Abstract
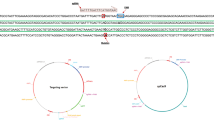
Angelman syndrome (AS) is a severe neurodevelopmental disorder caused by a mutation or deletion of the maternally inherited UBE3A allele. In neurons, the paternally inherited UBE3A allele is silenced in cis by a long non-coding RNA called UBE3A-ATS. Here, as part of a systematic screen, we found that Cas9 can be used to activate ('unsilence') paternal Ube3a in cultured mouse and human neurons when targeted to Snord115 genes, which are small nucleolar RNAs that are clustered in the 3′ region of Ube3a-ATS. A short Cas9 variant and guide RNA that target about 75 Snord115 genes were packaged into an adeno-associated virus and administered to a mouse model of AS during the embryonic and early postnatal stages, when the therapeutic benefit of restoring Ube3a is predicted to be greatest1,2. This early treatment unsilenced paternal Ube3a throughout the brain for at least 17 months and rescued anatomical and behavioural phenotypes in AS mice. Genomic integration of the adeno-associated virus vector into Cas9 target sites caused premature termination of Ube3a-ATS at the vector-derived polyA cassette, or when integrated in the reverse orientation, by transcriptional collision with the vector-derived Cas9 transcript. Our study shows that targeted genomic integration of a gene therapy vector can restore the function of paternally inherited UBE3A throughout life, providing a path towards a disease-modifying treatment for a syndromic neurodevelopmental disorder.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
All data generated or analysed during this study are included in this published article (and its Supplementary information files).
References
Silva-Santos, S. et al. Ube3a reinstatement identifies distinct developmental windows in a murine Angelman syndrome model. J. Clin. Invest. 125, 2069–2076 (2015).
Zylka, M. J. Prenatal treatment path for Angelman syndrome and other neurodevelopmental disorders. Autism Res. 13, 11–17 (2020).
Huang, H. S. et al. Topoisomerase inhibitors unsilence the dormant allele of Ube3a in neurons. Nature 481, 185–189 (2011).
Meng, L. et al. Truncation of Ube3a-ATS unsilences paternal Ube3a and ameliorates behavioral defects in the Angelman syndrome mouse model. PLoS Genet. 9, e1004039 (2013).
Zhu, S. et al. Genome-scale deletion screening of human long non-coding RNAs using a paired-guide RNA CRISPR–Cas9 library. Nat. Biotechnol. 34, 1279–1286 (2016).
Dindot, S. V., Antalffy, B. A., Bhattacharjee, M. B. & Beaudet, A. L. The Angelman syndrome ubiquitin ligase localizes to the synapse and nucleus, and maternal deficiency results in abnormal dendritic spine morphology. Hum. Mol. Genet. 17, 111–118 (2008).
Bortolin-Cavaillé, M. L. & Cavaillé, J. The SNORD115 (H/MBII-52) and SNORD116 (H/MBII-85) gene clusters at the imprinted Prader–Willi locus generate canonical box C/D snoRNAs. Nucleic Acids Res. 40, 6800–6807 (2012).
de Smith, A. J. et al. A deletion of the HBII-85 class of small nucleolar RNAs (snoRNAs) is associated with hyperphagia, obesity and hypogonadism. Hum. Mol. Genet. 18, 3257–3265 (2009).
Bieth, E. et al. Highly restricted deletion of the SNORD116 region is implicated in Prader–Willi syndrome. Eur. J. Hum. Genet. 23, 252–255 (2015).
Anderlid, B. M., Lundin, J., Malmgren, H., Lehtihet, M. & Nordgren, A. Small mosaic deletion encompassing the snoRNAs and SNURF-SNRPN results in an atypical Prader–Willi syndrome phenotype. Am. J. Med. Genet. A 164A, 425–431 (2014).
Hsiao, J. S. et al. A bipartite boundary element restricts UBE3A imprinting to mature neurons. Proc. Natl Acad. Sci. USA 116, 2181–2186 (2019).
Stein, J. L. et al. A quantitative framework to evaluate modeling of cortical development by neural stem cells. Neuron 83, 69–86 (2014).
Ran, F. A. et al. In vivo genome editing using Staphylococcus aureus Cas9. Nature 520, 186–191 (2015).
