Abstract
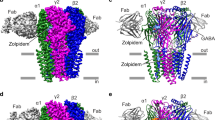
Most general anaesthetics and classical benzodiazepine drugs act through positive modulation of γ-aminobutyric acid type A (GABAA) receptors to dampen neuronal activity in the brain1,2,3,4,5. However, direct structural information on the mechanisms of general anaesthetics at their physiological receptor sites is lacking. Here we present cryo-electron microscopy structures of GABAA receptors bound to intravenous anaesthetics, benzodiazepines and inhibitory modulators. These structures were solved in a lipidic environment and are complemented by electrophysiology and molecular dynamics simulations. Structures of GABAA receptors in complex with the anaesthetics phenobarbital, etomidate and propofol reveal both distinct and common transmembrane binding sites, which are shared in part by the benzodiazepine drug diazepam. Structures in which GABAA receptors are bound by benzodiazepine-site ligands identify an additional membrane binding site for diazepam and suggest an allosteric mechanism for anaesthetic reversal by flumazenil. This study provides a foundation for understanding how pharmacologically diverse and clinically essential drugs act through overlapping and distinct mechanisms to potentiate inhibitory signalling in the brain.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
Atomic model coordinates for bicuculline methbromide, GABA + propofol, GABA + flumazenil, GABA + etomidate, GABA + phenobarbital, GABA + diazepam, GABA and GABA + picrotoxin-bound structures have been deposited in the Protein Data Bank with accession codes 6X3S, 6X3T, 6X3U, 6X3V, 6X3W, 6X3X, 6X3Z and 6X40, respectively. Cryo-EM density maps have been deposited in the Electron Microscopy Data Bank with accession codes EMD-22031, EMD-22032, EMD-22033, EMD-22034, EMD-22035, EMD-22036, EMD-22037 and EMD-22038, respectively.
References
Hemmings, H. C., Jr et al. Emerging molecular mechanisms of general anesthetic action. Trends Pharmacol. Sci. 26, 503–510 (2005).
Forman, S. A. & Miller, K. W. Mapping general anesthetic sites in heteromeric γ-aminobutyric acid type A receptors reveals a potential for targeting receptor subtypes. Anesth. Analg. 123, 1263–1273 (2016).
Sieghart, W. & Savić, M. M. International Union of Basic and Clinical Pharmacology. CVI: GABAA receptor subtype- and function-selective ligands: key issues in translation to humans. Pharmacol. Rev. 70, 836–878 (2018).
Sigel, E. & Ernst, M. The benzodiazepine binding sites of GABAA receptors. Trends Pharmacol. Sci. 39, 659–671 (2018).
Olsen, R. W. GABAA receptor: positive and negative allosteric modulators. Neuropharmacology 136, 10–22 (2018).
Meyer, H. Welche Eigenschaft der Anasthetica bedingt ihre narkotische Wirkung? Naunyn Schmiedebergs Arch. Exp. Pathol. Pharmakol. 42, 109–118 (1899).
Meyer, H. Zur Theorie der Alkoholnarkose: der Einfluss wechselnder Temperatur auf Wirkungsstärke und Theilungscoefficient der Narcotica. Naunyn Schmiedebergs Arch. Exp. Pathol. Pharmakol. 46, 338–346 (1901).
Overton, E. Studien über die Narkose Zugleich ein Beitrag zur allgemeinen Pharmakologie (Gustav Fischer, 1901).
Janoff, A. S., Pringle, M. J. & Miller, K. W. Correlation of general anesthetic potency with solubility in membranes. Biochim. Biophys. Acta 649, 125–128 (1981).
Franks, N. P. & Lieb, W. R. Molecular and cellular mechanisms of general anaesthesia. Nature 367, 607–614 (1994).
Krasowski, M. D. & Harrison, N. L. General anaesthetic actions on ligand-gated ion channels. Cell. Mol. Life Sci. 55, 1278–1303 (1999).
Krasowski, M. D. Contradicting a unitary theory of general anesthetic action: a history of three compounds from 1901 to 2001. Bull. Anesth. Hist. 21, 1–24 (2003).
Mihic, S. J. et al. Sites of alcohol and volatile anaesthetic action on GABAA and glycine receptors. Nature 389, 385–389 (1997).
Drexler, B., Antkowiak, B., Engin, E. & Rudolph, U. Identification and characterization of anesthetic targets by mouse molecular genetics approaches. Can. J. Anaesth. 58, 178–190 (2011).
Walters, R. J., Hadley, S. H., Morris, K. D. & Amin, J. Benzodiazepines act on GABAA receptors via two distinct and separable mechanisms. Nat. Neurosci. 3, 1274–1281 (2000).
