Abstract
Tropane alkaloids from nightshade plants are neurotransmitter inhibitors that are used for treating neuromuscular disorders and are classified as essential medicines by the World Health Organization1,2. Challenges in global supplies have resulted in frequent shortages of these drugs3,4. Further vulnerabilities in supply chains have been revealed by events such as the Australian wildfires5 and the COVID-19 pandemic6. Rapidly deployable production strategies that are robust to environmental and socioeconomic upheaval7,8 are needed. Here we engineered baker’s yeast to produce the medicinal alkaloids hyoscyamine and scopolamine, starting from simple sugars and amino acids. We combined functional genomics to identify a missing pathway enzyme, protein engineering to enable the functional expression of an acyltransferase via trafficking to the vacuole, heterologous transporters to facilitate intracellular routing, and strain optimization to improve titres. Our integrated system positions more than twenty proteins adapted from yeast, bacteria, plants and animals across six sub-cellular locations to recapitulate the spatial organization of tropane alkaloid biosynthesis in plants. Microbial biosynthesis platforms can facilitate the discovery of tropane alkaloid derivatives as new therapeutic agents for neurological disease and, once scaled, enable robust and agile supply of these essential medicines.
Similar content being viewed by others
Main
Tropane alkaloids (TAs) such as cocaine and atropine are present in plants from the nightshade (Solanaceae), coca (Erythroxylaceae) and bindweed (Convolvulaceae) families. Some TAs, including hyoscyamine and scopolamine, are used to treat neuromuscular disorders ranging from nerve agent poisoning to Parkinson’s disease1,2. Direct chemical syntheses of TAs are not economically viable owing to challenging stereochemistries9. Thus, intensive cultivation of Duboisia shrubs from the nightshade family in Australia, India, Brazil and Saudi Arabia undergirds the global supply for medicinal TAs2,10,11. This agriculture-based supply chain poses three risks to public health. First, overall increasing demand for TA-based medicines already results in recurring supply shortages3,4. Second, regional events, such as the 2019–2020 Australian wildfires, can threaten global supply5. Third, global crises, such as the ongoing COVID-19 pandemic, can threaten local availability owing to demand spikes and disruption to supply chains6,12. The urgency of having options for quickly scaling production of essential medicines to match regional and local demand, free of geopolitical dependencies and robust to environmental and socioeconomic upheaval, is widely recognized7,8.
Phytochemical production using engineered yeast can address many of the vulnerabilities associated with crop cultivation. The rapid generation times and high cell densities achieved in microbial fermentations enable production of target compounds with reduced time, space and resource requirements relative to plant extraction. Cultivation in closed bioreactors can also reduce supply chain susceptibility to environmental and geopolitical disruption, while providing improved batch-to-batch consistency and active ingredient purity.
However, biosynthesis of TAs in Solanaceae exhibits extensive intra- and intercellular compartmentalization, with enzymes active across specific sub-cellular compartments (cytosol, mitochondrion, chloroplast, peroxisome, ER membrane, vacuole), cell types (root pericycle, endodermis, cortex) and tissues (secondary roots)11. Reconstitution of such pathways in yeast is thus made challenging by incompatibilities of enzymes adapted for specific spatial or regulatory contexts, and metabolite transport strategies that are not readily realized in microbial hosts.
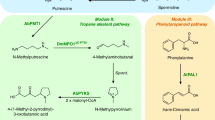
Hyoscyamine and scopolamine comprise an arginine-derived 8-azabicyclo[3.2.1]octane (‘tropine’) acyl acceptor esterified with a phenylalanine-derived phenyllactic acid (PLA) acyl donor (Fig. 1a). The identification of a type III polyketide synthase (PYKS) and cytochrome P450 (CYP82M3) catalysing the cyclization of N-methylpyrrolinium to tropinone in Atropa belladonna13 enabled us and others to engineer yeast strains for de novo production of tropine14,15. The recent report of a UDP-glucosyltransferase (UGT84A27) and serine carboxypeptidase-like (SCPL) acyltransferase (littorine synthase) catalysing the condensation of tropine and phenyllactate to littorine16 resolved a debate about the nature of the acyl transfer reaction9. However, functional expression of plant SCPL acyltransferases (SCPL-ATs) in non-plant hosts has not been reported. Also, although the cytochrome P450 (CYP80F1) that catalyses rearrangement of littorine to hyoscyamine aldehyde17,18 and the 2-oxoglutarate-dependent hydroxylase/dioxygenase (H6H) that catalyses epoxidation of hyoscyamine to scopolamine are established19,20, no enzymatic activity for reduction of hyoscyamine aldehyde to hyoscyamine is known, necessitating discovery of such an enzyme (Fig. 1a).
a, Modular pathway construction for scopolamine biosynthesis in yeast. Enzyme/protein colour scheme: orange, yeast (overexpressed); green, plant; purple, bacteria; red, other eukaryote; grey, spontaneous/non-enzymatic. Red boxes indicate disrupted yeast proteins; dotted or solid lines of vacuole membrane delineate functional biosynthetic modules. DsRed–AbLS, Discosoma sp. red fluorescent protein fused to the N terminus of A. belladonna littorine synthase. b, PLA production in yeast engineered for expression of PPRs or LDHs. Heterologous enzymes or negative control (BFP) were expressed from low-copy plasmids in strain CSY1251. c, Multiple reaction monitoring (MRM) and extracted ion chromatogram (EIC) traces from culture medium of yeast engineered for step-wise reconstitution of PLA glucoside biosynthesis via module III. Chromatogram traces are representative of three biological replicates. d, Relative titres of PLA glucoside in yeast engineered for overexpression of UDP-glucose biosynthetic enzymes. Enzymes or negative control (BFP) were expressed from low-copy plasmids in strain CSY1288. e, Relative PLA glucoside titres in CSY1288 with disruptions to endogenous glucosidases. In d and e, PLA glucoside accumulation was compared using relative titres owing to lack of an authentic chemical standard. Strains were cultured for 72 h before liquid chromatography–tandem mass spectrometry (LC–MS/MS) analysis of metabolites in culture supernatant. Data in b, d and e represent the mean of n = 3 biologically independent samples (open circles), error bars denote s.d. *P < 0.05, **P < 0.01, ***P < 0.001, Student’s two-tailed t-test. Statistical significance is shown relative to controls. Exact P values are in Supplementary Table 5.
