Abstract
Interferon-induced transmembrane protein 3 (IFITM3) has previously been identified as an endosomal protein that blocks viral infection1,2,3. Here we studied clinical cohorts of patients with B cell leukaemia and lymphoma, and identified IFITM3 as a strong predictor of poor outcome. In normal resting B cells, IFITM3 was minimally expressed and mainly localized in endosomes. However, engagement of the B cell receptor (BCR) induced both expression of IFITM3 and phosphorylation of this protein at Tyr20, which resulted in the accumulation of IFITM3 at the cell surface. In B cell leukaemia, oncogenic kinases phosphorylate IFITM3 at Tyr20, which causes constitutive localization of this protein at the plasma membrane. In a mouse model, Ifitm3−/− naive B cells developed in normal numbers; however, the formation of germinal centres and the production of antigen-specific antibodies were compromised. Oncogenes that induce the development of leukaemia and lymphoma did not transform Ifitm3−/− B cells. Conversely, the phosphomimetic IFITM3(Y20E) mutant induced oncogenic PI3K signalling and initiated the transformation of premalignant B cells. Mechanistic experiments revealed that IFITM3 functions as a PIP3 scaffold and central amplifier of PI3K signalling. The amplification of PI3K signals depends on IFITM3 using two lysine residues (Lys83 and Lys104) in its conserved intracellular loop as a scaffold for the accumulation of PIP3. In Ifitm3−/− B cells, lipid rafts were depleted of PIP3, which resulted in the defective expression of over 60 lipid-raft-associated surface receptors, and impaired BCR signalling and cellular adhesion. We conclude that the phosphorylation of IFITM3 that occurs after B cells encounter antigen induces a dynamic switch from antiviral effector functions in endosomes to a PI3K amplification loop at the cell surface. IFITM3-dependent amplification of PI3K signalling, which in part acts downstream of the BCR, is critical for the rapid expansion of B cells with high affinity to antigen. In addition, multiple oncogenes depend on IFITM3 to assemble PIP3-dependent signalling complexes and amplify PI3K signalling for malignant transformation.
Similar content being viewed by others
Main
IFITM3 is a small antiviral membrane protein (133 amino acids in size) with activity against influenza A virus, severe acute respiratory syndrome (SARS) coronavirus and many other viruses1,2,3,4,5. Endocytosis and preferential endosomal localization of IFITM3 are mediated by its N-terminal AP2 sorting motif, YEML1. Phosphorylation of this motif at Y20 by SRC kinases prevents not only endocytosis but also proteasomal degradation6,7. Individuals with the IFITM3 single-nucleotide polymorphism (SNP) rs12252-C allele show increased morbidity and mortality from H1N1 strains of influenza A virus8 and SARS coronavirus 24. IFITM3 SNP rs12252-C was predicted through mRNA splicing to encode a 21-amino-acid truncation of the N terminus (which includes Y20), although a recent transcript analysis found no evidence for such processed mRNA9. IFITM3 probably functions as an antiviral effector molecule by limiting viral fusion through increasing the ‘rigidity’ of endosomal cell membranes10,11. IFITM3 is overexpressed in multiple types of cancer, and its expression levels correlated with histopathological grading and staging12,13. Although IFITM3-dependent antiviral effector functions are restricted to endosomal compartments14, we report here on IFITM3-dependent functions at the cell surface of B cells, where IFITM3 functions as amplifier of PI3K signalling.
IFITM3 is essential for B cell affinity maturation
Although IFITM3 is only minimally expressed in normal resting B cells, BCR engagement and acute activation of the PI3K pathway strongly induced the expression of this protein (Extended Data Fig. 1a–c). Expression of IFITM proteins has recently been established as a diagnostic biomarker of high-risk B cell-lineage acute lymphoblastic leukaemia (B-ALL) that expresses the oncogenic BCR–ABL1 tyrosine kinase or related kinases15; we found that BCR–ABL1 kinase inhibition decreased levels of IFITM3 expression (Extended Data Fig. 1d). Levels of IFITM3 mRNA were generally higher in B cell malignancies than in their normal counterparts (Extended Data Fig. 1f, g) and elevated IFITM3 mRNA levels were associated with poor clinical outcomes in multiple clinical cohorts of patients with B cell malignancies (including paediatric and adult B-ALL and mantle cell lymphoma) (Extended Data Fig. 1h–l). IFITM3 was previously identified as a transcriptional target of the transcriptional repressor IKZF116, which suggests that high levels of IFITM3 expression may be the result of genetic lesions in IKZF1. Our results confirmed that reconstitution of IKZF1 suppresses IFITM3 expression, and that inducible deletion of Ikzf1 in mice relieves transcriptional repression (Extended Data Fig. 1m, n). However, a multivariate analysis of risk factors in B-ALL showed that high IFITM3 mRNA levels are an independent predictor of poor outcome, regardless of IKZF1 deletion (Extended Data Fig. 1o). We next examined how the genetic deletion of Ifitm3 affects normal B cell development in mice. Surface expression of CD19—a common B cell surface receptor that promotes SRC kinase and PI3K signalling17—was substantially diminished in Ifitm3−/− pre-B cells (Fig. 1a). Consistent with increased apoptosis and reduced proliferation, expression of MYC and BCL2 as well as PI3K signalling were reduced in Ifitm3−/− pre-B cells (Fig. 1a–c). The development of resting Ifitm3−/− B cell populations was largely unperturbed (Extended Data Fig. 2), whereas B1 cell and marginal-zone B cell populations were decreased (Fig. 1d, Extended Data Fig. 2). To assess B-cell-specific effects of IFITM3 during humoral immune responses, we performed adoptive transfer experiments: Ifitm3+/+ or Ifitm3−/− splenic B cells were transferred into congenic μMT mice that lack endogenous mature B cells. After reconstitution of mature B cell development, recipient μMT mice were immunized with the hapten 4-hydroxy-3-nitrophenyl acetyl (NP) conjugated to keyhole limpet haemocyanin (KLH) (NP–KLH). Upon immunization, μMT mice reconstituted with Ifitm3+/+ B cells developed about 5% NP-specific germinal-centre B cells, which were largely absent in μMT mice reconstituted with Ifitm3−/− B cells (Fig. 1e–h). Similarly, the number of follicles in the spleen, PNA+ germinal-centre B cells and production of IgM, IgG1 and IgG2b antibodies were reduced in the absence of Ifitm3 (Fig. 1i). These abnormalities in Ifitm3-deficient B cells phenocopy B cell defects seen in mice upon deletion of Pik3cd18 and Cd1919.
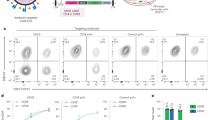
a, CD19 surface expression (top), cell cycle progression (middle) (percentages in S phase) and cell viability (bottom) (percentages of annexin V+7-AAD+ cells) were measured. b, Number of viable Ifitm3+/+ and Ifitm3−/− pre-B cells were counted at the times indicated. c, Levels of AKT phosphorylated at S473 (AKT-pS473), AKT, MYC, p53, p21 and BCL2 were assessed in Ifitm3+/+ and Ifitm3−/− pre-B cells. d, Splenic B cells from Ifitm3+/+ and Ifitm3−/− mice (n = 5) were analysed for CD21highCD23low or CD21highCD23− marginal-zone B cells (MZB), peritoneal-cavity B cells for MAC1+IgM+ B1 cells and CD5+IgM+ B1a cells. e, Splenic B cells from Ifitm3+/+ and Ifitm3−/− mice were adoptively transferred to μMT recipient mice (n = 10) followed by immunization with 0.5 mg of NP–KLH or vehicle. Spleens were collected on day 12 after immunization and subjected to immunofluorescence staining of tissue sections with B220, CD3 and peanut agglutinin (PNA). Scale bar, 500 μm. f–h, Splenocytes collected from μMT mice (n = 10) were analysed by flow cytometry 12 days after immunization for CD95, GL7 and NP to identify NP-specific germinal-centre B cells. Relative fractions (f), absolute numbers (g) and representative flow cytometry plots (h) are shown. i, Levels of serum immunoglobulin isotypes in μMT recipient mice transplanted with Ifitm3+/+ or Ifitm3−/− B cells are shown before and after immunization (n = 10; day 12). Serum levels of IgM, IgG1 and IgG2b were determined by enzyme-linked immunosorbent assay (ELISA). For gel source data for a–c, see Supplementary Fig. 1. In b, f, g, i, mean ± s.d. indicated; significance determined by two-tailed t-test.
Role of Ifitm3 in oncogenic signalling
To elucidate a potential contribution of Ifitm3 to oncogenic signalling, we studied the transformation of Ifitm3+/+ and Ifitm3−/− pre-B cells by BCR-ABL1 and NRASG12D (Fig. 2a, b). As in normal B cells, Ifitm3−/− leukaemia cells in a mouse model of B-ALL (Ifitm3−/− B-ALL cells) showed multiple defects in survival and proliferation, and a markedly reduced ability to form colonies in semi-solid methylcellulose (Fig. 2b, c). Consistent with a 70-fold reduction of the frequency of leukaemia-initiating cells (P = 3.7 × 10−7), Ifitm3−/− B-ALL cells did not initiate overt leukaemia in immunodeficient transplant recipient mice (Fig. 2d, e). Similar to B-ALL cells, deletion of IFITM3 in mantle cell lymphoma (MCL) cells reduced competitive fitness and proliferation (Fig. 2f). Both mouse Ifitm3−/− B cells and human IFITM3-deficient MCL cells showed defective Ca2+ release in response to BCR engagement, and partially lost CD19 surface expression (Extended Data Fig. 2c, d). In addition, BCR engagement in IFITM3−/− MCL cells did not induce processive signal amplification between CD19 and LYN17, as CD19 was not phosphorylated and did not interact with LYN. IFITM3-deficient MCL cells expressed lower levels of LYN and did not activate SRC kinases in response to BCR engagement (Extended Data Fig. 2e). RNA-sequencing gene-expression studies revealed upregulation of Ifitm1 in Ifitm3−/− B-ALL cells, presumably to compensate for loss of Ifitm3. Gene set enrichment analyses showed that Ifitm3 deficiency was associated with reduced expression of components of the BCR signalling pathway and a phenotype reminiscent of B cell anergy (Extended Data Fig. 3a–c).
a, CD19 surface expression (top), cell cycle progression (middle) (percentages in S phase) and cell viability (bottom) (percentages of annexin V+7-AAD+ cells) were measured. b, c, Ifitm3+/+ and Ifitm3−/− B-ALL cells transformed with human BCR-ABL1 or NRASG12D were assayed for levels of AKT-pS473, AKT, MYC, p53, p21 and BCL2 (n = 3) (b) and plated in semi-solid methylcellulose (c). Scale bars, 7 mm (c, top row), 2.5 mm (c, bottom row). d, Kaplan–Meier analyses of NOD-SCID-gamma (NSG) recipient mice injected with the indicated numbers of Ifitm3+/+ and Ifitm3−/− BCR-ABL1 (left) or NRASG12D (right) B-ALL cells (n = 5). e, Frequencies of leukaemia-initiating cells (LIC) estimated with extreme limiting dilution analysis (ELDA) (90% confidence interval; likelihood ratio test). f, Jeko1 MCL cells expressing doxycycline (Dox)-inducible Cas9 were transduced with IFITM3-targeting or non-targeting (NT) single-guide RNAs. Enrichment or depletion of targeted cells (Cas9+gRNA+) was monitored by flow cytometry upon treatment with doxycycline (mean ± s.d.), and IFITM3 levels were measured by western blot. g, Premalignant LSL-BcrBCR-ABL1 × Mb1-cre pre-B cells expressing IFITM3, IFITM3(Y20E) or empty vector (EV) were plated for colony-forming assays (7 days). Representative images shown at 1× (top) and 10× (bottom) magnification. h, Survival analyses (P = 0.0001, log-rank test) of congenic recipient mice transplanted with LSL-BcrBCR-ABL1 × Mb1-cre B cells transduced with empty vector, IFITM3 or IFITM3(Y20E) (n = 7). i, The engraftment and expansion of luciferase-labelled leukaemia cells were monitored by luciferase bioimaging at the times indicated. j, Effects of IFITM3 or IFITM3(Y20E) on oncogenic signalling in LSL-BcrBCR-ABL1 × Mb1-cre B cell precursors measured by western blot and compared to empty vector. Phosphorylation of CD19 at Y531 (CD19-pY531), of SRC at Y416 (SRC-pY416) and AKT at S473 were examined. Gel source data for a–c, f, g, j are shown in Supplementary Fig. 1. In c, g, two-tailed t-test.
Effect of phosphorylation of IFITM3 at Y20
Combinations of surface and intracellular staining revealed the cytoplasmic localization of the N terminus of IFITM3, with a short extracellular portion of the C terminus (Extended Data Fig. 3d). Previous studies have shown that phosphorylation of IFITM3 at Y20 prevents both endocytosis and degradation6,7. Phosphomimetic IFITM3(Y20E) was constitutively localized at the cell surface. Similarly, phosphorylation by BCR–ABL1 induced the accumulation of IFITM3 at the cell membrane, in a process that was sensitive to inhibition of BCR–ABL1 or SRC kinases (Extended Data Fig. 3e). Therefore, the phosphorylation of Y20 induces a dynamic switch from antiviral functions in endosomes to BCR or oncogene signalling at the cell surface (Extended Data Fig. 3e, f). To test whether phosphorylation of IFITM3 at Y20 and constitutive membrane localization are sufficient to enable oncogenic signalling, we overexpressed wild-type and IFITM3(Y20E) in premalignant B cells carrying a BCR-ABL1-knock-in allele, which did not readily give rise to leukaemia20. Expression of IFITM3(Y20E) increased the ability of pre-B cells to form colonies by 129-fold (P = 2.1 × 10−8) (Fig. 2g), induced oncogenic signalling and initiated fatal leukaemia in congenic transplant recipient mice (Fig. 2h–j). To elucidate how membrane-bound IFITM3(Y20E) promotes malignant transformation, we performed phosphoproteomic analyses of IFITM3 Y20 signalling in patient-derived B-ALL cells. Consistent with processive signal amplification17, engagement of IFITM3 induced phosphorylation of CD19 on multiple residues—including Y531, which mediates PI3K activation. In addition, Y20 of IFITM3 engaged multiple components of the BCR and integrin receptor signalling pathways (Fig. 3a, Extended Data Fig. 4a), which converge with CD19 at the level of PI3K activation. A comprehensive analysis of phosphosites showed a significant enrichment of phosphorylation in the PI3K, BCR and integrin receptor pathways, which we confirmed for SRC, CD19 and PI3K signalling by western blot (Fig. 3a–c). IFITM3(Y20E) required crosslinking by a full antibody, as the engagement of Y20 of monomeric IFITM3 using F(ab) fragments did not elicit PI3K activation (Extended Data Fig. 4b). Because PI3K signalling in mouse B-ALL cells increased Ifitm3 mRNA levels by 10-fold and protein levels by more than 50-fold, these observations suggest a feed-forward loop of signal amplification between IFITM3 and PI3K signalling (Extended Data Fig. 1b–e). Comparable to effects on BCR signalling and CD19 surface expression in B cells, genetic deletion of IFITM3 in human Jurkat T cells compromised T cell receptor signalling and CD3 surface expression (Extended Data Fig. 4c), which suggests that IFITM3 may have a similar role in T cells.
a, Levels of proteins phosphorylated at the indicated residues in PDX2 B-ALL cells transduced with HA-tagged IFITM3(Y20E) or empty vector (EV) control were identified by mass spectrometry upon IFITM3 crosslinking with anti-HA antibodies. b, Feature set enrichment analysis (FSEA) for phosphorylated proteins in the BCR (red), integrin (blue) and PI3K (grey) pathways, ranked by log2-transformed fold change (Kolmogorov–Smirnov). c, Levels of SRC-pY416, LYN, CD19-pY531, CD19, AKT-pS473 and AKT assessed in ICN12 cells expressing IFITM3–HA, IFITM3(Y20E)–HA or HA-tagged empty vector upon IFITM3 crosslinking. d, e, Interactomes of BirA–IFITM3(Y20E) or empty vector control expressed in PDX2 B-ALL (d) and Jeko1 MCL (e) cells. f, g, Validation of IFITM3-interacting proteins by anti-Flag co-immunoprecipitation (IP) in PDX2 cells transduced with Flag–IFITM3 or Flag-tagged empty vector, followed by western blotting for validation of interacting proteins, and STAT5 and eIF4E as specificity controls. WCL, whole-cell lysates. h, i, Proximity ligation assay (PLA) in Jeko1 cells upon BCR stimulation, assessed for proximity of CD79B to IFITM3. Representative images with PLA signal (red), nuclei (DAPI) and plasma membrane (WGA) (green). Scale bars, 5 μm (i). h, Quantification of PLA signals per cell (two-tailed t-test). j, Structural model of IFITM3-mediated signal amplification between CD19 and LYN. In resting B cells (left), IFITM3 is localized in endosomes. Upon antigen encounter (right), LYN-mediated phosphorylation of IFITM3 induces membrane translocation, acting as a scaffold for CD19 and LYN in proximity to BCR molecules. BCR–CD19–IFITM3 complexes form clusters for PI3K activation and accumulation of PIP3 in lipid rafts. In a–i, n = 3. For gel source data, see Supplementary Fig. 1. In a, d, e, row-scaled protein abundance ranked by fold change; key proteins of interest are highlighted.
