Abstract
Perisynaptic astrocytic processes are an integral part of central nervous system synapses1,2; however, the molecular mechanisms that govern astrocyte–synapse adhesions and how astrocyte contacts control synapse formation and function are largely unknown. Here we use an in vivo chemico-genetic approach that applies a cell-surface fragment complementation strategy, Split-TurboID, and identify a proteome that is enriched at astrocyte–neuron junctions in vivo, which includes neuronal cell adhesion molecule (NRCAM). We find that NRCAM is expressed in cortical astrocytes, localizes to perisynaptic contacts and is required to restrict neuropil infiltration by astrocytic processes. Furthermore, we show that astrocytic NRCAM interacts transcellularly with neuronal NRCAM coupled to gephyrin at inhibitory postsynapses. Depletion of astrocytic NRCAM reduces numbers of inhibitory synapses without altering glutamatergic synaptic density. Moreover, loss of astrocytic NRCAM markedly decreases inhibitory synaptic function, with minor effects on excitation. Thus, our results present a proteomic framework for how astrocytes interface with neurons and reveal how astrocytes control GABAergic synapse formation and function.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
Proteomics data are available in the MassIVE database under accession MSV000085821. The data that support the findings of this study are available from the corresponding author upon reasonable request.
References
Yu, X., Nagai, J. & Khakh, B. S. Improved tools to study astrocytes. Nat. Rev. Neurosci. 21, 121–138 (2020).
Lanjakornsiripan, D. et al. Layer-specific morphological and molecular differences in neocortical astrocytes and their dependence on neuronal layers. Nat. Commun. 9, 1623 (2018).
Araque, A., Parpura, V., Sanzgiri, R. P. & Haydon, P. G. Tripartite synapses: glia, the unacknowledged partner. Trends Neurosci. 22, 208–215 (1999).
Khakh, B. S. & Sofroniew, M. V. Diversity of astrocyte functions and phenotypes in neural circuits. Nat. Neurosci. 18, 942–952 (2015).
Ma, Z., Stork, T., Bergles, D. E. & Freeman, M. R. Neuromodulators signal through astrocytes to alter neural circuit activity and behaviour. Nature 539, 428–432 (2016).
Papouin, T., Dunphy, J., Tolman, M., Foley, J. C. & Haydon, P. G. Astrocytic control of synaptic function. Phil. Trans. R. Soc. Lond. B 372, 20160154 (2017).
Panatier, A. et al. Astrocytes are endogenous regulators of basal transmission at central synapses. Cell 146, 785–798 (2011).
Araque, A. et al. Gliotransmitters travel in time and space. Neuron 81, 728–739 (2014).
Stogsdill, J. A. et al. Astrocytic neuroligins control astrocyte morphogenesis and synaptogenesis. Nature 551, 192–197 (2017).
Stork, T., Sheehan, A., Tasdemir-Yilmaz, O. E. & Freeman, M. R. Neuron–glia interactions through the Heartless FGF receptor signaling pathway mediate morphogenesis of Drosophila astrocytes. Neuron 83, 388–403 (2014).
Sloan, S. A. & Barres, B. A. Mechanisms of astrocyte development and their contributions to neurodevelopmental disorders. Curr. Opin. Neurobiol. 27, 75–81 (2014).
Allen, N. J. & Lyons, D. A. Glia as architects of central nervous system formation and function. Science 362, 181–185 (2018).
Branon, T. C. et al. Efficient proximity labeling in living cells and organisms with TurboID. Nat. Biotechnol. 36, 880–887 (2018).
Schopp, I. M. et al. Split-BioID a conditional proteomics approach to monitor the composition of spatiotemporally defined protein complexes. Nat. Commun. 8, 15690 (2017).
De Munter, S. et al. Split-BioID: a proximity biotinylation assay for dimerization-dependent protein interactions. FEBS Lett. 591, 415–424 (2017).
Kinoshita, N. et al. Genetically encoded fluorescent indicator GRAPHIC delineates intercellular connections. iScience 15, 28–38 (2019).
Lee, Y., Messing, A., Su, M. & Brenner, M. GFAP promoter elements required for region-specific and astrocyte-specific expression. Glia 56, 481–493 (2008).