Friedland, A. E. et al. Characterization of Staphylococcus aureus Cas9: a smaller Cas9 for all-in-one adeno-associated virus delivery and paired nickase applications. Genome Biol. 16, 257 (2015).
Bengtsson, N. E. et al. Muscle-specific CRISPR/Cas9 dystrophin gene editing ameliorates pathophysiology in a mouse model for Duchenne muscular dystrophy. Nat. Commun. 8, 14454 (2017).
Avagliano Trezza, R. et al. Loss of nuclear UBE3A causes electrophysiological and behavioral deficits in mice and is associated with Angelman syndrome. Nat. Neurosci. 22, 1235–1247 (2019).
Johnston, S. et al. AAV ablates neurogenesis in the adult murine hippocampus. Preprint at https://doi.org/10.1101/2020.01.18.911362 (2020).
Kishore, S. & Stamm, S. The snoRNA HBII-52 regulates alternative splicing of the serotonin receptor 2C. Science 311, 230–232 (2006).
Doe, C. M. et al. Loss of the imprinted snoRNA mbii-52 leads to increased 5htr2c pre-RNA editing and altered 5HT2CR-mediated behaviour. Hum. Mol. Genet. 18, 2140–2148 (2009).
Sonzogni, M. et al. A behavioral test battery for mouse models of Angelman syndrome: a powerful tool for testing drugs and novel Ube3a mutants. Mol. Autism 9, 47 (2018).
Mandel-Brehm, C., Salogiannis, J., Dhamne, S. C., Rotenberg, A. & Greenberg, M. E. Seizure-like activity in a juvenile Angelman syndrome mouse model is attenuated by reducing Arc expression. Proc. Natl Acad. Sci. USA 112, 5129–5134 (2015).
Hanlon, K. S. et al. High levels of AAV vector integration into CRISPR-induced DNA breaks. Nat. Commun. 10, 4439 (2019).
Nelson, C. E. et al. Long-term evaluation of AAV-CRISPR genome editing for Duchenne muscular dystrophy. Nat. Med. 25, 427–432 (2019).
von Bartheld, C. S., Bahney, J. & Herculano-Houzel, S. The search for true numbers of neurons and glial cells in the human brain: a review of 150 years of cell counting. J. Comp. Neurol. 524, 3865–3895 (2016).
Friedel, R. H. & Soriano, P. Gene trap mutagenesis in the mouse. Methods Enzymol. 477, 243–269 (2010).
Landers, M. et al. Maternal disruption of Ube3a leads to increased expression of Ube3a-ATS in trans. Nucleic Acids Res. 33, 3976–3984 (2005).
King, I. F. et al. Topoisomerases facilitate transcription of long genes linked to autism. Nature 501, 58–62 (2013).
Sanjana, N. E., Shalem, O. & Zhang, F. Improved vectors and genome-wide libraries for CRISPR screening. Nat. Methods 11, 783–784 (2014).
Mabb, A. M. et al. Topoisomerase 1 regulates gene expression in neurons through cleavage complex-dependent and -independent mechanisms. PLoS One 11, e0156439 (2016).
Dickerson, A. S. et al. Autism spectrum disorder prevalence and associations with air concentrations of lead, mercury, and arsenic. Environ. Monit. Assess. 188, 407 (2016).
Kishore, S. et al. The snoRNA MBII-52 (SNORD 115) is processed into smaller RNAs and regulates alternative splicing. Hum. Mol. Genet. 19, 1153–1164 (2010).
Walantus, W., Castaneda, D., Elias, L. & Kriegstein, A. In utero intraventricular injection and electroporation of E15 mouse embryos. J. Vis. Exp. 239, 239 (2007).
Kim, J. Y., Grunke, S. D., Levites, Y., Golde, T. E. & Jankowsky, J. L. Intracerebroventricular viral injection of the neonatal mouse brain for persistent and widespread neuronal transduction. J. Vis. Exp. 91, e51863 (2014).
Bae, S., Park, J. & Kim, J. S. Cas-OFFinder: a fast and versatile algorithm that searches for potential off-target sites of Cas9 RNA-guided endonucleases. Bioinformatics 30, 1473–1475 (2014).
Martin, M. CUTADAPT removes adapter sequences from high-throughput sequencing reads. EMBnet.journal 17, 10–12 (2011).
Blazie, S. M. et al. Alternative polyadenylation directs tissue-specific miRNA targeting in Caenorhabditis elegans somatic tissues. Genetics 206, 757–774 (2017).
