Abstract
Myeloid malignancies, including acute myeloid leukaemia (AML), arise from the expansion of haematopoietic stem and progenitor cells that acquire somatic mutations. Bulk molecular profiling has suggested that mutations are acquired in a stepwise fashion: mutant genes with high variant allele frequencies appear early in leukaemogenesis, and mutations with lower variant allele frequencies are thought to be acquired later1,2,3. Although bulk sequencing can provide information about leukaemia biology and prognosis, it cannot distinguish which mutations occur in the same clone(s), accurately measure clonal complexity, or definitively elucidate the order of mutations. To delineate the clonal framework of myeloid malignancies, we performed single-cell mutational profiling on 146 samples from 123 patients. Here we show that AML is dominated by a small number of clones, which frequently harbour co-occurring mutations in epigenetic regulators. Conversely, mutations in signalling genes often occur more than once in distinct subclones, consistent with increasing clonal diversity. We mapped clonal trajectories for each sample and uncovered combinations of mutations that synergized to promote clonal expansion and dominance. Finally, we combined protein expression with mutational analysis to map somatic genotype and clonal architecture with immunophenotype. Our findings provide insights into the pathogenesis of myeloid transformation and how clonal complexity evolves with disease progression.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
Raw data are available on dbGAP (accession number phs002049.v1.p1) in the form of loom files and FASTQ files for each sample.
Code availability
All scripts and processed data files are available at https://github.com/bowmanr/scDNA_myeloid.
References
Genovese, G. et al. Clonal hematopoiesis and blood-cancer risk inferred from blood DNA sequence. N. Engl. J. Med. 371, 2477–2487 (2014).
Jan, M. et al. Clonal evolution of preleukemic hematopoietic stem cells precedes human acute myeloid leukemia. Sci. Transl. Med. 4, 149ra118 (2012).
Papaemmanuil, E. et al. Genomic classification and prognosis in acute myeloid leukemia. N. Engl. J. Med. 374, 2209–2221 (2016).
Patel, J. P. et al. Prognostic relevance of integrated genetic profiling in acute myeloid leukemia. N. Engl. J. Med. 366, 1079–1089 (2012).
Ley, T. J. et al. Genomic and epigenomic landscapes of adult de novo acute myeloid leukemia. N. Engl. J. Med. 368, 2059–2074 (2013).
Rampal, R. et al. Genomic and functional analysis of leukemic transformation of myeloproliferative neoplasms. Proc. Natl Acad. Sci. USA 111, E5401–E5410 (2014).
Jaiswal, S. et al. Age-related clonal hematopoiesis associated with adverse outcomes. N. Engl. J. Med. 371, 2488–2498 (2014).
Pellegrino, M. et al. High-throughput single-cell DNA sequencing of acute myeloid leukemia tumors with droplet microfluidics. Genome Res. 28, 1345–1352 (2018).
Levine, R. L. et al. X-inactivation-based clonality analysis and quantitative JAK2V617F assessment reveal a strong association between clonality and JAK2V617F in PV but not ET/MMM, and identifies a subset of JAK2V617F-negative ET and MMM patients with clonal hematopoiesis. Blood 107, 4139–4141 (2006).
Kralovics, R. et al. A gain-of-function mutation of JAK2 in myeloproliferative disorders. N. Engl. J. Med. 352, 1779–1790 (2005).
Xie, M. et al. Age-related mutations associated with clonal hematopoietic expansion and malignancies. Nat. Med. 20, 1472–1478 (2014).
Abdel-Wahab, O. et al. Genetic analysis of transforming events that convert chronic myeloproliferative neoplasms to leukemias. Cancer Res. 70, 447–452 (2010).
Ortmann, C. A. et al. Effect of mutation order on myeloproliferative neoplasms. N. Engl. J. Med. 372, 601–612 (2015).
Buscarlet, M. et al. DNMT3A and TET2 dominate clonal hematopoiesis and demonstrate benign phenotypes and different genetic predispositions. Blood 130, 753–762 (2017).
Coombs, C. C. et al. Therapy-related clonal hematopoiesis in patients with non-hematologic cancers is common and associated with adverse clinical outcomes. Cell Stem Cell 21, 374–382 (2017).
McMahon, C. M. et al. Clonal selection with RAS pathway activation mediates secondary clinical resistance to selective FLT3 inhibition in acute myeloid leukemia. Cancer Discov. 9, 1050–1063 (2019).
