Abstract
In mammals, telomere protection is mediated by the essential protein TRF2, which binds chromosome ends and ensures genome integrity1,2. TRF2 depletion results in end-to-end chromosome fusions in all cell types that have been tested so far. Here we find that TRF2 is dispensable for the proliferation and survival of mouse embryonic stem (ES) cells. Trf2−/− (also known as Terf2) ES cells do not exhibit telomere fusions and can be expanded indefinitely. In response to the deletion of TRF2, ES cells exhibit a muted DNA damage response that is characterized by the recruitment of γH2AX—but not 53BP1—to telomeres. To define the mechanisms that control this unique DNA damage response in ES cells, we performed a CRISPR–Cas9-knockout screen. We found a strong dependency of TRF2-null ES cells on the telomere-associated protein POT1B and on the chromatin remodelling factor BRD2. Co-depletion of POT1B or BRD2 with TRF2 restores a canonical DNA damage response at telomeres, resulting in frequent telomere fusions. We found that TRF2 depletion in ES cells activates a totipotent-like two-cell-stage transcriptional program that includes high levels of ZSCAN4. We show that the upregulation of ZSCAN4 contributes to telomere protection in the absence of TRF2. Together, our results uncover a unique response to telomere deprotection during early development.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
All raw sequencing reads for RNA-seq are available on NCBI Gene Expression Omnibus under accession number GSE156534. Source data are provided with this paper.
References
Celli, G. B. & de Lange, T. DNA processing is not required for ATM-mediated telomere damage response after TRF2 deletion. Nat. Cell Biol. 7, 712–718 (2005).
de Lange, T. Shelterin: the protein complex that shapes and safeguards human telomeres. Genes Dev. 19, 2100–2110 (2005).
Artandi, S. E. et al. Telomere dysfunction promotes non-reciprocal translocations and epithelial cancers in mice. Nature 406, 641–645 (2000).
Griffith, J. D. et al. Mammalian telomeres end in a large duplex loop. Cell 97, 503–514 (1999).
Doksani, Y., Wu, J. Y., de Lange, T. & Zhuang, X. Super-resolution fluorescence imaging of telomeres reveals TRF2-dependent T-loop formation. Cell 155, 345–356 (2013).
Okamoto, K. et al. A two-step mechanism for TRF2-mediated chromosome-end protection. Nature 494, 502–505 (2013).
Martínez, P., Ferrara-Romeo, I., Flores, J. M. & Blasco, M. A. Essential role for the TRF2 telomere protein in adult skin homeostasis. Aging Cell 13, 656–668 (2014).
Lobanova, A. et al. Different requirements of functional telomeres in neural stem cells and terminally differentiated neurons. Genes Dev. 31, 639–647 (2017).
White, D. E. et al. KAP1, a novel substrate for PIKK family members, colocalizes with numerous damage response factors at DNA lesions. Cancer Res. 66, 11594–11599 (2006).
Dimitrova, N. & de Lange, T. Cell cycle-dependent role of MRN at dysfunctional telomeres: ATM signaling-dependent induction of nonhomologous end joining (NHEJ) in G1 and resection-mediated inhibition of NHEJ in G2. Mol. Cell Biol. 29, 5552–5563 (2009).
Hockemeyer, D., Daniels, J. P., Takai, H. & de Lange, T. Recent expansion of the telomeric complex in rodents: two distinct POT1 proteins protect mouse telomeres. Cell 126, 63–77 (2006).
Wu, L. et al. Pot1 deficiency initiates DNA damage checkpoint activation and aberrant homologous recombination at telomeres. Cell 126, 49–62 (2006).
Gyuris, A. et al. The chromatin-targeting protein Brd2 is required for neural tube closure and embryogenesis. Biochim. Biophys. Acta 1789, 413–421 (2009).
