Abstract
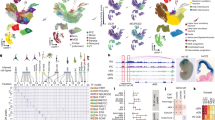
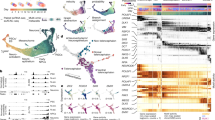
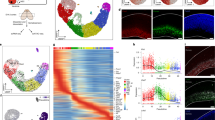
Lineage-specific epigenomic changes during human corticogenesis have been difficult to study owing to challenges with sample availability and tissue heterogeneity. For example, previous studies using single-cell RNA sequencing identified at least 9 major cell types and up to 26 distinct subtypes in the dorsal cortex alone1,2. Here we characterize cell-type-specific cis-regulatory chromatin interactions, open chromatin peaks, and transcriptomes for radial glia, intermediate progenitor cells, excitatory neurons, and interneurons isolated from mid-gestational samples of the human cortex. We show that chromatin interactions underlie several aspects of gene regulation, with transposable elements and disease-associated variants enriched at distal interacting regions in a cell-type-specific manner. In addition, promoters with increased levels of chromatin interactivity—termed super-interactive promoters—are enriched for lineage-specific genes, suggesting that interactions at these loci contribute to the fine-tuning of transcription. Finally, we develop CRISPRview, a technique that integrates immunostaining, CRISPR interference, RNAscope, and image analysis to validate cell-type-specific cis-regulatory elements in heterogeneous populations of primary cells. Our findings provide insights into cell-type-specific gene expression patterns in the developing human cortex and advance our understanding of gene regulation and lineage specification during this crucial developmental window.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
All datasets used in this study (PLAC-seq, ATAC-seq, RNA-seq) are available at the Neuroscience Multi-Omic Archive (NeMO Archive) under controlled access. Chromatin interactions, open chromatin peaks, and gene expression profiles for each cell type can be downloaded from the NeMO Archive using the following link: https://assets.nemoarchive.org/dat-uioqy8b. Cell-type-specific 3D epigenomes can be visualized on the WashU Epigenome Browser using the datahub at the following link: http://epigenomegateway.wustl.edu/browser/?genome=hg38&position=chr17:72918238-73349675&hub=https://shen-msong.s3-us-west-1.amazonaws.com/hfb_submission/hfb_datahub.json.
Code availability
All of the software used in this study are listed in the Reporting Summary along with their versions.
References
Nowakowski, T. J. et al. Spatiotemporal gene expression trajectories reveal developmental hierarchies of the human cortex. Science 358, 1318–1323 (2017).
Zhong, S. et al. A single-cell RNA-seq survey of the developmental landscape of the human prefrontal cortex. Nature 555, 524–528 (2018).
Hansen, D. V., Lui, J. H., Parker, P. R. & Kriegstein, A. R. Neurogenic radial glia in the outer subventricular zone of human neocortex. Nature 464, 554–561 (2010).
Pontious, A., Kowalczyk, T., Englund, C. & Hevner, R. F. Role of intermediate progenitor cells in cerebral cortex development. Dev. Neurosci. 30, 24–32 (2008).
Anderson, S., Mione, M., Yun, K. & Rubenstein, J. L. Differential origins of neocortical projection and local circuit neurons: role of Dlx genes in neocortical interneuronogenesis. Cereb. Cortex 9, 646–654 (1999).
Zheng, H. & Xie, W. The role of 3D genome organization in development and cell differentiation. Nat. Rev. Mol. Cell Biol. 20, 535–550 (2019).
Li, Y., Hu, M. & Shen, Y. Gene regulation in the 3D genome. Hum. Mol. Genet. 27 (R2), R228–R233 (2018).
Schoenfelder, S. & Fraser, P. Long-range enhancer-promoter contacts in gene expression control. Nat. Rev. Genet. 20, 437–455 (2019).
Won, H. et al. Chromosome conformation elucidates regulatory relationships in developing human brain. Nature 538, 523–527 (2016).
Thomsen, E. R. et al. Fixed single-cell transcriptomic characterization of human radial glial diversity. Nat. Methods 13, 87–93 (2016).
Fang, R. et al. Mapping of long-range chromatin interactions by proximity ligation-assisted ChIP-seq. Cell Res. 26, 1345–1348 (2016).