Demaree, B. et al. Joint profiling of proteins and DNA in single cells reveals extensive proteogenomic decoupling in leukemia. Preprint at https://doi.org/10.1101/2020.02.26.967133 (2020).
van Galen, P. et al. Single-cell RNA-seq reveals AML hierarchies relevant to disease progression and immunity. Cell 176, 1265–1281 (2019).
Petti, A. A. et al. A general approach for detecting expressed mutations in AML cells using single cell RNA-sequencing. Nat. Commun. 10, 3660 (2019).
Falini, B., Nicoletti, I., Martelli, M. F. & Mecucci, C. Acute myeloid leukemia carrying cytoplasmic/mutated nucleophosmin (NPMc+ AML): biologic and clinical features. Blood 109, 874–885 (2007).
Majeti, R., Park, C. Y. & Weissman, I. L. Identification of a hierarchy of multipotent hematopoietic progenitors in human cord blood. Cell Stem Cell 1, 635–645 (2007).
Goardon, N. et al. Coexistence of LMPP-like and GMP-like leukemia stem cells in acute myeloid leukemia. Cancer Cell 19, 138–152 (2011).
Levine, J. H. et al. Data-driven phenotypic dissection of AML reveals progenitor-like cells that correlate with prognosis. Cell 162, 184–197 (2015).
Shlush, L. I. et al. Identification of pre-leukaemic haematopoietic stem cells in acute leukaemia. Nature 506, 328–333 (2014).
Klco, J. M. et al. Functional heterogeneity of genetically defined subclones in acute myeloid leukemia. Cancer Cell 25, 379–392 (2014).
Paguirigan, A. L. et al. Single-cell genotyping demonstrates complex clonal diversity in acute myeloid leukemia. Sci. Transl. Med. 7, 281re2 (2015).
Tefferi, A. & Vardiman, J. W. Classification and diagnosis of myeloproliferative neoplasms: the 2008 World Health Organization criteria and point-of-care diagnostic algorithms. Leukemia 22, 14–22 (2008).
Cheng, D. T. et al. Memorial Sloan Kettering-Integrated Mutation Profiling of Actionable Cancer Targets (MSK-IMPACT): a hybridization capture-based next-generation sequencing clinical assay for solid tumor molecular oncology. J. Mol. Diagn. 17, 251–264 (2015).
Proellochs, N. & Feuerriegel, S. ReinforcementLearning: Model-Free Reinforcement Learning. R package v.1.0.4 (2019).
Oksanen, J. et al. vegan: Community Ecology Package. R package v.2.5-6 (2019).
Griffith, D. M., Veech, J. A. & Marsh, C. J. cooccur: probabilistic species co-occurrence analysis in R. J. Stat. Softw. 69, 1–17 (2016).
Konopka, T. umap: Uniform Manifold Approximation and Projection. R package v.0.2.4.1 (2020).
Chen, H. Rphenograph: R implementation of the phenograph algorithm. R package v.0.99.1 (2015).
Wickham, H. ggplot2: Elegant Graphics for Data Analysis (Springer, 2016).
Gu, Z., Eils, R. & Schlesner, M. Complex heatmaps reveal patterns and correlations in multidimensional genomic data. Bioinformatics 32, 2847–2849 (2016).
Conway, J. R., Lex, A. & Gehlenborg, N. UpSetR: an R package for the visualization of intersecting sets and their properties. Bioinformatics 33, 2938–2940 (2017).
Csardi, G. & Nepusz, T. The igraph software package for complex network research. InterJournal Complex Syst. 2005, 1695 (2005).