Kim, J. J. et al. Systematic bromodomain protein screens identify homologous recombination and R-loop suppression pathways involved in genome integrity. Genes Dev. 33, 1751–1774 (2019).
Kratz, K. & de Lange, T. Protection of telomeres 1 proteins POT1a and POT1b can repress ATR signaling by RPA exclusion, but binding to CST limits ATR repression by POT1b. J. Biol. Chem. 293, 14384–14392 (2018).
Denchi, E. L. & de Lange, T. Protection of telomeres through independent control of ATM and ATR by TRF2 and POT1. Nature 448, 1068–1071 (2007).
Gursoy-Yuzugullu, O., Carman, C. & Price, B. D. Spatially restricted loading of BRD2 at DNA double-strand breaks protects H4 acetylation domains and promotes DNA repair. Sci. Rep. 7, 12921 (2017).
Eckersley-Maslin, M. A. et al. MERVL/Zscan4 network activation results in transient genome-wide DNA demethylation of mESCs. Cell Rep. 17, 179–192 (2016).
Zhang, W. et al. Zscan4c activates endogenous retrovirus MERVL and cleavage embryo genes. Nucleic Acids Res. 47, 8485–8501 (2019).
Macfarlan, T. S. et al. Embryonic stem cell potency fluctuates with endogenous retrovirus activity. Nature 487, 57–63 (2012).
Falco, G. et al. Zscan4: a novel gene expressed exclusively in late 2-cell embryos and embryonic stem cells. Dev. Biol. 307, 539–550 (2007).
Zalzman, M. et al. Zscan4 regulates telomere elongation and genomic stability in ES cells. Nature 464, 858–863 (2010).
Dan, J. et al. Zscan4 inhibits maintenance DNA methylation to facilitate telomere elongation in mouse embryonic stem cells. Cell Rep. 20, 1936–1949 (2017).
Pucci, F., Gardano, L. & Harrington, L. Short telomeres in ESCs lead to unstable differentiation. Cell Stem Cell 12, 479–486 (2013).
Colla, S. et al. Telomere dysfunction drives aberrant hematopoietic differentiation and myelodysplastic syndrome. Cancer Cell 27, 644–657 (2015).
Ferron, S. R. et al. Telomere shortening in neural stem cells disrupts neuronal differentiation and neuritogenesis. J. Neurosci. 29, 14394–14407 (2009).
Vooijs, M., Jonkers, J. & Berns, A. A highly efficient ligand-regulated Cre recombinase mouse line shows that LoxP recombination is position dependent. EMBO Rep. 2, 292–297 (2001).
Chavez, A., George, V., Agrawal, V. & Johnson, F. B. Sumoylation and the structural maintenance of chromosomes (Smc) 5/6 complex slow senescence through recombination intermediate resolution. J. Biol. Chem. 285, 11922–11930 (2010).
Baerlocher, G. M., Vulto, I., de Jong, G. & Lansdorp, P. M. Flow cytometry and FISH to measure the average length of telomeres (flow FISH). Nat. Protoc. 1, 2365–2376 (2006).
Li, W. et al. MAGeCK enables robust identification of essential genes from genome-scale CRISPR/Cas9 knockout screens. Genome Biol. 15, 554 (2014).
Tzelepis, K. et al. A CRISPR dropout screen identifies genetic vulnerabilities and therapeutic targets in acute myeloid leukemia. Cell Rep. 17, 1193–1205 (2016).
Rucci, F. et al. Homozygous DNA ligase IV R278H mutation in mice leads to leaky SCID and represents a model for human LIG4 syndrome. Proc. Natl Acad. Sci. USA 107, 3024–3029 (2010).
Acknowledgements
We thank A. Nussenzweig, A. Sfeir, C. Attwooll and members of the Lazzerini Denchi laboratory for discussions and critical reading of the manuscript; F. Livak and the NCI Flow Cytometry Core Facility FACS for assistance with FACS analysis; and S. Shema and the CCR Genomics Core for assistance with next-generation sequencing experiments. M.M.-P. is a recipient of the William Guy Forbeck Research Foundation Award; T.O. is a recipient of the Helen Hay Whitney Postdoctoral Fellowship.
