Abstract
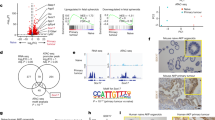
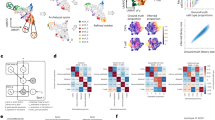
Blood vessels support tumours by providing nutrients and oxygen, while also acting as conduits for the dissemination of cancer1. Here we use mouse models of breast and lung cancer to investigate whether endothelial cells also have active ‘instructive’ roles in the dissemination of cancer. We purified genetically tagged endothelial ribosomes and their associated transcripts from highly and poorly metastatic tumours. Deep sequencing revealed that metastatic tumours induced expression of the axon-guidance gene Slit2 in endothelium, establishing differential expression between the endothelial (high Slit2 expression) and tumoural (low Slit2 expression) compartments. Endothelial-derived SLIT2 protein and its receptor ROBO1 promoted the migration of cancer cells towards endothelial cells and intravasation. Deleting endothelial Slit2 suppressed metastatic dissemination in mouse models of breast and lung cancer. Conversely, deletion of tumoural Slit2 enhanced metastatic progression. We identified double-stranded RNA derived from tumour cells as an upstream signal that induces expression of endothelial SLIT2 by acting on the RNA-sensing receptor TLR3. Accordingly, a set of endogenous retroviral element RNAs were upregulated in metastatic cells and detected extracellularly. Thus, cancer cells co-opt innate RNA sensing to induce a chemotactic signalling pathway in endothelium that drives intravasation and metastasis. These findings reveal that endothelial cells have a direct instructive role in driving metastatic dissemination, and demonstrate that a single gene (Slit2) can promote or suppress cancer progression depending on its cellular source.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
RNA-seq data have been deposited in the NCBI GEO accession GSE145319. Other data generated are available from the corresponding author upon reasonable request. Source data are provided with this paper.
Code availability
Code generated is available from the author upon reasonable request.
References
Strilic, B. et al. Tumour-cell-induced endothelial cell necroptosis via death receptor 6 promotes metastasis. Nature 536, 215–218 (2016).
Png, K. J., Halberg, N., Yoshida, M. & Tavazoie, S. F. A microRNA regulon that mediates endothelial recruitment and metastasis by cancer cells. Nature 481, 190–194 (2012).
Pencheva, N. et al. Convergent multi-miRNA targeting of ApoE drives LRP1/LRP8-dependent melanoma metastasis and angiogenesis. Cell 151, 1068–1082 (2012).
Pencheva, N. & Tavazoie, S. F. Control of metastatic progression by microRNA regulatory networks. Nat. Cell Biol. 15, 546–554 (2013).
Rafii, S., Butler, J. M. & Ding, B. S. Angiocrine functions of organ-specific endothelial cells. Nature 529, 316–325 (2016).
Kobayashi, H. et al. Angiocrine factors from Akt-activated endothelial cells balance self-renewal and differentiation of haematopoietic stem cells. Nat. Cell Biol. 12, 1046–1056 (2010).
Ding, B. S. et al. Endothelial-derived angiocrine signals induce and sustain regenerative lung alveolarization. Cell 147, 539–553 (2011).
Ding, B. S. et al. Inductive angiocrine signals from sinusoidal endothelium are required for liver regeneration. Nature 468, 310–315 (2010).
Tavora, B. et al. Endothelial-cell FAK targeting sensitizes tumours to DNA-damaging therapy. Nature 514, 112–116 (2014).
Sanz, E. et al. Cell-type-specific isolation of ribosome-associated mRNA from complex tissues. Proc. Natl Acad. Sci. USA 106, 13939–13944 (2009).
Wang, Y. et al. Ephrin-B2 controls VEGF-induced angiogenesis and lymphangiogenesis. Nature 465, 483–486 (2010).
Gibson, D. A. et al. Dendrite self-avoidance requires cell-autonomous Slit/Robo signaling in cerebellar purkinje cells. Neuron 81, 1040–1056 (2014).
Brose, K. et al. Slit proteins bind Robo receptors and have an evolutionarily conserved role in repulsive axon guidance. Cell 96, 795–806 (1999).
Wang, K. H. et al. Biochemical purification of a mammalian Slit protein as a positive regulator of sensory axon elongation and branching. Cell 96, 771–784 (1999).
Strickland, P., Shin, G. C., Plump, A., Tessier-Lavigne, M. & Hinck, L. Slit2 and netrin 1 act synergistically as adhesive cues to generate tubular bi-layers during ductal morphogenesis. Development 133, 823–832 (2006).
Svensson, K. J. et al. A secreted Slit2 fragment regulates adipose tissue thermogenesis and metabolic function. Cell Metab. 23, 454–466 (2016).
Ballard, M. S. & Hinck, L. A roundabout way to cancer. Adv. Cancer Res. 114, 187–235 (2012).