Chan, K. Y. et al. Engineered AAVs for efficient noninvasive gene delivery to the central and peripheral nervous systems. Nat. Neurosci. 20, 1172–1179 (2017).
Uezu, A. et al. Identification of an elaborate complex mediating postsynaptic inhibition. Science 353, 1123–1129 (2016).
Zhang, Y. et al. An RNA-sequencing transcriptome and splicing database of glia, neurons, and vascular cells of the cerebral cortex. J. Neurosci. 34, 11929–11947 (2014).
Zhang, Y. et al. Purification and characterization of progenitor and mature human astrocytes reveals transcriptional and functional differences with mouse. Neuron 89, 37–53 (2016).
Sakers, K. & Eroglu, C. Control of neural development and function by glial neuroligins. Curr. Opin. Neurobiol. 57, 163–170 (2019).
Incontro, S., Asensio, C. S., Edwards, R. H. & Nicoll, R. A. Efficient, complete deletion of synaptic proteins using CRISPR. Neuron 83, 1051–1057 (2014).
Custer, A. W. et al. The role of the ankyrin-binding protein NrCAM in node of Ranvier formation. J. Neurosci. 23, 10032–10039 (2003).
Feinberg, K. et al. A glial signal consisting of gliomedin and NrCAM clusters axonal Na+ channels during the formation of nodes of Ranvier. Neuron 65, 490–502 (2010).
Demyanenko, G. P. et al. Neural cell adhesion molecule NrCAM regulates Semaphorin 3F-induced dendritic spine remodeling. J. Neurosci. 34, 11274–11287 (2014).
Mohan, V. et al. Temporal regulation of dendritic spines through NrCAM-Semaphorin3F receptor signaling in developing cortical pyramidal neurons. Cereb. Cortex 29, 963–977 (2019).
Mauro, V. P., Krushel, L. A., Cunningham, B. A. & Edelman, G. M. Homophilic and heterophilic binding activities of Nr-CAM, a nervous system cell adhesion molecule. J. Cell Biol. 119, 191–202 (1992).
Derouiche, A., Anlauf, E., Aumann, G., Mühlstädt, B. & Lavialle, M. Anatomical aspects of glia-synapse interaction: the perisynaptic glial sheath consists of a specialized astrocyte compartment. J. Physiol. Paris 96, 177–182 (2002).
Lavialle, M. et al. Structural plasticity of perisynaptic astrocyte processes involves ezrin and metabotropic glutamate receptors. Proc. Natl Acad. Sci. USA 108, 12915–12919 (2011).
Scheiffele, P., Fan, J., Choih, J., Fetter, R. & Serafini, T. Neuroligin expressed in nonneuronal cells triggers presynaptic development in contacting axons. Cell 101, 657–669 (2000).
Graf, E. R., Zhang, X., Jin, S. X., Linhoff, M. W. & Craig, A. M. Neurexins induce differentiation of GABA and glutamate postsynaptic specializations via neuroligins. Cell 119, 1013–1026 (2004).
Chih, B., Gollan, L. & Scheiffele, P. Alternative splicing controls selective trans-synaptic interactions of the neuroligin–neurexin complex. Neuron 51, 171–178 (2006).
Tremblay, R., Lee, S. & Rudy, B. GABAergic interneurons in the neocortex: from cellular properties to circuits. Neuron 91, 260–292 (2016).
Miles, R., Tóth, K., Gulyás, A. I., Hájos, N. & Freund, T. F. Differences between somatic and dendritic inhibition in the hippocampus. Neuron 16, 815–823 (1996).
Wierenga, C. J. & Wadman, W. J. Miniature inhibitory postsynaptic currents in CA1 pyramidal neurons after kindling epileptogenesis. J. Neurophysiol. 82, 1352–1362 (1999).
Martell, J. D. et al. A split horseradish peroxidase for the detection of intercellular protein-protein interactions and sensitive visualization of synapses. Nat. Biotechnol. 34, 774–780 (2016).
Loh, K. H. et al. Proteomic analysis of unbounded cellular compartments: synaptic clefts. Cell 166, 1295–1307 (2016).