Author information
Authors and Affiliations
Contributions
E.L.D. and M.M.-P. conceived the study and designed the experiments; M.M.-P. and A.M.L. conducted experiments; E.L.D. and M.M.-P. analysed data; O.K. and A.L. generated TRF2 conditional-knockout ES cells. T.O. and S.R. provided critical reagents and assisted with the ES cell differentiation studies. E.L.D. and M.M.-P. wrote the manuscript with editorial help from A.M.L., T.O. and S.R.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature thanks Ylli Doksani and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Trf2−/− ES cells are not viable upon differentiation.
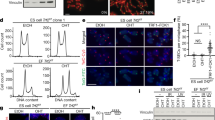
a, Schematic showing the hanging drops protocol used to generate embryoid bodies (EBs) and differentiated FLICs. b, Representative images of embryoid bodies derived from control and TRF2-depleted ES cells before adhesion (left) and 3 days after induction of differentiation (right). The experiment was performed three times with similar results. Scale bar, 400 μm. c, Growth properties of embryoid bodies derived from TRF2-proficient (control (cont.)) or TRF2-depleted ES cells (OHT) undergoing differentiation measured by confluence. Mean and s.d. are derived from the analysis of 47 images per condition. d, Immunofluorescence staining for the pluripotency marker OCT4 in ES cells and FLICs. The percentage of positive cells is shown. The experiment was performed three times with similar results. Scale bar, 10 μm. e, Schematics of the experimental approach used to derive single-cell isolates 3 weeks after OHT-mediated TRF2 deletion in ES cells and FLICs. The table summarizes the number of cells seeded, the number of colonies obtained and the TRF2 genotype of the isolated colonies. An example of a genotyping PCR performed on the resulting clones and the representation of the Trf2-floxed and Trf2-null alleles1 is shown below. The unprocessed image is provided in the Source Data for Extended Data Fig. 1. f, Growth property measured by confluency of 4 heterozygous ES cells clones (Trf2f/−) (dark blue) and 4 knockout clones (Trf2−/−) (red) derived from the experiment described in e, and the parental ES cells (Trf2f/f) (light blue). Mean and s.d. are derived from the analysis of 36 images per condition.
Extended Data Fig. 2 Reduced activation of DDR in ES cells following telomere deprotection.
a, Representative immunofluorescence for KAP1 phosphorylated at S824 (p-KAP1) in ES cells and FLICs either before (cont.) or after Cre induction (OHT). Scale bar, 10 μm. Percentages of p-KAP1-positive cells are indicated in the figure as well as in b. The experiment was performed twice for ES cells with similar results and once for FLICs. c, Representative immunofluorescence staining for γH2AX (red), p-KAP1 (pink) and FISH staining for telomeric DNA (green) of TRF2-proficient (cont.) and TRF2-deficient (OHT) ES cells. Percentage of TIF-positive cells (third panel from left) and percentage of cells with TIF and positive for p-KAP1 (last panel from left) are shown. Scale bar, 4 μm. d, Percentage of ES cells and FLICs with more than 10 γH2AX TIF. Where indicated, TRF2-proficient (cont.) or TRF2-deficient (OHT) cells were treated with an ATM inhibitor. Data panels in the figure are representative of three experiments. ns, non-significant (P ≥ 0.05), *P < 0.05, **P < 0.01, ***P < 0.001, one-way ANOVA. e, 53BP1 (red) localization at telomeric DNA (green) in TRF2-proficient (cont.) and TRF2-deficient (OHT) ES cells and FLICs. Quantification of these data are reported in Fig. 1f. Scale bar, 4 μm. f, Western blotting analysis for p-KAP1, γH2AX and—as a loading control—tubulin. Protein lysates were isolated from ES cells or FLICs either mock-treated or gamma irradiated (2.5 Gy) 30 min before collection. g, Western blotting analysis for the expression of p-KAP1 or—as loading control—tubulin. Protein lysates were derived from ES cells that were either untreated or treated with the ATM inhibitor, as indicated. Where indicated, cells were treated with OHT to induce TRF2 depletion, with UV (1,200 J per m2) or with gamma irradiation (2.5 Gy). h, Dissipation of 53BP1 DNA damage foci following IR-induced damage in FLICs (left) and ES cells (right). Cells were treated with gamma irradiation (2.5 Gy) and fixed at the indicated time points. 0, non-treated control samples. For each time point, at least 140 nuclei were scored and the percentage of cells with the indicated number of 53BP1 foci at any given time point is indicated; additional details can be found in Supplementary Table 2.