Lang, J. E. et al. RNA-seq of circulating tumor cells in stage II–III breast cancer. Ann. Surg. Oncol. 25, 2261–2270 (2018).
Gantier, M. P. & Williams, B. R. The response of mammalian cells to double-stranded RNA. Cytokine Growth Factor Rev. 18, 363–371 (2007).
Johnsen, I. B. et al. Toll-like receptor 3 associates with c-Src tyrosine kinase on endosomes to initiate antiviral signaling. EMBO J. 25, 3335–3346 (2006).
Itoh, K., Watanabe, A., Funami, K., Seya, T. & Matsumoto, M. The clathrin-mediated endocytic pathway participates in dsRNA-induced IFN-β production. J. Immunol. 181, 5522–5529 (2008).
Kawasaki, T. & Kawai, T. Toll-like receptor signaling pathways. Front. Immunol. 5, 461 (2014).
Shivapurkar, N. et al. Multiple regions of chromosome 4 demonstrating allelic losses in breast carcinomas. Cancer Res. 59, 3576–3580 (1999).
Dallol, A. et al. Frequent epigenetic inactivation of the SLIT2 gene in gliomas. Oncogene 22, 4611–4616 (2003).
Gröne, J. et al. Robo1/Robo4: differential expression of angiogenic markers in colorectal cancer. Oncol. Rep. 15, 1437–1443 (2006).
Huang, W. Y. et al. MethHC: a database of DNA methylation and gene expression in human cancer. Nucleic Acids Res. 43, D856–D861 (2015).
Macias, H. et al. SLIT/ROBO1 signaling suppresses mammary branching morphogenesis by limiting basal cell number. Dev. Cell 20, 827–840 (2011).
Macias, H. & Hinck, L. Mammary gland development. Wiley Interdiscip. Rev. Dev. Biol. 1, 533–557 (2012).
Marlow, R. et al. SLITs suppress tumor growth in vivo by silencing Sdf1/Cxcr4 within breast epithelium. Cancer Res. 68, 7819–7827 (2008).
Escamilla-Tilch, M. et al. The interplay between pathogen-associated and danger-associated molecular patterns: an inflammatory code in cancer? Immunol. Cell Biol. 91, 601–610 (2013).
Bakhoum, S. F. et al. Chromosomal instability drives metastasis through a cytosolic DNA response. Nature 553, 467–472 (2018).
Nabet, B. Y. et al. Exosome RNA unshielding couples stromal activation to pattern recognition receptor signaling in cancer. Cell 170, 352–366 (2017).
Redzic, J. S., Balaj, L., van der Vos, K. E. & Breakefield, X. O. Extracellular RNA mediates and marks cancer progression. Semin. Cancer Biol. 28, 14–23 (2014).
Rooney, M. S., Shukla, S. A., Wu, C. J., Getz, G. & Hacohen, N. Molecular and genetic properties of tumors associated with local immune cytolytic activity. Cell 160, 48–61 (2015).
Kassiotis, G. Endogenous retroviruses and the development of cancer. J. Immunol. 192, 1343–1349 (2014).
Zernecke, A. & Preissner, K. T. Extracellular ribonucleic acids (RNA) enter the stage in cardiovascular disease. Circ. Res. 118, 469–479 (2016).
Khakpour, S., Wilhelmsen, K. & Hellman, J. Vascular endothelial cell Toll-like receptor pathways in sepsis. Innate Immun. 21, 827–846 (2015).
Mantovani, A., Allavena, P., Sica, A. & Balkwill, F. Cancer-related inflammation. Nature 454, 436–444 (2008).
Tavazoie, S. F. et al. Endogenous human microRNAs that suppress breast cancer metastasis. Nature 451, 147–152 (2008).
Reynolds, L. E. & Hodivala-Dilke, K. M. Primary mouse endothelial cell culture for assays of angiogenesis. Methods Mol. Med. 120, 503–509 (2006).
Aslakson, C. J. & Miller, F. R. Selective events in the metastatic process defined by analysis of the sequential dissemination of subpopulations of a mouse mammary tumor. Cancer Res. 52, 1399–1405 (1992).
Fidler, I. J. Biological behavior of malignant melanoma cells correlated to their survival in vivo. Cancer Res. 35, 218–224 (1975).
May, T. et al. Establishment of murine cell lines by constitutive and conditional immortalization. J. Biotechnol. 120, 99–110 (2005).
Guy, C. T., Cardiff, R. D. & Muller, W. J. Induction of mammary tumors by expression of polyomavirus middle T oncogene: a transgenic mouse model for metastatic disease. Mol. Cell. Biol. 12, 954–961 (1992).
Wagner, K. U. et al. Cre-mediated gene deletion in the mammary gland. Nucleic Acids Res. 25, 4323–4330 (1997).
Lánczky, A. et al. miRpower: a web-tool to validate survival-associated miRNAs utilizing expression data from 2178 breast cancer patients. Breast Cancer Res. Treat. 160, 439–446 (2016).