Cijsouw, T. et al. Mapping the proteome of the synaptic cleft through proximity labeling reveals new cleft proteins. Proteomes 6, E48 (2018).
Li, J. et al. Cell-surface proteomic profiling in the fly brain uncovers wiring regulators. Cell 180, 373–386 (2020).
Cho, K. F. et al. Split-TurboID enables contact-dependent proximity labeling in cells. Proc. Natl Acad. Sci. USA 117, 12143–12154 (2020).
Elmariah, S. B., Oh, E. J., Hughes, E. G. & Balice-Gordon, R. J. Astrocytes regulate inhibitory synapse formation via Trk-mediated modulation of postsynaptic GABAA receptors. J. Neurosci. 25, 3638–3650 (2005).
Hughes, E. G., Elmariah, S. B. & Balice-Gordon, R. J. Astrocyte secreted proteins selectively increase hippocampal GABAergic axon length, branching, and synaptogenesis. Mol. Cell. Neurosci. 43, 136–145 (2010).
Turrigiano, G. G., Leslie, K. R., Desai, N. S., Rutherford, L. C. & Nelson, S. B. Activity-dependent scaling of quantal amplitude in neocortical neurons. Nature 391, 892–896 (1998).
O’Brien, R. J. et al. Activity-dependent modulation of synaptic AMPA receptor accumulation. Neuron 21, 1067–1078 (1998).
Spence, E. F. et al. In vivo proximity proteomics of nascent synapses reveals a novel regulator of cytoskeleton-mediated synaptic maturation. Nat. Commun. 10, 386 (2019).
Shin, J. H., Yue, Y. & Duan, D. Recombinant adeno-associated viral vector production and purification. Methods Mol. Biol. 798, 267–284 (2012).
Takano, T. et al. LMTK1 regulates dendritic formation by regulating movement of Rab11A-positive endosomes. Mol. Biol. Cell 25, 1755–1768 (2014).
Takano, T. et al. Discovery of long-range inhibitory signaling to ensure single axon formation. Nat. Commun. 8, 33 (2017).
Ippolito, D. M, Eroglu, C. Quantifying synapses: an immunocytochemistry-based assay to quantify synapse number. J. Vis. Exp. 16, 2270 (2010).
Dani, A., Huang, B., Bergan, J., Dulac, C. & Zhuang, X. Superresolution imaging of chemical synapses in the brain. Neuron 68, 843–856 (2010).
Acknowledgements
We thank H. Katsura for modifying the promoter of the surface and split TurboID plasmids for HEK 293T cell expression, and B. Duncan for technical support. This work was supported by Brain initiative RO1DA047258 from NIH (S.H.S. and C.E.), R01MH113280 from NIH (P.F.M.), Kahn Neurotechnology Award (S.H.S. and C.E.), a Grant-in-Aid for JSPS Fellows (PD) 20153173 from the Japan Society for the Promotion of Science (T.T.), The Uehara memorial Foundation (T.T.), and National Institute of Mental Health Fellowship F30MH117851 (J.L.C).
Author information
Authors and Affiliations
Contributions
T.T., C.E. and S.H.S. designed the study. T.T., J.T.W., A.P., C.E. and S.H.S. wrote the manuscript. T.T., J.T.W., A.U. and E.J.S. performed in vivo BioID-proteomics analysis. T.T., J.T.W., J.L.C., T.S. and P.F.M. produced the constructs. T.T., J.T.W. and K.T.B. performed imaging analysis and the morphological analysis of the astrocytes. A.P. performed electrophysiological analysis. T.T. and K.T.B. performed the biological experiments. All authors discussed the results and commented on the manuscript text.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature thanks Thomas Biederer, Peter Scheiffele and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 The reconstituted activity of Split-TurboID in neurons and astrocytes in vitro.
a, Schematics of constructs tested. b, Immunoblot analysis of construct expression and biotinylation activity. c, Schematic of neuron-astrocyte mixed-culture assay for Split-TurboID with cell-type-specific AAVs in vitro. d, Cultured neurons and astrocytes were infected with AAV1/2-GfaABC1D-TurboID-HA-surface, AAV1/2-hSynI-V5-N-TurboID and/or AAV1/2-GfaABC1D-C-TurboID-HA. Representative images of neuron and astrocyte at DIV14 after the treatment of 500 μM biotin for 6h are shown. n = 3 biological repeats.