IFITM3 enables CD19–PI3K signalling
By enriching for cell-surface proteins using N-linked glycosylation-site biotin labelling, we identified 65 surface receptors—most of which were associated with lipid rafts—that were downregulated in Ifitm3−/− B-ALL cells, including CD19, BCR-associated and adhesion receptors. Only five surface proteins were upregulated—including the IFITM family member IFITM2, which suggests a possible compensation mechanism (Extended Data Fig. 4d). We performed validation by flow cytometry for 17 surface receptors for IFITM3 loss-of-function (Ifitm3−/−) and gain-of-function (IFITM3(Y20E)) (Extended Data Fig. 4e–g). Continuous forced expression of CD19 (for more than one week) in Ifitm3−/− B cells partially restored defective survival and proliferation, which required the PI3K-activation motif Y531 of CD19 (ref. 21) (Extended Data Fig. 5a–e). However, inducible translocation of CD19 to the cell membrane did not restore any defects in Ifitm3−/− B cells, which suggests that IFITM3 is still required to integrate CD19 into signalling complexes for SRC kinase and PI3K signalling (Extended Data Fig. 5f–h). To study complexes that interact with IFITM3, we performed enzyme-catalysed proximity labelling studies on the basis of N-terminal fusions of the BirA biotin ligase with IFITM3(Y20E) carrying an extracellular haemagglutinin (HA) tag in B-ALL and IgM+ MCL cells. After engagement with IFITM3 (anti-HA) or BCR (anti-IgM), IFITM3-interacting proteins were biotinylated on the basis of their proximity to the cytoplasmic BirA moiety, collected using streptavidin-coated beads and identified by mass spectrometry. Consistent with phosphoproteomic analyses (Fig. 3a), and across different cell types (B-ALL and MCL) and stimulations (anti-HA and anti-IgM), IFITM3-interacting proteins included BCR, integrin receptor and PI3K signalling elements (Fig. 3d, e, Extended Data Fig. 6a–d). We transduced MCL cells with Flag–IFITM3 for validation by co-immunoprecipitation and western blotting (Fig. 3f, g). Using proximity ligation assays, we confirmed the inducible formation of a complex between IFITM3 and the BCR signalling chain CD79B upon engagement of the BCR. Interactions between BCR and IFITM3 were induced within minutes and dissociated after 30 min (Fig. 3h, i). Consistent with endosomal BCR functions22, interactions between BCR and IFITM3 occasionally colocalized with LAMP1+ endosomal compartments (Extended Data Fig. 6e). Therefore, BCR-mediated activation of SRC-mediated phosphorylation of Y20 of IFITM3 induces the accumulation of IFITM3 at the plasma membrane, where IFITM3 functions as a scaffold for CD19 and LYN in proximity to BCR molecules to enable processive signal amplification (Fig. 3j). In resting B cells, IgM BCRs are excluded from lipid rafts and recruited upon encounters with antigen23. By contrast, IgD BCRs are already in close proximity to CD19 within lipid rafts in resting B cells, which suggests that IFITM3 may specifically enable IgM BCRs.
IFITM3–PIP3 binding is needed to form lipid rafts
Reduced expression of integrins and adhesion receptors was associated with decreased homotypic adhesion of Ifitm3−/− pre-B cells, whereas membrane-bound IFITM3(Y20E) triggered the formation of large clusters (Extended Data Fig. 7). Because IKZF1 negatively regulates adhesion24,25, we examined whether this regulation is mediated by transcriptional repression of Ifitm316. Overexpression of a dominant-negative IKZF1-mutant (termed IK6) markedly increased colony formation, adhesion to stroma and surface expression of adhesion receptors in the presence—but not in the absence—of IFITM3. Although dominant-negative inhibition of IKZF1 relieved transcriptional repression of adhesion receptors, the normal expression and function of these receptors at the cell surface still required IFITM3 and its ability to link integrins and adhesion receptors to the PI3K pathway. Loss of IFITM3, and reduced activity of the SRC kinase LYN and CD19–PI3K signalling, could broadly affect lipid rafts and short-circuit BCR signalling. Consistent with this scenario, the lipid-raft components cholesterol and ganglioside GM1 were markedly reduced in Ifitm3−/− B cells (Fig. 4a). In addition, BCR engagement increased membrane rigidity in IFITM3+/+ MCL cells. However, membrane stiffening in response to BCR engagement was significantly reduced in IFITM3−/− MCL cells (Extended Data Fig. 8a). Consistent with defects in PI3K signalling, the ratios of PIP3 to the PI3K substrate PIP2 were reduced substantially in mouse Ifitm3−/− B cells and patient-derived IFITM3−/− B-ALL cells. By contrast, overexpression of IFITM3 and IFITM3(Y20E) increased PIP3:PIP2 ratios by about 3-fold and 5.5-fold, respectively (Fig. 4b, c). To elucidate how IFITM3 interacts with plasma membrane lipids, we probed 15 distinct lipid classes for binding to IFITM3 in vitro. IFITM3 bound selectively to PIP3, but not to any of the other 14 lipids (Fig. 4d). Because IFITM3 function is required to retain normal PIP3 levels, we tested whether exogenous delivery of PIP3 could rescue Ifitm3 deficiency in mice. Insertion of exogenous PIP3 into the cell membrane partially restored AKT and MYC activation and colony formation, but not CD19 and SRC phosphorylation or surface expression of lipid-raft-associated receptors (Extended Data Fig. 8b–d). Therefore, IFITM3 promotes PI3K signalling not only by increasing the amount of PIP3 but also by integrating PIP3 into signalling complexes in lipid rafts.
a, Ganglioside GM1 and cholesterol (stained with filipin III) levels measured in Ifitm3+/+ and Ifitm3−/− pre-B (top) and B-ALL (bottom) cells. b, c, Ratios of PIP3 and PIP2 were measured in Ifitm3+/+ and Ifitm3−/− mouse (b, left), and human B-ALL cells (PDX2) (b, right), and upon expression of empty vector, IFITM3 or IFITM3(Y20E) (c). d, In vitro lipid-binding assays for glutathione-S-transferase-tagged IFITM3 (GST–IFITM3) and 15 lipid classes. Recombinant GST tag was used as baseline. e, Lipid-binding assays were performed for biotin–IFITM3 N terminus (amino acids 1–57), the first intramembrane α-helix (IM-α) (amino acids 58–70), conserved intracellular loop (CIL) (amino acids 71–105) and transmembrane α-helix (TM-α) (amino acids 89–105). f, Lipid-binding assays to study interactions of PIP3 with biotinylated fragments of the IFITM3 conserved intracellular loop that carry mutations of basic residues. g–i, PIP3 binding to K83 and K104, and to R85, R87 and K88, basic residue patches (g). Dashed lines indicate PIP3 contacts within 3.5 Å. h, Interaction energies of PIP3 with basic residues K83 or R85 averaged over molecular dynamics simulation trajectories (median levels indicated by red dotted line) (Methods). i, Binding affinity for PIP3 as shown as a heat map of contact frequencies between PIP3 and each residue. j–l, Levels of AKT-pS473, AKT, S6K phosphorylated at T389 (S6K-pT389), S6K phosphorylated at T421 and S424 (S6K-pT421/S424), CD19-pY531, CD19, SRC-pY416, LYN, PAK1 and PAK3 phosphorylated at S199 and S204 (PAK1/3-pS199/204), PAK2 phosphorylated at S192 and S197 (PAK2-pS192/197), PAK1, FAK phosphorylated at Y397 (FAK-pY397), FAK phosphorylated at Y576 and Y577 (FAK-pY576/577), FAK, CXCR4 phosphorylated at S339 (CXCR4-pS339), CXCR4 and integrin-β1 in PDX2 cells expressing empty vector, IFITM3–HA, IFITM3(Y20E)–HA, and the R85A, K83A, R85A/R87A/K88A and K83A/K104A mutants of IFITM3, upon IFITM3 crosslinking with anti-HA antibodies. In a–f, j–l, n = 3, For gel source data, see Supplementary Fig. 1. In b, c, e, h, two-tailed t-test.
Structural basis of IFITM3 binding to PIP3
Proteins that contain a cluster of four or more basic residues located at the membrane–solution interface can laterally sequester PIP3, which does not require a PH domain or any other specific structure26. To map PIP3 binding to specific portions of IFITM3, we repeated lipid-binding assays with biotin-tagged fragments, including the N terminus, the first intramembrane α-helix and the conserved intracellular loop. PIP3 binding was detected only for the conserved intracellular loop (Fig. 4e), which contains five basic residues that are in close proximity in sequence and that spatially cluster into two distinct basic patches. To address why IFITM3 preferentially binds to PIP3 over PIP2, we performed multi-scale molecular dynamics simulations of IFITM3 in multi-component lipid bilayers that mimic the plasma membrane. All-atom molecular dynamics simulations revealed that the representative binding pose of PIP2 from its most populated conformation cluster showed a charge interaction with one single basic residue (R85) (−213 kJ mol−1), which was energetically weaker than the two charge interactions with PIP3 at K83 and K104 (−704 kJ mol−1) (P = 0.0008). Two-residue contacts were rare for PIP2 and more common for PIP3 (P = 0.011) (Extended Data Fig. 9). To experimentally test the predicted function of the basic residues of the conserved intracellular loop, we performed PIP3 binding assays for biotinylated fragments of the IFITM3 conserved intracellular loop that carried mutations of these residues. The R85A and R85A/R87A/K88A mutants largely retained the ability to bind PIP3, which was abrogated for the K83A/K104A mutant (Fig. 4f). To understand the structural basis of this difference, we modelled the interaction of PIP3 with each of the mutants of the basic residues (K83A/K104A, R85A/R87A/K88A and R85A/R87A) using all-atom molecular dynamics simulations. PIP3 shows an interaction energy that is three times stronger with K83 and K104 than with R85, R87 and K88 (Fig. 4g–i). PIP3 makes dual-pronged salt-bridge interactions with K83 and K104, whereas it interacts only with R85 and not with R87 and K88 in the other basic patch. The R87 and K88 residues form intra-protein salt bridges with D92 and D86, respectively, and are not available for interaction with PIP3.
IFITM3 amplifies PI3K, integrin and BCR signalling
To experimentally validate this model, we globally identified changes in signal transduction that depend on K83 and R85 residues in human B-ALL and MCL B cells. The most prominent differences were enrichments for phosphorylation sites associated with PI3K, BCR and adhesion receptor signalling (Extended Data Fig. 10a, b). Although most phosphorylation events induced by membrane-bound IFITM3(Y20E) were lost upon mutation of K83, this was not the case for substitution of the R85 residue (Extended Data Fig. 10c, d). Proximity ligation assays revealed IFITM3-dependent recruitment of multiple proteins in the PI3K, BCR and integrin signalling pathways. However, almost all of these interactions were lost upon mutation of K83 (Extended Data Fig. 10e–h). Biochemical studies revealed that crosslinking of IFITM3(Y20E) resulted in massive activation of PI3K and integrin receptor pathways, whereas CD19 and SRC were constitutively phosphorylated in the presence of IFITM3(Y20E) (Fig. 4j–l). The R85A mutation had only minor effects, whereas the substitution of K83 almost entirely abolished IFITM3-dependent signal transduction. In contrast to the triple-mutant IFITM3(R85A/R87A/K88A), the IFITM3(K83A/K104A) double mutant lost all activity in PI3K, BCR and adhesion receptor pathways (Fig. 4j–l).
In summary, we have revealed a function for IFITM3 as a PIP3 scaffold in the amplification of PI3K signalling. In Ifitm3−/− B cells, lipid rafts were depleted of PIP3, which resulted in marked defects in lipid-raft-associated BCR and integrin receptor signalling. IFITM3-dependent amplification of PI3K signalling, acting—in part—downstream of the BCR and adhesion receptors, is critical for the rapid expansion of B cells with high affinity to antigen. In addition, multiple oncogenes depend on IFITM3 to assemble PIP3-dependent signalling complexes and amplify PI3K signalling for malignant transformation.
Methods
No statistical methods were used to predetermine sample size. The experiments were not randomized and investigators were not blinded to allocation during experiments and outcome assessment.
Analysis of patient gene expression and outcome data
Gene expression microarray data from three large cohorts of patients with pre-B ALL were downloaded from GSE531427 (the Eastern Cooperative Oncology Group (ECOG) Clinical Trial E2993), GSE1187728 (the Children’s Oncology Group (COG) Clinical Trial P9906), St. Jude Research Hospital paediatric ALL29 (http://www.stjuderesearch.org/site/data/ALL3/). In ECOG E2993, pretreatment bone marrow or peripheral blood samples were obtained at diagnosis before any treatment from 83 patients with Ph+ B-ALL enrolled in the Medical Research Council (MRC) UKALLXII/Eastern Cooperative Oncology Group (ECOG) E2993 phase III trial. In data set from St. Jude childhood, 15 cases of Ph+ B-ALL were selected from the original 327 diagnostic bone marrow aspirates. Minimal residual disease (MRD) data were available for patients with paediatric high-risk B-ALL (COG P9906). IFITM3 expression levels were measured in patients with either a MRD positive (MRD+) status or a MRD negative (MRD-) status. In COG P9906 clinical outcome, expression profiles were obtained in pretreatment leukaemic samples from 207 uniformly treated children with high-risk ALL30, a risk category largely defined by pretreatment clinical characteristics. Patients had MRD tested by flow cytometry with two combinations (CD20/CD10/CD45 or CD9/CD19/CD34/CD45), and were defined as MRD positive or MRD negative at the end of induction therapy (day 29) using a threshold of 0.01% as previously described31. Then, RNA was purified from 207 pretreatment diagnostic samples with more than 80% blasts (131 bone marrow, 76 peripheral blood) and subjected to microarrays. Log-rank test was used to assess statistical significance.