Middendorp, S. J., Maldifassi, M. C., Baur, R. & Sigel, E. Positive modulation of synaptic and extrasynaptic GABAA receptors by an antagonist of the high affinity benzodiazepine binding site. Neuropharmacology 95, 459–467 (2015).
Votey, S. R., Bosse, G. M., Bayer, M. J. & Hoffman, J. R. Flumazenil: a new benzodiazepine antagonist. Ann. Emerg. Med. 20, 181–188 (1991).
Laverty, D. et al. Cryo-EM structure of the human α1β3γ2 GABAA receptor in a lipid bilayer. Nature 565, 516–520 (2019).
Masiulis, S. et al. GABAA receptor signalling mechanisms revealed by structural pharmacology. Nature 565, 454–459 (2019).
Löscher, W. & Rogawski, M. A. How theories evolved concerning the mechanism of action of barbiturates. Epilepsia 53, 12–25 (2012).
Zhu, S. et al. Structure of a human synaptic GABAA receptor. Nature 559, 67–72 (2018).
Gielen, M., Barilone, N. & Corringer, P.-J. The desensitization pathway of GABAA receptors, one subunit at a time. Preprint at https://doi.org/10.1101/2020.05.18.101287 (2020).
Chiara, D. C. et al. Specificity of intersubunit general anesthetic-binding sites in the transmembrane domain of the human α1β3γ2 γ-aminobutyric acid type A (GABAA) receptor. J. Biol. Chem. 288, 19343–19357 (2013).
Zeller, A., Arras, M., Jurd, R. & Rudolph, U. Identification of a molecular target mediating the general anesthetic actions of pentobarbital. Mol. Pharmacol. 71, 852–859 (2007).
Belelli, D., Callachan, H., Hill-Venning, C., Peters, J. A. & Lambert, J. J. Interaction of positive allosteric modulators with human and Drosophila recombinant GABA receptors expressed in Xenopus laevis oocytes. Br. J. Pharmacol. 118, 563–576 (1996).
Vuyk, J., Sitsen, E. & Reekers, M. in Miller’s Anesthesia 9th edn (eds Gropper, M. A. et al.) Ch. 23, 638–679 (Elsevier, 2020).
Forman, S. A. Clinical and molecular pharmacology of etomidate. Anesthesiology 114, 695–707 (2011).
Belelli, D., Lambert, J. J., Peters, J. A., Wafford, K. & Whiting, P. J. The interaction of the general anesthetic etomidate with the γ-aminobutyric acid type A receptor is influenced by a single amino acid. Proc. Natl Acad. Sci. USA 94, 11031–11036 (1997).
Siegwart, R., Jurd, R. & Rudolph, U. Molecular determinants for the action of general anesthetics at recombinant α2β3γ2γ-aminobutyric acidA receptors. J. Neurochem. 80, 140–148 (2002).
Li, G. D. et al. Identification of a GABAA receptor anesthetic binding site at subunit interfaces by photolabeling with an etomidate analog. J. Neurosci. 26, 11599–11605 (2006).
Krasowski, M. D. et al. Propofol and other intravenous anesthetics have sites of action on the γ-aminobutyric acid type A receptor distinct from that for isoflurane. Mol. Pharmacol. 53, 530–538 (1998).
Jayakar, S. S. et al. Multiple propofol-binding sites in a γ-aminobutyric acid type A receptor (GABAAR) identified using a photoreactive propofol analog. J. Biol. Chem. 289, 27456–27468 (2014).
Bali, M. & Akabas, M. H. Gating-induced conformational rearrangement of the γ-aminobutyric acid type A receptor β-α subunit interface in the membrane-spanning domain. J. Biol. Chem. 287, 27762–27770 (2012).
Jayakar, S. S. et al. Identifying drugs that bind selectively to intersubunit general anesthetic sites in the α1β3γ2 GABAAR transmembrane domain. Mol. Pharmacol. 95, 615–628 (2019).
Yip, G. M. et al. A propofol binding site on mammalian GABAA receptors identified by photolabeling. Nat. Chem. Biol. 9, 715–720 (2013).
Jurd, R. et al. General anesthetic actions in vivo strongly attenuated by a point mutation in the GABAA receptor β3 subunit. FASEB J. 17, 250–252 (2003).
Reynolds, D. S. et al. Sedation and anesthesia mediated by distinct GABAA receptor isoforms. J. Neurosci. 23, 8608–8617 (2003).
Chiara, D. C. et al. Mapping general anesthetic binding site(s) in human α1β3 γ-aminobutyric acid type A receptors with [3H]TDBzl-etomidate, a photoreactive etomidate analogue. Biochemistry 51, 836–847 (2012).