DeRose, Y. S. et al. Tumor grafts derived from women with breast cancer authentically reflect tumor pathology, growth, metastasis and disease outcomes. Nat. Med. 17, 1514–1520 (2011).
Sikora, M. J. et al. Invasive lobular carcinoma cell lines are characterized by unique estrogen-mediated gene expression patterns and altered tamoxifen response. Cancer Res. 74, 1463–1474 (2014).
Ponomarev, V. et al. A novel triple-modality reporter gene for whole-body fluorescent, bioluminescent, and nuclear noninvasive imaging. Eur. J. Nucl. Med. Mol. Imaging 31, 740–751 (2004).
Dhir, A. et al. Mitochondrial double-stranded RNA triggers antiviral signalling in humans. Nature 560, 238–242 (2018).
Acknowledgements
We thank V. Padmanaban, L. Noble, D. Hsu, D. Huh, R. Moy, S. Belkaya, B. Boyraz and D. Mucida for comments on previous versions of the manuscript; Rockefeller University resource centres (S. Mazel and S. Semova of the flow cytometry resource centre, C. Zhao from the genomics resource centre, V. Francis from the Comparative Bioscience Center veterinary staff for animal husbandry and care, and A. North, C. Pyrgaki and staff of the Bio-Imaging Resource Facility); S. J. Gendler for providing MMTV-PyMT mice; A. Chedotal for the generation and distribution of the Slit2-floxed mice; M. Tessier-Lavigne for transferring Slit2-floxed mice; R. Adams for providing the Cdh5(PAC)-creERT2 mice; and K. Svensson for providing the SLIT2 adenoviral particles. T.M., K.W., M.S., M.M. and J.-Y.K. are members of the German Academic Scholarship Foundation (Studienstiftung des deutschen Volkes). K.W. and S.R. were awarded a fellowship from Boehringer Ingelheim Foundation. B.T. was supported by the Lucy Lee Chiles Fellowship (HFCR-15-06-04) from the Hope Funds for Cancer Research. This work was supported by grants from the National Institutes of Health (RO1-CA236954-01A1 to S.F.T.; R01CA24098 to H.G.) and the Department of Defense Collaborative Scholars and Innovators award (W81-XWH-12-1-0301). S.F.T. was also supported by a Faculty Scholars grant from the Howard Hughes Medical Institute, the Breast Cancer Research Foundation award, the Black Family Foundation and the Reem-Kayden award.
Author information
Authors and Affiliations
Contributions
B.T. and S.F.T. designed the experiments. B.T performed all the experiments together with contributions from other authors. T.M. and K.W. performed in vivo experiments including tuSLIT2 deletion, poly(I:C) effect on circulating tumour cells as well as in vitro conditioned medium and cell migration assays and contributed to the manuscript writing. S.R. conducted tumour experiments such as ecSLIT2 deletion in PyMT tumours, as well as in vitro cell migration assays, human breast cancer immunostainings and CellProfiler analysis. M.S. carried out mouse tumour and in vitro studies including ecSLIT2 deletion. M.M. conducted tumour cell studies and characterized ecSLIT2-knockout tumours. B.N.O. performed in vivo experiments and RNAseq analysis. X.L. performed J2 immunoprecipitation. J.-Y.K. conducted analysis of PyMT metastasis data. A.L.W. collected and provided PDXs from patients with breast cancer. H.G. performed RiboTag and J2 pull-down RNA-seq analyses. O.O. provided the Slit2-floxed mice. B.T. and S.F.T. wrote the paper with input from the co-authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature thanks Andrew Oberst and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Endothelial cells upregulate SLIT2 upon treatment with conditioned medium from highly metastatic 4T1 cells.