Acknowledgements
We acknowledge the use of the MSKCC Integrated Genomics Core for all library sequencing, which is funded by MSKCC Support Grant NIH P30 CA008748. We thank members of the Levine laboratory for their critique of our work and assistance with revisions, and M. Roshal and W. Xiao for their input regarding AML stem and progenitor cell protein expression. L.A.M. is supported by a Career Development Program Fellowship of the Leukemia and Lymphoma Society (5479-19) and a Postdoctoral Fellowship from the MSKCC Marie-Josée Kravis Women in Science Endeavor (WiSE). R.L.B. is supported by the Sohn Foundation Fellowship of the Damon Runyon Cancer Research Foundation (DRG 22-17) and a National Cancer Institute grant (K99 CA248460). C.L.D. is supported by a Swiss National Science Foundation fellowship (grant no. 183853). K.L.B. is supported by grants including a National Institute of Health grant (K08 CA241318), an American Society of Hematology (ASH) grant, and an EvansMDS grant. A.D.V. is supported by the William Raveis Charitable Fund Fellowship of the Damon Runyon Cancer Research Foundation (DRG 117-15), an EvansMDS Young Investigator grant from the Edward P. Evans Foundation, and a National Cancer Institute career development grant (K08 CA215317). This work is supported by grants to S.E.M. including National Cancer Institute R37 CA226433, a Conquer Cancer Now Award from the Concern Foundation, and Sidney Kimmel Cancer Center (SKCC) Support Grant NIH P30 CA056036. This work was supported by grants to R.L.L. including a Cycle For Survival Innovation Grant, National Cancer Institute R35 CA197594, National Cancer Institute R01 CA173636, a grant from the Samuel Waxman Cancer Research Foundation, and SCOR grants from the Leukemia and Lymphoma Society.
Author information
Authors and Affiliations
Contributions
L.A.M., R.L.B., A.D.V., and R.L.L. conceptualized studies. L.A.M., R.L.B, A.T.O., R.D.-D., P.M., C.A., B.D., C.L.D., A.R.A., M.M., S.S., M.B., and R.L.L. designed and optimized experimental methodologies and bioinformatic workflow. L.A.M., R.L.B., T.R.M., I.S.C., and A.T.O. performed experiments. C.F., M.A.P., A.D.G., K.L.B., A.Z., R.R., M.P.C., S.E.M., A.D.V., and R.L.L. provided patient-associated resources and/or patient samples for use in the studies. L.A.M., R.L.B., and R.L.L wrote the original manuscript. L.A.M., R.L.B., S.E.M., and R.L.L assisted with review and editing of the manuscript. R.L.L supervised studies and manuscript preparation.
Corresponding author
Ethics declarations
Competing interests
L.A.M. and A.D.V. received travel support and honoraria from Mission Bio. A.T.O., R.D.-D., P.M., C.A., M.M., and S.S. are employed by Mission Bio and own equity in Mission Bio. A.R.A. is a cofounder and shareholder of Mission Bio. A.Z. has received honoraria from Illumina. M.P.C. has consulted for Janssen Pharmaceuticals. A.D.G. has served on advisory boards or as a consultant for AbbVie, Aptose, Celgene, Daiichi Sankyo, and Genentech, received research funding from AbbVie, ADC Therapeutics, Aprea, Aptose, AROG, Celularity, Daiichi Sankyo, and Pfizer, and received honoraria from Dava Oncology. R.R. has consulted for Constellation, Incyte, Celgene, Promedior, CTI, Jazz Pharmaceuticals, Blueprint, Stemline, Galecto, Pharmessentia, and Abbvie, and received research support from Incyte, Stemline, and Constellation. A.D.V. is on the Editorial Advisory Board of Hematology News. R.L.L. is on the supervisory board of QIAGEN and is a scientific advisor to Mission Bio, Loxo (until February 2019), Imago, C4 Therapeutics, and Isoplexis. He receives research support from and consulted for Celgene and Roche and has consulted for Lilly, Jubilant, Janssen, Astellas, Morphosys, and Novartis. He has received honoraria from Roche, Lilly, and Amgen for invited lectures and from Celgene and Gilead for grant reviews. R.L.B., T.R.M., I.S.C., C.F., M.A.P., M.B., B.D., C.L.D., K.B., and S.E.M. disclose no competing interests.
Additional information
Peer review information Nature thanks the anonymous reviewers for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Single-cell DNA-sequencing patient cohort.
a, Oncoprint of patient samples analysed by scDNA-seq. b, Table describing patient cohort characteristics. Standard deviation calculated for mean age of patients at sample collection date. Absolute number of samples denoted with percent of total samples in parentheses. c, Number of individual mutations identified for each gene covered on our custom amplicon panel by scDNA-seq (n = 146 biologically independent samples for c–f). Genes are ranked by the number of identified protein coding mutations from highest to lowest. Genes with zero identified mutations are not listed. d, Number of patients with protein coding mutations in a given gene. Genes are ranked by decreasing number of patients identified with mutations. e, Number of patients with a given number of identified mutant genes via single-cell sequencing. f, Number of patients with a given number of identified protein altering variants via single-cell sequencing. g, Correlation of bulk sequencing SNV data VAF versus single-cell SNV data VAF from MSKCC samples. Statistical significance was calculated by Pearson correlation coefficient. h, Violin plot of computed VAF from scDNA-seq for mutations found in both scDNA-seq and in bulk sequencing (identified; red), or mutations only identified in scDNA-seq (missed; blue) (top panel). Samples identified by scDNA-seq only were found to be low VAF mutations (P < 2.2 × 10−16; two-sample Mann–Whitney test). Bar plot of the number of new mutations in each sample identified by scDNA-seq only (bottom panel).