TA acyl acceptor and donor biosynthesis
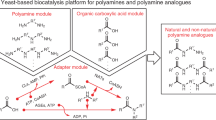
We designed a biosynthetic pathway comprising five functional modules for hyoscyamine and scopolamine production from simple precursors in yeast (Fig. 1a). Modules I/II and III enable de novo biosynthesis of the acyl acceptor and donor moieties; module IV enables TA scaffold modifications to produce hyoscyamine and scopolamine; module V comprises the central acyltransferase reaction linking upstream acyl acceptor/donor biosynthesis to downstream scaffold modifications. As a starting point, we used a yeast platform strain (CSY1251) that was previously engineered for de novo production of the acyl acceptor tropine via modules I and II14 (Extended Data Fig. 1). A putrescine biosynthesis module (I) designed to increase putrescine accumulation incorporated (i) overexpression of glutamate N-acetyltransferase (Arg2), arginase (Car1), ornithine decarboxylase (Spe1) and polyamine oxidase (Fms1); (ii) a parallel plant/bacterial pathway encoded by Avena sativa arginine decarboxylase (AsADC) and Escherichia coli agmatine ureohydrolase (speB); and (iii) disruptions to polyamine regulatory mechanisms encoded by methylthioadenosine phosphorylase (Meu1) and ornithine decarboxylase antizyme-1 (Oaz1). A tropine biosynthesis module (II) incorporated (i) seven enzymes: A. belladonna and Datura stramonium putrescine N-methyltransferases (AbPMT1 and DsPMT1), Datura metel N-methylputrescine oxidase engineered for improved peroxisomal activity (DmMPO1ΔC-PTS1), A. belladonna pyrrolidine ketide synthase (AbPYKS) and tropinone synthase (AbCYP82M3), Arabidopsis thaliana cytochrome P450 reductase (AtATR1) and D. stramonium tropinone reductase 1 (DsTR1); and (ii) disruptions to five aldehyde dehydrogenases (Hfd1, Ald2, Ald3, Ald4 and Ald5) to reduce loss of pathway intermediates.
We designed a third module (III) for production of the acyl donor 1-O-β-phenyllactoylglucose (PLA glucoside) from phenylalanine via aromatic aminotransferases Aro8 and Aro9, phenylpyruvate reductase (PPR) and PLA UDP-glucosyltransferase (UGT84A27)16. Yeast produce 3-phenylpyruvate from phenylalanine via Aro8 and Aro921, and wild-type yeast and CSY1251 produce trace levels of PLA, potentially via nonspecific activity of a lactate dehydrogenase (LDH) acting on 3-phenylpyruvate22. We screened PPRs from E. coli23, Lactobacillus (UniProt A0A2U9AUW1), A. belladonna24 and Wickerhamia fluorescens25 and LDHs from Bacillus and Lactobacillus with reported activity on 3-phenylpyruvate22,26,27 via expression from a plasmid in CSY1251. All screened enzymes yielded modest (1.3- to 3.5-fold) improvements in PLA production relative to control, except for W. fluorescens PPR, which resulted in a nearly 80-fold increase to approximately 250 mg l−1 (Fig. 1b) and was integrated into CSY1251 to make strain CSY1287.
In A. belladonna, PLA is activated for acyl transfer to tropine via glucosylation by UGT84A27 (AbUGT)16. Plant UGTs participate in the biosynthesis of diverse phenylpropanoids and often exhibit broad substrate scope28. We expressed AbUGT from a plasmid in CSY1251 and measured conversion of three phenylpropanoid acyl donors (PLA, cinnamic acid and ferulic acid) to their respective glucosides (Extended Data Fig. 2a, c). Whereas AbUGT glucosylated approximately 60% and 90% of cinnamic acid and ferulic acid, respectively, less than 3% of PLA was glucosylated (Extended Data Fig. 2b). AbUGT orthologues identified from transcriptomes of other TA-producing Solanaceae (Supplementary Note 1) and structure-guided active site mutants (Supplementary Note 2) exhibited poor activity on PLA (Extended Data Fig. 2b, d–f), which suggests that PLA glucosylation may constitute a key limitation in TA production. We constructed strain CSY1288 by integrating codon-optimized W. fluorescens 3-phenylpyruvate reductase (WfPPR) and AbUGT into the genome of CSY1251, and verified PLA production (66 mg l−1) and minimal PLA glucoside accumulation (Fig. 1c).
We increased PLA glucoside levels by incorporating genetic modifications that promote UDP-glucose accumulation and decrease glycoside degradation. We overexpressed the PGM2 and UGP1 genes, which encode proteins that catalyse the isomerization of glucose-6-phosphate to glucose-1-phosphate and the conversion of glucose-1-phosphate to UDP-glucose, respectively, from plasmids in CSY1288. Although overexpression of PGM2 resulted in no improvement relative to control, overexpression of UGP1 resulted in an approximately 1.8-fold increase in the production of PLA glucoside (Fig. 1d). We disrupted three native glucosidase genes—EXG1, SPR1 and EGH1—in CSY1288, as glucosidases have been shown to hydrolyse heterologous glucosides in yeast29. The disruption of EGH1 more than doubled PLA glucoside production (Fig. 1e), indicating that hydrolysis by Egh1 (steryl-β-glucosidase) constitutes a substantial loss of TA precursor from the pathway. We thus incorporated both UGP1 overexpression and EGH1 disruption into a complete TA production strain.
HDH discovery and scopolamine biosynthesis
We used a functional genomics approach to discover the enzyme, hyoscyamine dehydrogenase (HDH), which catalyses the reduction of hyoscyamine aldehyde to hyoscyamine. We searched for genes that co-express with TA biosynthetic genes in secondary root tissues by mining a publicly available A. belladonna transcriptome dataset30. Starting from more than 40,000 identified transcripts, we removed transcripts without putative dehydrogenase or reductase-like domains, and further filtered by clustering tissue-specific expression profiles with those of bait genes AbCYP80F1 (littorine mutase) and AbH6H (Extended Data Fig. 3a). Nearly all candidates exhibited the secondary root-specific expression pattern observed for TA biosynthetic genes. Owing to missing sequence regions, we repeated the de novo transcriptome assembly from raw RNA sequencing (RNA-seq) reads30 using the Trinity software package31 and reconstituted missing fragments for 12 HDH candidates via alignment of incomplete regions against the newly assembled transcriptome (Supplementary Table 1).
We identified the missing HDH activity by screening candidates generated via transcriptome mining in yeast. Lack of an authentic commercial standard for hyoscyamine aldehyde and insufficient yield from chemical syntheses, as well as similar chromatographic and mass spectrometric properties of littorine and hyoscyamine, necessitated screening of HDH candidates by detection of scopolamine (m/z 304 [M + H]+) from fed littorine (m/z 290 [M + H]+) via a three-step biosynthetic pathway (Fig. 1a). We constructed an HDH screening strain (CSY1292) by integrating codon-optimized AbCYP80F1 and an optimal H6H orthologue from D. stramonium (DsH6H) (Extended Data Fig. 4) into the genome of CSY1251, and expressed codon-optimized HDH candidates from a plasmid. One of the candidates, HDH2 (that is, AbHDH), exhibited a 35% decrease in hyoscyamine aldehyde levels and accumulation of scopolamine (7.2 μg l−1), indicating the missing HDH activity (Fig. 2a).
a, Production of hyoscyamine aldehyde and scopolamine in yeast engineered for expression of A. belladonna HDH candidates. Candidates or a negative control (BFP) were expressed from low-copy plasmids in CSY1292. Accumulation of hyoscyamine aldehyde was compared using relative titres owing to lack of an authentic chemical standard. Amino acid sequences are in Supplementary Table 1. Data represent the mean of n = 3 biologically independent samples (open circles), error bars denote s.d. *P < 0.05, **P < 0.01, ***P < 0.001, Student’s two-tailed t-test. Statistical significance is shown relative to control. Exact P values are in Supplementary Table 5. b, Homology model of AbHDH. NADPH and Zn2+ are shown in orange and pink, respectively. Box shows magnified view of AbHDH active site with NADPH and docked hyoscyamine aldehyde. Dashed lines indicate interactions important for catalysis. c, MRM traces from culture media of yeast engineered for step-wise reconstitution of module IV for conversion of littorine to scopolamine. Blue trace represents 125 nM (38 μg l−1) scopolamine standard. Chromatogram traces are representative of three biological replicates. In a and c, strains were cultured for 72 h with 1 mM littorine before LC–MS/MS analysis of metabolites in culture supernatant.