Primary human samples and cell lines
Patient samples (Supplementary Table 7) were sourced ethically from patients who gave informed consent, and were in compliance with the internal review boards of the Beckman Research Institute of City of Hope. We have complied with all relevant ethical regulations. Patient samples were harvested from biopsy of bone marrow from patients with ALL at the time of diagnosis or relapse. All samples were transplanted into sublethally irradiated NOD.Cg-Prkdcscid Il2rgtm1Wjl/SzJ mice (NSG mice, The Jackson Laboratory) through tail-vein injection. After samples were collected, patient-derived primary human pre-B ALL xenografts were cultured on OP9 stroma in Alpha Minimum Essential Medium (MEMα; Life Technologies) with GlutaMAX containing 20% fetal bovine serum (FBS), 100 IU ml-1 penicillin, 100 μg ml-1 streptomycin and 1 mM sodium pyruvate. The human cell lines (Supplementary Table 8) were cultured in RPMI-1640 (GIBCO) with GlutaMAX containing 20%FBS, 100 IU ml-1 penicillin, 100 μg ml-1 streptomycin at 37 °C in a humidified incubator with 5% CO2. All human primary samples and cell lines were tested negative for mycoplasma by detection kit (MycoAlert PLUS, LONZA).
Genetic mouse models
Genetic mouse models used in this study are listed in Supplementary Table 9. NOD.Cg-Prkdcscid Il2rgtm1Wjl/SzJ (NSG) mice were purchased from Jackson Laboratory. Ifitm3tm1Masu (ref. 32) mice were backcrossed to the C57BL/6J background for more than eight generations. B6.C(Cg)-Cd79atm1(cre)Reth/EhobJ (Mb1–Cre) mice were purchased from Jackson Laboratory. B6.129S2-Ighmtm1Cgn/J (μMT) mice were purchased from Jackson Laboratory. C;129S4-Ptentm1Hwu/J (Ptenfl/fl) mice were purchased from Jackson Laboratory. For mice bred in-house, littermates of the same sex were randomized to experimental groups. For the in vivo leukaemia initiation assay, 8- to 10-week-old female NSG mice were randomly allocated before injection. To generate a model for pre-leukaemic B cell precursors expressing BCR–ABL1, LSL-Bcr+/BCR-ABL1 mice33 were crossed with Mb1-cre strain (Mb1-cre × LSL-Bcr+/ BCR-ABL1) for excision of a stop-cassette in early pre-B cells. For in vivo oncogenic priming assay with Mb1-cre × LSL-Bcr+/ BCR-ABL1 B cell precursors, 8- to 10-week-old female NSG mice were randomly allocated before injection. Temperatures of 18–23 °C with 40–60% humidity were maintained with 14-h light/10-h dark cycle. The following scores were considered as end-point: (1) a failure to eat food or drink water for 24 h; (2) a failure to make normal postural adjustments or to display normal behaviour; and (3) when tumour burden reached 1.5 cm × 1.5 cm × 1.5 cm (tumour ulceration was not expected). If a mouse lost 25% of its initial body weight (or reached 16 g of body weight, regardless of the initial weight), or if we observed a weight loss of 15% on 2 sequential weight measurements, we euthanized the mouse immediately. All mouse experiments were subject to institutional approval by the Beckman Research Institute of City of Hope Animal Care and Use Committee.
Mouse primary and leukaemia cells
Bone marrow cells from 6–10-week-old mice were collected by flushing cavities of the femur and tibia with chilled PBS, followed by filtering through 40-μm strainer to yield a single-cell suspension. Spleen or thymus cells were directly extracted by forcing tissues through a 40-μm strainer into chilled PBS. Filtered cells were further incubated with lysis buffer (RBC Lysis Buffer, BioLegend) to lyse erythrocytes. After washing with PBS, cells were subjected to further experiments. For IL-7-dependent pre-B cell culture, bone marrow cells were collected and cultured in Iscove’s modified Dulbecco’s medium (IMDM) (GIBCO) with GlutaMAX containing 20% FBS, 50 μM 2-mercaptoethanol, 100 IU ml−1 penicillin, 100 μg ml−1 streptomycin in the presence of 10 ng ml−1 recombinant mouse IL-7 (Peprotech). For the BCR–ABL1 leukaemia model, pre-B cells were retrovirally transformed with BCR-ABL1 and then IL-7 was removed to select the transduced cells. NrasG12D ALL cells were selected with puromycin (GIBCO) and were maintained with IL-7-supplemented IMDM.
Retroviral and lentiviral transduction
Retroviral supernatant was generated by cotransfection of HEK 293FT cells with retroviral constructs together with pHIT60 (gag-pol) and pHIT123 (for mouse) or pHIT456 (for human ecotropic envelope) using Lipofectamine 2000 (Invitrogen). Lentiviral supernatant for CRISPR-mediated gene editing was produced by cotransfection of HEK 293FT cells with lentiviral constructs together with pCDNL/BH and EM140. Transfected HEK 293FT cells were cultured in high-glucose Dulbecco’s modified Eagle’s medium (DMEM) (GIBCO) with GlutaMAX containing 10% fetal bovine serum, 100 IU ml−1 penicillin, 100 μg ml1 streptomycin, 25 mmol l−1 HEPES, 1 mmol l−1 sodium pyruvate and 0.1 mmol l−1 non-essential amino acids for 16 h. After sodium butyrate (10 mM) induction for 8 h, the virus supernatant was collected and filtered through a 0.45-μm filter. The virus-containing supernatants were loaded by centrifugation (2,000g, 90 min at 32 °C) on 50 μg ml−1 retronectin (Takara)-coated non-tissue culture 6-well plates. For retroviral transduction, 3–5 × 106 cells were transduced per well by centrifugation at 600g for 30 min in the appropriate culture medium and maintained at 37 °C at 5% CO2 for 48 h. For lentiviral transduction, 3–5 × 106 cells per well were centrifuged at 600g for 30 min in the presence of lentiviral supernatant and maintained at 37 °C at 5% CO2. The lentiviral supernatant was replaced with fresh medium the next day.
Western blotting
PBS-washed cells were lysed in CelLytic buffer (Sigma-Aldrich) supplemented with 1% protease inhibitor cocktail (Roche Diagnostics), 1% phosphatase inhibitor cocktail (EMD Millipore) and 1 mM PMSF on ice. A total of 10 μg of cell lysates was separated on mini-precast gels (Bio-Rad) and transferred onto nitrocellulose membranes (Bio-Rad). Membranes were probed with the appropriate primary antibodies. Membranes were then incubated with alkaline-phosphatase conjugated secondary antibodies (Invitrogen) and chemiluminescent substrate (Invitrogen) and were further detected by film exposure, UVP BioSpectrum 810 Imaging System (Thermo Fisher Scientific) or by using the ChemiDoc MP Imaging System (BioRad). Antibodies used in this study are provided in Supplementary Table 10, and were used at 1:750 to 1:1,000 dilution in blocking buffer.
Flow cytometry
PBS-washed cells were blocked with Fc blocker for 10 min on ice and then stained with the appropriate antibodies listed in Supplementary Table 10, or with isotype control, for 25 min on ice. Cells were then washed and resuspended in chilled PBS containing 0.75 μg ml−1 of DAPI to exclude dead cells. Acquisition was performed by LSRFortessa flow cytometer (BD Biosciences) with BD FACSDIVA software. The fluorescence-based cell sorting was performed by FACSAria II (BD Biosciences) with BD FACSDIVA software. FACS data were analysed with FlowJo software (FlowJo). For apoptosis analyses, annexin V and 7-AAD (BD Biosciences) were used. For cell cycle analysis, the BrdU flow cytometry kit (BD Biosciences) was used according to manufacturer’s instructions. For intracellular staining of cytoplasmic proteins, cells were first stained for cell surface antigens and subsequently fixed in fixation and permeabilization solution (BD Biosciences) containing 4% paraformaldehyde and the detergent saponin. Cells were then washed and resuspended in Perm/Wash Buffer (BD Biosciences) and stained with the appropriate antibodies. For statistical quantification, data were plotted with GraphPad Prism 7 or SigmaPlot. FACS antibodies, listed in Supplementary Table 10, were pre-diluted 1:5–1:10. Two μl of diluted antibody was added to 1–2 million cells per 50 μl in PBS for final dilution of 1:50 for human cells or 1:200 for mouse cells.
Pharmacological inhibitors and reagents
Imatinib was purchased from LC Laboratories. Stock solutions were prepared in sterile water at 10 mmol l−1 and administered at 10 μmol l−1. Dasatinib was purchased from SelleckChem. Stock solutions were prepared in sterile DMSO at 25 μmol l−1 and administered at 25 nmol l−1 (Supplementary Table 11).
Analysis of chromatin immunoprecipitation with sequencing data
IKZF1 chromatin immunoprecipitation with sequencing (ChIP-seq) was performed as previously described34 with a patient-derived human B-ALL xenograft cell line (LAX2), which expresses wild-type full-length IKZF1 and no detectable level of the dominant-negative IKZF1 isoform. ChIP-seq tracks (GSE58825) for the IKZF1 antibody in LAX2 on IFITM3 gene promoter regions are shown in the bottom of Extended Data Fig. 1m. The y axis represents the normalized number of reads per million reads for peak summit for each track. ChIP-seq tracks (GSE86897) for the enrichment of RNAPII and H3K4me3 at the Ifitm3 locus in pre-B cells from Ikzf1exon5fl/fl mice upon Cre-mediated deletion of Ikzf1 are shown in the top of Extended Data Fig. 1m. The ChIP-seq peaks were called by the MACS peak-caller by comparing read density in the ChIP experiment relative to the input chromatin control reads, and are shown as bars under each wiggle track. Gene models are shown in UCSC genome browser hg19. Integrative Genomics Viewer (IGV) was used to visualize ChIP-seq tracks.
Inducible expression of IKZF1
Human Philadelphia chromosome-positive (Ph+) pre-B ALL cells (BV173) carrying deletions of IKZF1 were transduced with pRetroX-Tet3G-Neo. Neomycin-resistant cells were transduced with pRetroX-TRE3G-wild-type IKZF1-Puro. Expression of wild-type IKZF1 in puromycin-resistant cells was initiated by treatment with 1 μg ml−1 of doxycycline (TetOn).
Inducible reconstitution of CD19
For inducible CD19 reconstitution, mouse CD19 was fused to the ligand-binding domain of a mutant oestrogen receptor (ERT2) at the C terminus of CD19. BCR–ABL1-transformed mouse B-lineage ALL cells were retrovirally transduced with MSCV CD19-ERT2-Puro. Puromycin-resistant cells were treated with 4-hydroxytamoxifen (4-OHT) (Sigma-Aldrich) or vehicle control to induce reconstitution of CD19.
Cell viability assay
One hundred thousand BCR–ABL1-transformed mouse B-lineage ALL cells were seeded in a volume of 100 μl in medium in 1 well of a 96-well plate (BD Biosciences). Imatinib (LC Laboratories) was added at the indicated concentration in a total volume of 150 μl. After culturing for 3 days, 15 μl of resazurin (R&D) was added into each well and incubated for 4 h at 37 °C. Medium without cells was used as blank. The fluorescence was read at 535 nm and the reference wavelength was 590 nm. Relative viability was calculated using baseline values of vehicle-treated cells as a reference.
Colony-forming assay
The methylcellulose colony-forming assays were performed with 10,000 cells. Cells were resuspended in mouse MethoCult medium (without cytokines for BCR-ABL1-transformed cells; with IL-7 for NRASG12D-expressing cells) and cultured on 3-cm-diameter dishes, with an extra water supply dish to prevent evaporation. Colonies were imaged and counted after seven days using GelCount (Oxford Optronix), Olympus IX71 microscope and Q-Capture pro 7.
In vitro IFITM3–HA crosslinking
Eight million patient-derived B-ALL cells per sample were resuspended into complete medium and treated with either 2.5 μg ml−1 polyclonal anti-HA (Abcam) or isotype control for the indicated times. F(ab) fragments for the anti-HA antibody were generated by using the F(ab) preparation kit following the manufacturer’s instructions (Thermo Fisher Scientific). F(ab) fragmentation was confirmed by Criterion TGX Stain-Free Precast Gels (Bio-Rad).
Measurement of intracellular calcium mobilization
In Extended Data Fig. 2f, 1 million fresh splenocytes were incubated with the 4 μM Rhod-2-AM Ca2+-binding dye (Thermo Fisher Scientific) for 15 min at room temperature in dark. Cells were then resuspended in PBS and maintained at 37 °C, and Ca2+ response was induced by adding 10 μg ml−1 of polyclonal anti-IgM (Southern Biotech) at 50 s after acquisition of background fluorescence. Intracellular Ca2+ mobilization in response to crosslinked IgM was measured up to 300 s by flow cytometry. In Extended Data Fig. 4c, 1 × 106 viable Jurkat cells were incubated with the 4 μM Fluo4-AM Ca2+-binding dye (Thermo Fisher Scientific) for 15 min at room temperature in dark. Cells were then resuspended in PBS and maintained at 37 °C, and Ca2+ response was induced by adding 10 μg ml−1 of monoclonal (OKT3) purified NA/LE anti-human CD3 (Biolegend) at 50 s after acquisition of background fluorescence. Intracellular Ca2+ mobilization in response to crosslinked IFITM3 was measured up to 300 s by flow cytometry. The same procedure was performed for Extended Data Fig. 6f; 1 × 106 viable Jeko1 cells were incubated with the 4 μM Fluo4-AM (Thermo Fisher Scientific). Ca2+ response was induced by adding 10 μg ml−1 of polyclonal F(ab′)2 anti-human μ chain (Jackson Immunoresearch).
Homotypic aggregation assay
Ifitm3−/− B-ALL cells expressing C-terminal HA-tagged wild-type IFITM3, IFITM3 mutant vector (Y20E) or an empty vector were incubated with 2.5 μg ml−1 of monoclonal (1D3) purified LEAF anti-mouse CD19 (Biolegend) or 2.5 μg ml−1 polyclonal anti-HA (Abcam) or isotype controls at 37 °C at 5% CO2. After culturing for 24 h, homotypic aggregation was visualized using light microscopy.
In vivo oncogenic priming assay with Mb1-cre;LSL-Bcr +/BCR-ABL1 B cell precursors
To generate a model for pre-leukaemic B cell precursors expressing BCR–ABL1, BCR–ABL1 knock-in mice were crossed with Mb1–Cre strain (Mb1-cre × LSL-Bcr+/BCR-ABL1) for excision of a stop cassette in early pre-B cells. Bone marrow cells collected from Mb1-cre × LSL-Bcr+/BCR-ABL1 mice, cultured in the presence of 10 ng ml−1 recombinant mouse IL-7 (Peprotech). IL-7 dependent pre-leukaemic B cells were labelled with retroviral firefly luciferase and selected by blasticidin for 7 days. After selection, cells were further transduced with MSCV-IFITM3-HA-IRES-Puro, MSCV-IFITM3Y20E-HA-IRES-Puro or empty vector and selected by puromycin for 3 days.Viable cells (1 × 107) were injected via the tail vein into sublethally irradiated (200 cGy) NSG recipient mice. In vivo expansion and leukaemia burden were monitored by luciferase bioimaging (IVIS 100 bioluminescence and optical imaging system; Xenogen) at the indicated time points. In brief, d-luciferin (Promega) dissolved in PBS was injected intraperitoneally at a dose of 2.5 mg per mouse 15 min before measuring luminescence. All mice were anaesthetized by 5% isoflurane and continued during detection of light emission with 2% isoflurane introduced through a nose cone. A mouse was euthanized when they showed signs of leukaemia burden such as a hunched back, weight loss and inability to move. Kaplan–Meier survival analysis was performed using GraphPad Prism 7 (GraphPad Software) to compare overall survival. Mantel–Cox log-rank test was used as statistical analysis using GraphPad Prism 7 (Supplementary Table 4).