Juric, I. et al. MAPS: Model-based analysis of long-range chromatin interactions from PLAC-seq and HiChIP experiments. PLoS Comput. Biol. 15, e1006982 (2019).
Heinz, S. et al. Simple combinations of lineage-determining transcription factors prime cis-regulatory elements required for macrophage and B cell identities. Mol. Cell 38, 576–589 (2010).
Englund, C. et al. Pax6, Tbr2, and Tbr1 are expressed sequentially by radial glia, intermediate progenitor cells, and postmitotic neurons in developing neocortex. J. Neurosci. 25, 247–251 (2005).
Lim, L., Mi, D., Llorca, A. & Marín, O. Development and functional diversification of cortical interneurons. Neuron 100, 294–313 (2018).
Liu, S. J. et al. Single-cell analysis of long non-coding RNAs in the developing human neocortex. Genome Biol. 17, 67 (2016).
Luo, C. et al. Cerebral organoids recapitulate epigenomic signatures of the human fetal brain. Cell Rep. 17, 3369–3384 (2016).
Javierre, B. M. et al. Lineage-specific genome architecture links enhancers and non-coding disease variants to target gene promoters. Cell 167, 1369–1384 (2016).
Choudhary, M. N. et al. Co-opted transposons help perpetuate conserved higher-order chromosomal structures. Genome Biol. 21, 16 (2020).
Feschotte, C. Transposable elements and the evolution of regulatory networks. Nat. Rev. Genet. 9, 397–405 (2008).
Zhang, Y. et al. Transcriptionally active HERV-H retrotransposons demarcate topologically associating domains in human pluripotent stem cells. Nat. Genet. 51, 1380–1388 (2019).
Bailey, S. D. et al. ZNF143 provides sequence specificity to secure chromatin interactions at gene promoters. Nat. Commun. 2, 6186 (2015).
Ngondo-Mbongo, R. P., Myslinski, E., Aster, J. C. & Carbon, P. Modulation of gene expression via overlapping binding sites exerted by ZNF143, Notch1 and THAP11. Nucleic Acids Res. 41, 4000–4014 (2013).
Sundaram, V. & Wang, T. Transposable element mediated innovation in gene regulatory landscapes of cells: re-visiting the “gene-battery” model. BioEssays 40, (2018).
Davis, C. A. et al. The Encyclopedia of DNA elements (ENCODE): data portal update. Nucleic Acids Res. 46 (D1), D794–D801 (2018).
Miller, D. J., Bhaduri, A., Sestan, N. & Kriegstein, A. Shared and derived features of cellular diversity in the human cerebral cortex. Curr. Opin. Neurobiol. 56, 117–124 (2019).
Suzuki, I. K. et al. Human-specific NOTCH2NL genes expand cortical neurogenesis through Delta/Notch regulation. Cell 173, 1370–1384 (2018).
Rani, N. et al. A Primate lncRNA mediates Notch signaling during neuronal development by sequestering miRNA. Neuron 90, 1174–1188 (2016).
Carbon, S. et al. AmiGO: online access to ontology and annotation data. Bioinformatics 25, 288–289 (2009).
Reilly, S. K. et al. Evolutionary genomics. Evolutionary changes in promoter and enhancer activity during human corticogenesis. Science 347, 1155–1159 (2015).
Visel, A., Minovitsky, S., Dubchak, I. & Pennacchio, L. A. VISTA Enhancer Browser—a database of tissue-specific human enhancers. Nucleic Acids Res. 35, D88–D92 (2007).
Walker, R. L. et al. Genetic control of expression and splicing in developing human brain informs disease mechanisms. Cell 179, 750–771 (2019).
Hoffman, G. E. et al. CommonMind Consortium provides transcriptomic and epigenomic data for schizophrenia and bipolar disorder. Sci. Data 6, 180 (2019).
Bulik-Sullivan, B. K. et al. LD score regression distinguishes confounding from polygenicity in genome-wide association studies. Nat. Genet. 47, 291–295 (2015).
Finucane, H. K. et al. Partitioning heritability by functional annotation using genome-wide association summary statistics. Nat. Genet. 47, 1228–1235 (2015).
Jansen, I. E. et al. Genome-wide meta-analysis identifies new loci and functional pathways influencing Alzheimer’s disease risk. Nat. Genet. 51, 404–413 (2019).