Structural and phylogenetic analyses provided insight into the catalytic mechanism and evolutionary history of HDH (Supplementary Notes 3, 4). Homology modelling indicated that AbHDH is a zinc-dependent alcohol dehydrogenase of the medium-chain dehydrogenase/reductase (MDR) superfamily and probably uses NADPH as the hydride donor for hyoscyamine aldehyde reduction (Fig. 2b, Supplementary Note 3). Ligand docking simulations and active site mutants suggested a mechanism in which the oxyanion intermediate formed upon hydride attack of hyoscyamine aldehyde is stabilized by a catalytic Zn2+, which is bound by Cys52, His74, Cys168 and a displaceable water molecule positioned by polar interactions with Ser54 (Fig. 2b, Extended Data Fig. 3b, Supplementary Note 3). We identified orthologues of AbHDH from transcriptomes of Datura innoxia (DiHDH) and D. stramonium (DsHDH)32, and verified their activity via co-expression with an additional copy of DsH6H from plasmids in CSY1292. DsHDH showed the highest substrate depletion and product accumulation of the variants tested (Extended Data Fig. 3c, d).
We reconstituted the medicinal TA biosynthetic branch (module IV) comprising optimal enzyme variants and overexpression of a limiting enzyme into our platform strain. Strain CSY1294 was constructed by integrating codon-optimized WfPPR and AbUGT (module III), DsHDH, and an additional copy of DsH6H, which limits scopolamine accumulation (Extended Data Fig. 3d), into CSY1292. Scopolamine production from fed littorine was verified in CSY1294 (Fig. 2c). Strain CSY1294 incorporates the enzymes for producing the acyl acceptor (tropine; modules I/II) and acyl donor (PLA glucoside; module III) for littorine biosynthesis, and the enzymes for modification of the TA scaffold to scopolamine (module IV), leaving the central acyltransferase reaction catalysed by littorine synthase (module V) as the final enzymatic step to implement.
Engineering vacuolar littorine biosynthesis
Recently, littorine biosynthesis in A. belladonna was demonstrated to occur via esterification of glucosylated PLA with tropine by an acyltransferase of the SCPL family (littorine synthase, AbLS)16. Few plant SCPL-ATs have been studied and no reports of in vivo activity in non-plant hosts have emerged, owing to difficulties of extensive post-translational processing and trafficking in microbial hosts33. SCPL-ATs are expressed via the secretory pathway and localize to the plant tonoplast33 (Extended Data Fig. 6a). An N-terminal signal peptide directs the nascent polypeptide to the ER, where it undergoes processing steps for folding—signal peptide cleavage, disulfide bond formation, and, in some cases, proteolytic removal of propeptide sequences. The partially folded SCPL-AT protein is transported through the Golgi, where it acquires N-glycosylation on asparagine residues within N-X-S/T motifs (in which X is not proline). Recognition of cryptic signal sequences by vacuole-associated transport factors directs SCPL-AT to the vacuole lumen34. Although the yeast secretory pathway possesses much the same compartments and processing steps as in plants, it is unlikely that yeast transport factors recognize the same signal sequences and yeast protein glycosylation patterns differ from those of plants35. Our initial attempts to express wild-type AbLS in CSY1294 resulted in a severe growth defect and no detectable TA biosynthesis.
We then showed that terminal and internal peptide sequences impact processing and localization of SCPL-ATs in yeast. A putative N-terminal signal peptide in AbLS suggested that it follows the expected SCPL-AT ER-to-vacuole trafficking pathway in planta. Fluorescence microscopy of N- and C-terminal green fluorescent protein (GFP) fusions of AbLS expressed from plasmids in CSY1294 revealed that the N-terminal fusion (GFP–AbLS) co-localized with a vacuolar membrane stain (Fig. 3a, Extended Data Fig. 6b), whereas no fluorescence was detected for the C-terminal fusion (AbLS–GFP), consistent with reports that a native C terminus is crucial for SCPL-AT folding36. To identify possible failure points in AbLS expression, maturation and trafficking in yeast, we screened AbLS variants engineered for localization to subcellular compartments (Supplementary Note 5) and compared AbLS N-glycosylation patterns in tobacco and yeast (Extended Data Fig. 6c–h, Supplementary Note 6), which did not implicate mis-targeting or mis-glycosylation as primary factors impeding activity in yeast. Characterization of AbLS endoproteolytic processing based on identification of a putative internal propeptide sequence suggested that the enzyme may become stalled in the yeast secretion pathway upstream of the trans-Golgi network (TGN) (Extended Data Figs. 6g, h, 7, Supplementary Note 7). This potential disruption of TGN sorting may account for the lack of activity and growth defect observed in CSY1294 expressing wild-type AbLS.
a, Yeast epifluorescence microscopy showing N-terminal GFP-tagged AbLS (GFP–AbLS), vacuolar membrane stain FM4-64, and bright-field merged images. Microscopy was performed on CSY1294 expressing GFP–AbLS from a low-copy plasmid. 2D deconvolution was performed as described in the Methods. Scale bar, 5 μm. Images are representative of two independent experiments. b, De novo hyoscyamine and scopolamine production in engineered yeast expressing AbLS N-terminal fusions. Table shows expected oligomerization state of each N-terminal domain; half-circles (eGFP, mCherry) indicate monomer/weak dimer. Wild-type (control) or AbLS fusions were expressed from low-copy plasmids in CSY1294. Transformed strains were cultured for 96 h before LC–MS/MS analysis of metabolites in culture supernatant. No littorine was detected, indicating complete conversion to downstream TAs. Data represent the mean of n = 3 biologically independent samples (open circles), error bars denote s.d. *P < 0.05, **P < 0.01, ***P < 0.001, Student’s two-tailed t-test. Exact P values are in Supplementary Table 5.