a, Primary MLECs (ICAM2-positive) upregulate SLIT2 when treated with conditioned medium derived from 4T1 cells (n = 3). Dot plot represents Slit2 mRNA levels measured by qPCR for each biological replicate with mean ± s.e.m. Two-tailed Student’s t-test. b, Primary nonendothelial cells (ICAM2-negative) from the lung do not upregulate SLIT2 upon treatment with 4T1 conditioned medium (n = 3). Dot plot represents Slit2 mRNA levels measured by qPCR for each biological replicate with mean ± s.e.m. Two-tailed Student’s t-test. c, Treatment of endothelial cells with 5 μM dynasore inhibits SLIT2 expression upon treatment with conditioned medium from 4T1 cells (n = 3). Dot plot represents Slit2 mRNA levels measured by qPCR for each biological replicate with mean ± s.e.m. Two-tailed Student’s t-test. d, e, Dot plots represent Slit2 mRNA expression by qPCR in endothelial cells exposed to 4T1 conditioned medium treated with (e) DNase I (10 μg/ml; n = 3), and (d) heat treatment (95 °C, 10 min; n = 3). Data are mean ± s.e.m. Two-tailed Student’s t-test. f, TLR3 wild-type (Tlr3 WT) and TLR3-knockout (Tlr3 KO) endothelial cells were treated with conditioned medium from 67NR, 4T07 and 4T1 cells. Western blot analysis revealed that wild-type endothelial cells display increased phosphorylation of ERK1 and ERK2 upon treatment with the conditioned medium from highly metastatic 4T1 cells. TLR3-knockout endothelial cells displayed reduced phosphorylation of ERK1 and ERK2 relative to wild-type controls. Dot plot displays densitometry quantification for three independent experiments. Two-tailed Student’s t-test. g, RNase A treatment of the 4T1 conditioned medium blunted endothelial phosphorylation of ERK1 and ERK2. h, Supplementation of basal medium with synthetic TLR9 ligand (CpG ODN, 2.5 or 12.5 μg/ml) did not induce endothelial SLIT2 upregulation (n = 3). Dot plot represents Slit2 levels measured by qPCR for each biological replicate with mean ± s.e.m. Two-tailed Student’s t-test. i, j, Supplementation of basal medium with synthetic TLR9 ligand (CpG ODN, 2.5 or 12.5 μg/ml) induced (i) endothelial Il6 (n = 3) and (j) Ifng mRNA expression (n = 3). Dot plot represents Il6 and Ifng levels measured by qPCR for each biological replicate with mean ± s.e.m. Two-tailed Student’s t-test. k, l, Quantification of RNA isolated from conditioned medium of (k) B16F0 (n = 3) and B16F10 cells (n = 3) and (l) 67NR (n = 3) and 4T1 cells (n = 3). Dot plot represents RNA concentrations detected in conditioned medium normalized by the cell number with mean ± s.e.m. Two-tailed Student’s t-test. m, RNA detection in plasma isolated from mice with 67NR (n = 3) and 4T1 (n = 5) mammary gland tumours. Tumour-free mice (n = 5) were used as a negative control. Increased concentrations of RNA were detected in the plasma of mice with the metastatic 4T1 tumours. Dot plot represents the RNA concentrations detected in the plasma of each mouse, either with no tumour or with 67NR and 4T1 tumours. Two-tailed Student’s t-test.
Extended Data Fig. 2 Endothelial SLIT2 deletion does not impair primary tumour growth and angiogenesis.
a–c, Tumour growth rates (left) for (a) spontaneous MMTV-PyMT mammary gland tumours (total tumour burden) in wild-type (n = 8) and ecSLIT2-knockout mice (n = 7), (b) orthotopic 4T1 mammary tumours in wild-type (n = 11) and ecSLIT2-knockout mice (n = 8), and (c) subcutaneous LLC tumours in wild-type (n = 22) and ecSLIT2-knockout mice (n = 19). Mean tumour volume ± s.e.m. for each time point. Two-tailed t-test for last time point. d, Mammary gland tumours from tamoxifen-treated Cdh5(PAC)-creERT2;Slit2-floxed;MMTV-PyMT (ecSLIT2-knockout) or CreERT2-negative Slit2-floxed;MMTV-PyMT (ecSLIT2 wild-type) mice were sectioned and stained for endomucin. No significant difference in blood vessel density was observed between tumours growing in wild-type and ecSLIT2-knockout mice. Each dot represents the average of endomucin area relative to total DAPI area in sections for each tumour, measured with ImageJ. Mean ± s.e.m. ecSLIT2 wild type, n = 6; ecSLIT2 knockout, n = 6. Scale bar, 50 μm. Two-tailed Student’s t-test. e, The 4T1 tumour sections were stained for endomucin. No difference in vessel density was observed between tumours from wild-type and ecSLIT2-knockout mice. Dot plot depicts endomucin area relative to DAPI area for each tumour, quantified by ImageJ. Mean ± s.e.m. ecSLIT2 wild type, n = 6; ecSLIT2 knockout, n = 5. Scale bar, 50 