Demontis, D. et al. Discovery of the first genome-wide significant risk loci for attention deficit/hyperactivity disorder. Nat. Genet. 51, 63–75 (2019).
Grove, J. et al. Identification of common genetic risk variants for autism spectrum disorder. Nat. Genet. 51, 431–444 (2019).
Stahl, E. A. et al. Genome-wide association study identifies 30 loci associated with bipolar disorder. Nat. Genet. 51, 793–803 (2019).
Savage, J. E. et al. Genome-wide association meta-analysis in 269,867 individuals identifies new genetic and functional links to intelligence. Nat. Genet. 50, 912–919 (2018).
Howard, D. M. et al. Genome-wide meta-analysis of depression identifies 102 independent variants and highlights the importance of the prefrontal brain regions. Nat. Neurosci. 22, 343–352 (2019).
Pardiñas, A. F. et al. Common schizophrenia alleles are enriched in mutation-intolerant genes and in regions under strong background selection. Nat. Genet. 50, 381–389 (2018).
Gazal, S. et al. Linkage disequilibrium-dependent architecture of human complex traits shows action of negative selection. Nat. Genet. 49, 1421–1427 (2017).
Sey, N. Y. A. et al. A computational tool (H-MAGMA) for improved prediction of brain-disorder risk genes by incorporating brain chromatin interaction profiles. Nat. Neurosci. 23, 583–593 (2020).
Andersen, O. M. & Willnow, T. E. Lipoprotein receptors in Alzheimer’s disease. Trends Neurosci. 29, 687–694 (2006).
Akbarian, S. Epigenetic mechanisms in schizophrenia. Dialogues Clin. Neurosci. 16, 405–417 (2014).
Yang, X. et al. SMART-Q: an integrative pipeline quantifying cell-type-specific RNA transcription. PLoS One 15, e0228760 (2020).
Lieberman-Aiden, E. et al. Comprehensive mapping of long-range interactions reveals folding principles of the human genome. Science 326, 289–293 (2009).
Dixon, J. R. et al. Topological domains in mammalian genomes identified by analysis of chromatin interactions. Nature 485, 376–380 (2012).
Schmitt, A. D. et al. A compendium of chromatin contact maps reveals spatially active regions in the human genome. Cell Rep. 17, 2042–2059 (2016).
Sobhy, H., Kumar, R., Lewerentz, J., Lizana, L. & Stenberg, P. Highly interacting regions of the human genome are enriched with enhancers and bound by DNA repair proteins. Sci. Rep. 9, 4577 (2019).
Corces, M. R. et al. An improved ATAC-seq protocol reduces background and enables interrogation of frozen tissues. Nat. Methods 14, 959–962 (2017).
Reimand, J., Kull, M., Peterson, H., Hansen, J. & Vilo, J. g:Profiler—a web-based toolset for functional profiling of gene lists from large-scale experiments. Nucleic Acids Res. 35, W193–W200(2007).
Hnisz, D. et al. Super-enhancers in the control of cell identity and disease. Cell 155, 934–947 (2013).
Grant, C. E., Bailey, T. L. & Noble, W. S. FIMO: scanning for occurrences of a given motif. Bioinformatics 27, 1017–1018 (2011).
Diao, Y. et al. A tiling-deletion-based genetic screen for cis-regulatory element identification in mammalian cells. Nat. Methods 14, 629–635 (2017).
Xie, S., Duan, J., Li, B., Zhou, P. & Hon, G. C. Multiplexed engineering and analysis of combinatorial enhancer activity in single cells. Mol Cell 66, 285–299 (2017).
Datlinger, P. et al. Pooled CRISPR screening with single-cell transcriptome readout. Nat. Methods 14, 297–301 (2017).
Labun, K. et al. CHOPCHOP v3: expanding the CRISPR web toolbox beyond genome editing. Nucleic Acids Res. 47, W171–W174 (2019).