Functional expression of AbLS in yeast was achieved by engineering N-terminal fusions that may alter sorting from the TGN. Transport of soluble proteins from the TGN to the vacuole requires recognition of a typically N-terminal signal sequence by vacuole protein sorting (Vps) cargo transport proteins, whereas integral membrane proteins that reach the yeast TGN are sorted to the vacuole by default37,38. We hypothesized that conversion of AbLS into a transmembrane protein by masking the signal peptide with an N-terminally fused soluble domain might resolve the putative obstruction in TGN sorting (Fig. 3a, Supplementary Note 8). We constructed AbLS variants with N-terminally fused soluble domains, including fluorescent proteins from Aequoria (eGFP, tagBFP, mVenus) and Discosoma (mCherry, DsRed.T3); small ubiquitin-related modifier (Smt3) with a mutated protease cleavage site (SUMO*)39; and AbUGT. We expressed these variants and wild-type AbLS from plasmids in CSY1294. Enhancement of AbLS activity appeared to be correlated with the N-terminal domain oligomerization state, with scopolamine production increasing from monomeric or weakly dimeric (GFP, BFP, mVenus, mCherry and SUMO*) to homodimeric (AbUGT) and homotetrameric (DsRed) domains; reaching de novo hyoscyamine and scopolamine titres up to 10.3 μg l−1 and 0.87 μg l−1, respectively (Fig. 3b). To generate a strain containing all five metabolic modules for complete TA biosynthesis (modules I–V) (Fig. 1a), we integrated a codon-optimized DsRed–AbLS and an additional copy of UGP1 into the genome of CSY1294 at the disrupted EGH1 site to generate CSY1296. CSY1296 exhibited de novo hyoscyamine and scopolamine titres of 10.2 μg l−1 and 1.0 μg l−1, respectively.
Inter-compartment transport limitations were addressed by incorporation of plant transporters. Vacuolar compartmentalization of DsRed–AbLS in CSY1296 (Extended Data Fig. 8) necessitates import of cytosolic tropine and PLA glucoside to the vacuole lumen and export of vacuolar littorine to the cytosol. Several multidrug and toxin extrusion (MATE) transporters responsible for vacuolar alkaloid and glycoside sequestration have been identified in Solanaceae, including three with observed or predicted activity on TAs40,41. We expressed Nicotiana tabacum jasmonate-inducible alkaloid transporter 1 (NtJAT1) and two MATEs (NtMATE1, NtMATE2) from plasmids in CSY1296. Expression of NtJAT1 and NtMATE2 improved TA production; the former resulting in 74% and 18% increases in hyoscyamine and scopolamine titres, respectively (Fig. 4a). Fluorescence microscopy of CSY1296 expressing C-terminal GFP fusions of NtJAT1 or NtMATE2 from plasmids supports the hypothesis that NtJAT1 localizes to the vacuolar membrane (co-localizing with DsRed–AbLS), whereas NtMATE2 is partitioned between vacuolar and plasma membranes (Extended Data Fig. 8), which suggests that both transporters might function to alleviate vacuolar substrate transport limitations while the latter might also improve cellular TA export (Fig. 4b).
a, Production of tropine, hyoscyamine and scopolamine in CSY1296 engineered for expression of heterologous alkaloid transporters. NtJAT1, MATE transporters 1/2, or a negative control (BFP) were expressed from low-copy plasmids in CSY1296 and transformed strains were cultured for 96 h. b, Illustration of proposed DsRed–AbLS trafficking and alleviation of substrate transport limitations via heterologous transporter expression in engineered yeast. Putative transport activities based on microscopy studies are indicated; ‘???’ indicates unknown transport mechanism. Circled numbers indicate major proposed steps in DsRed-AbLS expression and activity, including maturation in (1) ER lumen and (2) Golgi, (3) trafficking to vacuole membrane, vacuolar (4) substrate import and (5) product export and (6) cellular TA export. c, Summary of strain and media optimization for de novo scopolamine production in engineered yeast. Strains were cultured in non-selective (CSY1296, CSY1297) or selective (CSY1298: leucine dropout) medium with or without cofactors (50 mM 2-oxoglutarate, 15 mg l−1 Fe2+) at 25 °C for 96 h. Strain CSY1298 is prototrophic for leucine and contains a blank plasmid with the LEU2 gene (pCS4213). In a and c, metabolite titres in culture supernatant were quantified by LC–MS/MS. Data indicate the mean of n = 3 biologically independent samples (open circles), error bars denote s.d. *P < 0.05, **P < 0.01, ***P < 0.001, Student’s two-tailed t-test. Exact P values are in Supplementary Table 5.
Improvements in TA production were achieved via overexpression of limiting enzymes and media optimization. Additional copies of WfPPR and DsH6H expressed from plasmids in CSY1296 resulted in 64% and 89% increases in hyoscyamine and scopolamine titres, respectively (Extended Data Fig. 9). Supplementation with iron and 2-oxoglutarate (2-OG), required for H6H activity19,42, resulted in 9.0- and 3.4-fold increases in hyoscyamine and scopolamine titres from CSY1296 (Fig. 4c). We constructed an optimized strain (CSY1297) by integrating NtJAT1 and additional copies of WfPPR and DsH6H into CSY1296, which showed 2.4- and 7.1-fold respective increases in hyoscyamine and scopolamine accumulation (Fig. 4c). Removing leucine auxotrophy by expressing 3-isopropylmalate dehydrogenase (Leu2) from a plasmid in CSY1297 (denoted CSY1298) increased conversion of hyoscyamine (85% decrease) to scopolamine (more than 3-fold increase) (Fig. 4c), potentially by improving access to Fe2+ via increased NADH regeneration43. Pseudo-fed-batch, high-density, shake-flask cultures grown in optimized media showed no littorine accumulation, hyoscyamine and scopolamine titres of approximately 30 μg l−1 in CSY1297 and CSY1298, respectively, and tropine and PLA accumulation up to 3 mg l−1 and 160 mg l−1 (Extended Data Fig. 10, Supplementary Note 9), suggesting incorporation of PLA into littorine is a major limitation and target for future improvement.
Discussion
Our final strain comprises 34 chromosomal modifications (26 genes, 8 gene disruptions), resulting in an integrated whole-cell system that expresses enzymes and transporters in diverse sub-cellular locations (cytosol, mitochondria, peroxisome, vacuole, ER and vacuolar membranes) (Supplementary Note 10). Combining functional genomics with our synthesis platform, we identified an oxidoreductase that catalyses the remaining uncharacterized step in the biosynthesis of hyoscyamine and scopolamine. We developed an N-terminal fusion strategy to achieve functional expression of the key TA scaffold-generating enzyme AbLS. Our strategy may improve folding and trafficking of the engineered transmembrane AbLS through the secretory pathway to the vacuole (Supplementary Note 8), potentially enabling heterologous expression of plant SCPL-ATs and expanding the diversity of natural product biosyntheses in yeast33. We used a plant vacuolar alkaloid importer to address import restrictions in that compartment. Although plasma membrane transporters have been used to improve cellular export and import of metabolites44,45, our work demonstrates that incorporation of plant transporters can facilitate intracellular transport and help reconstruct sub-cellular compartmentalization inherent to many plant biosynthetic pathways.
