Abstract
Despite its success in achieving the long-term survival of 10–30% of treated individuals, immune therapy is still ineffective for most patients with cancer1,2. Many efforts are therefore underway to identify new approaches that enhance such immune ‘checkpoint’ therapy3,4,5 (so called because its aim is to block proteins that inhibit checkpoint signalling pathways in T cells, thereby freeing those immune cells to target cancer cells). Here we show that inhibiting PCSK9—a key protein in the regulation of cholesterol metabolism6,7,8—can boost the response of tumours to immune checkpoint therapy, through a mechanism that is independent of PCSK9’s cholesterol-regulating functions. Deleting the PCSK9 gene in mouse cancer cells substantially attenuates or prevents their growth in mice in a manner that depends on cytotoxic T cells. It also enhances the efficacy of immune therapy that is targeted at the checkpoint protein PD1. Furthermore, clinically approved PCSK9-neutralizing antibodies synergize with anti-PD1 therapy in suppressing tumour growth in mouse models of cancer. Inhibiting PCSK9—either through genetic deletion or using PCSK9 antibodies—increases the expression of major histocompatibility protein class I (MHC I) proteins on the tumour cell surface, promoting robust intratumoral infiltration of cytotoxic T cells. Mechanistically, we find that PCSK9 can disrupt the recycling of MHC I to the cell surface by associating with it physically and promoting its relocation and degradation in the lysosome. Together, these results suggest that inhibiting PCSK9 is a promising way to enhance immune checkpoint therapy for cancer.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
PCSK9 and CD8A mRNA expression data from various human cancers were downloaded from the GENT (http://gent2.appex.kr/gent2/) database43. PCSK9 mRNA expression and overall survival data were from TCGA data sets included in cBioportal44,45 (https://www.cbioportal.org/) in November 2018. Western blot source data are provided in Supplementary Fig. 1. Source data for the quantitative graphs are provided for Figs. 1–4 and Extended Data Figs. 1–9. Other data in support of this study are available from the corresponding author upon reasonable request. Source data are provided with this paper.
References
Topalian, S. L. et al. Safety, activity, and immune correlates of anti-PD-1 antibody in cancer. N. Engl. J. Med. 366, 2443–2454 (2012).
Brahmer, J. R. et al. Safety and activity of anti-PD-L1 antibody in patients with advanced cancer. N. Engl. J. Med. 366, 2455–2465 (2012).
Manguso, R. T. et al. In vivo CRISPR screening identifies Ptpn2 as a cancer immunotherapy target. Nature 547, 413–418 (2017).
Pan, D. et al. A major chromatin regulator determines resistance of tumor cells to T cell-mediated killing. Science 359, 770–775 (2018).
Patel, S. J. et al. Identification of essential genes for cancer immunotherapy. Nature 548, 537–542 (2017).
Abifadel, M. et al. Mutations in PCSK9 cause autosomal dominant hypercholesterolemia. Nat. Genet. 34, 154–156 (2003).
Cohen, J. et al. Low LDL cholesterol in individuals of African descent resulting from frequent nonsense mutations in PCSK9. Nat. Genet. 37, 161–165 (2005); corrigendum 37, 328 (2005).
Cohen, J. C., Boerwinkle, E., Mosley, T. H., Jr & Hobbs, H. H. Sequence variations in PCSK9, low LDL, and protection against coronary heart disease. N. Engl. J. Med. 354, 1264–1272 (2006).
Yang, W. et al. Potentiating the antitumour response of CD8+ T cells by modulating cholesterol metabolism. Nature 531, 651–655 (2016).
Ma, X. et al. Cholesterol negatively regulates IL-9-producing CD8+ T cell differentiation and antitumor activity. J. Exp. Med. 215, 1555–1569 (2018).
Naslavsky, N., Weigert, R. & Donaldson, J. G. Characterization of a nonclathrin endocytic pathway: membrane cargo and lipid requirements. Mol. Biol. Cell 15, 3542–3552 (2004).
Benjannet, S. et al. NARC-1/PCSK9 and its natural mutants: zymogen cleavage and effects on the low density lipoprotein (LDL) receptor and LDL cholesterol. J. Biol. Chem. 279, 48865–48875 (2004).