Acknowledgements
This work was supported by the UCSF Weill Institute for Neuroscience Innovation Award (to Y.S. and A.R.K.), the National Institutes of Health (NIH) grants R01AG057497, R01EY027789, and UM1HG009402 (to Y.S.) and R35NS097305 (to A.R.K.), the Hillblom Foundation, and the American Federation for Aging Research New Investigator Award in Alzheimer’s Disease (to Y.S). This work was also supported by the NIH grants R01HL129132, U544HD079124, and R01MH106611 (to Y.L.), R01HG007175, U24ES026699, and U01HG009391 (to T.W.), and the American Cancer Society grant RSG-14-049-01-DMC (to T.W). M.S. is supported by T32GM007175. M.P. is supported by the National Science Foundation Graduate Research Fellowship Program grant 1650113. U.C.E. is supported by 5T32GM007618-42. This work was made possible in part by the NIH grants P30EY002162 to the UCSF Core Grant for Vision Research, P30DK063720, and S101S10OD021822-01 to the UCSF Parnassus Flow Cytometry Core.
Author information
Authors and Affiliations
Contributions
Y.S., A.A.P., and A.R.K. conceived the study. Y.S., M.H., A.A.P., and A.R.K. supervised the study. M.S., M.-P.P., X.Y., I.R.J., X.C., U.C.E., L.M., and J.L. performed experiments. M.S., A.A., S.B., J.D.R., B.L., I.R.J., and M.H. performed computational analysis. C.F. and M.N.K.C. performed TE analysis under the supervision of T.W. J.W., and W.L. performed SNP heritability enrichment analysis under the supervision of Y.L. M.S., M.-P.P., X.Y., and Y.S. analysed and interpreted the data. M.S., M.-P.P., X.Y., M.H., and Y.S. prepared the manuscript with input from all other authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature thanks Andrew Adey, Chongyuan Luo and Barbara Treutlein for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Representative contour plots depicting FACS gating strategy.
a, Cells were separated from debris of various sizes based on the forward scatter area (FSC-A) and side scatter area (SSC-A). b, c, Specifically, they were passed through two singlet gates using the width and height metrics of the side scatter (SSC-H versus SSC-W) (b) and forward scatter (FSC-H versus FSC-W) (c). d, SOX2+ and SOX2− and IPC populations were isolated by gating on EOMES-PE-Cy7 and SOX2-PerCP-Cy5.5 staining. e, RG and interneurons were isolated based on high PAX6/high SOX2 and medium SOX2/low PAX6 staining, respectively. f, excitatory neurons were isolated from the SOX2- population by gating on SATB2-Alexa Fluor 647 staining.
Extended Data Fig. 2 Reproducibility between RNA-seq, ATAC-seq and PLAC-seq replicates.
a, RNA-seq replicates were hierarchically clustered according to gene expression sample distances using DESeq2. b, Heat map showing correlations between gene expression profiles for the sorted cell populations and single-cell RNA sequencing (scRNA-seq) data in the developing human cortex. The sorted cell populations exhibit the highest correlation with their corresponding subtypes while exhibiting reduced correlation with the endothelial, mural, microglial, and choroid plexus lineages. c, Heat map showing correlations and hierarchical clustering for read densities at open chromatin peaks across all ATAC-seq replicates. d, Principal component analysis (PCA) was performed based on normalized contact frequencies across all PLAC-seq replicates (Methods). PCA was performed using interacting 5-kb bins in both 300- and 600-kb windows.
Extended Data Fig. 3 Identification of significant H3K4me3-mediated chromatin interactions.
a, Illustration of XOR and AND interactions in a representative PLAC-seq contact matrix. The blue tracks represent H3K4me3 peaks at anchor bins. Purple cells represent AND interactions where both of the interacting bins are anchor bins. Orange cells represent XOR interactions where only one of the interacting bins is an anchor bin. Grey cells represent NOT interactions where neither of the interacting bins are anchor bins. b, Venn diagram displaying cell-type-specificity for interactions in each cell type. c, Proportions of interactions occurring within and across topologically associated domains in the germinal zones and cortical plate for matching cell types.
Extended Data Fig. 4 Chromatin interactions influence cell-type-specific transcription.
a, GO enrichment analysis for genes participating in cell-type-specific interactions. The top annotation clusters from DAVID are reported along with their group enrichment scores for each cell type (Methods). b, Scatter plots showing the correlation between the difference in the number of interactions for each promoter and the difference in the expression of the corresponding genes across all cell types (Pearson product-moment correlation coefficient, two-tailed, n = 13,996 anchor bins with promoters). The trend line from linear regression is shown. c, Fold enrichment of open chromatin peaks over distance-matched background regions in 1-Mb windows around distal interacting regions in IPCs, excitatory neurons and interneurons.