Our demonstration of total biosynthesis of hyoscyamine and scopolamine via engineered yeast suggests that centralized, plantation-based supply of medicinal TAs can be complemented or replaced by industrial fermentation. Process improvements to increase productivities from titres reported here (around 30 to 80 μg l−1) (Fig. 4c), which are typical of first implementations of complex plant natural product pathways46,47,48, to commercial production (approximately 5 g l−1) are becoming routine49, and we anticipate would take 1–2 years of focused effort by a professional team (Supplementary Note 11). From a land-use perspective, we estimate that a fermentation-based process sourcing sugar from sugarcane would require at least 10-fold less land than the existing Duboisia farming-based approach (Supplementary Note 12). Transitioning from agriculture- to fermentation-based production could have many indirect effects ranging from land-use and natural biodiversity, to labour markets and livelihoods, to supply-chain decouplings and geopolitical interdependencies50. Practically, because a fermentation-based approach can be implemented where needed and operated with a process time of days, our results support development of flexible manufacturing platforms enabling robust and agile supply of essential medicines.
Methods
Data reporting
No statistical methods were used to predetermine sample size. The experiments were not randomized and investigators were not blinded to allocation during experiments and outcome assessment.
Chemical compounds and standards
Tropine, (S)-hyoscyamine hydrobromide and (S)-scopolamine hydrobromide were purchased from Santa Cruz Biotechnology. (R)-Littorine hydrochloride was purchased from Toronto Research Chemicals. All other chemicals were purchased from Sigma.
Plasmid construction
DNA oligonucleotides used in this study were synthesized by the Stanford Protein and Nucleic Acid Facility and are listed in Supplementary Data 1. Genes encoding biosynthetic enzymes used in this study are listed by source and accession number in Supplementary Table 2; for HDH genes newly identified in this work, full amino acid sequences are given in Supplementary Table 1. Endogenous yeast genes were amplified from Saccharomyces cerevisiae CEN.PK2-1D51 genomic DNA via colony PCR52. Gene sequences encoding heterologous enzymes were codon-optimized for expression in S. cerevisiae using GeneArt GeneOptimizer software (Thermo Fisher Scientific) and synthesized as double-stranded gene fragments (Twist Bioscience). Plasmids used in this study are listed in Supplementary Data 2. Three types of plasmids were used in this work: yeast expression plasmids, yeast integration plasmids, and Agrobacterium tumefasciens binary vectors.
Yeast expression plasmids harboured a gene of interest flanked by a constitutive promoter and terminator, an auxotrophic selection marker, and either a low-copy CEN6/ARS4 or a high-copy 2μ yeast origin of replication. These plasmids were constructed by addition of 5′ and 3′ restriction sites to genes of interest using PCR, restriction digestion of PCR amplicons or synthesized gene fragments, and ligation of digested inserts into similarly digested vectors pAG414GPD-ccdB, pAG415GPD-ccdB, pAG416GPD-ccdB, pAG424GPD-ccdB, pAG425GPD-ccdB or pAG426GPD-ccdB53 using T4 DNA ligase (New England Biolabs, NEB). Yeast expression plasmids expressing fusions of multiple proteins or enzymes were prepared by PCR amplification of each gene of interest with 15–25 bp of overlap to adjacent fragments, assembly of fragments into single inserts with 5′ and 3′ restriction sites using overlap-extension PCR, and ligation cloning into digested vectors as described.
Yeast integration plasmids comprised a gene of interest flanked by a constitutive promoter and terminator, but lacked a selection marker and origin of replication for yeast expression. These plasmids were constructed by PCR linearization of the empty holding vectors pCS2656, pCS2657, pCS2658, pCS2661 or pCS2663 using primers complementary to the 3′ and 5′ ends of the promoter and terminator, respectively. Genes intended for yeast genomic integration were PCR-amplified to append 5′ and 3′ overhangs with 35–40 bp of homology to the termini of the linearized holding vectors and then assembled using Gibson assembly.
For transient expression of littorine synthase variants in Nicotiana benthamiana, A. tumefasciens binary vectors contained a transfer-DNA (T-DNA) region comprising a gene of interest flanked by the constitutive Cauliflower Mosaic Virus (CaMV) 35S promoter/Cowpea Mosaic Virus (CPMV) 5′UTR and a nopaline synthase terminator, as well as an analogous expression cassette for the p19 RNAi-suppressor protein. These plasmids were constructed via addition of 5′ AgeI and 3′ XhoI restriction sites to a gene of interest via PCR, followed by digestion and ligation into the pEAQ-HT binary vector pCS335254.
All PCR amplification was performed using Q5 DNA polymerase (NEB) and linear DNA was purified using the DNA Clean and Concentrator-5 kit (Zymo Research). Assembled plasmids were transformed into chemically competent E. coli (TOP10, Thermo Fisher Scientific) via heat-shock and propagated with selection in Luria–Bertani (LB) broth or on LB-agar plates with either carbenicillin (100 μg ml−1) or kanamycin (50 μg ml−1) selection. E. coli plasmid DNA was isolated by alkaline lysis from overnight cultures grown at 37 °C and 250 rpm in selective LB media using Econospin columns (Epoch Life Science) according to the manufacturer’s protocol. Plasmid sequences were verified by Sanger sequencing (Quintara Biosciences).
Yeast strain construction
Yeast strains used in this study (Supplementary Table 3) were derived from our previously reported tropine-producing strain CSY125114, which is in turn derived from the parental strain CEN.PK2-1D51. Strains were grown non-selectively in yeast-peptone media supplemented with 2% w/v dextrose (YPD media), yeast nitrogen base (YNB) defined media (Becton, Dickinson and Company, BD) supplemented with synthetic complete amino acid mixture (YNB-SC; Clontech) and 2% (w/v) dextrose, or on agar plates of the aforementioned media. Strains transformed with plasmids bearing auxotrophic selection markers (URA3, TRP1 and/or LEU2) were grown selectively in YNB media supplemented with 2% w/v dextrose and the appropriate dropout solution (YNB-DO; Clontech) or on YNB-DO agar plates.
Yeast genomic modifications were performed using the CRISPRm method55. CRISPRm plasmids expressing Streptococcus pyogenes Cas9 and a single guide RNA (sgRNA) targeting a genomic locus were constructed by assembly PCR and Gibson assembly of DNA fragments encoding SpCas9 (pCS3410), tRNA promoter and HDV ribozyme (pCS3411), a 20-nucleotide guide RNA sequence oligonucleotide, and tracrRNA and terminator (pCS3414) (Supplementary Data 2). For gene insertions, integration fragments comprising one or more genes of interest flanked by unique promoters and terminators were PCR-amplified from yeast integration plasmids using Q5 DNA polymerase (NEB) with flanking 40 bp microhomology regions to adjacent fragments and/or to the yeast genome at the integration site (Extended Data Fig. 1, Supplementary Data 1). For gene disruptions, integration fragments comprised 6–8 stop codons in all three reading frames flanked by 40 bp of microhomology to the disruption site, which was located within the first half of the open reading frame. Approximately 0.5–1 μg of each integration fragment was co-transformed with 500 ng of multiplex CRISPR plasmid targeting the desired genomic site. Positive integrants were identified by yeast colony PCR52, Sanger sequencing, and/or functional screening by LC–MS/MS.