Cell adhesion assay
Ifitm3+/+ or Ifitm3−/− BCR-ABL1 B-ALL cells were transduced with MSCV-IK6-IRES-GFP or MSCV-IRES-GFP as negative control. GFP+ cells (1 × 105) sorted by FACS were cultured on 1.5 × 105 OP9 stroma on a 6-well plate with IMDM-GlutaMAX containing 20% FBS, 50 μM 2-mercaptoethanol, 100 IU ml−1 penicillin and 100 μg ml−1 streptomycin. For calculation of the ratio of adherent cells to nonadherent cells, nonadherent cells were collected and counted with the trypan blue exclusion method using the Countess II FL Automated Cell Counter (Life Technologies). After washing the plate with PBS twice, adherent cells were detached with trypsinization and GFP+ B-ALL cells and were counted with the trypan blue exclusion method using the Countess II FL Automated Cell Counter (Life Technologies) and ratios were calculated.
Adoptive transfer of purified B cells into μMT mice
For the adoptive transfer of B cells, splenocytes from 7- to 10-week-old Ifitm3+/+ or Ifitm3−/− mice were negatively selected by MojoSort Mouse Pan B Cell Isolation Kit II (Miltenyi Biotec) using immunomagnetic beads against CD3, CD4, CD8a, CD11c, CD49b, Ly6G/Ly6C (GR1) and TER119. Ten million flow-sorted splenic B cells were intravenously injected into μMT mice (B6.129S2-Ighmtm1Cgn/J) that lack the Cμ exon and mature B cell development as a result of defective surface IgM expression. Reconstitution of donor B cells was determined by flow cytometry 20 days after injection. Recipient μMT mice were immunized with the hapten NP coupled to a carrier protein (KLH) on day 0 and day 7, and spleens were collected on day 12.
Immunization and immunohistology
Ifitm3+/+ and Ifitm3−/− littermate mice were immunized with 0.5 mg NP–KLH (Biosearch Technologies) in alum (Sigma-Aldrich) intraperitoneally. Seven days later, mice were immunized a second time for five days. Spleens were isolated at day 12 after immunization. The spleen was embedded in optimum cutting temperature compound and 5-μm-thick cryosections were used for staining. Sections were fixed with acetone for 10 min, and nonspecific antigens were blocked in DPBS containing 2% FBS for 15 min. Sections were stained with 1:200 diluted polyclonal (RA3-6B2) anti-CD45R (B220, BD Biosciences), monoclonal anti-mouse CD3 (17A2, BioLegend) antibodies and biotinylated peanut agglutinin (B-1075, Vector Laboratories) for 45 min. Sections were washed and further stained with Alexa Fluor 647 streptavidin (BioLegend) antibody for 45 min. All antibodies were diluted 1:100 to their original concentration in blocking buffer. After washing with blocking buffer, sections were mounted in ProLong Diamond Antifade Mountant (ThermoFisher Scientific). Images were acquired on a ZEISS LSL 880 confocal microscope and analysed on ZEN 2.3 (Zeiss) software.
ELISA measurements
For determination of the concentrations of immunoglobulin isotypes in sera, ELISAs were carried out according to the manufacturer’s protocol (Ig Isotyping Mouse Instant ELISA Kit for IgG1, IgG2b and IgM) (Supplementary Table 11). NP-specific antibodies were measured by ELISA using 10 μg/ml of NP(24)–BSA (Biosearch Technologies) as the coating reagent. NP-specific IgM and IgG1 was detected using goat anti–mouse IgM and IgG1 Fc-specific antibodies conjugated to horseradish peroxidase and developed with tetramethylbenzidine (Sigma). Optical densities were determined by using an ELISA reader at 450 nm (SpectraMax M3, Molecular Devices).
RNA-sequencing analysis
Total mRNA from Ifitm3+/+ and Ifitm3−/− BCR-ABL1 or NRASG12D B-ALL cells was extracted using RNeasy Kit (Qiagen) according to the manufacturer’s instructions. Sequencing was performed on an Illumina Hiseq 2500 (Illumina) instrument using the TruSeq SR Cluster Kit v.4-cBot-HS (Illumina) with v.4 chemistry. Quality control of RNA-sequencing reads was performed using FastQC. For analysis, raw sequence reads were mapped to the mouse genome (mm10) using STAR v.2.5.335, and the frequency of genes was counted using featureCounts v.1.5.136. The raw counts were then normalized using the trimmed mean of M values method and compared using Bioconductor package ‘edgeR’37. Reads per kilobase per million (RPKM) mapped reads were also calculated from the raw counts. For differential expression analysis, transcripts were quantified using Salmon v.1.1.0 against gencode GRCm38 v.M24 transcript annotations; normalization and statistical analysis were done in R using DESeq2 v.1.28.1. Differentially expressed genes were identified if RPKM ≥ 1 in at least one sample, fold change ≥ 2 and P ≤ 0.05 (Extended Data Fig. 3b). RPKM data were later used in the gene set enrichment analysis (GSEA). GSEA was performed using the DOSE package in R38; genes were ranked by log2-transformed fold change; and gene sets were obtained from MSigDB or from internal data, as indicated.
CRISPR-mediated gene deletion
All lentiviral constructs expressing Cas9 nuclease and guide RNA were purchased from Transomic Technologies. For gene deletion, cells were transduced with pTOL-hCMV-Tet3G-Hygromycin. Hygromycin-resistant cells were subsequently transduced with pCLIP-Tre3g-hCMV-Cas9-P2A-zsGreen. Expression of Cas9–P2A–zsGreen was induced at 1 μg ml−1 of doxycycline for 16 h, and zsGreen+ cells were sorted by flow cytometry. Cells were washed out to remove doxycycline to turn off Cas9 expression (TetOff) and then subsequently transduced with pCLIP-gRNA-hCMV-RFP-gRNA. Sorted RFP+ cells were subjected to further experiments. Gene deletion was initiated by treatment with 1 μg ml−1 of doxycycline (TetOn). Non-targeting guide RNA was used as control.
Gene deletion by non-viral genome targeting
Chemically synthesized CRISPR RNAs (160 μM) and trans-activating CRISPR RNAs (160 μM) were mixed 1:1 by volume and annealed by incubation at 37 °C for 30 min. Recombinantly produced Cas9 (40 μM) was then mixed 1:1 by volume with gRNA to produce RNA ribonucleoprotein (RNP) complexes. RNPs were freshly complexed before electroporation. Electroporation was performed by using pulse code EH-115 on a Lonza 4D 96-well electroporation system. Predesigned Alt-R CRISPR–Cas9 guide RNAs were purchased from IDT. Non-targeting control guide RNAs were purchased from IDT.
Co-immunoprecipitation
Co-immunoprecipitation was performed with the Pierce Crosslink Magnetic IP/Co-IP kit according to manufacturer’s instructions (Thermo Fisher Scientific). In brief, patient-derived Ph+ ALL cells (PDX2) were transduced with MSCV Flag-IRES-Puro or MSCV Flag-IFITM3-IRES-Puro and selected in puromycin for 3 days. Viable cells (5 × 107) were collected and washed by PBS before lysis using IP lysis/Wash buffer (Thermo Fisher Scientific). Then, 5 μg of anti-Flag antibody M2 (F1804, Sigma-Aldrich) per sample was coupled to protein A/G magnetic beads and covalently crosslinked with 20 μM disuccinimidyl suberate. The antibody crosslinked beads were incubated with cell lysate, washed to remove non-bound material and eluted in a low-pH elution buffer that dissociates bound antigen from the antibody crosslinked beads. The enriched antigen in low pH was immediately neutralized and subjected to western blotting. Jeko1 cells electroporated with either non-targeting RNP or IFITM3-targeting RNP complex were stimulated with 10 μg ml−1 F(ab′)2 fragment goat anti-human μ chain (Jackson Immunoresearch) for the indicated time points at 37 °C and 5% CO2. Cells were immediately washed with chilled PBS and subjected to co-immunoprecipitation with anti-CD19 antibody (no. 90176, CST).
PLA
For PLA of pre-BCR or BCR–IFITM3, cells were incubated with 10 μg ml−1 F(ab′)2 fragment goat anti-human μ chain (Jackson Immunoresearch) for 5 min at 37 °C and 5% CO2. Cells were immediately washed with chilled PBS and subsequently fixed in fixation buffer (Biolegend) containing 4% paraformaldehyde for 25 min on ice and then washed with chilled PBS. For cellular membrane staining, cells were labelled with 5 μg ml−1 WGA conjugated to Alexa Fluor 488 (Thermo Fisher Scientific) for 5 min at room temperature. Cells were then permeabilized in Perm/Wash Buffer (BD Biosciences) and then blocked in Duolink Blocking buffer for 30 min at room temperature. Cells were incubated with 1:150 diluted primary antibodies listed in Supplementary Table 10 overnight at 4 °C. For late endosome staining, cells were incubated with anti-human LAMP1 conjugated to Alexa Fluor 488 (R&D Systems) with the primary antibodies. Cells were washed and settled on Cell-Tak (Corning)-coated Shandon Single Cytoslide by cytospin at 400g for 5 min. PLA reactions were carried out according to the manufacturer’s protocol (Duolink, Sigma). In brief, primary antibodies were coupled with Duolink in situ PLA PLUS or MINUS probes (Sigma-Aldrich), and then probes were visualized with Duolink Detection Reagent Red (Sigma-Aldrich). Cells were mounted with Duolink in situ mounting medium with DAPI (Sigma-Aldrich). Microscopy images were acquired using an Olympus IX3-55 and analysed by CellSens imaging software (Olympus) and ImageJ. For quantification of PLA signals, one dot was defined as pixel size of 5 × 5 by BlobFinder software. Statistical significance was calculated using unpaired Student’s t-test with Excel and plotted with GraphPad Prism 7.
Phosphoproteomic analysis
Patient-derived B-ALL (PDX2) cells transduced with HA-tagged IFITM3(Y20E) or empty vector control were incubated with 2.5 μg ml−1 of polyclonal anti-HA (Abcam) for 5 min at 37 °C at 5% CO2. Cellular extracts were prepared in urea lysis buffer, sonicated, centrifuged, reduced with DTT and alkylated with iodoacetamide. Fifteen mg total protein for each sample was digested with trypsin, and 500 μg total protein for each sample was digested with LysC and trypsin mix for IMAC analysis. Samples were purified over C18 columns and dried in a lyophilizer. Dried samples were resuspended and enriched with Fe-IMAC beads, purified over C18 STAGE tips (Rappsilber). Replicate injections of each sample were run nonsequentially on the instrument. Peptides were eluted using a 150-min (IMAC) linear gradient of acetonitrile in 0.125% formic acid delivered at 280 nl/min. Tandem mass spectra (MS/MS) were collected in a data-dependent manner with a Thermo Orbitrap Fusion Lumos Tribrid mass spectrometer using a top-twenty MS/MS method, a dynamic repeat count of one and a repeat duration of 30 s. Real-time recalibration of mass error was performed using lock mass (Olsen) with a singly charged polysiloxane ion of m/z = 371.101237. MS/MS spectra were evaluated using SEQUEST and the Core platform from Harvard University (by J. K. Eng, E. L. Huttlin and J. Villen). Files were searched against the SwissProt Homo sapiens FASTA database. A mass accuracy of ±5 ppm was used for precursor ions and 0.02 Da for product ions. Enzyme specificity was limited to trypsin, with at least one tryptic (K- or R-containing) terminus required per peptide and up to four miscleavages allowed. Cysteine carboxamidomethylation was specified as a static modification, and oxidation of methionine and phosphorylation on serine, threonine and tyrosine residues were allowed as variable modifications. Reverse decoy databases were included for all searches to estimate false discovery rates, and filtered using a 1% false-discovery rate in the linear discriminant module of Core. Peptides were also manually filtered using a ±5ppm mass error range and presence of a phosphorylated residue. All quantitative results were generated using Skyline (MacLean) to extract the integrated peak area of the corresponding peptide assignments. Accuracy of quantitative data was ensured by manual review in Skyline or in the ion chromatogram files.
Bio-ID proteomics
Patient-derived PDX2 B-ALL cells and Jeko1 MCL cells were transduced with Flag-tagged IFITM3(Y20E) or IFITM3(K83A/K104A) mutant constructs with N-terminal Turbo-ID BirA (engineered biotin ligase) and selected by puromycin for three days. Turbo-ID BirA-expressing construct tagged with Flag was used as negative control. To induce biotinylation of proteins proximal to IFITM3, cells were treated with 50 μmol l−1 biotin and with anti-HA antibody for simultaneous induction of IFITM3 crosslinking for 10 min. Cells were washed three times with chilled PBS and lysed in IP/WASH buffer (Thermo Fisher Scientific) with 1× HALT protease inhibitor (Thermo Fisher Scientific). The lysates were incubated by streptavidin C1 MyOne Dynabeads (Invitrogen) for 16 h at 4 °C. Unbound proteins were washed 3 times by 2% SDS–PBS, 3 times by PBS and 3 times with pure water. The eluted proteins were gel-purified, followed by in-gel digestion and subjected to mass spectrometry. For liquid chromatography (LC)–MS/MS analysis, peptides were analysed using a Dionex UltiMate 3000 Rapid Separation LC system and a Orbitrap mass spectrometer (ThermoFisher Scientific). Six-μl peptide samples were loaded onto the trap column, which was 150 μm × 3 cm and in-house-packed with 3-μm C18 beads. The analytical column was a 75 μm × 10.5-cm PicoChip column packed with 3-μm C18 beads (New Objectives). The flow rate was kept at 300 nl/min. Solvent A was 0.1% formic acid (FA) in water and solvent B was 0.1% FA in ACN. The peptide was separated on a 120-min analytical gradient from 5% ACN and 0.1% FA to 40% ACN and 0.1% FA. The mass spectrometer was operated in data-dependent mode. The source voltage was 2.40 kV and the capillary temperature was 275 °C. MS1 scans were acquired from 400–2,000 m/z at 60,000 resolving power and automatic gain control (AGC) set to 1 × 106. The 15 most abundant precursor ions in each MS1 scan were selected for fragmentation. Precursors were selected with an isolation width of 1 Da and fragmented by collision-induced dissociation at 35% normalized collision energy in the ion trap; previously selected ions were dynamically excluded from reselection for 60 s. The MS2 AGC was set to 3 × 105. For data analysis, proteins were identified from the mass spectrometry raw files using Mascot search engine (Matrix science). MS/MS spectra were searched against the SwissProt human database. All searches included carbamidomethyl cysteine as a fixed modification and oxidized Met, deamidated Asn and Gln, and acetylated N terminus as variable modifications. Three missed tryptic cleavages were allowed. The MS1 precursor mass tolerance was set to 10 ppm and the MS2 tolerance was set to 0.6 Da. A 1% false discovery rate cut-off was applied at the peptide level. Only proteins with a minimum of two peptides above the cut-off were considered for further study. For comparison to empty vector control, background peptide abundances for missing values were imputed from a Gaussian distribution centred around the minimum observed abundance using the MinProb method from MSnbase package in R.