Extended Data Fig. 2 Split-TurboID maps excitatory and inhibitory perisynaptic proteins.
a–d, Representative images demonstrating that proteins biotinylated by astrocytic TurboID-surface or Split-TurboID (cyan) are adjacent to excitatory presynaptic marker VGLUT1 (a), postsynaptic marker HOMER1 (b), inhibitory presynaptic marker VGAT (c), and postsynaptic marker gephyrin (d). Astrocytes were visualized with GfaABC1D-mCherry-CAAX. n = 3 biological repeats.
Extended Data Fig. 3 Brain-wide transduction of astrocytes and neurons.
a, Schematic of AAV PHP.eB viruses for neuronal-EGFP or astrocyte-mCherry-CAAX and retro-orbital injection. b, Sagittal section of mouse brain showing expression throughout the cortex and other structures. c, Representative image from cortex, hippocampus or cerebellum showing high coverage of neuronal and astrocytic expression.
Extended Data Fig. 4 Mapping and identification of tripartite synaptic cleft proteins by Split-TurboID in vivo.
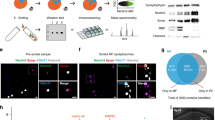
a, Biotinylation activity of Split-TurboID in vivo. Lysates of mouse brain infected with cell-type-specific TurboID-surface-HA, V5-N-TurboID and/or C-TurboID-HA. Brain lysates were analysed by immunoblotting with anti-Streptavidin, anti-V5, anti-HA and anti-Tubulin antibodies. b, The graph indicates the ratio of botinylation activity in vivo (n = 4 brains per each condition). c, d, The biotinylation of Split-TurboID in mouse cortex. e, f, Quantification of average number of excitatory or inhibitory synaptic colocalized puncta in layer 2/3 of the visual cortex. n = 15 slices per each condition from 3 mice. g, Chart summarizing proteomic data set identified by mass spectrometry and filters used to identify top candidates. h, Venn diagram comparing proteome list of Split-TurboID and TurboID-surface. i, Scale-free network of Split-TurboID (green) and TurboID-surface (blue) identified proteins. High-confidence proteins enriched in both Split-TurboID and TurboID-surface fractions are shown in red. Neuronal enriched proteins (RNA-seq expression ratio <1) and astrocyte enriched proteins (RNA-seq expression ratio≧1.0) are represented as circle or diamond, respectively. At least n = 4 biological repeats. One-way ANOVA (Dunnett’s multiple comparison, P < 0.0001, 0.001). Data are means ± s.e.m.
Extended Data Fig. 5 The validation of candidate proteins with CRISPR-based astrocytic candidate gene depletion strategy.
a, Schematic of CRISPR-based deletion of astrocytic NrCAM in vitro. b, Immunoblots showing loss of NrCAM with sgRNA. AAV1/2-U6-empty sgRNA or AAV1/2-U6-NrCAM sgRNA was co-infected with AAV1/2- GfaABC1D-Cas9 to cultured neurons and astrocytes at DIV14. The cells were subjected to immunoblot analysis with an anti-NrCAM antibody. Tubulin was used as a loading control. c, The bar graph indicates the expression level of NrCAM from 3 independent experiments. d, Schematic of CRISPR-based deletion strategy of candidate gene. e, Experimental timeline of AAV-mediated CRISPR-based astrocytic gene deletion strategy in Flex-TdTomato mice. f, AAV PHP.eB-U6-NrCAM sgRNA was co-infected with AAV PHP.eB-GfaABC1D-Cas9 in Flex-TdTomato mice at P21. Coronal sections were prepared and immunostained with an anti-TdTomato antibody. g, A High-magnification image is shown. h, Images of Tenm2-, Tenm4- or NrCAM-deleted astrocytes (cyan) and their territories (red outlines) in visual cortexes of juvenile mice. i, Average territory volumes at P42 of Tenm2-, Tenm4- or NrCAM-deleted astrocytes. Between 20-25 cells per condition from 3 mice. j, Images of Tenm2-, Tenm4- or NrCAM-deleted astrocytes (cyan) and their NIV reconstructions (orange) in visual cortexes of juvenile mice. k, Average NIV at P42 of Tenm2-, Tenm4- or NrCAM-deleted astrocytes. 51 cells per each condition from 3 mice. n = 3 biological repeats. One-way ANOVA (Dunnett’s multiple comparison, P < 0.0001, 0.01). Data are means ± s.e.m.