μm. Two-tailed Student’s t-test. f, LLC tumour sections were stained for endomucin. No difference in blood vessel density was observed between tumours growing in ecSLIT2-knockout and wild-type mice. Mean ± s.e.m. ecSLIT2 wild type, n = 4; ecSLIT2 knockout, n = 4. Scale bar, 50 μm. Two-tailed Student’s t-test. g, h, Immunofluorescence staining for PyMT in lung sections of MMTV-PyMT ecSLIT2 wild type or ecSLIT2-knockout mice reveals reduction in both micrometastasis (g) and macrometastasis (h). Dot plot displays the number of lung nodules per mouse, divided into micrometastases or macrometastases. ecSLIT2 wild type, n = 9; ecSLIT2 knockout, n = 9. Data are mean ± s.e.m. Two-tailed Mann–Whitney test. Arrowheads indicate macrometastasis and arrows indicate micrometastasis. i, Wild-type and ecSLIT2-knockout mice bearing 4T1 primary tumours were intravenously injected with PE–PECAM antibody and Hoechst. The 4T1 tumour sections were prepared, and vessel permeability was quantified. Representative images of tumour sections showing Hoechst nuclear staining and perfused PE–PECAM vessels. Scale bar, 50 μm. Dot plot represents the mean ratio of Hoechst signal relative to PE–PECAM signal ± s.e.m.; ecSLIT2 wild type, n = 5; ecSLIT2 knockout, n = 5. j, Tumour sections from wild-type and ecSLIT2-knockout mice bearing 4T1 primary tumours were injected via tail vein with PE–PECAM antibody and stained for PECAM to quantify the proportion of perfused vessels relative to total tumour vessels. Representative images of tumour sections showing PE–PECAM perfused vessels (functional vessels) relative to total vessels stained with PECAM. White arrows indicate nonperfused blood vessels. Scale bar, 50 μm. Bar chart represents the mean ratio of Hoechst relative to endomucin staining ± s.e.m. ecSLIT2 wild type, n = 5; ecSLIT2 knockout, n = 5. i, j, Two-tailed Student’s t-test. k, Tumour growth rates for the MMTV-PyMT tumours in tuSLIT2-knockout (n = 12) or wild-type (n = 10) control mice. Tumour burden was calculated by adding individual tumours in each mouse. Data are mean ± s.e.m. Two-tailed t-test for last time point. l, Blood vessel density was measured by immunostaining for endomucin in sections of mammary gland tumours from MMTV-PyMT mice (tuSLIT2 wild type or tuSLIT2 knockout). Bar chart represents the average endomucin area relative to DAPI area ± s.e.m. Scale bar, 50 μm. tuSLIT2 wild type, n = 5; tuSLIT2 knockout, n = 5. m, Tumoural SLIT2 deletion was confirmed by immunostaining of tumours for SLIT2. Fluorescent quantification revealed a significant reduction in SLIT2 levels in tuSLIT2-knockout tumours. Bar chart with each dot representing the average of fluorescent quantification of different tumour sections for each mouse ± s.e.m. tuSLIT2 wild type, n = 5; tuSLIT2 knockout, n = 5. l, m, Two-tailed Student’s t-test. Scale bar, 50 μm.
Extended Data Fig. 3 Endothelial SLIT2 deletion does not affect metastatic colonization upon tail vein injection.
a, 4T1 cells were injected intravenously into the tail veins of ecSLIT2-knockout or wild-type littermate controls. Survival is depicted as the number of days until each mouse was euthanized owing to metastatic disease. ecSLIT2 wild type, n = 11; ecSLIT2 knockout, n = 12. log-rank (Mantel–Cox) test. b, Metastatic burden was measured by quantification of mean luminescence relative to day 0 ± s.e.m. ecSLIT2 wild type, n = 11; ecSLIT2 knockout, n = 12. Two-tailed Student’s t-test. c, H&E-stained lung sections were used for quantification of lung nodules 17 d after injection of cells. Dot plot represents the average number of lung nodules per mouse ± s.e.m. ecSLIT2 wild type, n = 6; ecSLIT2 knockout, n = 3. Scale bar, 0.5 cm. Two-tailed Student’s t-test. d, LLC cells were injected into the tail veins of wild-type or ecSLIT2-knockout littermate controls. Survival is depicted as in a. ecSLIT2 wild type, n = 11; ecSLIT2 knockout, n = 14. log-rank (Mantel–Cox) test. e, Metastatic burden was measured by mean bioluminescence quantification relative to day 0 ± s.e.m. ecSLIT2 wild type, n = 6, ecSLIT2 knockout, n = 6. Two-tailed Student’s t-test. f, H&E-stained lung sections revealed no significant difference in lung nodule numbers between groups, 15 d after injection. Data are mean ± s.e.m. ecSLIT2 wild type, n = 6; ecSLIT2 knockout, n = 6. Scale bar, 0.5 cm. Two-tailed Student’s t-test.
Extended Data Fig. 4 Time course of endothelial SLIT2 upregulation upon conditioned medium treatment.