Krasowski, M. D., Hong, X., Hopfinger, A. J. & Harrison, N. L. 4D-QSAR analysis of a set of propofol analogues: mapping binding sites for an anesthetic phenol on the GABAA receptor. J. Med. Chem. 45, 3210–3221 (2002).
Krasowski, M. D., Nishikawa, K., Nikolaeva, N., Lin, A. & Harrison, N. L. Methionine 286 in transmembrane domain 3 of the GABAA receptor beta subunit controls a binding cavity for propofol and other alkylphenol general anesthetics. Neuropharmacology 41, 952–964 (2001).
Eaton, M. M. et al. Multiple non-equivalent interfaces mediate direct activation of GABAA receptors by propofol. Curr. Neuropharmacol. 14, 772–780 (2016).
Ritchie, T. K. et al. Chapter eleven - reconstitution of membrane proteins in phospholipid bilayer nanodiscs. Methods Enzymol. 464, 211–231 (2009).
Damgen, M. A. & Biggin, P. C. A refined open state of the glycine receptor obtained via molecular dynamics simulations. Structure 28, 130–139.e2 (2020).
Gielen, M., Thomas, P. & Smart, T. G. The desensitization gate of inhibitory Cys-loop receptors. Nat. Commun. 6, 6829 (2015).
Dahaba, A. A. et al. Effect of flumazenil on bispectral index monitoring in unpremedicated patients. Anesthesiology 110, 1036–1040 (2009).
Safavynia, S. A. et al. Effects of γ-aminobutyric acid type A receptor modulation by flumazenil on emergence from general anesthesia. Anesthesiology 125, 147–158 (2016).
Ueno, S., Bracamontes, J., Zorumski, C., Weiss, D. S. & Steinbach, J. H. Bicuculline and gabazine are allosteric inhibitors of channel opening of the GABAA receptor. J. Neurosci. 17, 625–634 (1997).
Baumann, S. W., Baur, R. & Sigel, E. Individual properties of the two functional agonist sites in GABAA receptors. J. Neurosci. 23, 11158–11166 (2003).
Rosen, A., Bali, M., Horenstein, J. & Akabas, M. H. Channel opening by anesthetics and GABA induces similar changes in the GABAA receptor M2 segment. Biophys. J. 92, 3130–3139 (2007).
Kim, J. H. et al. High cleavage efficiency of a 2Å peptide derived from porcine teschovirus-1 in human cell lines, zebrafish and mice. PLoS ONE 6, e18556 (2011).
Morales-Perez, C. L., Noviello, C. M. & Hibbs, R. E. Manipulation of subunit stoichiometry in heteromeric membrane proteins. Structure 24, 797–805 (2016).
Lyons, J. A., Bøggild, A., Nissen, P. & Frauenfeld, J. Chapter three - saposin–lipoprotein scaffolds for structure determination of membrane transporters. Methods Enzymol. 594, 85–99 (2017).
Zivanov, J. et al. New tools for automated high-resolution cryo-EM structure determination in RELION-3. eLife 7, e42166 (2018).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Zhang, K. Gctf: real-time CTF determination and correction. J. Struct. Biol. 193, 1–12 (2016).
Rohou, A. & Grigorieff, N. CTFFIND4: Fast and accurate defocus estimation from electron micrographs. J. Struct. Biol. 192, 216–221 (2015).
Wagner, T. et al. SPHIRE-crYOLO is a fast and accurate fully automated particle picker for cryo-EM. Commun. Biol. 2, 218 (2019).
Bai, X. C., Rajendra, E., Yang, G., Shi, Y. & Scheres, S. H. Sampling the conformational space of the catalytic subunit of human γ-secretase. eLife 4, e11182 (2015).
Kucukelbir, A., Sigworth, F. J. & Tagare, H. D. Quantifying the local resolution of cryo-EM density maps. Nat. Methods 11, 63–65 (2014).
Schwede, T., Kopp, J., Guex, N. & Peitsch, M. C. SWISS-MODEL: an automated protein homology-modeling server. Nucleic Acids Res. 31, 3381–3385 (2003).
Miller, P. S. & Aricescu, A. R. Crystal structure of a human GABAA receptor. Nature 512, 270–275 (2014).
Pettersen, E. F. et al. UCSF Chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D 66, 486–501 (2010).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D 66, 213–221 (2010).
Laskowski, R. A. & Swindells, M. B. LigPlot+: multiple ligand-protein interaction diagrams for drug discovery. J. Chem. Inf. Model. 51, 2778–2786 (2011).