Maxwell, K. N., Fisher, E. A. & Breslow, J. L. Overexpression of PCSK9 accelerates the degradation of the LDLR in a post-endoplasmic reticulum compartment. Proc. Natl Acad. Sci. USA 102, 2069–2074 (2005).
Zhang, D. W. et al. Binding of proprotein convertase subtilisin/kexin type 9 to epidermal growth factor-like repeat A of low density lipoprotein receptor decreases receptor recycling and increases degradation. J. Biol. Chem. 282, 18602–18612 (2007).
Lagace, T. A. et al. Secreted PCSK9 decreases the number of LDL receptors in hepatocytes and in livers of parabiotic mice. J. Clin. Invest. 116, 2995–3005 (2006).
Poirier, S. et al. Dissection of the endogenous cellular pathways of PCSK9-induced low density lipoprotein receptor degradation: evidence for an intracellular route. J. Biol. Chem. 284, 28856–28864 (2009).
Poirier, S. et al. The proprotein convertase PCSK9 induces the degradation of low density lipoprotein receptor (LDLR) and its closest family members VLDLR and ApoER2. J. Biol. Chem. 283, 2363–2372 (2008).
Canuel, M. et al. Proprotein convertase subtilisin/kexin type 9 (PCSK9) can mediate degradation of the low density lipoprotein receptor-related protein 1 (LRP-1). PLoS One 8, e64145 (2013).
Demers, A. et al. PCSK9 induces CD36 degradation and affects long-chain fatty acid uptake and triglyceride metabolism in adipocytes and in mouse liver. Arterioscler. Thromb. Vasc. Biol. 35, 2517–2525 (2015).
Jonas, M. C., Costantini, C. & Puglielli, L. PCSK9 is required for the disposal of non-acetylated intermediates of the nascent membrane protein BACE1. EMBO Rep. 9, 916–922 (2008).
Blom, D. J. et al. A 52-week placebo-controlled trial of evolocumab in hyperlipidemia. N. Engl. J. Med. 370, 1809–1819 (2014).
Robinson, J. G. et al. Efficacy and safety of alirocumab in reducing lipids and cardiovascular events. N. Engl. J. Med. 372, 1489–1499 (2015).
Cong, L. et al. Multiplex genome engineering using CRISPR/Cas systems. Science 339, 819–823 (2013).
Ran, F. A. et al. Genome engineering using the CRISPR-Cas9 system. Nat. Protocols 8, 2281–2308 (2013).
Ishibashi, S. et al. Hypercholesterolemia in low density lipoprotein receptor knockout mice and its reversal by adenovirus-mediated gene delivery. J. Clin. Invest. 92, 883–893 (1993).
Wolchok, J. D. et al. Nivolumab plus ipilimumab in advanced melanoma. N. Engl. J. Med. 369, 122–133 (2013).
Kühnast, S. et al. Alirocumab inhibits atherosclerosis, improves the plaque morphology, and enhances the effects of a statin. J. Lipid Res. 55, 2103–2112 (2014).
Chan, J. C. et al. A proprotein convertase subtilisin/kexin type 9 neutralizing antibody reduces serum cholesterol in mice and nonhuman primates. Proc. Natl Acad. Sci. USA 106, 9820–9825 (2009).
Kim, K. et al. Eradication of metastatic mouse cancers resistant to immune checkpoint blockade by suppression of myeloid-derived cells. Proc. Natl Acad. Sci. USA 111, 11774–11779 (2014).
Chandramohan, V. et al. Improved efficacy against malignant brain tumors with EGFRwt/EGFRvIII targeting immunotoxin and checkpoint inhibitor combinations. J. Immunother. Cancer 7, 142 (2019).
Hogquist, K. A. et al. T cell receptor antagonist peptides induce positive selection. Cell 76, 17–27 (1994).
Shan, L. et al. PCSK9 binds to multiple receptors and can be functionally inhibited by an EGF-A peptide. Biochem. Biophys. Res. Commun. 375, 69–73 (2008).