Cell-surface proteome analyses
Cell-surface proteins were labelled with biotin using the N-linked glycosylation-site biotin labelling method39. In brief, 40 million Ifitm3+/+ or Ifitm3−/− B-ALL cells were washed twice and resuspended in 1 ml of ice-cold PBS, and treated with 1.6 mM sodium metaperiodate (VWR) at 4 °C for 20 min to oxidize the vicinal diols of sugar residues linked to surface proteins. The cells were then washed twice in PBS to remove excess sodium metaperiodate. Cells were resuspended in 1ml of ice-cold PBS and treated with 1 mM biocytin hydrazide (Biotium) and 10 mM aniline (Sigma-Aldrich) at 4 °C for 90 min with gentle mixing to biotinylate free aldehydes exposed on the sugar residues. After labelling, cells were washed 3 times with ice-cold PBS to removed excess biotin, frozen in liquid nitrogen and stored at −80 °C until further processing for mass spectrometry. All experiments were performed in biological triplicates with replicates collected from consecutive passages. Frozen cell pellets were thawed on ice in 1 ml of RIPA buffer (Millipore) with the addition of 1× HALT protease inhibitors (Pierce). After incubation on ice for 10 min, cells were disrupted by sonication and the lysates were clarified by centrifugation at 17,000 rcf at 4 °C for 10 min. Clarified lysate was mixed with 500 μl of neutravidin agarose resin (Thermo Fisher Scientific) and incubated at 4 °C for 2 h with end-over-end mixing. Neutravidin beads with captured biotinylated surface proteins were washed extensively by gravity flow to remove unbound proteins using 50 ml of 1× RIPA + 1 mM EDTA, followed by 50 ml of PBS + 1 M NaCl and finally 50 ml of 50 mM ABC + 2 M urea buffer. Washed beads were resuspended in digestion buffer (50 mM Tris pH 8.5, 10 mM TCEP, 20 mM 2-iodoacetamide and 1.6 M Urea) with 10 μg of added trypsin protease (Pierce, 90057) to perform simultaneous disulfide reduction, alkylation and on-bead peptide digestion at room temperature overnight (16–20 h). After digestion, the pH was dropped to about 2 with neat trifluoroacetic acid (Sigma-Aldrich) and the peptide mixture was desalted using a SOLA-HRP column (Thermo Fisher Scientific) on a vacuum manifold. Desalted peptides were eluted with 50% acetonitrile (Sigma-Aldrich) and 50% water with 0.1% TFA and dried down completely in a speedvac. Dried peptides were resuspended in LC–MS grade water (Fisher) with 2% ACN and 0.1% FA. Peptide concentration was measured using 280 nm absorbance on a Nanodrop, and the peptide concentration was adjusted to 0.2 μg μl−1 for mass spectrometry runs.
LC–MS and data analysis for cell-surface proteome
For each replicate, 1 μg of peptide was injected onto a Dionex Ultimate 3000 NanoRSLC instrument with a 15-cm Acclaim PEPMAP C18 (Thermo Fisher Scientific, 164534) reverse-phase column. The samples were separated on a 3.5-h nonlinear gradient using a mixture of buffer A (0.1% FA) and B (80% ACN/0.1% FA), from 2.4% ACN to 32% ACN. Eluted peptides were analysed with a Thermo Q-Exactive Plus mass spectrometer. The mass spectrometry survey scan was performed over a mass range of 350–1500 m/z with a resolution of 70,000, with a maximum injection time of 100 ms. We performed a data-dependent MS2 acquisition at a resolution of 17,500, AGC of 5 × 104, and injection time of 150 ms. The 15 most intense precursor ions were fragmented in the higher-energy collisional dissociation at a normalized collision energy of 27. Dynamic exclusion was set to 20 s to avoid over-sampling of highly abundant species. The raw spectral data files are available at the ProteomeXchange PRIDE repository (accession number PXD014691). Raw spectral data were analysed using MaxQuant v.1.5.1.240 to identify and quantify peptide abundance and searched against the human Swiss-Prot annotated human proteome from Uniprot (downloaded with 20,303 entries). The ‘match-between-runs’ option was selected to increase peptide identifications and the ‘fast LFQ’ option was selected to calculate label-free quantification values of identified proteins. All other settings were left at the default MaxQuant values. The MaxQuant output data were analysed using Perseus41 and the R program (version 3.4.0). Proteins annotated as ‘reverse’, ‘only identified by site’ and ‘potential contaminant’ were filtered out, as were proteins that were quantified in less than two out of three biological replicates in at least one experimental group. Missing values were imputed on the basis of the normal distribution of the dataset as implemented by Perseus. Volcano plots were generated using output from a two-sample t-test comparing the log2-transformed label-free quantification protein abundance values from different cell lines with a false-discovery rate set to 0.01. Validation was performed by flow cytometry (Supplementary Table 1).
Cholesterol and lipid raft measurement
For the depletion of cholesterol, cells were preincubated with 5 mM of methy-β-cyclodextrin for 30 min at 37 °C before filipin staining. For ganglioside GM1 staining, cells were labelled with cholera toxin B using Vybrant lipid raft labelling kit (Molecular Probes), according to the manufacturer’s protocol. In brief, cells were labelled with cholera toxin unit B conjugated with Alexa Fluor 594 on ice for 15 min, washed twice with PBS. Cholera-toxin-B-labelled lipid rafts were crosslinked with anti-cholera-toxin-B antibody on ice for 15 min, washed twice with PBS and analysed by flow cytometry.
In vivo transplantation of leukaemia cells
Mouse pre-B ALL cells transformed by BCR–ABL1 or NRAS(G12D) were injected into sublethally irradiated (200 cGy) NOD-scid IL2Rgnull (NSG) mice via the tail vein. Eight- to ten-week-old female NSG mice were randomly allocated before injection. A mouse was euthanized when they showed signs of leukaemia burden, such as a hunched back, weight loss and inability to move, and then the bone marrow and/or spleen were collected to test leukaemia infiltration by flow cytometry. Kaplan–Meier survival analysis was performed using GraphPad Prism 7 (GraphPad Software) to compare overall survival. The Mantel–Cox log-rank test was used as statistical analysis using GraphPad Prism 7 (Supplementary Table 3). The minimal number of mice in each group was calculated by using the ‘cpower’ function in Hmisc package of R. No blinding was used. All mouse experiments were subject to institutional approval by the Beckman Research Institute of City of Hope Animal Care and Use Committee.
PIP3 quantification
For Fig. 4, 60 million viable cells were resuspended in chilled 0.5 M trichloroacetic acid (TCA) in a total volume of 1 ml and incubated on ice for 5 min. Cells were centrifuged at 3,000 rpm for 7 min at 4 °C and resuspended in 5% TCA and 1 mM EDTA in a total volume of 1 ml. After vortexing for 30 s, cells were washed again with 5% TCA and 1 mM EDTA. To extract neutral lipids, cells were resuspended in 1 ml of MeOH:CHCl3 (2:1) and vortexed for 10 min at room temperature and centrifuged at 3,000 rpm for 5 min. After 1 more extraction of neutral lipids, the acidic lipids were extracted by adding 750 μl MeOH:CHCl3:12 M HCl (80:40:1) with vigorous vortexing for 25 min at room temperature. After centrifugation at 3,000 rpm for 5 min, supernatant was transferred to a 2-ml centrifuge tube and mixed with 250 μl of CHCl3 and 450 μl of 0.1 M HCl, vortexed for 30 s, and centrifuged at 3,000 rpm for 5 min to separate organic and aqueous phases. Five hundred μl of the lower organic phases was collected into a clean 1.5-ml vial and dried in a vacuum dryer for 1 h. Dried lipid was stored at −20 °C. On the day of assay, lipid samples were reconstituted with 200 μl of PBS containing 0.25% protein stabilizer; 20 μl was used for PI(4,5)P2 measurement and the rest (180 μl) was used for PI(3,4,5)P3 measurement using an ELISA kit (Echelon Biosciences) according to the manufacturer’s instructions.
Exogenous delivery of PIP3
Phosphatidylinositol 3,4,5-trisphosphate diC16 (Echelon Biosciences) was freshly reconstituted with PBS at 2 mM. Unlabelled shuttle PIP carrier 2 (histone H1) was freshly reconstituted with water at 2 mM. PIP3 in a volume of 75 μl was mixed with 25 μl of PIP carrier and incubated at room temperature for 10 min. One hundred μl of PIP3–carrier complex was added into Ifitm3+/+ or Ifitm3−/− B-ALL cells in a total volume of 5 ml and incubated for 24 h. PIP3-loaded Ifitm3+/+ or Ifitm3−/− B-ALL cells were subjected to colony-forming assay, as shown in Extended Data Fig. 8.
Lipid-binding assay
To assess the direct binding of IFITM3 to lipids, a lipid-binding assay was performed using Membrane Lipid Strips (Echelon Biosciences) according to the manufacturer’s instructions. In brief, membranes were blocked in 5% fatty acid-free BSA (Sigma-Aldrich) in TBST (50 mM Tris-HCl,150 mM NaCl, and 0.1% Tween 20) for 1 h at room temperature in the dark followed by overnight incubation with 0.5 μg/ml of recombinant proteins in blocking buffer at 4 °C with gentle agitation. After washing membranes 3 times for 30 min in TBST, membranes were incubated for 1 h with anti-GST tag polyclonal Ab (Thermo Fisher Scientific) or anti-biotin monoclonal antibody (Cell Signaling Technology) antibodies, listed in Supplementary Table 10. Membranes were incubated with alkaline-phosphatase-conjugated secondary antibodies (Invitrogen) and chemiluminescent substrate (Invitrogen) and were further detected by UVP BioSpectrum 810 Imaging System (Thermo Fisher Scientific). Recombinant GST-tagged protein was purchased from Sigma-Aldrich. Recombinant human GST–IFITM3 protein was purchased from Abnova. Recombinant human IFITM3 fragments listed in Supplementary Table 5 were synthesized at LifeTein.
Preferential PIP3 accumulation and binding to IFITM3 using multi-scale molecular dynamics simulations
The IFITM3 protein structure from residues 58 to 128 was modelled by coarse-grained molecular dynamics simulation method in GROMACS, in explicit cell membrane mimic bilayer. There is no structure was available for IFITM3 or for any close homologue. We calculated the hydrophobicity for each position using the hydrophobic index of amino acids42 and averaged hydrophobicity over a sliding window of seven amino acids. Previous structural studies on IFITM3 structure43 have shown there is only one transmembrane domain 2 that is inserted into the membrane. On the basis of this evidence, we used the topology shown in Supplementary Fig. 9 as the starting structure. TM1 was generated as an α-helix structure, and the CRAC motifs and linker region were built as fully extended chain. The linker region was relaxed using 5,000 steps of coarse-grained simulation using GROMACS program44 with Martini force field45 to remove stress in the system. We then added the TM2 region as a helix and inserted TM2 into a cell-membrane-mimicking bilayer46. The composition and ratio of lipids for the mixed bilayer is POPC:DOPC:POPE:DOPE:CHOL:Sph:GM1 = 0.2:0.2:0.05:0.05:0.25:0.15:0.1 for the outer leaflet of the bilayer, and 0.05:0.05:0.20:0.2:0.08:0.08:0.25:0.03:0.03:0.03 of POPC: DOPC: POPE: DOPE: POPS: DOPS: CHOL: PIP1:PIP2:PIP3 for the inner leaflet. We generated three possible starting setup for the mixed lipid bilayer with water in the upper and inner regions with CHARMM-GUI47,48. The charges in the system were neutralized with 0.15 M NaCl. We used nine replicates of the protein IFITM3 structure in the simulation box (Supplementary Fig. 9a). The initial configuration was minimized using steepest decent method for 5,000 steps, and then equilibrated in NPT ensemble for 5 ns, at 303-K temperature and 1 bar pressure. The temperature was controlled by Berendsen thermostat49 with coupling constant of 1 ps, and the pressure was controlled by Berendsen barostat49 in semi-isotropic type with a coupling constant of 5 ps and a compressibility of 3 × 10−4 bar-1. The reaction field coulomb50 was applied to describe electrostatics, with cut-off at 1.1 nm. The cut-off for van der Waals interactions used was also 1.1 nm. We used the Leap-frog integrator, and a 20-fs time step for integration. All systems were solvated in coarse-grained water model adopted in the Martini forcefield water, neutralized with 0.15 M NaCl. Ten μs of production simulation was performed for each of the three starting conformations of the lipid bilayer. The final 1 μs of the trajectory from each of the 3 setups was used to analyse the folded structure of the CRAC1–basic patch–CRAC2 region of the IFITM3 model. We clustered the conformations of IFITM3 by the root mean square deviation cut-off of 0.3 nm in the backbone coarse-grained particles in the coarse-grained simulations. We analysed the binding patterns of PIP2 and PIP3 in this most populated conformational cluster. We observed that both PIP2 and PIP3 compete for the ‘basic patch’ of residues shown in the amino acid sequence in Extended Data Fig. 9. We extracted three snapshots from this conformational cluster that represent the most diverse patterns in PIP2 and PIP3 binding.
Details of the all-atom molecular dynamics simulations for phospholipid binding
To better understand the detailed binding conformation of PIP2 and PIP3 to IFITM3 and to calculate their binding energies to IFITM3, we converted the three snapshots extracted from the coarse-grained simulations to an all-atom system using Martini tools51. The three chosen snapshots were cut into 9 × 9-nm2 box centred at the IFITM3 unit from the coarse-grained simulations to preserve the local optimized lipid environment. The lipid and protein were converted to an all-atom model, and were resolvated into a 9 × 9 × 9-nm3 simulation box, neutralized by 0.15 M NaCl. Extended Data Figure 9 shows the detailed interaction between PIP2 or PIP3 with IFITM3 protein in an all-atom resolution base on these snapshots. The all-atom simulations were performed using GROMACS package44 and CHARMM36 force field52, with the TIP3 water model. The nonbond interactions were calculated with a cut-off of 12 Å, and the particle-mesh Ewald method51 was applied to solve long-range van der Waals interactions. Each system was gradually heated to 310 K with random initial velocities sampled from a Boltzmann distribution. The heating process lasted for 1ns, with temperature controlled by a Nosé–Hoover thermostat53, followed by 30-ns equilibration in NPT ensemble with harmonic position restraints on protein heavy atoms. The pressure was maintained at 1 bar in semi-isotropic environment, controlled by the Parrinello–Rahman method. The restraint force was gradually reduced from 5 kcal/mol to 0 kcal/mol with a −1 kcal/mol step per 5-ns window. The final frame of equilibration was taken for a production run. The production run was performed twice for 50 ns each, with 2 different random initial velocities assigned to the equilibrated structure. An integration time step of 2 fs was used. The interaction energy between PIP3 with IFITM3 protein was calculated as the sum of electrostatic Coulombic energy and van der Waals potential energy averaged over the two 50-ns trajectories, totalling 100 ns.
Interaction of PIP3 with the two basic residue patches
The PIP3-binding and AKT-signalling activation assays showed that mutation of the K83/K104 patch to K83A/K104A had higher effect than mutating the R85/R87/K88 patch to R85A/R87A/K88A. To further validate the interaction strength difference between PIP3 and the two basic patches, we converted all PIP1 and PIP2 in the simulation box to PIP3. We further performed simulations on the IFITM3(K83A/K104A) and on IFITM3(R85A/R87A/K88A) in the PIP3-only simulation box. Starting from this structure, we performed equilibration of the system and 5 all-atom molecular dynamics production simulations (each 200-ns long), using the protocol for all-atom molecular dynamics simulations described in ‘Details of the all-atom molecular dynamics simulations for phospholipid binding’. We aggregated the final 100 ns of simulation trajectories from each of the 5 runs, which we added to 500 ns of molecular dynamics simulation trajectories for analysis. We calculated the interaction energy as the sum of electrostatic Coulombic and van der Waals potential energies between PIP3 and residues forming the basic patch only (K83/K104 or R85/R87/K88). The interaction energies were averaged over the 500 ns of aggregated molecular dynamics simulation trajectories for each system. We repeated these interaction energy calculations for the wild-type IFITM3 and for the mutant K83A/K104A and R85A/R87A/K88A constructs.