Acknowledgements
We thank E. McCoy, G. Salazar, E. Hopkins, T. Ptacek and B. Taylor-Blake for technical assistance; and the UNC Catalyst for Rare Diseases for use of their high-throughput screening equipment. This work was supported by grants to M.J.Z. from the Angelman Syndrome Foundation, the Simons Foundation (SFARI, award ID 631904), the National Institute of Neurological Disorders and Stroke (NINDS; 1R01NS109304-01A1) and the Eshelman Institute for Innovation. J.M.W. was supported by grants from the National Institute for Child Health and Human Development (NICHD; T32HD040127) and a Pfizer-NCBiotech Distinguished Postdoctoral Fellowship in Gene Therapy. H.M. was supported by the NICHD (T32HD040127). J.L.S. was supported by grants from the National Institute of Mental Health (R01MH118349, R00MH102357 and R01MH120125). The microscopy core and J.M.S. in the bioinformatics core were supported by the NICHD (P50HD103573) and the NINDS (P30NS045892). The UNC Flow Cytometry Core Facility is supported in part by the National Cancer Institute (P30CA016086), awarded to the UNC Lineberger Comprehensive Cancer Center. The UNC Mouse Behavioral Phenotyping Core is supported by the NICHD (P50HD103573).
Author information
Authors and Affiliations
Contributions
J.M.W., G.F. and M.J.Z. conceived the study. J.M.S. provided bioinformatic support for gRNA design. J.M.W. cloned the gRNA library, performed all of the experiments in mouse primary neuron cultures, phNPCs and RT–PCR from cortical samples, and analysed behavioural data. J.M.W. and G.F. performed and analysed the CRISPR screen in primary neuron cultures. H.M. performed all viral injections, behavioural experiments, western blots and histology. H.O.B. performed vDNA tropism experiments. J.L.K. and J.M.W. performed the vDNA integration experiments. J.L.K. performed the DNA amplicon sequencing experiments. B.O., J.M.W. and J.M.S. analysed the data. J.L.S. provided reagents and protocols for performing human neural progenitor culture and differentiation. J.M.W. and M.J.Z. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
M.J.Z. serves as a consultant to AskBio, to which technologies evaluated in this paper have been licensed. J.M.W., H.M., G.F., J.M.S. and M.J.Z. are inventors of the technology and could receive royalties. These relationships have been disclosed to and are under management by UNC-Chapel Hill. The remaining authors have no competing interests.
Additional information
Peer review information Nature thanks Jeremy Day, Fyodor Urnov, Charles Williams and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Genomic map of Spjw33 targets and functional outcomes on Ube3a and Snord115 locus.
a, Mouse genome browser view. b, Zoom in showing four Spjw33 target sites. Snord115 genes (blue peaks) are conserved (phyloP track) relative to other regions of Ube3a-ATS. c, Systematic testing of pairs of gRNAs in Ube3am+/pYFP primary cortical neurons. Top panel: locations of gRNAs positioned to selectively target the Snord115 cluster (gRNA sets A and B), the Snord116 cluster (gRNA sets B and C), or the Ube3a-ATS promoter (gRNA sets A/C and D). Bottom panel: Percentage patYFP+/Cas9+ neurons for each pair of gRNAs relative to the positive control (SpCas9+Spjw33 alone). d, Location of primers used to quantify transcript expression and discriminate between maternal Ube3a and paternal Ube3a-YFP alleles. Expected band size indicated. e, Agarose gel showing RT–PCR products amplified from Ube3am+/pYFP cortical neuron cultures. All bands are of the expected size. f, g, Expression (qPCR) of maternal or paternal Ube3a alleles in cortical neuron cultures transduced with lentivirus carrying SpCas9 and a gRNA targeting human SNORD115 (neg. control), Spjw33 (f), or treated with topotecan (g). Data normalized to Eif4a2, n = 3. h, i, Expression of the indicated genes in wild-type cortical neuron cultures treated with topotecan (h), or lentivirally transduced with SpCas9 and a gRNA targeting human SNORD115 (neg. control) (i). Dashed line marks vehicle (h) or neg. control gRNA (i) expression levels. Normalized to Eif4a2, n = 3. *P < 0.01. j, Ube3am+/pYFP neurons transfected with gRNAs containing the indicated number of base mismatches relative to Spjw33. n = 4, *P < 0.05. k, Table summarizing mutations identified at the Spjw33 target site. Primary cortical neurons were lentivirally transduced with SpCas9 and either a neg. control gRNA or Spjw33. Genomic DNA spanning the Spjw33 target site was PCR amplified and individual clones were analysed by Sanger sequencing. Red arrow marks SpCas9 cleavage site. Red letters denote mutations that were introduced by SpCas9, based on the observation that these mutations were not found in neurons treated with the neg. control gRNA nor were they found in other Snord115 genes (mm9 genome build). l, Percentage YFP colocalization in neurons transfected with SpCas9 and gRNAs targeting the indicated number of genomic sites in the Snord115 locus.