Fasano, T., Sun, X. M., Patel, D. D. & Soutar, A. K. Degradation of LDLR protein mediated by ‘gain of function’ PCSK9 mutants in normal and ARH cells. Atherosclerosis 203, 166–171 (2009).
Duff, C. J. et al. Antibody-mediated disruption of the interaction between PCSK9 and the low-density lipoprotein receptor. Biochem. J. 419, 577–584 (2009).
Yu, Y. Y. et al. Definition and transfer of a serological epitope specific for peptide-empty forms of MHC class I. Int. Immunol. 11, 1897–1906 (1999).
Zhang, L. et al. Intratumoral T cells, recurrence, and survival in epithelial ovarian cancer. N. Engl. J. Med. 348, 203–213 (2003).
Carstens, J. L. et al. Spatial computation of intratumoral T cells correlates with survival of patients with pancreatic cancer. Nat. Commun. 8, 15095 (2017).
Labun, K. et al. CHOPCHOP v3: expanding the CRISPR web toolbox beyond genome editing. Nucleic Acids Res. 47 (W1), W171–W174 (2019).
Shalem, O. et al. Genome-scale CRISPR-Cas9 knockout screening in human cells. Science 343, 84–87 (2014).
Borowicz, S. et al. The soft agar colony formation assay. J. Vis. Exp. 92, e51998 (2014).
Moore, M. W., Carbone, F. R. & Bevan, M. J. Introduction of soluble protein into the class I pathway of antigen processing and presentation. Cell 54, 777–785 (1988).
Curtsinger, J. M., Lins, D. C. & Mescher, M. F. CD8+ memory T cells (CD44high, Ly-6C+) are more sensitive than naive cells to (CD44low, Ly-6C−) to TCR/CD8 signaling in response to antigen. J. Immunol. 160, 3236–3243 (1998).
Park, S. J., Yoon, B. H., Kim, S. K. & Kim, S. Y. GENT2: an updated gene expression database for normal and tumor tissues. BMC Med. Genomics 12 (Suppl 5), 101 (2019).
Cerami, E. et al. The cBio cancer genomics portal: an open platform for exploring multidimensional cancer genomics data. Cancer Discov. 2, 401–404 (2012).
Gao, J. et al. Integrative analysis of complex cancer genomics and clinical profiles using the cBioPortal. Sci. Signal. 6, pl1 (2013).
Acknowledgements
We thank J. M. Cook and colleagues at the Flow Cytometry Facility of Duke University School of Medicine for their assistance. We also thank the Duke University Light Microscopy Core Facility for professional help with confocal microscopy. We thank I. Li for proofreading our manuscript. We also thank S. Coffman of MedMedia Solutions for help with illustrations. C.-Y.L. is supported by US National Institutes of Health (NIH) grants ES024015, CA208852 and CA216876, and by a Cancer Center Support Grant (CCSG, CA014236) to Duke University. X.L. is supported by Guangdong Basic and Applied Basic Research Foundation grant 2020B1515020054 and Shenzhen Science and Technology Program grant JCYJ20190807154813511.
Author information
Authors and Affiliations
Contributions
X.L., X.B. and C.-Y.L. designed the study. X.L., X.B. and H.C. carried out CRISPR–Cas9-mediated gene knockouts in tumour cells. X.L., X.B. and H.C. performed western blot analyses. X.L. generated PCSK9 and H2-K1 mutants and carried out immunoprecipitation/western blot experiments. X.L., X.B. and M.H. carried out mouse tumour-growth experiments. X.L. and X.B. characterized tumour cells in vitro and in vivo and intratumoral lymphocytes in vivo using flow cytometry. X.B. maintained OVA-specific T-cell culture, performed CTL assays and analysed the results, X.B., L.X. and X.L. analysed TCGA data for PCSK9 expression and its relationship to the expression of CD8A and prognosis of patients with cancer. M.H., M.J., J.C. and Q. H. carried out immunofluorescence and immunohistochemistry analyses. F.L. advised on CRISPR knockouts and provided material support. X.L., X.B. and C.L. wrote the manuscript with help from all co-authors. C.-Y.L. provided funding and study supervision.