Extended Data Fig. 3 Identification of synthetically lethal genes in TRF2-deficient ES cells.
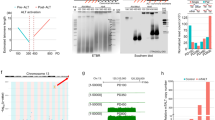
a, Scatter plots from three independent CRISPR–Cas9 screens from Fig. 2b. Genes that are synthetically lethal with TRF2 knockout are labelled with dark blue dots. Genes that are essential for ES cells are labelled with magenta dots. b, Heat map displaying the essentiality score (β-score) of the 13 candidate synthetically lethal genes (dark blue), and for 8 genes that have previously been described as essential in ES cells31 (purple), across 3 independent experiments. c, Schematic of the Pot1b and Brd2 genomic loci showing genome organization, the predicted CRISPR–Cas9 cut site (scissors, red) and key domains required for protein function (blue). Sanger sequencing of Pot1b−/− (exon 2) and Brd2−/− (exon 4) for different clones isolated is shown. For Pot1b, a repair template cassette containing multiple STOP codons was used. gRNA sequences are highlighted in red.
Extended Data Fig. 4 Codepletion of TRF2 and POT1A or POT1B result in LIG4-dependent end-to-end fusions.
a, Immunofluorescence staining for the pluripotency marker OCT4 (green) in Pot1b−/− and Brd2−/− ES cells. The percentage of positive cells is shown. Scale bar, 10 μm. b, Schematic of the Lig4 genomic locus, showing the predicted CRISPR–Cas9 cut site (scissors, red), and the sequence coding the essential R278 residue32 (blue). Sanger sequencing results showing successful gene editing of the Lig4 locus in four independent clones; the gRNA sequence is highlighted in red. c, Metaphases from ES cells of indicated genotypes. The percentage of telomeres engaging in telomere fusions are shown. In total, three independent experiments were performed. For a detailed summary of the data, see Supplementary Table 1 and Source Data for Extended Data Fig. 4.
Extended Data Fig. 5 Pot1b, Pot1a and Brd2 depletion in TRF2-null cells induce the DDR pathway.
a, Representative IF–FISH for γH2AX (red) and telomeres (green) of control (cont.) and TRF2-depleted (OHT) ES cells with the indicated Pot1b genotype. Quantifications are provided in Fig. 3e. Scale bar, 4 μm. b, Scatter plot representing the distribution of telomeric fusions in control (cont.) and TRF2-depleted (OHT) ES cells of the indicated Pot1a genotype. Each dot represents the percentage of telomeres fused in one metaphase spread. For detailed information, see Supplementary Table 1 and Source Data for Extended Data Fig. 5 (n = 3 independent experiments). c, Scatter plot representing the distribution of telomeric fusions in control (cont.) and TRF2-depleted (OHT) Pot1a−/− ES cells that were either left untreated or treated with the ATR inhibitor (ATRi). Each dot represents the percentage of telomeres fused in one metaphase spread. For detailed information, see Supplementary Table 1 and Source Data for Extended Data Fig. 5. d, Metaphases derived from TRF2-proficient (cont.) or TRF2-deficient (OHT) Pot1b−/− ES cells treated with ATMi (left) or ATRi (right). Percentage of telomere fusions were calculated from a total of >19 metaphases per genotype. e, Graph representing the percentage of telomeric fusions in FLICs of the indicated genotype and treatment. In total, >30 metaphases per genotype and treatment were analysed. f, Representative immunofluorescence for p-KAP1 (red) in ES cells of the indicated genotype in the presence (cont.) or absence of TRF2 (OHT). Scale bar, 10 μm. g, Percentage of p-KAP1-positive cells in response to TRF2 depletion (OHT) in ES cells of indicated genotype and in control cells. Additional information for b–e is provided in Supplementary Table 1 and Source Data for Extended Data Fig. 5.