Krissinel, E. & Henrick, K. Inference of macromolecular assemblies from crystalline state. J. Mol. Biol. 372, 774–797 (2007).
Smart, O. S., Neduvelil, J. G., Wang, X., Wallace, B. A. & Sansom, M. S. HOLE: a program for the analysis of the pore dimensions of ion channel structural models. J. Mol. Graph. 14, 354–360 (1996).
Pei, J., Kim, B. H. & Grishin, N. V. PROMALS3D: a tool for multiple protein sequence and structure alignments. Nucleic Acids Res. 36, 2295–2300 (2008).
Morin, A. et al. Collaboration gets the most out of software. eLife 2, e01456 (2013).
Jo, S., Kim, T., Iyer, V. G. & Im, W. CHARMM-GUI: a web-based graphical user interface for CHARMM. J. Comput. Chem. 29, 1859–1865 (2008).
Abraham, M. J. et al. GROMACS: High performance molecular simulations through multi-level parallelism from laptops to supercomputers. SoftwareX 1–2, 19–25 (2015).
Periole, X. & Marrink, S. J. The Martini coarse-grained force field. Methods Mol. Biol. 924, 533–565 (2013).
Wassenaar, T. A., Pluhackova, K., Böckmann, R. A., Marrink, S. J. & Tieleman, D. P. Going backward: a flexible geometric approach to reverse transformation from coarse grained to atomistic models. J. Chem. Theory Comput. 10, 676–690 (2014).
Best, R. B. et al. Optimization of the additive CHARMM all-atom protein force field targeting improved sampling of the backbone φ, ψ and side-chain χ1 and χ2 dihedral angles. J. Chem. Theory Comput. 8, 3257–3273 (2012).
Vanommeslaeghe, K. et al. CHARMM general force field: a force field for drug-like molecules compatible with the CHARMM all-atom additive biological force fields. J. Comput. Chem. 31, 671–690 (2010).
Bussi, G., Donadio, D. & Parrinello, M. Canonical sampling through velocity rescaling. J. Chem. Phys. 126, 014101 (2007).
Parrinello, M. & Rahman, A. Crystal structure and pair potentials: a molecular-dynamics study. Phys. Rev. Lett. 45, 1196–1199 (1980).
Essmann, U. et al. A smooth particle mesh Ewald method. J. Chem. Phys. 103, 8577–8593 (1995).
Hess, B. P-LINCS: a parallel linear constraint solver for molecular simulation. J. Chem. Theory Comput. 4, 116–122 (2008).
Humphrey, W., Dalke, A. & Schulten, K. VMD: visual molecular dynamics. J. Mol. Graph. 14, 33–38 (1996).
Michaud-Agrawal, N., Denning, E. J., Woolf, T. B. & Beckstein, O. MDAnalysis: a toolkit for the analysis of molecular dynamics simulations. J. Comput. Chem. 32, 2319–2327 (2011).
McGibbon, R. T. et al. MDTraj: a modern open library for the analysis of molecular dynamics trajectories. Biophys. J. 109, 1528–1532 (2015).
Orellana, L., Yoluk, O., Carrillo, O., Orozco, M. & Lindahl, E. Prediction and validation of protein intermediate states from structurally rich ensembles and coarse-grained simulations. Nat. Commun. 7, 12575 (2016).
Lindahl, V., Gourdon, P., Andersson, M. & Hess, B. Permeability and ammonia selectivity in aquaporin TIP2;1: linking structure to function. Sci. Rep. 8, 2995 (2018).
Phulera, S. et al. Cryo-EM structure of the benzodiazepine-sensitive α1β1γ2S tri-heteromeric GABAA receptor in complex with GABA. eLife 7, e39383 (2018).
Miller, P. S. et al. Heteromeric GABAA receptor structures in positively-modulated active states. Preprint at https://doi.org/10.1101/338343 (2018).
Ingólfsson, H. I. et al. Computational lipidomics of the neuronal plasma membrane. Biophys. J. 113, 2271–2280 (2017).
Nury, H. et al. X-ray structures of general anaesthetics bound to a pentameric ligand-gated ion channel. Nature 469, 428–431 (2011).
Fourati, Z. et al. Structural basis for a bimodal allosteric mechanism of general anesthetic modulation in pentameric ligand-gated ion channels. Cell Rep. 23, 993–1004 (2018).
Hibbs, R. E. & Gouaux, E. Principles of activation and permeation in an anion-selective Cys-loop receptor. Nature 474, 54–60 (2011).