Corresponding author
Ethics declarations
Competing interests
X.L. and C.-Y.L. are inventors on a patent application filed by Duke University that covers the use of anti-PCSK9 antibodies in cancer immunotherapy. The other authors declare no competing interests.
Additional information
Peer review information Nature thanks the anonymous reviewers for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 CRISPR–Cas9-mediated knockout of PCSK9 and its effect on tumour-cell growth in vitro and in vivo.
a, Western blot analysis of the expression of PCSK9 in murine tumour lines (B16F10, 4T1, MC38 and CT26) with and without PCSK9 knockout (PCSK9KO). GAPDH was used as a protein loading control. The analysis was done twice with biologically independent samples. αGAPDH, anti-GAPDH antibody; αPCSK9, anti-PCSK9 antibody. b, Cell growth of vector control or PCSK9KO B16F10 tumour cells. Results (means ± s.e.m.) are from five biologically independent samples; P-values were calculated by unpaired two-sided t-test. c, Soft agar analysis of the colony-formation ability of B16F10 tumour cells transduced with vector control or PCSK9 sgRNA. d, Quantitative representation of soft agar formation in c. n = 4 biologically independent samples, showing means ± s.e.m; P-values calculated by unpaired two-sided t-test. e, Details of the in vivo competition assay. f, Change in ratios of mixed control–tdTomato and PCSK9KO–EGFP B16F10 cells after 12 days of growth in vivo (subcutaneously) in C57BL/6 mice, as determined by flow cytometry. g, Quantitative representation of the flow analysis in f, showing means ± s.e.m. n = 2 and 4 biologically independent tumour samples for in vitro and in vivo groups, respectively. P-values determined by unpaired two-sided t-test.
Extended Data Fig. 2 Effect of PCSK9 re-expression and the host immune system on tumour formation by PCSK9-knockout cells.
a, Western blot analysis of the expression of exogenously transduced, HA-tagged PCSK9 in PCSK9KO B16F10 cells. The analysis was done once. b, c. Tumour formation from B16F10 PCSK9KO cells transduced with either vector control or PCSK9 (b), and Kaplan–Meier survival curves of host mice (c). About 2 × 105 tumour cells were injected subcutaneously into C57BL/6 mice and observed for tumour formation. n = 5 tumours per group. Error bars show means ± s.e.m.; P-values were determined by two-way ANOVA in b and log-rank test in c. d–i, Growth rate (d, g), host survival (e, h) and endpoint tumour weight (f, i) of vector control and PCSK9KO 4T1 (d–f) and B16F10 (g–i) tumours. In each case, about 1 × 105 tumour cells were injected subcutaneously and observed for tumour formation in NCG mice. n = 6 mice for d, e, g, h; and n = 5 tumours for f, i. Error bars in d, f, g, i represent means ± s.e.m.; ns, not significant, as determined by two-way ANOVA (d, g), log-rank test (e, h), or unpaired two- sided t-test (f, i). j, k, Tumour growth from vector control and PCSK9KO B16F10 cells in Rag1−/− C57BL/6 mice (j) and Kaplan–Meier survival curve of tumour-bearing host mice (k). About 1 × 105 vector control or PCSK9KO B16F10 tumour cells were injected into Rag1−/− C57BL/6 mice and observed for tumour formation. n = 5 tumours per group. Error bars in j show means ± s.e.m. P-values were calculated by two-way ANOVA in j and by log-rank test in k.
Extended Data Fig. 3 The influence of tumour or host cell LDLR and host cholesterol levels on tumour growth from control or PCSK9KO tumour cells in immunocompetent hosts.
a, Western blot analysis of CRISPR–Cas9-mediated knockdown (KD) of LDLR in B16F10 cells. The analysis was done once. b, Tumour growth from vector control and LDLR KD B16F10 cells in C57BL/6 mice. n = 5 tumours per group. Error bars show means ± s.e.m. P-values were calculated by two-way ANOVA. c, Kaplan–Meier survival curve of mice (from b) bearing control and LDLR KD B16F10 tumours. n = 5 mice per group. P-values calculated by log-rank test. d, Tumour growth from vector control and PCSK9-knockout B16F10 cells in wild-type (C57) and LDLR−/− mice fed on a high-fat diet. n = 12, 12, 5 and 5 tumours in, respectively, wild-type mice inoculated with control or PCSK9KO tumour cells, and in LDLR−/− mice inoculated with control and PCSK9KO tumour cells. Error bars show means ± s.e.m. P-values were calculated by two-way ANOVA with multiple comparisons. e, Kaplan–Meier survival curve for wild-type and LDLR−/− mice (from d) bearing vector control and PCSK9-knockout tumours. P-values were calculated by log-rank test.