Extended Data Fig. 6 ZSCAN4 contributes to end protection in TRF2-depleted cells.
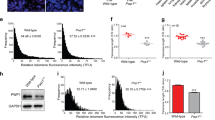
a, Differentially expressed genes in response to TRF2 depletion in ES cells (left) and differentiated cells (FLICs) (right). Volcano plots represent the log2-transformed fold change (FC) (x axis) and the −log2(P value) (y axis). Transcriptional profiling was performed three days after OHT-mediated TRF2 deletion. In the top panels, genes downregulated (P < 0.005 and FC < −2) upon OHT treatment are labelled with dark blue dots, and upregulated (P < 0.005 and FC >2) are labelled with red dots. Bottom panels, genes expressed in the 2-cell stage are labelled in purple, shelterin components are labelled in green. b, Quantifications of telomeric fusions detected in ES cells following ZSCAN4C downregulation (relative to Fig. 4d). Scatter plot representing the distribution of telomeric fusions. Each dot represents the percentage of telomeres fused in one metaphase spread. For detailed analysis information, see Supplementary Table 1 and Source Data for Extended Data Fig. 6. c, ZSCAN4C downregulation does not induce ES cell differentiation. Representative immunofluorescence staining for the pluripotency marker OCT4 in control and OHT-treated ES cells following ZSCAN4C downregulation. The percentage of OCT4-positive cells is shown. Scale bar, 10 μm. d, Representative immunofluorescence staining for ZSCAN4 in FLICs with stable integration of a doxycycline-inducible ZSCAN4C expression construct (iZSCAN4). Cells were either left untreated (top) or treated with doxycycline to induce Zscan4c expression (bottom) for two days before collection. e, The FLICs described in d either induced with doxycycline (+) or not (−) were treated with OHT and collected 4 days later for western blot analysis. As a control (cont.), FLICs lacking the inducible cassette were used. f, Growth of Trf2f/f FLICs before (cont.) or after Cre induction (OHT), untreated (circle) or treated (inverted triangle) with doxycycline to induce Zscan4c. Mean and s.d. are derived from the analysis of 49 images per condition. g, Representative IF–FISH staining for 53BP1 (red) and telomeric DNA (green) in TRF2-proficient (cont.) or TRF2-deficient FLICs. Where indicated, cells were treated with doxycycline (Dox) 7 days before collection to induce expression of exogenous Zscan4c. Scale bar, 4 μm. h, Quantification of data shown in g, cells with more than 10 53BP1 foci at telomeres were scored as positive for TIF.