Acknowledgements
We thank R. Cabuco and L. Baxter for baculovirus production, and all members of the Hibbs laboratory for discussion. Single-particle cryo-EM data were collected at the University of Texas Southwestern Medical Center Cryo-Electron Microscopy Facility, which is supported by the CPRIT Core Facility Support Award RP170644, at the Harvard Cryo-Electron Microscopy Center for Structural Biology, and at the Pacific Northwest Cryo-EM Center at Oregon Health & Science University, which is supported by NIH grant U24GM129547, accessed through EMSL (grid.436923.9) a DOE office of Science User Facility sponsored by the Office of Biological and Environmental Research. Computational resources were provided by the Swedish National Infrastructure for Computing. J.J.K. and S.Z. acknowledge support from the American Heart Association grants 20POST35200127 and 18POST34030412, respectively. This work was supported by Vetenskapsrådet VR and the Knut and Alice Wallenberg foundation to E.L. and by The Welch Foundation (I-1812) and grants from the NIH (DA037492, DA042072, and NS095899) to R.E.H.
Author information
Authors and Affiliations
Contributions
J.J.K. and R.E.H. conceived the project. J.J.K. performed the construct design, protein production, purification, EM sample preparation and structural analysis including the EM data processing. J.J.K., A.G. and R.E.H. built the atomic models. J.T. performed the mutagenesis and electrophysiology experiments. S.Z., C.M.N. and R.M.W. collected the EM data. Y.Z., R.J.H. and E.L. performed and analysed simulations. J.J.K., A.G. and R.E.H. wrote the manuscript with input from all other authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature thanks Margot Ernst and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Biochemistry, sample condition screening, and stability of atomistic molecular dynamics simulations in brain lipids.
In 2018, our group reported the structure of the α1β2γ2 receptor in complex with GABA and flumazenil in detergent21. Although this initial study revealed details of the classical neurotransmitter and benzodiazepine binding sites, the structures showed an unanticipated asymmetric occluded state in the transmembrane region, where we observed the γ2 TMD collapsed into the pore or structurally disordered. Structures in complex with GABA85 or with a nanobody modulator86, also in detergent, exhibited very low resolution in the membrane domain that precluded detailed analysis. Structures of the α1β3γ2 receptor in lipid nanodiscs were reported more recently, with a well-ordered and approximately symmetric transmembrane domain18,19. We first sought to improve order and prevent collapse of the symmetric transmembrane domain (TMD) quaternary structure by optimizing lipid reconstitution of the GABA plus flumazenil receptor complex as a benchmark. a, Analytical size-exclusion chromatography of the α1β2γ2 receptor at different stages of preparation of the GABA plus flumazenil complex, which we used to benchmark the reconstitution approach: receptor in detergent, increasing in size after exchange into nanodiscs, then a further increase in size after addition of Fab. Inset, SDS–PAGE shows relatively pure nanodisc–Fab–receptor complex, which was used for grid preparation. b–e, TMD z-slices of 3D reconstructions from preparations with GABA, flumazenil and various membrane mimetics. Inset numbers are resolution values from the reconstructions and white dashed lines highlight subunit boundaries. b is from the dataset published in 201821; c is from the sample purified in DDM supplemented with brain lipids, more symmetric but very low resolution; d is from protein purified in DDM supplemented with soy polar lipid extract (Avanti) and cholesteryl hemisuccinate (CHS, Anatrace) and exchanged in MSP1E3 nanodiscs containing soy lipids, highly asymmetric; e is the condition used to obtain the GABA plus flumazenil complex in this study. We applied this purification and nanodisc reconstitution approach to all other complexes. f, Results from atomistic molecular dynamics simulations validating the stability of these complexes in a brain-lipid environment, as well as differential dynamics in the presence of different ligands. After embedding our models in mixed membranes with expected brain-lipid proportions87 and equilibrating with coarse-grained simulations72, cholesterol and PtdIns(4,5)P2 were found to accumulate at the protein surface, particularly at subunit interfaces (Supplementary Videos 9 and 10, respectively). Such interactions could contribute to the symmetrizing effect of brain lipids relative to detergent or other lipid mixtures. Subsequent quadruplicate 500-ns all-atom molecular dynamics simulations of all 8 structures reported in this work were largely stable, converging to ≤3 Å r.m.s.d. for all protein Cα atoms. This panel shows deviations from starting conformations (r.m.s.d., Å) of protein Cα atoms in α1β2γ2 receptor structures. Each trace represents one of four 500-ns replicates. g, An alternative conformation observed in multiple exploratory simulations of the flumazenil-bound structure (grey) with flumazenil removed. Within 200 ns, the γ M2-helix spontaneously translocates to block the pore (snapshot at 500 ns, coloured), supporting a flexible conformational repertoire for this subunit. Transition is tracked over time (red–blue) by the position of P-2′ in α and γ. h, Simulation results for propofol stability at all five interfacial TMD sites, with probability distributions at left, and raw data (n = 500 samples from 4 simulations, see Methods) plus box plots indicating sample median, interquartile range (25th–75th percentiles), minimum–maximum range, and outliers at right. Propofol was inserted at the α–β, α–γ and γ–β sites by symmetry superposition of the resolved β–α propofol. In quadruplicate simulations of >400 ns each, the inserted propofol molecules were not stably bound, sampling a broad distribution up to 8 Å r.m.s.d. from initial poses. By contrast, propofol at the β–α interfaces remained within 4 Å r.m.s.d. of its initial poses. Thus, simulations support a preference for propofol binding at the β–α interface over other interfaces.