Extended Data Fig. 4 Additional data on anti-PD1 treatment in murine tumours.
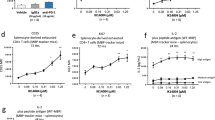
a, Treatment schedule for PCSK9KO 4T1 cells. Balb/c mice were implanted subcutaneously with PCSK9KO 4T1 tumour cells, treated with an anti-PD1 antibody at the indicated times, and observed for tumour formation. Animals that were implanted with PCSK9KO 4T1 tumour cells but did not form visible tumours by day 9 after inoculation were excluded from treatment with anti-PD1 antibodies. b, Tumour-growth delay in mice bearing PCSK9KO 4T1 tumours with or without anti-PD1 treatment. n = 5 tumours per group. Error bars show means ± s.e.m. P-values were calculated by two-way ANOVA. c, Kaplan–Meier survival curves for tumour-bearing mice from b. P-values were calculated by log-rank test. d, Treatment schedule for PCSK9KO CT26 tumours. Balb/c mice were implanted subcutaneously with PCSK9KO CT26 tumour cells, treated with an anti-PD1 antibody at the indicated times, and observed for tumour formation. e, Tumour-growth delay in mice bearing PCSK9KO CT26 tumours with or without anti-PD1 treatment. n = 5 tumours per group. Error bars show means ± s.e.m. P-values were determined by two-way ANOVA. f, Kaplan–Meier survival curves for tumour-bearing mice from e. Error bars show means ± s.e.m. P-values were determined by log-rank test. g, A scheme to develop anti-PD1-resistant MC38R tumour cells. h, Scheme for treating anti-PD1-resistant MC38R tumours with evolocumab and an anti-PD1 antibody. i, Tumour growth kinetics from anti-PD1-resistant MC38R tumours treated with anti-PD1 antibody and/or evolocumab. n = 5 tumours per group. Error bars show means ± s.e.m. P-values were determined by two-way ANOVA. j, Kaplan–Meier survival curve for mice bearing MC38R tumours from i. P-values were determined by log-rank test. k, Treatment schedule for PCSK9KO MC38 tumours. l, m, Tumour growth delay (l) and host mouse survival (m) among isotype (iso)- or evolocumab-treated mice bearing MC38-PCSK9KO tumours. n = 5 tumours per group. P-values were calculated by two-way ANOVA test in l and log-rank test in m.
Extended Data Fig. 5 Rechallenge of mice that were tumour free after initial tumour inoculation, and gating strategy for intratumoral immune effector cells.
a–c, Treatment scheme (a), tumour growth (b) and survival of host mice (c) after rechallenge with wild-type 4T1 tumour cells in Balb/c mice that remained tumour-free 43 days after initial challenge with PCSK9-knockout 4T1 cells. The control group consisted of tumour-naive Balb/c mice challenged with wild-type 4T1 cells. n = 5 and 12 mice for naive and rechallenged groups, respectively. Error bars in b show means ± s.e.m. P-values in b, c were calculated by two-way ANOVA test and log-rank test, respectively. d–f, Treatment scheme (d), tumour growth (e) and survival of host mice (f) after rechallenge with wild-type B16F10 tumour cells in C57BL/6 mice that remained tumour-free 26 days after initial challenge with PCSK9-knockout B16F10 cells and treatment with anti-PD1 antibody. The control group consisted of tumour-naive C57BL/6 mice challenged with wild-type B16F10 cells. n = 5 and 13 mice for tumour-naive and rechallenge groups, respectively. Error bars in e show means ± s.e.m. P-values in e, f were calculated by two-way ANOVA and log-rank test, respectively. g–i, Treatment scheme (g), tumour growth (h) and survival of host mice (i) after rechallenge with parental MC38 tumour cells in C57BL/6 mice that remained tumour-free 34 days after initial challenge with PCSK9-knockout MC38 cells and treatment with anti-PD1 antibody. The control group consisted of tumour-naive C57BL/6 mice challenged with wild-type MC38 cells. n = 5 mice per group. Error bars in h show means ± s.e.m. P-values were calculated by two-way ANOVA (h) and log-rank test (i). j, Representative flow-cytometry gating strategy for quantifying the numbers of various immune effector cell subsets in murine tumours.