Extended Data Fig. 6 NrCAM is a novel tripartite synaptic protein.
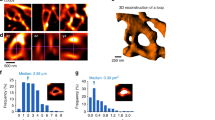
a, A high magnification STED image showing that endogenous NrCAM was enriched at biotinylated proteins in vivo. b, Immunoblot analysis of endogenous NrCAM, astrocyte marker GFAP, neuronal marker b-Tubulin III or loading control α-Tubulin from mouse brain or purified astrocyte lysate. c, Schematic of the visualization of astrocytic membrane and endogenous NrCAM in vivo. d, STED images demonstrating the localization of endogenous NrCAM in vivo. Coronal sections were immunostained with anti-NrCAM antibody (cyan). High magnification image was shown (right panel). e, Schematic of the visualization of both astrocytic and neuronal NrCAM in vivo. f, STED images demonstrating that the colocalization of astrocytic NrCAM with neuronal NrCAM in vivo. Coronal sections were prepared and co-immunostained with an anti-V5 (cyan) and anti-HA (magenta) antibody. A high-magnification image is shown in the right. n = 3 biological repeats. Data represent means ± s.e.m.
Extended Data Fig. 7 The role of astrocytic NrCAM in astrocytic morphogenesis in vivo.
a, Schematic of CRISPR-based NrCAM deletion in vivo. b, Schematic of hNrCAM domains and fragments. SP, signal peptide; IG, immunoglobulin; FN, fibronectin; TMD, transmembrane domain; ECD, extracellular domains. c, Immunoblots showing the expression of each NrCAM fragments in HEK293T cells. d, f, h, j, Images of astrocytes following deletion of astrocyte NrCAM alone (NrCAM sgRNA), with coexpression with indicated constructs of sgRNA-resistant human NrCAM, neuronal NrCAM deletion (neuroNrCAM sgRNA), or following neuronal NrCAM deletion alone. Images at indicated ages represent. e, i, Analysis of astrocyte territory, 15–29 cells per each condition from 3 mice; g, k, Analysis of neuropil infiltration volume. 50–51 cells per each condition from 3 mice. n = 3 biological repeats. One-way ANOVA (Dunnett’s multiple comparison, P < 0.0001). Data represent means ± s.e.m.
Extended Data Fig. 8 NrCAM controls inhibitory synaptic specializations through binding the gephyrin.
a, Immunoblot analysis of endogenous NrCAM, astrocyte marker GFAP, Neuroligin 2, Neuroligin 3, Kir4.1 or EAAT2 (GLT1) from purified astrocyte lysate. b, The bar graph indicates the expression level. c, The interaction of NrCAM with PSD95 and gephyrin in HEK293T cells. Cell lysates coexpressing NrCAM-HA with GFP, PSD95-GFP or GFP-gephyrin were incubated with anti-GFP-bound beads. Immunoprecipitated (right) or total (left) NrCAM, GFP, PSD95-GFP or GFP-gephyrin were detected by immunoblotting with anti-HA and anti-GFP antibodies. d, Schematic of HEK293T/neuronal mixed-cultured assay in vitro. e–h, Images of in vitro inhibitory synapse formation assays. The graph shows average of the total integrated intensity of VGAT (Cont = 258, NL2 = 222, NrCAM = 242, NrCAM-ΔIG = 288, NrCAM-ΔECD = 303 cells) or GABAA receptor (Cont = 313, NRX1β4(-) = 310, NrCAM = 300, NrCAM-ΔIG = 278, NrCAM-ΔECD = 278 cells) clusters that contact transfected HEK293T cells. i–l, Images of in vitro excitatory synapse formation assay. The graph shows average of the total integrated intensity of VGLUT1 (Cont = 259, NL2 = 306, NrCAM = 286, NrCAM-ΔIG = 321, NrCAM-ΔECD = 196 cells) or HOMER1 (Cont = 471, NRX1β4(-) = 214, NrCAM = 247, NrCAM-ΔIG = 387, NrCAM-ΔECD = 251 cells) clusters that contact transfected HEK293T cells. n = 3 biological repeats. One-way ANOVA (Dunnett’s multiple comparison, P < 0.0001). Data are means ± s.e.m.