Extended Data Fig. 2 Detailed cryo-EM processing flowchart for GABA plus flumazenil complex.
a, A representative cryo-EM image. b, Projection images from the final selected 2D classes. c, 3D classification results; good classes selected for further processing are boxed in red and in lower row have TMD z-slices shown. Note fuzzy nanodisc appearance adjacent to γ2 subunit, consistent with conformational heterogeneity in this region. d–f, 3D maps from a second round of 3D classification (d), from which particle from four classes (red boxes) were selected and used to generate map shown in e. Signal subtraction and γ2 subunit focused 3D classification resulted in the map in f.
Extended Data Fig. 3 Overall and local map resolution and global map–model agreement.
For each structure, the sharpened map is coloured by local resolution, and map FSC (upper right) and map-model FSC (lower right) plots are shown. For the flumazenil complex, two maps were used in building: a higher resolution map that had weak γ-TMD density, and a lower resolution map with strong γ-TMD signal. Shown here, for this structure, is the lower resolution map with strong signal for the whole receptor. Both maps will be deposited for this flumazenil complex, and relevant statistics for these maps are shown in Extended Data Tables 1, 2.
Extended Data Fig. 4 Map quality and ligand binding sites.
a–h, Each panel shows a side view and a TMD slice from the experimental density map, accompanied by the chemical structure of the ligand in that complex. Note, GABA is present in all structures except the bicuculline complex. Solid boxes highlight GABA binding sites; dashed boxes highlight allosteric ligands (including picrotoxin) binding sites. Propofol binding sites at subunit interfaces in f are distinct from the intrasubunit sites identified initially in the prokaryotic GLIC channel88, and similar in location but distinct in pose compared to the intersubunit site mutants of GLIC89.
Extended Data Fig. 5 Lipid interactions in TMD.
a, An atomic model overview of the TMD sites for possible lipid binding in the GABA plus propofol complex; densities for putative lipids are shown in tan. A subset of these are consistent with those modelled as POPC in the α1β3γ2 structures18,19. b–e, Side views of lipid density at the different subunit interfaces. The lipid density maps shown were generated using the unsharpened map. f–h, Structure of the GABA plus diazepam complex. f, An atomic model overview of the TMD sites for possible lipid binding; densities for putative lipids are shown in tan. g, h, Side views of potential lipid density at the subunit interfaces.
Extended Data Fig. 6 Representative map quality and model fit and structural analysis of GABA alone, diazepam and flumazenil complexes.
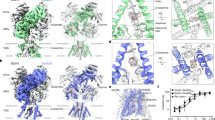
Semitransparent surface is shown for central ligand and contacting side chains for a–d. a, The GABA site at chain A–B β–α interface in the GABA-alone structure. The two β–α GABA sites from the structure superimpose nearly perfectly and do not shed light on the differences in functional contributions found in electrophysiology studies with concatamers48. Structures of apo receptor may be essential in identifying structural differences in the two GABA sites. b, Flumazenil site at the α–γ interface. c, Diazepam at the same ECD interface. d, Bicuculline site at the same interface as in a. e, The picrotoxin site in TMD; here, density is shown for ligand and all nearby protein structure elements. f, Superposition of two GABA plus flumazenil complexes, one from the detergent condition21 and one from this study in brain lipids, to illustrate absence of differences in backbone conformation. Note, loops that interact with the TMD do vary in conformation. g, Detail of flumazenil site from the superposition in f. h, i, Superpositions of three structures from the current study: GABA alone, GABA plus diazepam and GABA plus flumazenil, focused on the two GABA-binding sites. j, Calculated interface areas and interaction energies for each subunit pair, for each of the benzodiazepine-related structures.