Contact-frequency heat-map calculations from molecular dynamics simulations
The important information that can be extracted from molecular dynamics simulations is the temporal frequency of PIP3 contacts with the two basic residue patches. The persistence of these interactions has an important role in the accumulation of PIP3 by IFITM3. We calculated the percentage of molecular dynamics snapshots that show contacts between each of the residues in the basic patch. This is termed the contact frequency. We also calculated the percentage of molecular dynamics snapshots that show simultaneous contacts made by PIP3 with two or more residues in the two basic residue patches. We generated a heat map using the frequencies calculated for the wild-type IFITM3 and in the alanine mutants.
Quantitative real-time PCR
Bone marrow from a healthy donor was stained and sorted as previously described54. Total RNA from cells was extracted using the RNA isolation kit from Macherey–Nagel. Complementary DNA was generated with the qScript cDNA SuperMix (Quanta Biosciences). Quantitative real-time PCR was performed with FAST SYBR Green Master Mix (Applied Biosystems) and the Vii7 real-time PCR system (Applied Biosystems) according to standard PCR conditions. COX6B was used as a reference gene.
Acoustic scattering measurements
Single-cell size-normalized acoustic scattering (SNACS) was measured using a previously established microfluidic method, which has been shown to be specifically sensitive to cell-surface stiffness55. Full measurement details can be found in a previous publication55. In brief, cells were flowed through a standing acoustic wave generated inside a vibrating suspended microchannel resonator (SMR). The SMR is a cantilever-based mass measurement tool56, which can also detect acoustic scattering from cells when the cells transit through the acoustic wave. The cantilever vibration frequency was monitored, and its shift was used to quantify the acoustic scattering from the cells as well as the buoyant mass of the cells. Before taking a set of measurements, the SMR was cleaned with 0.25% trypsin–EDTA for 20 min, followed by 5% bleach for 3-min and then a 5-min rinse with deionized H2O, to remove persistent biological debris. After cleaning, the SMR was passivated with 1 mg/ml PLL-g-PEG in H2O for 10 min at room temperature, followed by a 5-min rinse with normal cell culture medium. During the measurement, all the samples were loaded into the SMR through 0.005-inch-inner-diameter fluorinated ethylene propylene tubing. The fluid flow across the SMR was driven by three independent electronic pressure regulators and three solenoid valves. A consistent differential pressure was applied across the SMR to maintain constant shear and data rate for cell measurement. The data displayed in Extended Data Fig. 8a were obtained using a 350-μm-long cantilever with a 15 × 20-μm-sized channel inside of the cantilever and an approximately 200-ms transit time through the cantilever. All the regulators, valves and data acquisition were controlled by custom software coded in LabVIEW 2017 (National Instruments) as detailed in a previous publication55. A parallel volume measurement using Coulter Counter (Beckman Coulter) was carried out to quantify average cell volume, which was used together with the single-cell buoyant mass measurements to calculate SNACS for each cell, as previously reported55. All measurements were carried out in normal cell culture medium at room temperature within 10 min of taking cells out of the cell culture incubator. After a measurement of the untreated Jeko1 cells, a new patch of same cells was obtained from the incubator, treated with 10 μg ml−1 F(ab′)2 fragment goat anti-human μ chain (Jackson Immunoresearch) and the cells were immediately (within about 1 min) loaded into the SMR for measurement. Antibody-treated Jeko1 cells were measured for about 7 min to ensure that the anti-human μ chain stimulated changes were not reversed during the experiment. The SMR was briefly washed with PBS between each experiment. Wild-type Jeko1 cells were also measured after fixation to obtain a positive control for cell stiffness. For fixation, the cells were washed twice with PBS, mixed with 8% PFA for 30 min, washed twice with PBS and stored at 4 °C before the acoustic scattering measurements.
Quantification and statistical analysis
Data are shown as mean ± s.d. unless stated. Statistical analysis was performed by GraphPad Prism 7 (GraphPad Software) using unpaired two-tailed t test, or as indicated in figure legends. Significance was considered at P < 0.05. For in vivo transplantation experiments, the minimal number of mice in each group was calculated through use of the ‘cpower’ function in the Hmisc package of R. Kaplan–Meier survival analysis was used to estimate overall survival with GraphPad Prism 7. Mantel–Cox log-rank test was used to compare the difference between two groups. No mice were excluded. For patient overall survival analysis, patients in each dataset were divided into two groups on the basis of whether their expression was above or below the median level of IFITM3 and Kaplan–Meier survival analysis was used to estimate overall survival. The datasets used include patient-outcome data for B-ALL (Children’s Oncology Group (COG) P9906, n = 207; Eastern Cooperative Oncology Group (ECOG) E2993, n = 83; and St Jude, n = 15), mantle cell lymphoma (Lymphoma/Leukemia Molecular Profiling Project (LLMPP), n = 92) and acute myeloid leukaemia (The Cancer Genome Atlas (TCGA), n = 200). A log-rank test was used to compare survival differences between patient groups. The R package ‘survival’ (version 2.35-8) was used for the survival analysis and Cox proportional hazards regression model in the R package for the multivariate analysis (https://www.r-project.org/). The investigators were not blinded to allocation during experiments and outcome assessment. Experiments were repeated to ensure reproducibility of the observations.
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this paper.
Data availability
Patient-outcome data for B-ALL were obtained from the National Cancer Institute TARGET DATA Matrix of the COG Clinical Trial P9906 (GSE11877) (refs. 28,30), ECOG Clinical Trial E2993 (GSE5314) (ref. 57) and St Jude Children’s Research Hospital (https://www.stjuderesearch.org/site/data/ALL3/) (ref. 29). Patient-outcome data for mantle cell lymphoma were obtained from https://llmpp.nih.gov/MCL/ (ref. 58). Patient-outcome data for AML were obtained from TCGA Acute Myeloid Leukaemia Project (http://www.cbioportal.org/study/summary?id=laml_tcga_pub#clinical)59. Proteomics data have been deposited to the ProteomeXchange Consortium via the PRIDE partner repository with the following accession numbers: cell-surface proteome, PXD014691; phosphoproteome, PXD020696; and IFITM3 interactomes, PXD020697. Levels of IFITM3 mRNA across human normal and malignant B-lymphoid samples were obtained from http://Amazonia.transcriptome.eu/. All other data are available from the corresponding author upon reasonable request. Genome binding and occupancy profiling from wild-type and IKDN stromal adherent pre-B cells were obtained from GSE86897. Immunohistochemistry images for IFITM3 levels in normal or malignant B cells were obtained from The Human Protein Atlas https://www.proteinatlas.org/. ChIP-seq data of the genome-wide mapping of IKZF1 binding in patient-derived B-ALL xenograft cells were obtained from GSE58825. ChIP-seq data of the genetic analysis of IKZF1 target genes and tumour suppressor function in Ph+ pre-B ALL cells were obtained from GSE90656. RNA-sequencing data for Ifitm3+/+ and Ifitm3−/− BCR-ABL1 or NRASG12D B-ALL cells are available at GSE155305. RNA-sequencing data for Ptenfl/fl pre-B cells carrying 4-OHT-inducible Cre-ERT2 or ERT2 are available at GSE155618. Supplementary Table 12 summarizes the accession numbers and publicly deposited data from this study. All other data needed to evaluate the conclusions in the paper are available within the Article and its Supplementary Information. Source data are provided with this paper.
Change history
12 March 2021
A Correction to this paper has been published: https://doi.org/10.1038/s41586-021-03388-4
References
Bailey, C. C., Zhong, G., Huang, I. C. & Farzan, M. IFITM-family proteins: the cell’s first line of antiviral defense. Annu. Rev. Virol. 1, 261–283 (2014).
Diamond, M. S. & Farzan, M. The broad-spectrum antiviral functions of IFIT and IFITM proteins. Nat. Rev. Immunol. 13, 46–57 (2013).
Brass, A. L. et al. The IFITM proteins mediate cellular resistance to influenza A H1N1 virus, West Nile virus, and dengue virus. Cell 139, 1243–1254 (2009).
Zhang, Y. et al. Interferon-induced transmembrane protein 3 genetic variant rs12252-C associated with disease severity in coronavirus disease 2019. J. Infect. Dis. 222, 34–37 (2020).
Zhao, X. et al. Identification of residues controlling restriction versus enhancing activities of IFITM proteins on entry of human coronaviruses. J. Virol. 92, e01535-17 (2018).
Jia, R. et al. The N-terminal region of IFITM3 modulates its antiviral activity by regulating IFITM3 cellular localization. J. Virol. 86, 13697–13707 (2012).
Chesarino, N. M., McMichael, T. M., Hach, J. C. & Yount, J. S. Phosphorylation of the antiviral protein interferon-inducible transmembrane protein 3 (IFITM3) dually regulates its endocytosis and ubiquitination. J. Biol. Chem. 289, 11986–11992 (2014).
Everitt, A. R. et al. IFITM3 restricts the morbidity and mortality associated with influenza. Nature 484, 519–523 (2012).
Makvandi-Nejad, S. et al. Lack of truncated IFITM3 transcripts in cells homozygous for the rs12252-C variant that is associated with severe influenza infection. J. Infect. Dis. 217, 257–262 (2018).
Kohn, A. Early interactions of viruses with cellular membranes. Adv. Virus Res. 24, 223–276 (1979).
Liu, S. Y., Sanchez, D. J. & Cheng, G. New developments in the induction and antiviral effectors of type I interferon. Curr. Opin. Immunol. 23, 57–64 (2011).
Li, D. et al. KLF4-mediated negative regulation of IFITM3 expression plays a critical role in colon cancer pathogenesis. Clin. Cancer Res. 17, 3558–3568 (2011).
Zhang, D., Wang, H., He, H., Niu, H. & Li, Y. Interferon induced transmembrane protein 3 regulates the growth and invasion of human lung adenocarcinoma. Thorac. Cancer 8, 337–343 (2017).
Spence, J. S. et al. IFITM3 directly engages and shuttles incoming virus particles to lysosomes. Nat. Chem. Biol. 15, 259–268 (2019).
Harvey, R. C. et al. Identification of novel cluster groups in pediatric high-risk B-precursor acute lymphoblastic leukemia with gene expression profiling: correlation with genome-wide DNA copy number alterations, clinical characteristics, and outcome. Blood 116, 4874–4884 (2010).
Witkowski, M. T. et al. Conserved IKAROS-regulated genes associated with B-progenitor acute lymphoblastic leukemia outcome. J. Exp. Med. 214, 773–791 (2017).
Fujimoto, M. et al. CD19 regulates Src family protein tyrosine kinase activation in B lymphocytes through processive amplification. Immunity 13, 47–57 (2000).
Okkenhaug, K. et al. Impaired B and T cell antigen receptor signaling in p110δ PI 3-kinase mutant mice. Science 297, 1031–1034 (2002).
Engel, P. et al. Abnormal B lymphocyte development, activation, and differentiation in mice that lack or overexpress the CD19 signal transduction molecule. Immunity 3, 39–50 (1995).
Chan, L. N. et al. Metabolic gatekeeper function of B-lymphoid transcription factors. Nature 542, 479–483 (2017).
Wang, Y. et al. The physiologic role of CD19 cytoplasmic tyrosines. Immunity 17, 501–514 (2002).
Phelan, J. D. et al. A multiprotein supercomplex controlling oncogenic signalling in lymphoma. Nature 560, 387–391 (2018).
Kläsener, K., Maity, P. C., Hobeika, E., Yang, J. & Reth, M. B cell activation involves nanoscale receptor reorganizations and inside-out signaling by Syk. eLife 3, e02069 (2014).
Schwickert, T. A. et al. Stage-specific control of early B cell development by the transcription factor Ikaros. Nat. Immunol. 15, 283–293 (2014).
Joshi, I. et al. Loss of Ikaros DNA-binding function confers integrin-dependent survival on pre-B cells and progression to acute lymphoblastic leukemia. Nat. Immunol. 15, 294–304 (2014).
McLaughlin, S. & Murray, D. Plasma membrane phosphoinositide organization by protein electrostatics. Nature 438, 605–611 (2005).
Geng, H. et al. Integrative epigenomic analysis identifies biomarkers and therapeutic targets in adult B-acute lymphoblastic leukemia. Cancer Discov. 2, 1004–1023 (2012).
Harvey, R. C. et al. Identification of novel cluster groups in pediatric high-risk B-precursor acute lymphoblastic leukemia with gene expression profiling: correlation with genome-wide DNA copy number alterations, clinical characteristics, and outcome. Blood 116, 4874–4884 (2010).
Ross, M. E. et al. Classification of pediatric acute lymphoblastic leukemia by gene expression profiling. Blood 102, 2951–2959 (2003).
Kang, H. et al. Gene expression classifiers for relapse-free survival and minimal residual disease improve risk classification and outcome prediction in pediatric B-precursor acute lymphoblastic leukemia. Blood 115, 1394–1405 (2010).
Borowitz, M. J. et al. Clinical significance of minimal residual disease in childhood acute lymphoblastic leukemia and its relationship to other prognostic factors: a Children’s Oncology Group study. Blood 111, 5477–5485 (2008).
Poddar, S., Hyde, J. L., Gorman, M. J., Farzan, M. & Diamond, M. S. The interferon-stimulated gene IFITM3 restricts infection and pathogenesis of arthritogenic and encephalitic alphaviruses. J. Virol. 90, 8780–8794 (2016).
Foley, S. B. et al. Expression of BCR/ABL p210 from a knockin allele enhances bone marrow engraftment without inducing neoplasia. Cell Rep. 5, 51–60 (2013).
Schjerven, H. et al. Genetic analysis of Ikaros target genes and tumor suppressor function in BCR-ABL1+ pre-B ALL. J. Exp. Med. 214, 793–814 (2017).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Liao, Y., Smyth, G. K. & Shi, W. featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30, 923–930 (2014).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140 (2010).
Yu, G., Wang, L.-G. & He, Q.-Y. ChIPseeker: an R/Bioconductor package for ChIP peak annotation, comparison and visualization. Bioinformatics 31, 2382–2383 (2015).
Wollscheid, B. et al. Mass-spectrometric identification and relative quantification of N-linked cell surface glycoproteins. Nat. Biotechnol. 27, 378–386 (2009).
Tyanova, S., Temu, T. & Cox, J. The MaxQuant computational platform for mass spectrometry-based shotgun proteomics. Nat. Protocols 11, 2301–2319 (2016).
Tyanova, S. et al. The Perseus computational platform for comprehensive analysis of (prote)omics data. Nat. Methods 13, 731–740 (2016).
Monera, O. D., Sereda, T. J., Zhou, N. E., Kay, C. M. & Hodges, R. S. Relationship of sidechain hydrophobicity and α-helical propensity on the stability of the single-stranded amphipathic α-helix. J. Pept. Sci. 1, 319–329 (1995).
Ling, S. et al. Combined approaches of EPR and NMR illustrate only one transmembrane helix in the human IFITM3. Sci. Rep. 6, 24029 (2016).
Berendsen, H. J. C., van der Spoel, D. & van Drunen, R. GROMACS: a message-passing parallel molecular dynamics implementation. Comput. Phys. Commun. 91, 43–56 (1995).
de Jong, D. H. et al. Improved parameters for the Martini coarse-grained protein force field. J. Chem. Theory Comput. 9, 687–697 (2013).