Extended Data Fig. 5 SIPs are enriched for lineage-specific genes.
a, Scatter plots showing the correlation between interaction counts and gene expression at promoters for each cell type (Pearson product-moment correlation coefficient, two-tailed, n = 13,996 anchor bins with promoters). b, CDF plots of the numbers of interactions for shared versus cell-type-specific genes for each cell type. P values determined by two-tailed two-sample t-test, two-tailed. c, Anchor bins were ranked according to their cumulative interaction scores in RG, IPCs and interneurons. SIPs are located past the point in each curve where the slope is equal to 1. d, Venn diagram displaying cell-type-specificity for SIPs in each cell type. e, f, Enrichment of super-enhancers and DNA methylation valleys at SIPs versus non-SIPs (left) and distal interacting regions for SIPs versus non-SIPs (right), P values determined by two-tailed Fisher’s exact test. Super-enhancers were based on data in the fetal brain and adult cortex, and DNA methylation valleys were based on data in 40- and 60-day cerebral organoids with closely matched gene expression profiles to mid-fetal cortex samples. g, Forrest plot showing that SIPs identified in haematopoietic cells are analogously enriched for cell-type-specific over shared genes. Odds ratios and 95% confidence intervals are shown. We identified 554, 709, 460, 712 and 401 SIPs in neutrophils, naive CD4+ T cells, monocytes, megakaryocytes, and erythroblasts, respectively.
Extended Data Fig. 6 Transposable elements in SIP formation.
a–c, Enrichment of TEs at the class (a), family (b), and subfamily (c) levels in SIPGs for each cell type. Only TE families occupying more than 1% of the genome are shown in b. Only TE subfamilies from the MIR and ERVL-MaLR TE families occupying more than 0.1% of the genome are shown in c. d, Both ERVL-MaLR TEs (left, 32% versus 19% of sequences, P < 2.2 × 10−16, binomial test, two-tailed) and THE1C TEs (right, 73% versus 19% of sequences, P < 2.2 × 10−16, binomial test, two-tailed) are enriched over background sequences for ZNF143 motifs in excitatory neurons. e, ZNF143 motifs are enriched at SIPGs in excitatory neurons (left, P = 5.39 × 10−82, two-sample t-test, two-tailed, n = 8,894 distal interacting regions). Data are mean and s.e.m. Distributions comparing the number of ZNF143 motifs per bin for actual versus shuffled SIPGs are shown (right, P < 2.2 × 10−16, Kolmogorov–Smirnov test, two-tailed, n = 638 SIPGs). f, ERVL-MaLR TEs in SIPGs are enriched over background sequences for ZNF143 motifs in excitatory neurons (31% versus 17% of sequences, P = 4.3 × 10−98, binomial test, two-tailed). g, Box plots showing elevated ADRA2A gene expression in excitatory neurons. The median, upper and lower quartiles, minimum and maximum are indicated. h, Illustration of the 12 distal interacting regions containing ERVL-MaLR TE-localized ZNF143 motifs in the ADRA2A SIPG. ZNF143 motifs are coloured by strand. The bin numbers correspond to Fig. 3g. i, Conservation of ERVL-MaLR TEs in the ADRA2A SIPG. Blue bars indicate consensus sequences, yellow bars indicate ERVL-MaLR TEs, and red bars indicate ZNF143 motifs. j, Alignment of THE1C TEs in the human genome to their consensus sequence. The THE1C subfamily contains two ZNF143 motifs, one at positions 47–61 (P1), and another at positions 96–110 (P2).
Extended Data Fig. 7 Developmental trajectories and mapping complex disorder- and trait-associated variants to their target genes.
a, Box plots showing the distributions of gene expression and cumulative interaction scores for the groups identified in Fig. 4a. The median, upper and lower quartiles, minimum, and maximum are indicated. b, Groups 4 and 5 are enriched for interactions with transcription factors containing domains associated with transcriptional repression. c, d, Counts of the numbers of GWAS SNPs (P < 10−8) interacting with their nearest gene only, with both their nearest and more distal genes, and with more distal genes only across all diseases (c) and specific disorders and traits (d).