Extended Data Fig. 2 Analysis of clonal architecture by disease type and gene mutation.
a, scDNA-seq data processing and analysis workflow. FASTQ sequencing files for each sample were uploaded and processed through Mission Bio Tapestri Insights platform for variant calling and cell finding (Commercial Platform). Included samples for further analysis harboured ≥ 1 variant which leads to a protein sequence change (non-synonymous/insertion/deletion) and included 50 cells with definitive genotyping for all protein coding variants within the sample (n = 146). This data was used for analysis in Fig. 1. Clones present in each sample were identified and samples removed if they contained less than 2 clones for clonal analysis studies. Samples were subjected to random resampling of cells using a bootstrapping approach to identify the stability of identified clones (n = 132). Following bootstrapping, clones with lower 95% confidence intervals <10 were removed as were variants identified only within those clones. Samples which harboured only 1 variant or presented with <2 clones after bootstrapping analysis were removed (n = 111). The number of samples at each step of processing is shown below the different steps of the workflow. b, Number of mutations in the most dominant clone identified in each sample (n = 111 biologically independent samples) stratified by cohort. Mean value for each cohort shown by height of bar with standard error of measurement (SEM) depicted with error bars. A two-sided t-test with FDR correction was used to determine statistical significance pairwise between all groups. For clarity, only significant P values referenced in the text are shown. *P < 0.1; **P < 0.01; ***P < 0.001. c, Association between clone size and the number of mutant alleles in the clone. Every clone (n = 111 biologically independent samples) identified in clinical cohort is depicted by black circle. Centre line: median; box: IQR; whiskers 1.5 × IQR. d, Bar plot depicting the prevalence of dominant clones for each DTAI gene across patient cohorts. Colour of bar plot annotates if mutation occurs in the dominant clone (red) or subclone (grey). Absence of bar denotes no clones were identified with the indicated mutation in a given cohort. e, Association of VAF with presence of mutation in either the dominant clone (red) or subclone (grey) for select genes (n = 101 biologically independent samples). Standard error of measurement depicted with error bars. A two-sided t-test with FDR correction was used to determine statistical significance pairwise between all groups. *P < 0.1; **P < 0.01; ***P < 0.001. Absence of P value for IDH2 and JAK2 due to lack of samples with subclonal mutations. f, Pairwise interaction matrix of mutually exclusive (red square) and inclusive (blue square) on a per-sample basis. Pairwise interactions with no colour did not garner a significant P value.
Extended Data Fig. 3 Clonal dominance, initiating mutation, and co-mutation patterns in patients with myeloid malignancies.