Yeast transformations
Yeast strains were chemically transformed using the Frozen-EZ Yeast Transformation II Kit (Zymo Research) as per the manufacturer’s instructions, with the following modifications. For competent cell preparation, individual colonies were inoculated into YPD media and grown overnight at 30 °C and 460 rpm. Saturated cultures (~14–18 h) were back-diluted between 1:10 and 1:50 in fresh YPD media and grown to exponential phase (~5–7 h). Cultures were pelleted by centrifugation at 500g for 4 min, washed twice with 50 mM Tris-HCl buffer (pH 8.5), and then resuspended in 20–50 μl of EZ2 solution per transformation. For transformation, competent cells were mixed with 250–1,000 ng of total DNA and 200–500 μl of EZ3 solution. Cell suspensions were incubated at 30 °C with slow rotation for 1–1.5 h. For plasmid transformations, the transformed yeast were directly plated onto YNB-DO agar plates. For CRISPRm genomic modifications, yeast suspensions were instead mixed with 1 ml YPD media, pelleted by centrifugation at 500g for 4 min, and then resuspended in 300–500 μl of fresh YPD medium. Suspensions were incubated at 30 °C with gentle rotation for 2–3 h to allow expression of geneticin resistance and then spread on YPD plates supplemented with 200–400 mg l−1 G418 (geneticin) sulfate. Plates were incubated at 30 °C for 72 h to allow sufficient colony formation before downstream applications.
Growth conditions for metabolite assays
Small-scale metabolite production assays were performed in YNB-SC or YNB-DO media supplemented with 2% dextrose and 5% glycerol (YNB-G) for optimal tropine production14 in at least three replicates. Our previous work showed that tropine biosynthesis is significantly enhanced by higher starting cell densities14. Therefore, yeast colonies were initially inoculated in triplicate into 1 ml YPD or YNB-DO and grown to saturation (~18–22 h) at 30 °C and 460 rpm, pelleted by centrifugation at 500g for 4 min and 3,000g for 1 min, resuspended in 1 ml of fresh selective or non-selective YNB-G media (for some experiments, additionally supplemented with 15 mg l−1 Fe2+ from iron (ii) sulfate and 50 mM 2-oxoglutarate56), and then 300 μl transferred into 2 ml deep-well 96-well plates sealed with AeraSeal gas-permeable film (Excel Scientific). Cultures were grown for 72–96 h at 25 °C, 460 rpm, and 80% relative humidity in a Lab-Therm LX-T shaker (Adolf Kuhner).
Growth conditions for time courses
To simulate high-density batch culture conditions, strains were inoculated in triplicate into 10 ml of YPD media or selective YNB-G media and grown overnight to saturation at 30 °C and 250 rpm. Saturated cultures were pelleted by centrifugation at 500g for 4 min and 3,000g for 1 min and then resuspended in 10 ml of fresh selective or non-selective YNB-G media supplemented with 50 mM 2-oxoglutarate and 15 mg l−1 Fe2+, and grown in 50-ml shake flasks with 10 ml starting volume in triplicates at 25 °C and 300 rpm for 120 h. Where indicated, fed-batch conditions were approximated by supplementing cultures after 72 h of growth with appropriate carbon sources and amino acids at 2% and 1× final concentrations, respectively. At appropriate time points, 250 μl samples were removed from cultures for analysis; 100 μl of culture was diluted 10× and used for optical density measurement at 600 nm on a Nanodrop 2000c spectrophotometer, and 150 μl of culture was used for metabolite quantification.
Analysis of metabolite production
Yeast cultures were pelleted by centrifugation at 3,500g for 5 min at 12 °C and 150 μl aliquots of supernatant were removed for analysis. Metabolites were analysed by LC–MS/MS using an Agilent 1260 Infinity Binary HPLC and an Agilent 6420 Triple Quadrupole mass spectrometer. Chromatography was performed using a Zorbax EclipsePlus C18 column (2.1 × 50 mm, 1.8 μm; Agilent Technologies) with 0.1% (v/v) formic acid in water as mobile phase solvent A and 0.1% (v/v) formic acid in acetonitrile as solvent B. The column was operated with a constant flow rate of 0.4 ml min−1 at 40 °C and a sample injection volume of 10 μl. Chromatographic separation was performed using the following gradient14: 0.00–0.75 min, 1% B; 0.75–1.33 min, 1–25% B; 1.33–2.70 min, 25–40% B; 2.70–3.70 min, 40–60% B; 3.70–3.71 min, 60–95% B; 3.71–4.33 min, 95% B; 4.33–4.34 min, 95–1% B; 4.34–5.00 min, equilibration with 1% B. For separation and detection of phenylpropanoid acyl donors (PLA, cinnamate and ferulate) and corresponding glucosides, the final equilibration step at 1% B was extended to 4.34–7.50 min. The LC eluent was directed to the MS from 0.01–5.00 min operating with electrospray ionization (ESI) in positive mode, source gas temperature 350 °C, gas flow rate 11 l min−1, and nebulizer pressure 40 psi. Data collection was performed using MassHunter Workstation LC/MS Data Acquisition software (Agilent). Metabolites were identified and quantified by integrated peak area in MassHunter Workstation Qualitative Analysis Navigator software (Agilent) using the mass fragment/transition parameters in Supplementary Table 4 and standard curves. Primary MRM transitions were identified by analysis of 0.1–1 mM aqueous standards using MassHunter Workstation Optimizer software (Agilent) and corroborated against published mass transitions if available, and/or against predicted transitions determined using the CFM-ID fragment prediction utility57 and the METLIN database58. As both PLA and its glucoside formed strong ammonium adducts, these metabolites were detected and quantified in positive mode using the corresponding [M + H + 17]+ ions, m/z 184 (PLA) and 346 (PLA glucoside) (Supplementary Table 4).
Fluorescence microscopy
Individual colonies of yeast strains transformed with plasmids encoding biosynthetic enzymes fused to fluorescent protein reporters were inoculated into 1 ml selective or non-selective YNB-G media and grown overnight (~14–18 h) at 30 °C and 460 rpm. Overnight cultures were back-diluted between 1:2 and 1:4 into fresh YNB-G media and grown to exponential phase at 30 °C and 460 rpm for an additional 6–8 h to allow slow-maturing fluorescent proteins to fold before imaging.
Yeast vacuoles were co-imaged with fluorescent reporter-fused biosynthetic enzymes using the FM4-64 stain (Thermo Fisher) and pulse-chase fluorescence microscopy. FM4-64 is a red-fluorescent lipophilic styryl dye that intercalates into the yeast plasma membrane and is endocytosed during growth on rich media, accumulating in vacuolar membranes59. Transformed yeast colonies were inoculated into 1 ml selective or non-selective YNB-G and grown overnight (~14–18 h) at 30 °C and 460 rpm, then back-diluted between 1:10 and 1:3 into 1 ml of fresh YNB-G and grown for an additional 2–4 h until OD600 value of 0.5–0.8. Cultures were pelleted by centrifugation at 5,000g for 5 min, resuspended in 500 μl fresh YPD with 8 μM (5 ng μl−1) FM4-64, and incubated at 30 °C for 30 min with gentle rotation. Stained cells were pelleted by centrifugation at 3,000g for 5 min (pellets were visibly red), washed twice with 1 ml YPD, resuspended in 5 ml YPD, and then incubated at 30 °C and 460 rpm for 90–120 min to allow endocytosis and vacuolar accumulation of the dye. Cultures were pelleted by centrifugation at 500g for 4 min followed by 3,000g for 1 min, then resuspended in 250 μl of 40 mM MES buffer (pH 6.5) and imaged immediately.