Extended Data Fig. 2 Targeting SNORD115 unsilences paternal UBE3A in human neurons.
a, Alignment of RNA-seq reads from a phNPC line before (top panel) and after 8 week neuronal differentiation (bottom panel). Coloured lines mark polymorphisms not present in the reference genome. b, Zoom in of the region between dashed lines in a. Arrow denotes SNV (rsID:61734190) used to quantify allele specific expression in o, p. c, Allelic expression of UBE3A and UBE3A-ATS before and after differentiation of phNPCs into neurons, calculated as the percentage of stranded RNA-seq reads containing a SNV at chr15:25,371,697. d, Guide RNA locations in the human SNORD115 cluster. e, Experimental design for transducing differentiated human neurons with lentiviruses containing SpCas9 and gRNAs. f, Sorting strategy to isolate differentiated neuron populations from non-neuronal cells. Neurons were labelled and identified using AAV2:hSyn1:EGFP (green), cells lentivirally transduced with Cas9 were identified based on mCherry (red = non-neuronal cells, green/yellow = neurons). Side scatter counts (SSC), forward scatter counts (FSC). g–n, Expression of the indicated genes in different cell populations using qPCR and SYBR green, normalized to EIF4A2. Values on Y-axis are presented as ∆Ct values (not normalized to a specific sample), so expression values can be directly compared across multiple genes. n = 3 for each qPCR reaction. o, Raw qPCR curves using allele specific TaqMan probes (rsID:61734190) from RNA extracted from populations of progenitors and neurons containing the indicated fluorescent markers. Spjw33 negative control (mouse-specific). Presumed maternal (T; red line) and paternal (A; blue line) allele. ∆Rn is change in reporter dye intensity per cycle, normalized to passive reference dye. One line per replicate, n = 4. p, Allele specific expression after lentiviral transduction of human neurons with SpCas9 and the indicated gRNAs, relative to neurons from the same cultures that were not transduced with Cas9. Number in parentheses refers to the number of gRNA target sites in SNORD115 locus. q, Expression of the indicated genes from Cas9+ neurons transduced with gRNA:hsa#3 versus Cas9- human neurons (dashed line). qPCR, normalized to EIF4A2, n = 3. *P < 0.01.
Extended Data Fig. 3 SaCas9 vector development and testing.
a, Sequence alignment between Spjw33 and Sajw33 gRNA and genomic target site. Underlined sequence marks a portion of Snord115. Red arrow marks Cas9 cleavage site. b, Percentage of UBE3A-YFP+ neurons from Ube3am+/pYFP cortical neurons cultures transiently transfected with SpCas9 or SaCas9 and their respective gRNAs; relative to neg. control gRNAs and relative to topotecan treatment (300 nm, 72 h). c, Map showing SaCas9 expression cassette in AAV backbone (not to scale). Human synapsin-1 promoter (hSyn1), simian virus 40 (SV40) nuclear localization sequence (NLS), nucleoplasmin (NP), hemagglutinin tag (HA), bovine growth hormone polyadenylation sequence element (bGH polyA), U6 promoter. d, Immunofluorescence for indicated proteins in the cerebral cortex of P30 mice injected i.c.v. with AAV9 SaCas9:Sajw33 at E15.5 or P1 (bilateral 1.5 × 1010 AAV particles per ventricle). Note bias for deeper layer neurons when AAV was injected at E15.5 and bias for upper layer neurons when AAV was injected at P1.