Extended Data Fig. 6 Additional data on the characterization of lymphocyte infiltration into murine tumours.
a, Immunofluorescence staining (left) and quantitative estimates (right) of CD45+ leukocytes in control and PCSK9KO tumours grown in syngeneic C57BL/6 mice. Scale bar, 50 μm; n = 3 biologically independent samples; four fluorescent fields for each of the three samples were counted. Error bars show means ± s.e.m.; P-values were calculated using unpaired two-sided t-tests. b, Immunofluorescence staining (left) and quantitative estimates (right) of CD8a+ cells in control and PCSK9KO B16F10 tumours. Scale bar, 20 μm. n = 3 biologically independent samples; four fluorescent fields for each of the three samples were counted; error bars show means ± s.e.m.; P-values were calculated using unpaired two-sided t-tests. c, Quantitative estimates of CD4+ and CD8+ T cells in the spleens of mice bearing control and PCSK9KO B16F10 tumours, as determined by flow cytometry. n = 3 mice per group; error bars show means ± s.e.m.; P-values were calculated using unpaired two-sided t-tests. d, Flow-cytometric determination of the percentage of intratumoral CD8+ T cells that expressed IFN-γ. n = 6, 5 tumours in the two groups. Error bars show means ± s.e.m.; P-values were calculated by unpaired two-sided t-test. e, f, qRT–PCR analysis of intratumoural IFNG (e) and GZMB (f) mRNA levels in control and PCSK9KO tumours. n = 3 and 4 tumours for INFG and GZMB groups, respectively. Error bars show means ± s.e.m.; P-values were determined by unpaired two-sided t-test. g–i, Flow-cytometric characterization of the cell-surface expression levels of exhaustion markers for intratumoral CD8+ T cells in vector control and PCSK9KO tumours. n = 6 and 5 tumours for control and PCSK9-knockout conditions. Error bars show means ± s.e.m.; P-values were determined by unpaired two-sided t-test. j, Schedule for treating Balb/c mice, injected with 4T1 tumour cells, with evolocumab (αPCSK9 Ab) and anti-PD1 antibodies. k, Growth of 4T1 tumours treated with anti-PD1 antibodies and/or evolocumab. n = 5 mice per group. P-values were determined by two-way ANOVA. l, Kaplan–Meier survival curves for mice in k. P-values were determined by log-rank tests. m, Frequency of CD8+ T cells in 4T1 tumours treated with anti-PD1 antibodies and/or evolocumab. n = 5 tumours per group. Error bars show means ± s.e.m.; P values were determined by unpaired two-sided t-test. n, Frequency of IFNγ+ CD8+ T cells in 4T1 tumours treated with anti-PD1 antibodies and/or evolocumab. n = 5 tumours per group. Error bars show means ± s.e.m.; P-values were determined by unpaired two-sided t-test.