C-terminal SLIT2 fragment is insufficient to promote 4T1 tumour cell migration. a, Endothelial cells overexpressing SLIT2-C–Flag were used to generate conditioned medium or lysed for protein extraction. Anti-Flag antibody was used to detect SLIT2 in either cell lysates or secreted SLIT2 in the conditioned medium. b, Western blot for SLIT2 to detect the full-length SLIT2 or its C-terminal fragment. HSC70 was used as a loading control. c, Endothelial cells were treated with conditioned medium from 4T1 cells for 3, 6, 12 and 24 h, and SLIT2 levels were assessed. Dot plot represents Slit2 mRNA levels for each biological replicate with mean ± s.e.m. n = 3 for each group. Two-tailed Student’s t-test. d, Western blot of SLIT2 protein upon treatment of wild-type endothelial cells or TLR3-knockout endothelial cells with conditioned medium from 4T1 cells or basal medium (control). e, The 4T1 tumour cells displayed enhanced migration towards increasing concentrations of recombinant SLIT2-N, but not SLIT2-C. Dot plot represents the number of migrated cells per optical field of view (10× objective) with mean ± s.e.m. Panel displays representative images from migrated cells (10× objective). n = 4 for each group. Two-tailed Student’s t-test. f, Adenoviral expression of full-length SLIT2 in ecSLIT2-knockout endothelial cells promoted tumour cell migration in a transwell assay while adenoviral expression of C-terminal SLIT2 (ecSLIT2 KO-C) or LacZ (ecSLIT2 KO-LacZ) did not enhance tumour cell migration. Mean ± s.e.m. Two-tailed Student’s t-test. Western blot with antibody against C-terminal region of SLIT2 detected full-length and C-terminal forms of SLIT2 in ecSLIT2-knockout endothelial cells transduced with adenovirus. n = 4 for each condition. g, SLIT2 depletion in 4T1 tumour cells using two independent shRNAs increased migration of tumour cells towards endothelial cells in a transwell migration assay. Dot plot represents the number of migrated 4T1 cells per optical field of view (10× objective) with mean ± s.e.m. P value between control and shRNA no. 1 and no. 2 corresponds to a one-tailed Student’s t-test. n = 4 for each condition. h, SLIT2 expression in 4T1 cells transduced with either scrambled shRNA or two independent shRNAs targeting SLIT2. Dot plot represents Slit2 mRNA levels for each biological replicate with mean ± s.e.m. Two-tailed Student’s t-test. n = 3 for each condition.
Extended Data Fig. 5 Endothelial and tumoural SLIT2 and tumoural ROBO1 levels are associated with cancer progression in patients with breast cancer.
a, b, Dot plots represent relative Robo1 mRNA expression by qPCR in mouse metastatic 4T1 (n = 4) and nonmetastatic 67NR cells (n = 4) (a) and parental 4T1 cells (primary n = 5) and 4T1 cells derived from lung metastases (lung mets., n = 5) (b). a, b Two-tailed Student’s t-test. c, SLIT2 quantification in endomucin-positive vessels of PDX tumours from patients with breast cancer revealed that high levels of endothelial SLIT2 correlate with worse prognosis. Survival data for the patients from whom the PDXs were isolated were stratified into high and low endothelial SLIT2 expression. High ecSLIT2 expression, n = 10 patients; low ecSLIT2 expression, n = 10 patients. log-rank (Mantel–Cox) test. d, RNA-seq data analysis from primary tumours and CTCs of patients with breast cancer from Gene Expression Omnibus GSE11184218 revealed reduced or undetectable SLIT2 expression in CTCs from patients with stage II or III breast cancer when compared with SLIT2 levels of matched primary tumours. Dot plot represents the mean SLIT2 levels ± s.e.m. Primary tumour samples, n = 12; CTC samples, n = 16. Two-tailed Mann–Whitney test. e, f, Kaplan–Meier survival analysis for SLIT2 (e) and ROBO1 (f) expression in human breast tumours generated using KMPLOT, with auto select best cut-off selected. SLIT2, n = 1,660; ROBO1, n = 3,951. log-rank (Mantel–Cox) test.
Extended Data Fig. 6 TLR3 knockout in the tumour stroma reduces tumour cell intravasation, whereas synthetic dsRNA induces intravasation by tumour cells.
a, 4T1-Luc-zsGreen tumours were established in the mammary fat pads of wild-type and TLR3-knockout mice. Circulating tumour cells were isolated from the whole blood of mice, and quantified by identifying luciferase-positive colonies. Dot plot represents measured bioluminescence (photons s−1). Representative images of luciferase-positive colonies growing on 10-cm tissue culture dishes. TLR3 wild type, n = 10; TLR3 knockout, n = 11. b, Quantification of SLIT2 expression in the blood vessels of 4T1 mammary tumours in either wild-type or TLR3-knockout mice. Dot plot represents mean fluorescence intensities of SLIT2 in endomucin-positive vessels of 4T1 tumours ± s.e.m. TLR3 wild type, n = 9; TLR3 knockout, n = 9 tumours. a, b, Data are mean ± s.e.m. Two-tailed Student’s t-test. c, Injection of poly(I:C) (25 μg) into NSG mice promoted intravasation by tumour cells, measured by quantification of circulating tumour cells through detection of luminescence (photons s−1) from luciferase-positive colonies. Dot plot with each dot representing measured bioluminescence (photons s−1), for the whole-blood-derived colonies for each mouse. Control group (ctrl), n = 7; poly(I:C), n = 8. Representative images of luciferase-positive colonies growing on a 10-cm tissue culture dish. d, ImageJ quantification of immunofluorescent SLIT2 staining that colocalized with endomucin-positive vessels in 4T1 tumours injected with either PBS (control) or poly(I:C). Dot plot represents fluorescent intensities of SLIT2 in the vasculature of 4T1 tumours ± s.e.m. Control, n = 7; poly(I:C), n = 8 tumours. e, PE–PECAM antibody and Hoechst perfusion did not reveal changes in vascular permeability by poly(I:C) treatment. Representative images of tumour sections showing Hoechst nuclear staining and perfused PE–PECAM vessels. Scale bar, 50 μm. Bar chart represents the average ratio of Hoechst signal relative to PE–PECAM signal normalized to the control group ± s.e.m.; n = 5 tumours for each group. c–e, Data are mean ± s.e.m. Two-tailed Student’s t-test. f, Robo1 knockdown in tumour cells with a second shRNA (Robo1 shRNA no. 2) inhibited poly(I:C)-induced intravasation. Dot plot with each dot representing measured bioluminescence (photons s−1), for the whole-blood-derived luciferase-positive colonies for each mouse with mean ± s.e.m. Control shRNA: control, n = 5; poly(I:C), n = 4; Robo1 shRNA no. 2: control, n = 5; poly(I:C), n = 6. One-tailed Student’s t-test. g, ROBO1 expression in 4T1-Luc-zsGreen cells transduced with either scrambled shRNA (control shRNA) or shRNA no. 2 targeting Robo1. Dot plot represents Robo1 mRNA levels for each replicate with mean ± s.e.m. Control shRNA, n = 3; Robo1 shRNA no. 2, n = 3. Two-tailed Student’s t-test.