a, Upset plot of co-occurring DTAI mutations in CH samples with more than one DTAI variant. Bar graph (top panel) depicts the number of samples with each mutant gene(s) and colour of bar annotating whether mutation(s) occur in the dominant clone (red) or subclones (grey). Black circles and connecting line in bottom panel demark the combination of mutations in each corresponding bar plot. b, Divergent frequency of co-mutated cells for epigenetic modifier genes (red) and signalling genes (blue). Individual samples (n = 6 samples) shown with black square. Centre line: median; box: IQR; whiskers 1.5 × IQR. A two-sided Student’s t-test was used to determine statistical significance *P < 0.1; **P < 0.01; ***P < 0.001. c, Fraction of mutant samples harbouring a homozygous mutation for the indicated given gene (at least >10% of cells). Homozygous sample denoted in blue. d, Correlation of VAF computed by scDNA sequencing to fraction of a mutant sample explained by the genetic trajectory starting with an initiating mutation in a given gene. Genes used as the initiating mutation for a given sample are denoted by colored squares (colours described in figure). Statistical significance calculated by Spearman’s rank correlation coefficient test (ρ = 0.93; P ≤ 2.2 × 10−16). e, Number of samples where a monoallelic clone for a given gene is observed. Dark blue denotes total number of mutant samples where single-mutant clone is present for a given gene and grey represents mutant samples where single-mutant clone is unobserved. f, Number of DNMT3A mutant samples where single-mutant clones are observed (red) or unobserved (grey) with samples categorized by DNMT3A R882 hotspot mutations, nonsense mutations, or missense mutations. A two-sided Fisher’s exact test was used to determine statistical significance (P ≤ 0.04) between DNMT3AR882 and other missense mutations. g, Differences in dominant and subclone size in DNMT3A mutant samples (n = 61 biologically independent clones). Fraction of sample in the dominant clone or subclone(s) for DNMT3A nonsense (red), R882-missense (green), and non-R882 missense (blue) mutations shown. Centre line: median; box: IQR; whiskers 1.5 × IQR. Each mutant clone denoted by black square. A two-sided t-test correction was used to determine statistical significance pairwise between all groups. For clarity, only significant P values referenced in the text are shown. *P < 0.1. h, As in Fig. 3e, fraction of sample in single- and double-mutant clones in DNMT3A/IDH2 mutant samples. Each sample is indicated by a connecting line, absence of a line for single mutants indicates absence of clone.
Extended Data Fig. 4 Clonal evolution in patients with myeloid malignancies.
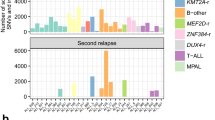
a, Paired samples from patients (n = 6) that underwent MPN to AML transformation were analysed. Samples with significant changes in clonal architecture or clonal sweeps were evaluated using a two-sided two proportions z-test; ***P < 0.001. Sample A (red) denotes the MPN sample and sample B (blue) denotes the AML sample. Clonotype plot depicts the frequency of a clone with given genotype in Sample A and B ranked by decreasing frequency based on Sample A (top panel). Heat map (bottom panel) shows the genotype of each identified protein coding mutation in the given clone with zygosity (wild type = light pink, heterozygous = orange, homozygous = red). Paired samples MSK75/76 are highlighted in Fig. 3f. b, Clonal sweeps, or significant clonal architecture alterations, following gilteritinib therapy of FLT3-mutant patients (n = 3). Line graphs for each pair of samples depict individual clones and the change in clone frequency between pre- (left) and post- (right) therapy samples. Clones harbouring FLT3 mutations (red), RAS mutations (blue), or wild-type (WT) clones (light blue) are significantly altered after gilteritinib therapy in each patient. FLT3–RAS mutations (orange) and clones harbouring additional mutations (Other; grey) are also included. Statistical significance was assessed using a two-sided two proportions z-test; ***P < 0.001 (a, b). c, As in a, clonotype plot of paired sample (n = 1 sample/time point) from a patient with AML (MSK95/96) who underwent gilteritinib therapy: sample A (red, pre-therapy) and sample B (blue, post-therapy).
Extended Data Fig. 5 Contribution of clonal haematopoiesis mutations to mature cell lineages.
Bar graphs of the mutant cell percentage found in myeloid (CD11b high; green), B (CD19 high; orange), and T (CD3 high; purple) cells in samples from patients with CH. DNMT3A and/or TET2 mutations found in each sample are listed above each graph. Double-mutant samples are shown on the left and single-mutant samples are depicted on the right.
Extended Data Fig. 6 Simultaneous molecular and immunophenotypic profiling of samples from patients with AML.
a, UMAP plot of MSK54 with cells clustered by immunophenotype. Genotype (wild type = grey; DNMT3A = red; IDH2 = green; DNMT3A/IDH2 double mutant = blue) overlaid onto each cell. b, UMAP from a with protein expression (high expression = red; low expression = blue) for each of the six antibody targets (CD3, CD11b, CD34, CD38, CD45RA, CD90) overlaid onto each cell. Relative protein expression is normalized across individual samples by CLR. c, Immunophenotype changes based on co-occurring mutations in clones. Heat map of normalized protein expression of CD34 (top panel) and CD11b (bottom panel) in DNMT3A and IDH1/2 single-mutant clones versus DNMT3A and IDH1/2 mutant clones with co-occurring NRAS or FLT3 mutations. High protein expression depicted in red and low protein expression depicted in blue.
Extended Data Fig. 7 Clonal architecture analysis using single-cell DNA+Protein sequencing of select AML samples.