Extended Data Fig. 7 Agonist and benzodiazepine complexes.
a–c, ECD binding sites viewed from the synaptic perspective, a, Overview of the diazepam complex. b, Position of diazepam with ligand map quality shown; side chains shown for residues contacting diazepam. c, Superposition of flumazenil and diazepam complexes. d, The three TMD sites identified for diazepam. e, f, Binding site details for diazepam at the β–α and γ–β interfaces. g, The two enantiomeric conformations of diazepam identified in the TMD sites. h, i, Snapshots from molecular dynamics simulations viewed from the extracellular side. Extracellular GABA and benzodiazepines are shown as sticks, coloured by frame (red–blue scale). h, Flumazenil-bound simulation with GABA in the upper site unbinding within 100 ns (pink–blue peripheral sticks). i, Diazepam-bound simulation with GABA retained in both orthosteric sites. Subunit subscripts denote chain ID. Stick representation is shown for residues within the van der Waals contact range.
Extended Data Fig. 8 Ligand site comparisons among α1β2γ2, α1β3γ2 and GluCl structures, and panel of pore conformations.
a, b, Superpositions of the GABA and diazepam ECD binding sites from the α1β2γ2 receptor (this study; subunits and ligands are coloured) and the α1β3γ2 receptor (in grey)19, respectively. c, A superposition similar to those in a and c but for the bicuculline complexes (N,N-dimethyl is the higher-affinity form from this study; bicuculline (single N-methyl) for α1β3γ2 in grey). d, e, Comparison of picrotoxin binding sites from three structures: this study, the α1β3γ2 structure and GluCl90. The results suggest that picrotoxin can bind to multiple conformations at different depths of the pore. GluCl is most widely open and picrotoxin binds most deeply; in that study, picrotoxin was used as a probe for an open-state conformation90. The pore is more tightly closed in α1β3γ2 than in α1β2γ2, which may allow picrotoxin to bind more deeply in the latter structure. In GluCl and in α1β2γ2, the picrotoxin isoprenyl tail orients towards the cytosol; in α1β3γ2, tail orients towards extracellular surface. This orientation allows in GluCl for favourable interactions between the ‘basket’ oxygens and the polar 2′ residues. The α1β2/β3γ2 receptors are more hydrophobic at the 2′ position, which might also explain favourable positioning of picrotoxin higher in the pore, where in the α1β2γ2 structure these oxygens are likely to make hydrogen-bonding interactions with conserved 6′ threonine hydroxyls. f, A sequence alignment of GABAA subunit M2 helices. Red boxes highlight residues potentially important in picrotoxin binding; in bold are the 15′ residues that have a role in anaesthetic selectivity and sensitivity. g, Pore conformational states for all ligand complexes, with opposing β1 and γ2 M2 α-helices shown as ribbons with pore-lining side chains shown as sticks. Purple and green spheres illustrate shape of the pore. Boxed distances in the pore are diameters at the desensitization gate (−2′) and resting gate (9′) positions. h, Free energies for chloride ion permeation along the pore axis (cytoplasmic side down, with −2′ gate at 0 nm), for representative α1β2γ2 complexes. Overlaid plots show the energy barrier at the 9′ hydrophobic gate (around 2 nm) in the bicuculline complex (orange) to be partially relieved in the GABA complex (green), and further relieved in complexes with GABA + phenobarbital or GABA + propofol (light or dark blue, respectively). i, All α1β2γ2 structures reported in this work (n = 8 independent structures), plotted along dominant principal components calculated for the TMD. Snapshots of a simulated transition78 between the GABA and bicuculline complexes (light-to-dark crosses) show that the GABA + picrotoxin complex maps along this pathway. GABA + diazepam and intravenous-anaesthetic-bound structures (GABA + diazepam, dark blue; etomidate, grey; phenobarbital, orange; propofol, purple) cluster at the lower left, distinct from GABA-alone or flumazenil- or inhibitor-bound states.
Extended Data Fig. 9 Ion-pore conformation and TMD subunit interface packing in α1β2γ2 compared with α1β3γ2 structures.
a, b, Pore conformations for α1β2γ2 (this study) (a) and α1β3γ219 (b) structures bound by GABA plus diazepam, with opposing β1 and γ2 M2 α-helices shown as ribbons and pore-lining side chains shown as sticks. Purple and green spheres illustrate the shape of the pore; purple is for radii >2.8 Å; green is 1.4–2.8 Å; red is < 1.4 Å. Distances on the right side of pore are radii at the desensitization gate (−2′) and resting gate (9′) positions. c, A comparison of these two structures in the form of a pore radius versus distance along the pore plot. Structures were aligned at y = 0 at the level of the −2′ desensitization gate. d–f and g–i make the same comparisons, but for the bicuculline (d–f) and GABA plus picrotoxin (g–i) complexes. j, Comparison of the interface area buried per subunit interface (Å2, ECD+TMD) for representative anion-selective receptors; top three are homopentamers for which the area given is the average from all interfaces, whereas for the two bicuculline structures the area comes from the average of the two β–α interfaces. Comparison is limited to anion-selective receptors owing to the absence of ordered intracellular domains; eukaryotic cation-selective receptors contain intracellular domains that contribute to interface surface area. k, Buried TMD subunit interface areas between pairs of GABAA receptor structures, to illustrate tighter packing in the α1β3γ2 receptor structures.