Extended Data Fig. 7 SLIT2 promoter is hypermethylated in breast cancer in humans.
a, SLIT2 promoter methylation in normal breast tissues and invasive breast carcinomas, reproduced from the Human Cancer database (MethHC)26. Dot plot represents the mean SLIT2 promoter methylation ± s.e.m. Breast tissue, n = 92; Breast cancer, n = 735. b, Slit2 expression by real-time qPCR in 67NR and 4T1 tumour cells. Dot plot represents Slit2 mRNA levels for each biological replicate with mean ± s.e.m. 67NR, n = 3; 4T1, n = 3. c, Treatment of 4T1 tumour cells with the demethylating agent 5-azacytidine (5-aza) upregulated SLIT2 expression. Dot plot represents Slit2 mRNA levels for each biological replicate with mean ± s.e.m. Control group (ctrl) n = 3; 5-aza group, n = 3. d, Pre-mRNA levels were measured by real-time qPCR using primers that span the exon–intron junction. Dot plot represents pre-mRNA Slit2 levels for each biological replicate (n = 3) with mean ± s.e.m. e, Genomic copy number for Slit2 in 67NR and 4T1 tumour cells relative to mouse lung endothelial cells (MLEC). Dot plot represents Slit2 copy number (TaqMan assay) for each replicate (n = 3) with mean ± s.e.m. a–e, Two-tailed Student’s t-test.
Extended Data Fig. 8 Tumoural SLIT2 deletion does not significantly affect apoptosis and expression of SLIT2-related cytokines.
a, No difference in cleaved caspase 3 staining in MMTV-PyMT tumour sections of wild-type versus tuSLIT2-knockout mice. Dot plot represents fluorescent intensity of cleaved caspase 3 in tumour sections. Mean ± s.e.m. tuSLIT2 wild type, n = 5; tuSLIT2 knockout, n = 5 tumours. Scale bar, 50 μm. b, Deletion of SLIT2 in the tumour compartment of MMTV-PyMT tumours did not significantly alter tumour netrin 1 expression. Dot plot represents fluorescent intensity of netrin 1 in tumour sections. Mean ± s.e.m. tuSLIT2 wild type, n = 5; tuSLIT2 knockout, n = 5 tumours. Scale bar, 50 μm. c, Deletion of SLIT2 in the tumour compartment of MMTV-PyMT tumours did not significantly affect tumour SDF1 expression; Dot plot represents fluorescent intensity of SDF1 in tumour sections ± s.e.m. tuSLIT2 wild type, n = 4; tuSLIT2 knockout, n = 6 tumours. Scale bar, 50 μm. d, Deletion of SLIT2 in the tumour compartment of MMTV-PyMT tumours did not significantly change tumour MCP1 expression; Dot plot represents fluorescent intensity of MCP1 in tumour sections ± s.e.m. tuSLIT2 wild type, n = 4; tuSLIT2 knockout, n = 4 tumours. Scale bar, 50 μm. e, Sdf1 (also known as Cxcl12) expression was measured by qPCR in endothelial cells treated with conditioned medium from the poorly metastatic 67NR or highly metastatic 4T1 cells, or with basal medium (control). Bar chart represents the mean expression levels of SDF1 ± s.e.m. Each group, n = 3. f, Mcp1 (also known as Mcpt1) expression was measured by qPCR in endothelial cells treated with conditioned medium from poorly metastatic 67NR or highly metastatic 4T1 cells, or basal medium (control). Bar chart depicts the mean expression of MCP1 ± s.e.m. Each group, n = 3. g, Cxcr4 expression was assessed by qPCR in endothelial cells treated with conditioned medium from poorly metastatic 67NR or highly metastatic 4T1 cells, or basal medium (control). Bar chart represents the mean expression values of CXCR4 ± s.e.m. Each group, n = 3. a–g, Two-tailed Student’s t-test.