Extended Data Fig. 7 Additional data on the effect of PCSK9 inhibition on immune effector function and antigen presentation.
a, Injection schedule for antibody-mediated depletion of CD4+, CD8+ and natural killer (NK) immune cells. b, c, Growth rates (b) and host mouse survival (c) for PCSK9KO tumours in mice administered with control or anti-CD4 antibodies. n = 5 tumours per group. Error bars show means ± s.e.m.; P-values were determined by two-way ANOVA (b) and log-rank test (c). d, e, Growth rates (d) and host mouse survival (e) for PCSK9KO tumours in mice administered with control or anti-NK1.1 antibody. n = 5 tumours per group. Error bars show means ± s.e.m.; P-values were determined by two-way ANOVA (d) and log-rank test (e). f, Fluorescence images of tdTomato-labelled tumour cells with or without the OVA antigen in the presence or absence of OVA-specific T cells. The experiments were repeated twice with similar results. Scale bar, 200 μm. g, Enhanced presentation of OVA antigen (SIINFEKL) by MHC I in cultured B16F10 cells following PCSK9 deficiency. Control and PCSK9KO B16F10 cells transduced with the OVA gene were treated with IFN-γ and assayed for the amount of cell-surface H-2Kb–SIINFEKL complex using flow cytometry. Shown are representative results from analyses of four sets of biologically independent samples. h, i, Flow-cytometric analysis of MHC II (h) and PD-L1(i) expression in control and PCSK9KO B16F10 cells. n = 5 and 4 biologically independent samples, respectively. P-values were determined by unpaired two-sided t-test. j, Western blot analysis of PCSK9 expression in control or PCSK9KO MDA-MB-231 cells. The analyses were carried out twice. k, Effects of evolocumab and alirocumab on HLA-ABC expression on the surface of MDA-MB-231 human breast cancer cells. n = 6, 6 and 5 biologically independent samples from left to right. Data represent means ± s.e.m.; P-values were determined by unpaired two-sided t-test. l, H2-Kd/Dd expression levels for 4T1 tumour cells that were exposed to anti-PD1 antibodies and/or evolocumab in vivo. n = 5 mice per group. Error bars show means ± s.e.m.; P-values were determined by unpaired two-sided t-test.
Extended Data Fig. 8 Additional data on the analysis of PCSK9, H2-K1 and LDLR in murine tumour cells.
a, Lentivirus-mediated overexpression of HA-tagged H2-K1 in B16F10 cells as determined by western blot analysis. The analysis was done once. b, c, Tumour-growth delay (b) and Kaplan–Meier survival curve (c) of tumour-bearing C57BL/6 mice implanted with vector control or H2-K1-overexpressing B16F10 cells. Error bars show means ± s.e.m.; n = 5 tumours in each group; P-values were determined by two-way ANOVA (b) and log-rank test (c). d, e, Tumour-growth delay (d) and Kaplan–Meier survival curves (e) in mice injected with vector control, H2-K1-knockdown, PCSK9-knockout, or PCSK9-knockout plus H2-K1-knockdown B16F10 cells. n = 5 mice per group. Error bars show means ± s.e.m.; P-values were determined by two-way ANOVA (d) and log-rank test (e). f, Western blot analysis of LDLR knockdown in control and PCSK9KO B16F10 tumour cells. The analysis was done once. g, h, Tumour-growth delay (g) and Kaplan–Meier survival curves (h) from LDLR KD and LDLR KD/PCSK9KO B16F10 tumours. n = 5 mice per group. Error bars show means ± s.e.m.; P values were determined by two-way ANOVA (h) and log-rank test (i). i, Flow-cytometric analysis of MHC I expression in tumours formed from tdTomato-labelled control and LDLRKD B16F10 cells. n = 6 biologically independent tumours. Error bars show means ± s.e.m.; P-values were calculated by unpaired two-sided t-test. j, Flow-cytometric analysis of MHC I expression in tumours formed from tdTomato-labelled LDLR KD (n = 6) and LDLR KD/PCSK9KO cells (n = 4). Error bars show means ± s.e.m.; P-values were calculated by unpaired two-sided t-test.