Yen, H.-Y. et al. PtdIns(4,5)P2 stabilizes active states of GPCRs and enhances selectivity of G-protein coupling. Nature 559, 423–427 (2018).
Hsu, P.-C. et al. CHARMM-GUI Martini maker for modeling and simulation of complex bacterial membranes with lipopolysaccharides. J. Comput. Chem. 38, 2354–2363 (2017).
Qi, Y. et al. CHARMM-GUI Martini maker for coarse-grained simulations with the Martini force field. J. Chem. Theory Comput. 11, 4486–4494 (2015).
Berendsen, H. J. C., Postma, J. P. M., van Gunsteren, W. F., DiNola, A. & Haak, J. R. Molecular dynamics with coupling to an external bath. J. Chem. Phys. 81, 3684–3690 (1984).
Tironi, I. G., Sperb, R., Smith, P. E. & van Gunsteren, W. F. A generalized reaction field method for molecular dynamics simulations. J. Chem. Phys. 102, 5451–5459 (1995).
Wassenaar, T. A., Pluhackova, K., Böckmann, R. A., Marrink, S. J. & Tieleman, D. P. Going backward: a flexible geometric approach to reverse transformation from coarse grained to atomistic models. J. Chem. Theory Comput. 10, 676–690 (2014).
Huang, J. & MacKerell, A. D. Jr. CHARMM36 all-atom additive protein force field: validation based on comparison to NMR data. J. Comput. Chem. 34, 2135–2145 (2013).
Martyna, G. J., Klein, M. L. & Tuckerman, M. Nosé–Hoover chains: the canonical ensemble via continuous dynamics. J. Chem. Phys. 97, 2635–2643 (1992).
Buchner, M. et al. Identification of FOXM1 as a therapeutic target in B-cell lineage acute lymphoblastic leukaemia. Nat. Commun. 6, 6471 (2015).
Kang, J. H. et al. Noninvasive monitoring of single-cell mechanics by acoustic scattering. Nat. Methods 16, 263–269 (2019).
Burg, T. P. et al. Weighing of biomolecules, single cells and single nanoparticles in fluid. Nature 446, 1066–1069 (2007).
Juric, D. et al. Differential gene expression patterns and interaction networks in BCR-ABL-positive and -negative adult acute lymphoblastic leukemias. J. Clin. Oncol. 25, 1341–1349 (2007).
Rosenwald, A. et al. The proliferation gene expression signature is a quantitative integrator of oncogenic events that predicts survival in mantle cell lymphoma. Cancer Cell 3, 185–197 (2003).
The Cancer Genome Atlas Research Network. Genomic and epigenomic landscapes of adult de novo acute myeloid leukemia. N. Engl. J. Med. 368, 2059–2074 (2013).
Acknowledgements
We thank M. A. Lemmon for critical discussion and advice, and current and former members of the Müschen laboratory for their support and helpful discussions. Research in the Müschen laboratory is funded by the NIH/NCI through R35CA197628, R01CA157644, R01CA213138 and P01CA233412, the Dr. Ralph and Marian Falk Medical Research Trust Transformational Awards Program through MCG-18447-19, the Leukemia and Lymphoma Society through MCL-7000-18 and Blood Cancer Discoveries Grant BCDG-20327-20, as well the Howard Hughes Medical Institute HHMI-55108547 (to M.M.). This work was partly supported by NIH grants R35CA200422, R01CA251275, AI116585, AI116585S, AI140718, AI140705, DE023926 and DE028521 (to J.U.J.), R01 AI104002 and R01 AI127513 (to M.S.D.), R01 GM117923 and R01 GM097261 (to N.V.), NCI R35 CA231958 (to D.M.W.), Wellcome Trust 110275/Z/15/Z (to T.P.M.) and the Paul G. Allen Frontiers Group and the Cancer Systems Biology Consortium funding (U54-CA217377) from the National Cancer Institute (to S.R.M.). Relevant research in the A.T.-T. laboratory was supported by the Cancer Moonshot grant U01 CA232563. T.S. is a fellow of the Lymphoma Research Foundation.V.K. is supported by a Career Development Fellow grant (5491-20) from the Leukemia and Lymphoma Society, and by a Young Investigator Award from Alex’s Lemonade Stand Foundation. Proteomics services were performed by the Northwestern Proteomics Core Facility, generously supported by NCI CCSG P30 CA060553 awarded to the Robert H Lurie Comprehensive Cancer Center, instrumentation award (S10OD025194) from NIH Office of Director, and the National Resource for Translational and Developmental Proteomics supported by P41 GM108569. M.M. is a Howard Hughes Medical Institute (HHMI) Faculty Scholar. J.C. is a Leukemia and Lymphoma Society (LLS) Scholar.
Author information
Authors and Affiliations
Contributions
J.L. performed experiments and contributed to all aspects of the study; in particular, western blotting, flow cytometry analysis, viable cell counts, growth competition assays, immunization assays, ELISA, colony-forming assays, CRISPR-mediated gene deletion, cell sorting, in vivo transplantation experiments, bio-imaging, bio-ID, immunoprecipitation assays, lipid strip assays, lipid raft analysis, calcium flux measurements and data analysis. M.E.R. and H.G. performed all bioinformatic and biostatistical analyses and power calculations for experimental design. N.M. and N.V. performed molecular dynamics simulations and structural analysis of IFITM3–PIP3 interactions. D.A. performed western blotting, flow cytometry analysis, viable cell counts, cell adhesion assays, CRISPR-mediated gene deletion and immunization assays. G.D., J.W., K.N.C., L.N.C., K.K. and V.K. performed flow cytometry analysis and viable cell counts. L.K. performed gene cloning, virus generation and flow cytometry analysis and cell sorting. M.A.A. performed RNA isolation for RNA sequencing, western blotting and CRISPR-mediated gene deletion. G.X. performed RNA-sequencing analysis with Ptenfl/fl ERT2 and Cre-ERT2 mouse models and measured PIP3:PIP2 ratios. T.S. performed PLA. V.K. also performed immunofluorescence staining after immunization assays. C.W.C. and J.C. provided expertise in gene-editing experiments. M.A.N. and A.P.W. provided expertise in cell-surface proteomics. A.T.-T. provided expertise in CD19 signalling, and M.F., M.S.D. and J.U.J. contributed expertise and reagents in antiviral effector functions of IFITM3. T.P.M., Y.Z., D.M.W. and S.R.M. performed SNACS cell-stiffness assays. M.M. secured funding, developed the amplification loop concept and wrote the manuscript. All co-authors reviewed and edited the manuscript.
Corresponding author
Ethics declarations
Competing interests
M.S.D. is a consultant for Inbios, Vir Biotechnology and NGM Biopharmaceuticals, and on the Scientific Advisory Board of Moderna and Immunome. The Diamond laboratory at Washington University School of Medicine has received unrelated sponsored research agreements from Moderna, Vir Biotechnology, and Emergent BioSolutions.
Additional information
Peer review information Nature thanks Marcus Clark, Paul Kellam, Michael Reth and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Ifitm3 expression is induced by oncogenic PI3K signalling and repressed by IKZF1.
a, Changes of Ifitm3 mRNA levels were monitored in mouse splenic B cells upon BCR engagement (mean ± s.e.; n = 2). b, c, Ptenfl/fl pre-B ALL cells carrying 4-OHT-inducible Cre-ERT2 or ERT2 were treated with 4-OHT and studied by RNA sequencing for Ifitm3 transcript expression (b) and western blot for protein levels of PTEN, AKT-pS473, AKT and IFITM3 (c; n = 3). d, STAT5 phosphorylated at Y694 (STAT5-pY694), STAT5 and IFITM3 levels in patient-derived B-ALL cells (PDX2) measured by western blotting upon imatinib treatment (n = 3). e, Scenario of the PI3K pathway as a positive regulator of IFITM3, which in turn amplifies BCR and oncogenic signalling. f, IFITM3 mRNA levels across human normal and malignant haematopoietic and B-lymphoid samples (source data and statistics are given in Supplementary Table 2). Two-tailed t-test. g, IFITM3 mRNA levels in pre-B cells from healthy donors and samples from patients with B-ALL were compared for two clinical cohorts (ECOG E2993 and St Jude). In ECOG E2993, bone marrow samples were obtained at diagnosis before treatment from 83 adults with B-ALL with a confirmed purity of >90% leukaemic blasts. For the St Jude datasets, 15 samples from children with B-ALL before treatment were compared to flow-sorted pre-B cells from bone marrow aspirates of healthy donors. Two-tailed Wilcoxon. h, Minimal residual disease (MRD) was determined in the COG P9906 cohort, IFITM3 mRNA levels were compared in patients who were positive (n = 67) and negative (n = 124) for MRD. Two-tailed Wilcoxon. i–l, Patients with leukaemia and lymphoma from five clinical cohorts were segregated into two groups based on higher (IFITM3high) or lower (IFITM3low) than median IFITM3 mRNA levels. Overall survival was compared by two-tailed log-rank test. m, ChIP-seq enrichment of RNAPII and H3K4me3 at the Ifitm3 locus in pre-B cells (top) from Ikzf1exon5fl/fl mice upon Cre-mediated deletion of Ikzf1 (GSE86897). Binding of IKZF1 to the promoter region of IFITM3 was also analysed in ChIP-seq data from patient-derived B-ALL cells (bottom, LAX2, GSE58825). n, Human B-ALL cells (BV173) carrying IKZF1 deletions were reconstituted with doxycycline-inducible IKZF1 or EV. Levels of IFITM3 were assessed by western blotting upon doxycycline-induction (n = 3). o, Multivariate analysis of established risk factors in children with B-ALL (COG P9906, n = 207) including mutation or deletion of IKZF1. Patients (n = 207) were separated into IKZF+/+ or IKZF-deleted groups, then further segregated based on higher or lower than median expression levels of IFITM3. The comparison of these four groups established IFITM3 mRNA levels as an independent risk factor regardless of IKZF1-deletion status (two-sided log-rank test; P = 0.0045). c, d, n, For gel source data, see Supplementary Fig. 1.
Extended Data Fig. 2 Ifitm3 is essential for the development of B1 and germinal centre B cells.
a, Hardy fractions of B cell subsets isolated from bone marrow of Ifitm3+/+ and Ifitm3−/− littermates analysed by flow cytometry (n = 3). b, Surface expression of IgM, CD20, CD19, IgD, CD2 and CD21 measured by flow cytometry in enriched bone marrow (GR1−, Nk1.1− and B220+) and splenic B cells (CD3− and B220+) from Ifitm3+/+ or Ifitm3−/− mice (n = 7; mean ± s.d.). Mean fluorescence intensities (MFI) values for individual measurement compared by two-tailed t-test. c, Ca2+ mobilization from cytoplasmic stores in response to BCR (IgM) engagement was measured in Ifitm3+/+ and Ifitm3−/− splenic B cells. Ca2+ release was induced by addition of 10 μg ml−1 anti-mouse IgM 60 s after acquisition of background fluorescence. Ca2+ release was measured over 300 s with cell-permeant Rhod-2 dye; MFI compared between replicates (n = 3). d, Ca2+ mobilization in response to BCR engagement measured upon CRISPR–Cas9-mediated deletion of IFITM3 in Jeko1 MCL cells. Ca2+ release upon addition of 10 μg ml−1 of polyclonal F(ab′)2 anti-human IgM was measured for 300 s with cell-permeant Fluo-4 dye; MFI compared between replicates (left; n = 3). Surface expression of CD19 following deletion of IFITM3 in Jeko1 MCL cells, MFIs for CD19 indicated (right; n = 3). e, Jeko1 MCL cells were electroporated with non-targeting RNP (Cas9-gRNA ribonucleoproteins, gNT) or IFITM3-targeting RNP complex (gIFITM3). Following electroporation, MCL cells were treated with vehicle (DMSO) or 25 nmol l−1 of dasatinib for 3 h. Cells were stimulated with 10 μg ml−1 of anti-human IgM F(ab′)2 for the indicated time points and subjected to co-immunoprecipitation with an anti-CD19 antibody. Immunoblots were performed to measure levels of CD19 tyrosine phosphorylation and binding of LYN to CD19. Levels of IFITM3, SRC-pY416 and LYN were assessed in whole-cell lysates (10% input) with β-actin as loading control. (n = 3; gel source data in Supplementary Fig. 1). f–h, Relative fractions (left) and absolute cell counts (right) of total B1 (f) and B1a (g) cells in the peritoneal cavity and marginal-zone B cells (h) in spleen of Ifitm3+/+ and Ifitm3−/− littermates (n = 5) are shown (means ± s.d.; two-tailed t-test).
Extended Data Fig. 3 Ifitm3-deficient B-ALL cells exhibit an anergic phenotype and compensatory upregulation of Ifitm1 and PI3K signalling molecules.
a, Numbers of viable Ifitm3+/+ and Ifitm3−/− BCR-ABL1 or NRASG12D B-ALL cells were counted by Trypan blue dye exclusion (n = 3; mean ± s.d.; two-tailed t-test). b, RNA sequencing was performed for Ifitm3+/+ and Ifitm3−/− BCR-ABL1 and NRASG12D B-ALL cells. Relative rlog-normalized gene expression values for all strongly differentially expressed genes (P < 1 × 10−5 and log2-transformed fold change >1; Wald test with Benjamini–Hochberg correction) in both BCR-ABL1 and NRASG12D conditions plotted as a heat map with row scaling. B-cell-signalling-related genes are labelled in red, anergy-related genes are labelled in blue and PI3K-signalling-related genes are labelled in grey. c, Gene set enrichment analysis for genes ranked by ratio of Ifitm3−/− to Ifitm3+/+ as log2-transformed fold change; red lines indicate running enrichment score (right axis), grey bars indicate fold change (left axis). Statistical significance was determined by two-tailed Kolmogorov–Smirnov test. d, Patient-derived B-ALL cells (PDX2) were transduced with N-terminally Flag-tagged or C-terminally HA-tagged IFITM3 constructs. Combinations of intracellular and surface staining were performed to examine IFITM3 topology at the cell membrane. e, Patient-derived B-ALL cells (PDX2) were transduced with C-terminal HA-tagged IFITM3 or the IFITM3(Y20E) phosphomimetic. Combinations of intracellular and surface staining, with or without SRC-kinase inhibition by dasatinib, were performed to examine IFITM3 topology at the cell membrane and its regulation by SRC kinases. d, e, Representative plots from three independent experiments. f, A scenario of the topology of IFITM3 regulated by SRC (LYN) or oncogenic tyrosine kinases (BCR–ABL1) at the plasma membrane is shown. Phosphorylation of Y20 hinders the recognition of 20YEM23 endocytosis motif by the AP-2 complex, thereby antagonizing endocytosis and endosomal trafficking of IFITM3.