Extended Data Fig. 9 Increased detection of dsRNA and ERVs in highly metastatic tumours.
dsRNA was detected by immunostaining of tumours with the J2 antibody. a, b, Fluorescent quantification revealed a significant increase in dsRNA signal in highly metastatic B16F10 relative to B16F0 tumours (a) and in metastatic 4T1 relative to nonmetastatic 67NR tumours (b). Bar chart with each dot representing the mean relative fluorescent quantification of different tumour sections for each tumour normalized to the low-metastatic B16F0 or 67NR tumours ± s.e.m. B16F0, n = 6; B16F10, n = 6; 67NR, n = 5; 4T1, n = 5. Representative images are shown for the immunostaining of dsRNA (J2), endomucin and DAPI in tumour sections of B16F0 and B16F10, and 67NR and 4T1, tumours. Scale bar, 50μm. a, b, Two-tailed Student’s t-test. c, d, Volcano plot displays the log2-transformed fold differences in expression of ERVs between B16F10 relative to B16F0 cells (c) as well as 4T1 relative to 67NR cells (d). Out of 12,332 annotations, 123 ERV sequences were detected in our RNA-seq libraries. n = 3 biological replicates. e, Cell-free RNA was isolated from the conditioned medium of 67NR and 4T1 cell cultures, and RNA-seq libraries were generated for analysis of the aforementioned 123 ERVs. Volcano plot displays the log2-transformed fold differences in detected ERVs in the supernatant of 4T1 cells relative to 67NR cells. n = 3 biological replicates. f, Pull-down of dsRNA with the J2 antibody followed by RNA-seq detected ERVs secreted by 4T1 cells. Table shows the total counts of ERV reads detected for two immunoprecipitation replicates (4T1-1 and 4T1-2).
Extended Data Fig. 10 ROBO1 knockdown in human MDA cells reduces orthotopic lung metastasis.
a, Two independent shRNAs were used to knockdown ROBO1 in human breast cancer MDA cells. Both ROBO1 knockdown (shRNA no. 1 and shRNA no. 2) and control shRNA (control) cells were injected into the mammary fat pads of NSG mice. Tumours were measured at days 7, 10, 21, 28, 35 and 38, and surgically resected on day 38. P values for the last time point (control–shRNA no. 1 and control–shRNA no. 2) are shown. Two-tailed Student’s t-test. b, Lung metastasis was assessed by bioluminescence imaging (photons per second). n = 5 for each group. P values for the last time point (control–shRNA no. 1 and control–shRNA no. 2) are shown. Two-tailed Student’s t-test. Representative images of lung bioluminescence are shown with colour scale (photons per s per cm2 per sr) for each independent group (ctrl and shRNA no. 1 or no. 2) c, Robo1 knockdown levels were confirmed by qPCR (n = 3). Two-tailed Student’s t-test. d, Blood vessel density was assessed by staining tumour sections with endomucin and DAPI. No significant difference in blood vessel density was detected between the control and Robo1-knockdown tumour groups (shRNA no. 1 or no. 2). Dot plot represents endomucin area relative to DAPI area for each tumour, quantified using ImageJ. Mean ± s.e.m. n = 5 each group. Two-tailed Student’s t-test. Scale bar, 50 μm.
Supplementary information
Supplementary Information
This file contains Supplementary Fig.1 (uncropped gel source data for figures) and Supplementary Table 1 (Genotyping of transgenic mouse lines – Primer sequences).
Source data
Rights and permissions
About this article
Cite this article
Tavora, B., Mederer, T., Wessel, K.J. et al. Tumoural activation of TLR3–SLIT2 axis in endothelium drives metastasis. Nature 586, 299–304 (2020). https://doi.org/10.1038/s41586-020-2774-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-020-2774-y
This article is cited by
-
Harnessing innate immune pathways for therapeutic advancement in cancer
Signal Transduction and Targeted Therapy (2024)
-
Cancer stem cell-immune cell crosstalk in the tumor microenvironment for liver cancer progression
Frontiers of Medicine (2024)
-
Gastric Cancer Mesenchymal Stem Cells Trigger Endothelial Cell Functional Changes to Promote Cancer Progression
Stem Cell Reviews and Reports (2024)
-
Lymphatic endothelial-like cells promote glioblastoma stem cell growth through cytokine-driven cholesterol metabolism
Nature Cancer (2024)
-
SEVs-mediated miR-6750 transfer inhibits pre-metastatic niche formation in nasopharyngeal carcinoma by targeting M6PR
Cell Death Discovery (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.