Extended Data Fig. 9 Additional data on the mapping and functional characterization of interacting domains in PCSK9 and MHC I, and on the association of PCSK9 expression with the prognosis of TCGA cohorts.
a, Domain structure of mouse PCSK9. Catalytic, catalytic domain; CRD; C-terminal domain; Pro, propeptide; SP, signal peptide. b, Immunoprecipitation/western blot analysis of the interaction between full-length FLAG-labelled H2-K1 and full-length or partially deleted mouse HA-labelled PCSK9. Plasmids encoding the two genes were transfected into 293T cells in pairs, and lysates from transduced cells were immunoprecipitated with an anti-HA antibody and probed with an anti-FLAG antibody by western blot analysis. The analyses were repeated twice with biologically independent samples with similar results. c, Immunoprecipitation/western blot analysis of the interaction between full-length HA-labelled mouse PCSK9 and full-length or partially deleted FLAG-labelled H2-K1 (amino acids 66–202) (α1–α2 domains). The analyses were repeated twice with biologically independent samples, with similar results. d, Immunoprecipitation/western blot analysis of the interaction of HA-labelled mouse PCSK9 with full-length H2-K1 or H2-K1 with more limited deletions (amino acids 66–100, α1 domain; or amino acids 68–70). The analyses were repeated twice with biologically independent samples, with similar results. e, f, Tumour growth rates (e) and Kaplan–Meier survival curves (f) for mice inoculated with PCSK9KO B16F10 tumour cells, with re-expressed wild-type or partially (ΔM2) deleted PCSK9. n = 5 tumours per group. Error bars show means ± s.e.m.; P-values were determined by two-way ANOVA (e) and log-rank test (f). g, h, Tumour growth rates (g) and Kaplan–Meier survival curve (h) for mice inoculated with H2-K1KO or H2-K1/PCSK9 double-knockout (DKO) B16F10 tumour cells re-expressed with wild-type or partially deleted (Δ68–70) H2-K1. n = 5 tumours per group. Error bars show means ± s.e.m.; P-values were determined by two-way ANOVA (g) and log-rank test (h). i, Higher levels of PCSK9 expression correlate with worse survival in nine cohorts of patients with cancer, including liver hepatocellular carcinoma (LIHC), pancreatic adenocarcinoma (PAAD), skin cutaneous melanoma (SKCM), uveal melanoma (UVM), bladder urothelial carcinoma (BLCA), lung adenocarcinoma (LUAD), kidney renal clear cell carcinoma (KIRC), kidney renal papillary cell carcinoma (KIRP) and ovarian carcinoma (OV). P-values were calculated by log-rank test. Data are from TCGA data sets.
Extended Data Fig. 10 Diagram illustrating PCSK9-mediated degradation of MHC I in the lysosome.
Left, in the presence of PCSK9, MHC I is transported into lysosomes and degraded. Right, in the absence of PCSK9 (through genetic deletion or antibody-mediated neutralization), MHC I levels on the surface remain high and can thus present tumour-specific peptide antigens more efficiently to T cells. Illustration by S. Coffman.
Supplementary information
Supplementary Information
This file contains Supplementary Figure 1. Raw images of immunoblots. Uncropped images of scanned immunoblots shown in Fig. 4g, Fig 4i, Fig. 4j, Fig 4k, and 4l, Extended Data Fig. 1a, Extended Data Fig. 2a, Extended Data Fig. 3a, Extended Data 7j, Extended Data Fig. 8a, Extended Data Fig. 8f, Extended Data Fig. 9b, Extended Data Fig. 9c, and Extended Data Fig. 9d.
Source data
Rights and permissions
About this article
Cite this article
Liu, X., Bao, X., Hu, M. et al. Inhibition of PCSK9 potentiates immune checkpoint therapy for cancer. Nature 588, 693–698 (2020). https://doi.org/10.1038/s41586-020-2911-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-020-2911-7
This article is cited by
-
Inhibition of autophagy-related protein 7 enhances anti-tumor immune response and improves efficacy of immune checkpoint blockade in microsatellite instability colorectal cancer
Journal of Experimental & Clinical Cancer Research (2024)
-
Targeting PCSK9 to upregulate MHC-II on the surface of tumor cells in tumor immunotherapy
BMC Cancer (2024)
-
Pharmaceutical targeting of OTUB2 sensitizes tumors to cytotoxic T cells via degradation of PD-L1
Nature Communications (2024)
-
Targeting proprotein convertase subtilisin/kexin type 9 (PCSK9): from bench to bedside
Signal Transduction and Targeted Therapy (2024)
-
Carnosine regulation of intracellular pH homeostasis promotes lysosome-dependent tumor immunoevasion
Nature Immunology (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.