Biofilms and Microbiomes (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.