Samples shown have significant differences in community representation between the dominant clone and subclones further discussed in Extended Data Fig. 8. MSK71 (depicted with ***) is highlighted in Fig. 4c–f. Clonotype plot depicts the number of cells identified with a given genotype and ranked by decreasing frequency (top panel). Mean cell counts for each clone are depicted with 95% confidence intervals derived from random resampling analysis. Heat map (middle panel) shows the genotype of each identified protein coding mutation in the given clone with zygosity (wildtype = light pink, heterozygous = orange, homozygous = red). Heat map of the relative protein expression for each cell-surface protein (n = 7) in each identified clone (purple = high expression; green = low expression).
Extended Data Fig. 8 Neighbourhood analysis of all single-cell DNA+Protein AML samples.
a, Divergences in cell-surface protein expression of CD34, CD38, CD11b, and CD45RA determined by presence of signalling effector mutation. Density plots of cells from MSK71 (further detailed in Figure 4c–f and Extended Data Figure 7) of DNMT3A mutant cells (yellow = single-mutant) with co-occurring FLT3 (black), KRAS (orange), or NRAS (light blue) mutations. Concentration of cells with a given immunophenotype depicted by the density of lines. b, UMAP plot of samples (n = 17) analysed by DNA+Protein single-cell sequencing with cells clustered by cell-surface protein expression of 6 antibody targets (CD3, CD11b, CD34, CD38, CD45RA, CD90). Cells from the same sample are denoted with same colour. c, Neighbourhood analysis of all samples from UMAP from b with communities of cells identified by neighbourhood analysis in overlaid colours.
Extended Data Fig. 9 Clone- and gene- specific alterations to cell-surface protein expression and community representation in AML samples.
a, Column normalized heat map of cell-surface protein expression for each community identified in phenoGraph analysis on UMAP from Extended Data Figure 8b, c. Expression is depicted by colour with blue being low expression and red annotating high expression. b, Community representation changes across all samples (n = 14) in the wild type, the dominant clone, and all subclones. The fraction of each sample within each community is shown with communities depicted by corresponding colour. Samples without communities shown for wild-type cells were found to not have any wild-type cells present in analysis. Changes in immunophenotype due to community representation changes for samples MSK94 (P ≤ 9.95 × 10−3) and MSK130 (P ≤ 2.45 × 10−8) are highlighted in c. A two proportions z-test for each sample was used to determine statistical significance between dominant clone communities and communities present in subclone ***P < 0.001. c, Cell-surface protein expression of CD11b, CD34, and CD38 between dominant clone (red) and subclones (black) in an FLT3-ITD mutant sample (MSK130; right panel; n = 2274 total cells) and JAK2 mutant sample (MSK94; left panel; n = 6012 total cells). Each error bar represents a distinct community that is significantly expanded or contracted, (error bar indicates ± standard error of measure, from the mean expression of indicated protein in a given community). A Student’s t-test was used to determine statistical significance *P < 0.1; **P < 0.01; ***P < 0.001.
Supplementary information
Supplementary Information
This file contains a Supplementary Discussion.
Supplementary Table
Supplementary Table 1. Amplicon Coverage for Custom Mission Bio single cell DNA sequencing panel. Each amplicon is listed with genomic location (hg19) and gene name.
Supplementary Table
Supplementary Table 2. Sample characteristics from clinical cohort. Patient information from date of sample collection and sample characteristics for each sample are denoted.
Rights and permissions
About this article
Cite this article
Miles, L.A., Bowman, R.L., Merlinsky, T.R. et al. Single-cell mutation analysis of clonal evolution in myeloid malignancies. Nature 587, 477–482 (2020). https://doi.org/10.1038/s41586-020-2864-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-020-2864-x
This article is cited by
-
Decoding leukemia at the single-cell level: clonal architecture, classification, microenvironment, and drug resistance
Experimental Hematology & Oncology (2024)
-
Uncovering the cellular and omics characteristics of natural killer cells in the bone marrow microenvironment of patients with acute myeloid leukemia
Cancer Cell International (2024)
-
Mutation order in acute myeloid leukemia identifies uncommon patterns of evolution and illuminates phenotypic heterogeneity
Leukemia (2024)
-
Tissue mosaicism following stem cell aging: blood as an exemplar
Nature Aging (2024)
-
Genetic variation across and within individuals
Nature Reviews Genetics (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.