Extended Data Fig. 4 AAV delivery of SaCas9 and Sajw33 unsilences paternal Ube3a with no detectable AAV mediated toxicity.
a–c, Histological staining at P30 for UBE3A and NEUN in the cortex, hippocampus, and cerebellum of P30 Ube3am-/p+ mice, injected at E15.5+P1 with AAV SaCas9 vector containing neg. control gRNA (a) or Sajw33 (b). Zoom-in view shows UBE3A protein in neurons (NEUN+) (c). In a and b, cortex and hippocampus, scale bar, 200 μm. cerebellum, scale bar, 100 μm. In c, scale bar, 50 μm. d, e, Western blot quantification of UBE3A levels in the cortex (cor.), hippocampus (hip.), and cerebellum (cblm.) of P30 Ube3am-/p+ (AS) mice treated with neg. control gRNA or Sajw33 (n = 3 per group; dual E15.5+P1 injections). WT = wild-type mice, age P30. *P < 0.05, **P < 0.01. f, g, Histological staining at P90 for UBE3A and NEUN in the cortex, hippocampus, and cerebellum of Ube3am-/p+ mice, injected at E15.5+P1 with AAV SaCas9 vector containing neg. control gRNA (f) or Sajw33 (g). White boxes are regions shown in Fig. 2a, b. Cortex and hippocampus, scale bar, 200 μm. Cerebellum and spinal cord, scale bar, 100 μm. h–k, Representative images of hippocampus from indicated genotypes and treatments at P30 from untreated or E15.5/P1 i.c.v. injected embryos. Immunofluorescence for progenitors (TBR2), immature neurons (DCX), astrocytes (GFAP), and microglia (IBA1). l, m, Quantification of TBR2+ (l) and IBA1+ (m) cells showed no significant difference of cell numbers in dentate gyrus among indicated genotype and treatment groups. n, Representative images of dorsal cortices stained for microglia (IBA1) of indicated genotypes and treatments at P30. o, Quantification of IBA1+ cells in a 200 μm column of each dorsal cortex imaged. n = 3 animals per genotype with treatment with 2 imaged sections each. Scale bar, 100 μm. n.s., not significant.
Extended Data Fig. 5 Expression and alternative splicing of Snord115 target genes are not affected in the brain.
a, RT–qPCR for Snord115 from cortex of P60 mice either untreated or dual injected (i.c.v.) at E15.5+P1 with AAV carrying SaCas9 and Sajw33. b–f, Expression of Snord115 target genes in cortex of P60 mice either untreated or dual injected (i.c.v.) at E15.5+P1 with AAV carrying SaCas9 and Sajw33. g, RT–PCR for Snord115 target genes with published primers31. Total RNA extracted from cortex of P60 mice dual injected at E15.5+P1 (i.c.v.) with AAV carrying SaCas9 and Sajw33. h, Quantification of alternate splicing.
Extended Data Fig. 6 Single injection of AAV containing SaCas9 and Sajw33 at P1 enduringly unsilences paternal UBE3A in 17-month-old mice.
Representative images from three 17-month-old Ube3am-/p+ animals treated with AAV9 carrying SaCas9 and Sajw33 at P1 through i.c.v. injection (bilateral 1.5 × 1010 AAV particles per ventricle). Brains stained for UBE3A (green) showed extensive unsilencing in neurons (NEUN, magenta). Zoom in images of indicated region in hippocampus CA1 (a), dentate gyrus (b), and cerebellum (c).
Extended Data Fig. 7 Evaluation of viral transduction in tissues from injected mice and their dams.
a, qPCR quantification of viral DNA (SaCas9 TaqMan probes) in cortex and liver of age matched untreated animals, P60 dams whose pups were injected i.c.v. at E15.5, and P60 mice that were dual injected i.c.v. at E15.5+P1. Data normalized to Eif4a2 (representing a gene with 2 copies per diploid genome). Limit of detection (LOD) determined by performing serial dilutions with known quantities of AAV particles spiked into gDNA samples from untreated mice. *P < 0.05, **P < 0.01, ***P < 0.001. b, Expression of SaCas9 mRNA in cortex and liver from same animals as a. ∆∆Ct method, normalized to β-actin.
Extended Data Fig. 8 Additional behavioural assays.
a, Body weight of male mice measured monthly over 9 months. b, c, Open field data at 11 weeks of age, first 0-15 min of experiment. Time spent in centre in seconds (b), distance travelled in meters (c). *P < 0.05, **P < 0.01, ***P < 0.001. d, e, Rotarod training data (average of three trials) at 8 weeks of age (d) and 28 weeks of age (e). f, g, Contextual (f) and cue-based (g) learning at 18 weeks of age. No statistically significant phenotypes were observed.