Extended Data Fig. 10 Nanodisc sizes correspond to the lipid ratio used in reconstitution.
a–c, Comparison of experimental EM maps (with docked structures), low-pass-filtered to 10 Å resolution, between matched α1β2γ2 and α1β3γ2 ligand complexes. d, Comparison of the reconstitution approach from the current study with the on-column approach used to obtain the α1β3γ2 receptor structures18,19. Asterisks indicate steps we propose give rise to the observed different nanodisc sizes: washing with lipid-free detergent buffer removes lipids, and the step of collecting affinity resin by centrifugation removes excess lipids, such that when the MSP2N2 scaffold and Bio-Beads are added, there are no extra lipids to fill the large scaffold.
Extended Data Fig. 11 Example electrophysiological recordings with cryo-EM construct.
All recordings were made in whole-cell voltage-clamp mode at −75 mV with transiently-transfected HEK cells. a, Wild-type, full-length receptor compared to cryo-EM construct, response to application of GABA. All remaining recordings are with the EM construct. b, A representative response is shown for application of GABA, then GABA plus diazepam, then GABA plus flumazenil, then GABA plus diazepam plus flumazenil. c, Application of GABA, then GABA plus phenobarbital. d, Application of GABA, then GABA plus etomidate. e, Application of GABA, then GABA plus propofol. f, Application of GABA, then GABA plus the methylated form of bicuculline. The patch-clamp experiments were repeated 3 times independently.
Supplementary information
Supplementary Information
This file contains supplementary video legends, uncropped gel from Extended Data Fig. 1a, expanded discussion of GABA alone, GABA plus flumazenil and GABA plus diazepam complexes, and an expanded analysis and discussion of GABA plus picrotoxin and bicuculline complex structures.
Video 1
Rocking movie to illustrate structural details of phenobarbital binding site at α-β interface.
Video 2
Rocking movie to illustrate structural details of phenobarbital binding site at γ-β interface.
Video 3
Rocking movie to illustrate structural details of etomidate binding site at one β-α interface.
Video 4
Rocking movie to illustrate structural details of propofol binding site at one β-α interface.
Video 5
Morphing movie between diazepam and flumazenil complex structures to illustrate conformational differences that give rise to the more expanded nature of the flumazenil complex.
Video 6
Rocking movie of structural superposition of diazepam complex onto flumazenil complex, focusing on details of benzodiazepine site. Colored model is diazepam complex; flumazenil complex is in grey. Superposition was of principal (α) subunits at the α-γ interface.
Video 7
Rocking movie to illustrate structural details of diazepam binding site at β-α interface.
Video 8
Rocking movie to illustrate structural details of diazepam binding site at γ-β interface.
Video 9
Cholesterol accumulation at protein-lipid interfaces. Representative converged snapshot of the GABA + phenobarbital model after 20 μs coarse-grained simulation in brain-lipid mixture, showing restrained protein subunits (α1, green; β2, blue; γ2, yellow) and proximal cholesterol molecules (cyan), viewed from the membrane plane. All other lipids are hidden for clarity.
Video 10
Simulation and coloring as in Supplementary Video 5, showing proximal PIP2 molecules (cyan).
Rights and permissions
About this article
Cite this article
Kim, J.J., Gharpure, A., Teng, J. et al. Shared structural mechanisms of general anaesthetics and benzodiazepines. Nature 585, 303–308 (2020). https://doi.org/10.1038/s41586-020-2654-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-020-2654-5
This article is cited by
-
A release of local subunit conformational heterogeneity underlies gating in a muscle nicotinic acetylcholine receptor
Nature Communications (2024)
-
Lipid nanodisc scaffold and size alter the structure of a pentameric ligand-gated ion channel
Nature Communications (2024)
-
Neurosteroids and their potential as a safer class of general anesthetics
Journal of Anesthesia (2024)
-
Structural interplay of anesthetics and paralytics on muscle nicotinic receptors
Nature Communications (2023)
-
Structural insights into opposing actions of neurosteroids on GABAA receptors
Nature Communications (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.