Extended Data Fig. 8 Partitioning SNP heritability for complex disorders and traits using alternative epigenomic annotations.
a, Forrest plot showing the enrichment of fetal and adult brain eQTL–TSS pairs in our interactions compared to n = 50 sets of distance-matched control interactions. P values determined by two-tailed Fisher’s exact test. Odds ratios and 95% confidence intervals are shown. The increased significance of adult brain eQTLs can be attributed to the larger sample size of the CommonMind Consortium (CMC) study (n = 1,332,863), while larger odds ratios were observed for the more closely matched fetal brain eQTLs (n = 6,446). b, c, Histograms displaying the numbers of adult and fetal brain eQTL–TSS pairs recapitulated by n = 50 sets of distance-matched control interactions in each cell type. The numbers of eQTL–TSS pairs recapitulated by our interactions are indicated by red lines. P values determined by two-tailed Fisher’s exact test. d, LDSC enrichment scores for each disease and cell type, conditioned on the baseline model previously described43 and stratified by PLAC-seq anchor and target bins. Non-significant enrichment scores are shown as striped bars. e, f, LDSC enrichment scores for each disease and cell type, conditioned on the baseline model previously described42 and using either distal open chromatin peaks (e) or cell-type-specific genes (f). Non-significant enrichment scores are shown as striped bars.
Extended Data Fig. 9 Enriched biological processes for genes interacting with non-coding variants for each disease and cell type.
Gene Ontology enrichment analysis for genes interacting with non-coding variants for each disease and cell type using H-MAGMA and gProfileR. P values determined by two-tailed Fisher’s exact test, BH method. The full results can be found in Supplementary Table 12.
Extended Data Fig. 10 Characterization of RG- and excitatory neuron-specific loci using CRISPRview.
a, b, Validation of distal interacting regions at the IDH1 locus in RG and excitatory neurons. Silencing region 1, which interacts with the IDH1 promoter only in excitatory neurons, results in the significant downregulation of IDH1 expression in excitatory neurons but not in RG. Silencing region 2, which interacts with the IDH1 promoter only in RG, results in the significant downregulation of IDH1 expression in RG but not in excitatory neurons. Silencing region 3, which interacts with the IDH1 promoter in both RG and excitatory neurons, results in the significant downregulation of IDH1 expression in both cell types. Interactions between the promoter of IDH1 and distal interacting regions containing open chromatin peaks that were targeted for silencing are highlighted. Box plots show results for experimental (red) and control (green) sgRNA-treated cells for each region. P values determined by two-tailed two-sample t-test. The median, upper and lower quartiles and 10% to 90% range are indicated. Open circles represent single cells. Sample sizes are indicated above each box plot. c–h, Validation of distal interacting regions at the TNC and HES1 loci in RG. Interactions between the promoters of TNC and HES1 and distal interacting regions containing open chromatin peaks that were targeted for silencing are highlighted. Representative images show staining for intronic RNAscope probes (white), DAPI (blue), GFAP (light blue), GFP (green), and mCherry (red). Scale bar, 50 μm.
Supplementary information
Supplementary Information
This file contains a guide to Supplementary Tables 1-13.
Supplementary Tables
This zipped file contains Supplementary Tables 1-13.
Rights and permissions
About this article
Cite this article
Song, M., Pebworth, MP., Yang, X. et al. Cell-type-specific 3D epigenomes in the developing human cortex. Nature 587, 644–649 (2020). https://doi.org/10.1038/s41586-020-2825-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-020-2825-4
This article is cited by
-
Annotating genetic variants to target genes using H-MAGMA
Nature Protocols (2023)
-
Joint epigenome profiling reveals cell-type-specific gene regulatory programmes in human cortical organoids
Nature Cell Biology (2023)
-
Transposable elements in mammalian chromatin organization
Nature Reviews Genetics (2023)
-
Identification of brain cell types underlying genetic association with word reading and correlated traits
Molecular Psychiatry (2023)
-
Developmental mechanisms underlying the evolution of human cortical circuits
Nature Reviews Neuroscience (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.