Extended Data Fig. 4 IFITM3 amplifies PI3K signalling downstream of BCR and integrin receptors.
a, Volcano plot of differentially phosphorylated proteins in patient-derived B-ALL (PDX2) cells transduced with IFITM3(Y20E) compared to empty vector (EV) control (n = 3; Wald test with Benjamini–Hochberg correction). b, F(ab) fragments of the anti-HA antibody or isotype control were purified and their identity confirmed by western blot (left). Eight million patient-derived B-ALL (PDX2) cells carrying IFITM3–HA, IFITM3(Y20E)–HA or EV control were resuspended in complete medium and treated with either 2.5 μg ml−1 of full antibodies, F(ab) fragments of anti-HA or isotype control for the indicated times. Levels of AKT-pS473, AKT and HA-tagged IFITM3 were assessed by western blots using β-actin as loading control (right). Data from three independent experiments. For gel source data, see Supplementary Fig. 1. c, Ca2+ mobilization in response to TCR engagement using CD3ɛ-specific antibodies was measured upon CRISPR–Cas9-mediated deletion of IFITM3 in T-ALL cells (Jurkat) (left). Ca2+ release from cytoplasmic stores was induced by adding 10 μg ml−1 of monoclonal (OKT3) anti-human CD3ɛ at 50 s after acquisition of background fluorescence. Surface expression of CD3 was measured following deletion of IFITM3 in Jurkat cells (right). MFIs for CD3 are indicated. Representative plots from three independent biological experiments are shown. MFI values for individual measurement were compared by two-tailed t-test. d, Surface proteins on Ifitm3+/+ and Ifitm3−/− B-ALL cells were labelled with biotin and enriched with streptavidin affinity pull-down followed by on-bead trypsin digestion, mass spectrometry and quantified with label-free quantification. Differentially expressed cell surface proteins on Ifitm3+/+ and Ifitm3−/− B-ALL cells are shown with the mean difference of label-free quantification plotted against the P value (Welch’s t-test). All experiments were performed in biological triplicates. e–g, Validation of differential expression of surface receptors between Ifitm3+/+, Ifitm3−/− and Ifitm3Y20E-overexpressing B-ALL cells. Flow cytometry analyses show surface expression of BCR signalling components (e), integrins and adhesion receptors (f) and other surface receptors (g) in Ifitm3+/+, Ifitm3−/− and B-ALL cells expressing IFITM3(Y20E) (n = 3).
Extended Data Fig. 5 Inducible membrane translocation of CD19 does not rescue defective SRC and PI3K signalling in Ifitm3-deficient B cells.
a, Surface expression of CD19 was assessed by flow cytometry following forced expression of CD19 for >1 week in Ifitm3+/+ and Ifitm3−/− B-ALL cells. b, Western blot analyses of AKT-pS473, AKT, MYC and BCL2 upon forced expression of CD19 for >1 week in mouse Ifitm3+/+ and Ifitm3−/− B-ALL cells. c, d, Colony-forming ability (c) and cell cycle progression (d) of Ifitm3+/+ and Ifitm3−/− B-ALL cells upon forced expression of CD19 for >1 week was examined. c, Colony numbers for individual measurement were compared by two-tailed t-test. d, Numbers indicate percentage of cells in S phase. e, Numbers of viable Ifitm3+/+ and Ifitm3−/− B-ALL cells following forced expression of CD19 were counted using the Trypan blue dye exclusion method (left). Ifitm3−/− B-ALL cells were transduced with GFP-tagged constructs for expression of CD19 with an intact (Y531) or mutant (Y531F) PI3K-activation motif in its cytoplasmic tail. Relative changes of GFP+ cells (transduced with CD19 or CD19(Y531F)) were plotted over time (means ± s.d.). a–e, Data from three independent experiments. b, For gel source data, see Supplementary Fig. 1. f, Mouse Cd19−/− B-ALL clones were generated by electroporation of mouse B-ALL cells with CD19-targeting RNP (Cas9-gRNA ribonucleoproteins, gCd19 ALL) and single-cell clones with biallelic deletion were selected. Cd19−/− B-ALL cells were transduced with CD19–ERT2, a fusion of the ER-ligand binding domain to the C terminus of CD19, or ERT2 as empty vector control. Reconstitution of Cd19−/− B-ALL cells with CD19–ERT2 resulted in stable expression of the fusion proteins that were retained in complex with cytoplasmic heatshock proteins. Addition of 4-OHT released CD19–ERT2 from its cytoplasmic heatshock chaperone and enable cell surface expression within 30 min of 4-OHT addition. g, To test the effect of inducible CD19 membrane translocation in Ifitm3−/− B-ALL cells, Ifitm3+/+ and Ifitm3−/− B-ALL cells were transduced with CD19–ERT2 or ERT2 empty vector control. 4-OHT-mediated translocation of CD19 to the cell surface was assessed by flow cytometry for the indicated times (0 to 3 h). h, Ifitm3+/+ and Ifitm3−/− B-ALL cells were transduced with CD19–ERT2 or ERT2 empty vector control. Cells were treated for 0, 1 and 3 h with 4-OHT for surface-translocation of CD19. Cell lysates from these populations were analysed by western blot for CD19-pY531, CD19, SRC-pY416, LYN, AKT-pS473, AKT, MYC and BCL2. While CD19–ERT2 reconstitutes CD19 protein levels in Ifitm3−/− B-ALL cells and restores rapid translocation to the cell surface (g), this change alone was not sufficient to induce proper phosphorylation of CD19, SRC kinases and PI3K signalling via AKT. f–h, Data from three independent experiments. h, For gel source data, see Supplementary Fig. 1.
Extended Data Fig. 6 IFITM3 links components of the BCR and integrin receptor pathways to PI3K signalling.
a, Schematic of HA-tagged-IFITM3(Y20E)–BirA fusion proteins used for TurboID interactome analyses. BirA (biotin ligase) was fused to N-terminal IFITM3 carrying the phosphomimetic Y20E mutation for membrane localization. b–d, HA–IFITM3(Y20E)–BirA or HA–BirA controls were expressed in PDX2 B-ALL (b) or Jeko1 MCL cells (c, d). c, d, Cells were incubated with exogenous biotin for 10 min upon IFITM3 (anti-HA) or BCR (anti-IgM) engagement. IFITM3(Y20E) interactome analyses identified interacting proteins by mass spectrometry, plotted based on significance and log2-transformed fold enrichment over EV control. Essential interactors as BCR component (red), integrin (blue) and PI3K signalling (grey) are highlighted. Data from three independent biological replicates. e, PLAs were performed with Jeko1 MCL cells upon engagement of BCR. Jeko1 MCL cells were stimulated by BCR engagement for 0, 3 and 30 min, then fixed, permeabilized and assessed for the proximity of CD79B to IFITM3. Representative microscopic images with PLA signal (red dot) and nuclei stained with DAPI as blue are shown. LAMP1 was used as a marker for endosomes to distinguish plasma-membrane-bound from endosomal localization of CD79B:IFITM3 complexes. Scale bars, 5 mm. Data from three independent replicates.
Extended Data Fig. 7 Ifitm3 functions as a central effector of B cell adhesion.
a, Homotypic aggregation was studied in Ifitm3−/− B-ALL cells that were reconstituted with C-terminal HA-tagged IFITM3, IFITM3(Y20E) or empty vector (EV) and incubated with anti-CD19, anti-HA antibodies or isotype control for 24 h. Data from three independent experiments. b, Ifitm3+/+ or Ifitm3−/− B-ALL cells were transduced with GFP-tagged IK6 or GFP alone. IK6 levels in flow-sorted GFP+ cells assessed by western blot analysis using β-actin as loading control (right). Ten thousand Ifitm3+/+ or Ifitm3−/− B-ALL cells carrying IK6 or EV were plated for colony forming assays. Colonies were imaged and counted after 7 days. Representative images are shown with colony numbers. Data from three independent experiments and assessed by two-tailed t-test (means ± s.d.). For gel source data, see Supplementary Fig. 1. c, One hundred thousand Ifitm3+/+ or Ifitm3−/− B-ALL cells carrying IK6 or EV were cultured on OP9 stroma cells. Ratios of adherent cells to nonadherent cells were calculated. Data from three independent biological experiments (right) were analysed and assessed by two-tailed t-test (means ± s.d.). d, Representative images of adherent B-ALL cells on OP9 stroma are shown. Round and light-refracting cells are adherent B-ALL cells attached to stroma cells. Dark and round cells are adherent B-ALL cells incorporated into stromal layer. Data from three independent biological replicates. e, Surface expression levels of integrins on adherent B-ALL cells were measured by flow cytometry. MFI values are indicated for individual measurements. Data from three independent biological replicates.
Extended Data Fig. 8 IFITM3 functions as a PIP3 scaffold and mediated cell-membrane stiffening upon BCR engagement.
a, SNACS was measured using a previously established microfluidic method as a metric for cell surface stiffness. Jeko1 mantle cell lymphoma cells were flown through a standing acoustic wave generated inside a vibrating suspended microchannel resonator. The cantilever vibration frequency was monitored, and its shift was used to quantify the acoustic scattering from the cells as well as the buoyant mass of the cells. The data displayed was obtained using a 350-μm-long cantilever with 15 × 20-μm-sized channel inside of the cantilever and an approximately 200-ms transit time through the cantilever. All the regulators, valves and data acquisition were controlled by custom software coded in LabVIEW 2017 (National Instruments). A parallel volume measurement using Coulter Counter was carried out to quantify average cell volume, which was used together with the single-cell buoyant mass measurements to calculate SNACS for each cell. Jeko1 mantle cell lymphoma cells without (IFITM3+/+) and with (IFITM3−/−) CRISPR–Cas9-mediated deletion of IFITM3 were kept under cell culture conditions and treated with an anti-IgM antibody. Fixation with paraformaldehyde (PFA) was used as a positive control. Representative plots from three independent experiments are shown (median levels in red dotted line). Statistical significance was determined by two-tailed t-test. Numbers indicated cells studied for SNACS measurement. b, Ifitm3+/+ and Ifitm3−/− B-ALL cells were incubated with 30 μM of the PIP3 carrier histone H1 or PIP3–histone H1 complex for 30 min. Levels of CD19-pY531, CD19, SRC-pY416, LYN, AKT-pS473, AKT and MYC were measured by western blot using β-actin as loading control. Data from three independent replicates. For gel source data, see Supplementary Fig. 1. c, Colony-formation assays were performed for Ifitm3+/+ and Ifitm3−/− B-ALL cells that were treated with 30 μM of PIP3–histone H1 or the shuttle protein histone H1 (H1) alone. Photomicrographs and colony numbers per 10,000 plated cells are shown. Data are presented as means ± standard derivation (s.d.) from three independent experiments. Statistical significance was determined by two-tailed t-test. d, Flow cytometry analyses of surface expression of CD19, CD44, CD25 and CD44 on Ifitm3+/+ and Ifitm3−/− B-ALL cells treated with 30 μM of PIP3–histone H1 or the shuttle protein histone H1 (H1) alone for 72 h. Data from three independent replicates.
Extended Data Fig. 9 Modelling of binding of IFITM3 to PIP2 and PIP3.
a–c, The hierarchical scheme of molecular dynamics simulations to delineate the structural and dynamical basis of PIP2 and PIP3 binding to IFITM3 (yellow). a, Coarse grained simulations of IFITM3 in composite cell membrane. The side views of the lipid bilayer are shown. The lipid raft markers are shown in colour with GM1 (pink), cholesterol (green) and sphingomyelin (blue). The grey surface represents all the other lipids. b, The simulation cell extracted from the coarse grained simulation as the starting structure for all-atom molecular dynamics simulations. The IFITM3 protein is shown in yellow and the PIP2 and PIP3 are shown in green and red stick representations. c, Close up view of one of the predicted binding poses of PIP2 and PIP3 in the most populated conformation of IFITM3. The dashed lines shown are the PIP2 or PIP3 contacts with the basic residues in IFITM3 (top). The amino acid sequence of IFITM3 in the stretch of residues between TM1 and TM2 from 57 to 128 modelled in this work is shown. The CIL region is boxed and basic amino acids are highlighted with blue. The average interaction energy of PIP2 or PIP3 with the two basic batches measured in all-atom molecular dynamics simulations is indicated (Supplementary Table 6). d, e, The average interaction energy of PIP2 (green) or PIP3 (red) with IFITM3 (residues 57–128) was measured in all-atom molecular dynamics simulations. The one-residue contact indicates the conformation of PIP2 or PIP3 binding with only one basic residue in the basic amino patch KSRDRK of IFITM3. The two-residue contact refers to the binding conformation of PIP2 or PIP3 that show contacts with two basic residues. Representative plots from at least four independent experiments are shown (mean ± s.d.). In d, P values were determined by two-tailed t-test. Populations showing one- or two-residue contacts of PIP2 (green) or PIP3 (red) to IFITM3 were quantitated from all-atom molecular dynamics simulations. The population density was assessed by the normalization of number of events with the total number of frames. Representative plots from three independent experiments are shown (mean ± s.d.). In e, statistical significance was determined by two-tailed t-test.
Extended Data Fig. 10 IFITM3-mediated PI3K signalling downstream of BCR and integrin receptors depends on K83–K104 but not on R85, R87 and K88 residues.
a–c, Levels of differentially phosphorylated proteins in patient-derived B-ALL (PDX2) cells transduced with IFITM3(Y20E), IFITM3(Y20E/K83A), IFITM3(Y20E/R85A) or empty vector (EV) control were identified by mass spectrometry (n = 3). Relative abundance values are plotted for all sites ranked by fold change as indicated; phosphosites of interest are highlighted. b, FSEA ranked by log2-transformed fold change are shown for phosphosites in PI3K signalling (grey), BCR signalling (red) and integrin and adhesion receptor elements (blue). Statistical significance was determined by two-tailed Kolmogorov–Smirnov test. d, Cumulative distribution frequencies for log2-transformed fold changes in phosphosite abundance between IFITM3-transduced and EV conditions were calculated for all sites and globally increased in IFITM3(Y20E) over EV. The analysis was repeated for the K83A (red) and R85A (green) mutants of IFITM3. The light grey line indicates background variance observed between EV replicates. Shifts that were caused by the K83A and R85A mutants are indicated by arrows. Overall changes in distribution (shifts) between IFITM3(Y20E) and IFITM3(Y20E/K83A) and IFITM3(Y20E/R85A) mutants were measured by two-tailed Kolmogorov–Smirnov test. e, Schematic of BirA (engineered biotin ligase) was fused at N-terminal ends of HA-tagged IFITM3(Y20E) or its K83A mutant and expressed in PDX2 B-ALL or Jeko1 MCL cells. e–g, Interactomes of IFITM3(Y20E) and its K83A mutant were compared in PDX2 B-ALL cells upon IFITM3 engagement (anti-HA) (e) or in Jeko1 MCL cells upon IFITM3 (anti-HA) (f) or BCR (anti-IgM) (g) engagement. Phosphosites of interest, including BCR signalling, PI3K signalling and integrins and adhesion receptor elements are highlighted.
Supplementary information
Supplementary Tables
This file contains Supplementary Tables S1-S12.
Supplementary Figure
Supplementary Figure 1: Source Data (Immunoblots).
Source data
Rights and permissions
About this article
Cite this article
Lee, J., Robinson, M.E., Ma, N. et al. IFITM3 functions as a PIP3 scaffold to amplify PI3K signalling in B cells. Nature 588, 491–497 (2020). https://doi.org/10.1038/s41586-020-2884-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-020-2884-6
This article is cited by
-
Gene expression profiling of host lipid metabolism in SARS-CoV-2 infected patients: a systematic review and integrated bioinformatics analysis
BMC Infectious Diseases (2024)
-
FOXP3+ regulatory T cell perturbation mediated by the IFNγ-STAT1-IFITM3 feedback loop is essential for anti-tumor immunity
Nature Communications (2024)
-
IFITM3 promotes glioblastoma stem cell-mediated angiogenesis via regulating JAK/STAT3/bFGF signaling pathway
Cell Death & Disease (2024)
-
Role of interferon-induced transmembrane protein family in cancer progression: a special focus on pancreatic cancer
Medical Oncology (2024)
-
Comprehensive characterization of IFNγ signaling in acute myeloid leukemia reveals prognostic and therapeutic strategies
Nature Communications (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.