For imaging, approximately 5–10 μl of cell suspension was spotted onto a glass microscope slide and covered with a glass coverslip (Thermo Fisher) and then imaged using an upright Zeiss AxioImager Epifluorescence/Widefield microscope with a × 64 oil immersion objective. Fluorescence excitation was performed using an EXFO X-Cite 120 illumination source and the following Semrock Brightline filter settings: GFP, 472/30 excitation and 520/35 emission; mCherry/DsRed/Cy3/TexasRed, 562/40 excitation and 624/40 emission. Emitted light was captured with a Zeiss Axiocam 503 mono camera and Zen Pro software, and subsequent image analysis was performed in ImageJ/Fiji (NIH). Images were converted to pseudocolor using the ‘merge channels’ and ‘split channels’ functions (Image→ Colour→Merge/Split Channels). For each sample, linear histogram stretching was applied across all images for a given channel to improve contrast.
To reduce the interference of light from other focal planes when imaging sub-cellular organelles, we performed 2D digital deconvolution analysis, a common computational technique used for removing out-of-focus light distortion from 2D images of 3D structures60. First, a theoretical point-spread function (PSF), which mathematically describes the diffraction of light from a point source in a specific imaging setup, was computed using the ‘Diffraction PSF 3D’ plugin for ImageJ (available from http://fiji.sc/Diffraction_PSF_3D) for the green and red channels using the following parameters: index of refraction of the media, 1.518 (lens oil); numerical aperture, 1.40; wavelength (nm), 520 (green) or 624 (red); longitudinal spherical aberration at max. aperture (nm), 0.00 (default); image pixel spacing (nm), 72; slice spacing (nm), 0; width (pixels), 240; height (pixels), 242; depth (slices), 1; normalization, sum of pixel values = 1. Next, green and red channel images were separately deconvolved against the corresponding PSFs using the ‘Parallel Spectral Deconvolution 2D’ plugin for ImageJ (available from http://fiji.sc/Parallel_Spectral_Deconvolution) with default settings and auto regularization.
Identification of HDH candidates
Tissue-specific abundances (fragments per kilobase of contig per million mapped reads, FPKM) and putative protein structural and functional annotations for each of 43,861 unique transcripts identified from the A. belladonna transcriptome were obtained from the MSU Medicinal Plant Genomics Resource30. Transcripts encoding hyoscyamine dehydrogenase candidates were identified based on clustering of tissue-specific expression profiles with those of the bait genes CYP80F1 (littorine mutase) and H6H (hyoscyamine 6β-hydroxylase/dioxygenase), which respectively precede and follow the dehydrogenase step in the TA biosynthetic pathway, using a custom R script which is described below.
First, the complete list of 43,861 transcripts was filtered for those annotated with any of the following oxidoreductase protein family (PFAM) IDs: PF00106, PF13561, PF08659, PF08240, PF00107, PF00248, PF00465, PF13685, PF13823, PF13602, PF16884 and PF00248; or any of the following functional annotation keywords: alcohol dehydrogenase, aldehyde reductase, short chain, aldo/keto. In addition, any transcripts with functional annotations containing the keywords putrescine, tropinone and tropine were included in the filter as positive control TA-associated genes to validate clustering with bait genes. Next, mean tissue-specific expression profiles were generated for the CYP80F1 and H6H bait genes. For each of the two bait genes, linear regression models were constructed to express the bait gene expression profile (in FPKM) as a linear function of each candidate gene profile and correlation P values were computed for each candidate. The candidates identified using each of the two bait genes were pooled and duplicates were removed. Combined P values for each candidate were computed as the sum of the log10(P values) of the correlations with each of the two bait genes. Transcripts matching known dehydrogenases in the TA biosynthetic pathway (that is, tropinone reductases I and II) were removed, and the remaining candidates were ranked by combined P value and by distance from bait genes via hierarchical clustering of tissue-specific expression profiles.
De novo transcriptome assembly
All data pre-processing, analysis, and de novo transcriptome assembly was performed on the Sherlock2.0 high-performance computing cluster (HPCC) at Stanford University. Paired-end raw RNA-seq reads corresponding to A. belladonna secondary roots (accession SRX060267, run SRR192881) and sterile seedling tissue (accession SRX060269, run SRR192882) were retrieved from the NCBI Sequence Read Archive (SRA) using the SRA Toolkit (NIH). The paired-end raw reads were analysed via FastQC (Babraham Bioinformatics) and then trimmed using the BBDuk.sh utility (Joint Genome Institute, Department of Energy) using the following parameters: k-mer trimming, right end only (‘ktrim=r’); k-mer length, 23 (‘k=23’); minimum k-mer for end-trimming, 11 (‘mink=11’), Hamming distance for k-mer matching, 1 (‘hdist=1’); trim paired reads to same length (‘tpe’), trim adapters using pair overlap detection (‘tbo’); quality trimming, both right and left ends (‘qtrim=rl’); quality cut-off, 5 (‘trimq=5’); minimum permissible read length after trimming, 25 (‘minlen=25’). Two independent de novo transcriptome assemblies were generated from the processed paired-end reads from secondary root (SRR192881) and seedling (SRR192882), respectively, using the Trinity software suite with default settings31,61.
Transcript functional annotation for each of the two assemblies (secondary root and seedling) was performed using the Trinotate package62. Following coding region prediction using the TransDecoder.LongOrfs and TransDecoder.Predict commands, annotations were generated using a BLASTp search against the UniProt/SwissProt databases and a protein homology search using HMMER. Complete ORF sequences for each of the candidate transcripts identified from co-expression analysis were generated by performing tBLASTn and tBLASTx searches against the Trinity transcriptome assemblies; hits with protein percent identity of at least 98%, accounting for sequencing errors, were assumed to be identical.
Identification of orthologues from transcriptome databases
Orthologues of A. belladonna UGT84A27 (UGT) were identified using tBLASTn searches of the transcriptomes of Brugmansia sanguinea and Datura metel in the 1000Plants (1KP) database63. This search yielded two unique, full-length amino acid sequences (that is, within 5% of the length of the query sequence) and with expectation value 0.0: scaffold-AIOU-2012986-Brugmansia_sanguinea (B. sanguinea, BsUGT) and scaffold-JNVS-2051323-Datura_metel (D. metel, DmUGT).
Orthologues of HDH were identified using tBLASTn searches of the transcriptomes of several Datura species in the Medicinal Plant RNA-seq database32. This search yielded two unique, full-length amino acid sequences (that is, within 5% of the length of the query sequence) and with expectation value 0.0: medp_datin_20101112|6354 (DiHDH) and medp_datst_20101112|10433 (DsHDH).