Extended Data Fig. 9 The effect of NrCAM on excitatory synapse formation and function in vivo.
a, Images of postsynapse PSD95 and presynapse VGLUT1 within NrCAM-deletion astrocytes in L1 of the visual cortex. High magnification images (bottom) correspond to boxes (above). b, Quantification of average number of excitatory synaptic colocalized puncta within astrocyte territories. n = 15 cells per each condition from 3 mice. c, mEPSC traces from L2/3 pyramidal neurons following astrocyte control empty sgRNA or NrCAM sgRNA expression. d–g, Quantification of mEPSC amplitude (d, e, Cont = 16, NrCAM sgRNA = 14 cells from 4 mice) and frequency (f, g, Cont = 14, NrCAM sgRNA = 17 cells from each of 4 mice). At least n = 3 biological repeats. Student’s t-test (paired, P < 0.05). Data represent means ± s.e.m.
Extended Data Fig. 10 In vivo chemogenetics method, Split-TurboID, reveals a novel astrocytic cell adhesion molecule, NrCAM, that controls inhibitory synaptic organization.
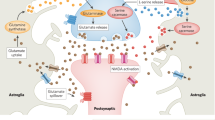
Development of in vivo chemo-affinity codes, Split-TurboID, and a working model of astrocytic NrCAM influencing inhibitory synaptic function. Split-TurboID can map the molecular composition of such intercellular contacts, even within the highly complex structure of the tripartite synapse in vivo. Mapping this interface, we discovered a new molecular mechanism by which astrocytes influence inhibitory synapses within the tripartite synaptic cleft via NrCAM. NrCAM is expressed in cortical astrocytes where it interacts with neuronal NrCAM that is coupled to gephyrin at inhibitory postsynapses. Loss of astrocytic NrCAM dramatically alters inhibitory synaptic organization and function in vivo.
Supplementary information
Supplementary Information
This file contains Supplementary Figure 1 (uncropped blots) and Supplementary Table 4 (Primers used for constructions or sequences).
Supplementary Table 1
Split-TurboID Enriched Proteins.
Supplementary Table 2
Astrocyte TurboID-Surface Enriched Proteins.
Supplementary Table 3
High Confidence Tri-partite Synapse Proteome.
Supplementary Table 5
Raw Data of Split-TurboID and Astrocyte TurboID-surface Proteome.
Supplementary Table 6
Contaminating Proteins.
Supplementary Table 7
Summary of Statistical Analysis.
Rights and permissions
About this article
Cite this article
Takano, T., Wallace, J.T., Baldwin, K.T. et al. Chemico-genetic discovery of astrocytic control of inhibition in vivo. Nature 588, 296–302 (2020). https://doi.org/10.1038/s41586-020-2926-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-020-2926-0
This article is cited by
-
Overcoming genetic and cellular complexity to study the pathophysiology of X-linked intellectual disabilities
Journal of Neurodevelopmental Disorders (2024)
-
Cross-link assisted spatial proteomics to map sub-organelle proteomes and membrane protein topologies
Nature Communications (2024)
-
Astrocytes in the adult dentate gyrus—balance between adult and developmental tasks
Molecular Psychiatry (2024)
-
Neuronal DSCAM regulates the peri-synaptic localization of GLAST in Bergmann glia for functional synapse formation
Nature Communications (2024)
-
The development of proximity labeling technology and its applications in mammals, plants, and microorganisms
Cell Communication and Signaling (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.