Extended Data Fig. 7 TRF2-depleted cells show increased telomere length.
a, Representative FACS plots (y axes, cell count; x axes, PI staining) showing the cell-cycle profiles of TRF2-proficient (cont.) and TRF2-deficient (OHT) FLICs. Where indicated, cells were treated with doxycycline (Dox) for 7 days before collection to induce expression of exogenous Zscan4c. G1 phase of the cell cycle is marked in blue; S phase in yellow; G2 phase in green. The unmarked peak indicates presence of polyploid cells. b, Q-FISH histograms of telomere length distributions of chromosomes derived from TRF2-proficient (Trf2f/−) and TRF2-deficient (Trf2−/−) ES cells clones, derived as described in Extended Data Fig. 1e. Telomere length was calculated on the basis of the florescence intensity of a telomeric FISH probe (PNA-TelC). c, Flow-FISH quantification of telomere length was performed on TRF2-proficient (Trf2f/−) and TRF2-deficient (Trf2−/−) ES cells clones. Top panels show the distribution of telomeric signal intensity (PNA-TelC) (y axis) and the DNA content (DAPI) (x axis). The bottom panels show the distribution of cells (y axis) on the basis of telomeric signal intensity (x axis). Cells were gated on the basis of DNA content into G1 (light blue) and S/G2 (orange). Note the shift in telomere length between the TRF2-proficient and TRF2-deficient cells.
Extended Data Fig. 8 Model of the unique telomere protection mechanism in ES cells.
a, Overview of the DDR in response to TRF2 depletion in somatic cells. In response to uncapped telomeres, ATM kinase initiates DDR, leading to end-to-end fusions via the NHEJ pathway. b, POT1A, POT1B and BRD2ensure genomic stability in pluripotent mouse ES cells by inhibiting the activities of ATM and ATR kinases. As a result, mild DDR is not enough to cause end-to-end fusions. c, Increased expression of Zscan4—a cluster of two-cell-signature genes—might result in recombination-based telomere elongation. Alternatively, the ZSCAN4 -dependent inhibition of the ATM and ATR kinases at TRF2-depleted telomeres of ES cells ensures telomere stability.
Supplementary information
Supplementary Table 1
: List of number of chromosomes analyzed for each experiment performed in this study and relative number telomere fusions detected. In this table statistical analyzes was performed on the total number of chromosomes analyzed. The table contains data relative to Figures: 1b-c, 3a-b, 4d-f and Extended Data Figures: 4c, 5b-e, 6b. Raw data used for graph generation are in Source Data File.
Supplementary Table
Supplementary Table 2: Number of cells examined for IF, IF-FISH and TIFs analysis. The table contains all data points of independent repeats for data shown for Figures: 1e-f, h-i, 3e-f, 4b and Extended Data Figures: 1d, 2b,d,h, 4a, 5g, 6c,g.
Supplementary Table
Supplementary Table 3 Essentiality score (β-score) across three independent experiments of the highest ranked genes identified in this study as synthetically lethal with TRF2 (synthetically lethal) and essential in ES cells regardless of the TRF2 status (essential in ES). β-scores were calculated using MAGeCK-MLE module. Essentiality scores for all the remaining genes can be found in the Source Data File.
Supplementary Table
Supplementary Table 4: List of genes detected as differentially expressed (DE) upon OHT-mediated TRF2 depletion in ESCs (DE_ESCs) and in FLICs (DE_FLICs). Fold change and p-values were calculated using DESeq2.
Rights and permissions
About this article
Cite this article
Markiewicz-Potoczny, M., Lobanova, A., Loeb, A.M. et al. TRF2-mediated telomere protection is dispensable in pluripotent stem cells. Nature 589, 110–115 (2021). https://doi.org/10.1038/s41586-020-2959-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-020-2959-4
This article is cited by
-
A non-genetic switch triggers alternative telomere lengthening and cellular immortalization in ATRX deficient cells
Nature Communications (2023)
-
Critically short telomeres derepress retrotransposons to promote genome instability in embryonic stem cells
Cell Discovery (2023)
-
Revolutionizing DNA repair research and cancer therapy with CRISPR–Cas screens
Nature Reviews Molecular Cell Biology (2023)
-
Connecting telomere maintenance and regulation to the developmental origin and differentiation states of neuroblastoma tumor cells
Journal of Hematology & Oncology (2022)
-
The landscape of aging
Science China Life Sciences (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.