Extended Data Fig. 9 Analysis of mutations and AAV integration in 10-month-old mice.
a, Analysis of mutations found in Sajw33 treated mice by high-throughput gDNA amplicon sequencing (no mutations found in controls). Capital letters denote Sajw33 target site, red arrow refers to SaCas9 cleavage site. Red nucleotides represent nucleotides not present in AS animals treated with neg. gRNA nor in the reference genome (mm9). b, Percentage of specific mutation types identified in 10-month-old Sajw33 treated mice (n = 3). c, qPCR from gDNA isolated from 10-month-old animals of the indicated genotypes. Data normalized to a region of Ube3a-ATS which contains two copies per diploid genome by ∆∆Ct method. d, PCR strategy to detect AAV integration events. e–j, All DNA/RNA extractions performed from cortices of 10-month-old mice, injected at E15.5+P1 with AAV containing SaCas9 and the negative control gRNA or Sajw33. e, PCR of genomic DNA with the indicated primers. f, g, DNA amplicon sequencing of AAV integration events, detailing the position in the AAV ITR or hSyn1 promoter that was immediately adjacent to endogenous gDNA at the Sajw33 target site. h, gDNA PCR at predicted Sajw33 off target sites using primers to interrogate AAV integration in both orientations. SaCas9 and Snord115/AAV integrations are positive controls. PCR was performed for 40 cycles. The absence of bands suggests no AAV integration at the top 10 predicted off-target sites. i, qPCR of genomic DNA. Forward primer anneals in Snord115 forward orientation, reverse primer anneals to both viral inverted terminal repeats (ITRs) to quantify the number of AAV integration events in the genome irrespective of orientation. *P < 0.05. j, RT–PCR with primers specific for the indicated genes/gene fusion. cDNA synthesized with random hexamers, no reverse transcriptase control (-RT) with Ube3a-ATS/AAV fusion primer pair.
Extended Data Fig. 10 AAV integration disrupts Ube3a-ATS via distinct mechanisms.
a, AAV integration in the forward orientation gene traps Ube3a-ATS, causing premature transcription termination at the AAV vector-derived polyadenylation sequence element (red box). b, In the reverse orientation, convergent transcription with the AAV vector-derived Cas9 transcript disrupts Ube3a-ATS. c, Paternal Ube3a (red line) is similarly disrupted (“silenced”) by convergent transcription.
Supplementary information
Supplementary Information
This file contains the Supplementary Discussion.
Supplementary Figure 1
Original confocal images used in Fig. 2. P90 Ube3am-/p+ animals treated with either control gRNA (a-e) or Sajw33 (f-j). Whole brain sagittal sections (a and f) and spinal cords (e and j) were imaged at 10x. The cortices (b and g), hippocampus (c and h), and cerebellum (Cblm, d and i) were imaged at 20x. All sections were stained for UBE3A (green), NEUN (grey), and DAPI (blue).
Supplementary Figure 2
Raw western blot data related to Fig. 2.
Supplementary Table 1
gRNA library and screen results.
Supplementary Table 2
gDNA mutation analysis.
Supplementary Table 3
Pathology report.
Supplementary Table 4
Morphological and behavioural data.
Supplementary Table 5
Off target amplicon sequencing.
Supplementary Table 6
A list of primers.
Rights and permissions
About this article
Cite this article
Wolter, J.M., Mao, H., Fragola, G. et al. Cas9 gene therapy for Angelman syndrome traps Ube3a-ATS long non-coding RNA. Nature 587, 281–284 (2020). https://doi.org/10.1038/s41586-020-2835-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-020-2835-2
This article is cited by
-
Transcription regulation by long non-coding RNAs: mechanisms and disease relevance
Nature Reviews Molecular Cell Biology (2024)
-
Chromatinopathies: insight in clinical aspects and underlying epigenetic changes
Journal of Applied Genetics (2024)
-
Advances in CRISPR therapeutics
Nature Reviews Nephrology (2023)
-
Techniques for investigating lncRNA transcript functions in neurodevelopment
Molecular Psychiatry (2023)
-
Therapeutic strategies for autism: targeting three levels of the central dogma of molecular biology
Translational Psychiatry (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.