Coding sequences for all putative orthologues were optimized for yeast expression and then cloned into expression vectors as described in ‘Plasmid construction’.
Protein structural analysis and substrate docking
Homology models of AbUGT, AbHDH and AbLS were constructed using RaptorX with default modelling parameters64. For docking simulations, the binding of cosubstrates (UDP-glucose for AbUGT) or cofactors (NADPH for AbHDH) was first predicted based on structural alignment with the crystal structures of A. thaliana salicylate UDP-glucosyltransferase UGT74F2 with bound UDP (PDB: 5V2K) and Populus tremuloides sinapyl alcohol dehydrogenase with bound NADPH (PDB: 1YQD) respectively, as the binding pockets for these cosubstrates are tightly conserved. Geometry optimizations of substrate structures (PLA or hyoscyamine aldehyde) before docking simulations were conducted using the Gaussian 16 software package on the Stanford Sherlock2.0 HPCC (run parameters: DFT, B3LYP, LANL2DZ). The energy-minimized ligand structures were then docked into the corresponding cosubstrate/cofactor-bound homology models using the Maestro and Glide XP software packages (Schrodinger) with default parameters; for the docking of hyoscyamine aldehyde into AbHDH, a spatial constraint of maximum 6 Å separation between the aldehyde carbon and the catalytic Zn2+ was additionally imposed65. Enzyme structures and docking results were visualized using PyMOL software (Schrodinger).
Phylogenetic analysis of HDH orthologues
Phylogenetic tree construction was based on a BLASTp search using AbHDH as a query against the UniProt/SwissProt database (annotated sequences only). Sequences chosen for tree construction included the top 50 BLASTp hits based on E-value, as well as 10 additional hits selected from among the next 100 ranks. Phylogenetic relationships were derived via bootstrap neighbour-joining with n = 1,000 trials in ClustalX2 and the resulting tree was visualized with FigTree software.
Expression of littorine synthase HA-tagged variants in tobacco
Binary vector (pEAQ-HT-based) plasmids were transformed into A. tumefasciens (GV3101) using the freeze-thaw method66. Transformants were grown on LB-agar plates supplemented with 50 μg ml−1 kanamycin and 30 μg ml−1 gentamicin at 30 °C for 48 h. Colonies were inoculated into 5 ml liquid cultures of LB with 50 μg ml−1 kanamycin and 30 μg ml−1 gentamicin and grown for 18–24 h at 30 °C and 250 rpm in a shaking incubator. Saturated cultures were pelleted by centrifugation at 5,000g for 5 min. Pellets were resuspended in the same volume (~5 ml) of freshly prepared infiltration buffer (10 mM MES buffer, pH 5.6, 10 mM MgCl2, 150 μM acetosyringone), incubated at room temperature for 2–3 h with gentle rocking to prevent settling, and then diluted in infiltration buffer to OD600 of 0.8–1.0. N. benthamiana plants were grown for 4 weeks under a 16 h light/8 h dark cycle before infiltration. Approximately 1–2 ml of Agrobacterium cell suspension was infiltrated into the underside of each of the three largest leaves of each plant using a needleless 1 ml syringe. Leaves were harvested four days after infiltration, flash-frozen in liquid nitrogen, and stored at −80 °C for downstream processing. All infiltrations were performed in triplicate, in which one biological replicate comprised three infiltrated leaves from a single plant.
Deglycosylation of yeast- and tobacco-expressed littorine synthase
Removal of N- and O-linked glycosylation from littorine synthase in yeast and N. benthamiana crude cell lysate was performed using PNGase F and O-glycosidase (NEB), respectively, following the manufacturer’s protocols. In brief, approximately 30 μg of total protein containing LS in crude cell lysate was denatured in 1× glycoprotein denaturing buffer at 100 °C for 10 min, followed by immediate chilling on ice. Denatured lysates were deglycosylated using PNGase F or O-glycosidase as per manufacturer instructions at 37 °C for 1 h, then stored at −20 °C until analysis.
Analysis of protein expression by western blot
For immunoblot analysis of yeast-expressed proteins, strain CSY1294 was transformed with HA-tagged AbLS expression vectors as described in ‘Yeast transformations’. Three days after transformation, transformed colonies were inoculated into 2 ml YNB-DO media and grown overnight (~16–20 h) to stationary phase at 30 °C and 460 rpm. Cells were pelleted by centrifugation at 3,000g for 5 min, resuspended in 200 μl H2O, mixed with 200 μl of 0.2 M NaOH, and incubated at room temperature for 5 min to allow hydrolysis of cell wall glycoproteins67. Cells were re-pelleted at 3,000g for 5 min, resuspended in 75 μl H2O, mixed with 25 μl of 4× NuPAGE LDS sample buffer (Thermo Fisher), and then boiled at 95 °C for 3 min to lyse cells. Suspensions were pelleted by centrifugation at 16,000g for 5 min to remove insoluble debris and supernatants were transferred to pre-chilled tubes. Samples were stored at −20 °C until further analysis.
For analysis of tobacco-expressed proteins, all three infiltrated leaves from a single plant were ground together to a fine powder under liquid nitrogen and resuspended in 4–5 ml of 25 mM potassium phosphate buffer (pH 8.0) with HALT protease inhibitor cocktail (Thermo Fisher). Leaf homogenate slurries (final volume 7–8 ml) were incubated at 4 °C with gentle rotation for 45–60 min and then clarified by centrifugation at 9,000g for 10 min. Supernatant fractions were transferred to new tubes and re-clarified. Lysate protein concentrations were estimated using the Bio-RAD Protein Assay kit. Samples were stored at −80 °C until further analysis.
For analysis under reducing conditions, protein lysates were mixed with β-mercaptoethanol (final concentration 10%) and incubated at 70 °C for 10 min. Approximately 20–40 μg of total protein was loaded onto NuPAGE Bis-Tris 4–12% acrylamide gels (Thermo Fisher) with Precision Plus Dual Colour protein molecular mass marker (BioRad). Electrophoresis was conducted in 1× NuPAGE MOPS SDS running buffer at 150 V for 90 min. Transfer of protein to nitrocellulose membranes was performed at 15 V for 15 min using a Trans-Blot Semi-Dry apparatus (BioRad) and NuPAGE transfer buffer (Thermo Fisher) per manufacturer instructions. For reducing conditions, NuPAGE antioxidant (Thermo Fisher) was added to a final concentration of 1× to both the running buffer and transfer buffer. Membranes with transferred protein were washed for 5 min in Tris-buffered saline with Tween (TBS-T; 137 mM NaCl, 2.7 mM KCl, 19 mM Tris base, 0.1% Tween20, pH 7.4) and then blocked with 5% skim milk in TBS-T for 1 h at room temperature. Membranes were incubated overnight at 4 °C with a 1:1,500 dilution of chimaeric rabbit IgGκ anti-HA HRP-conjugated antibody (Absolute Antibody, 16.43/Ab00828-23.0) in TBS-T with 5% milk, washed three times for 5 min each with TBS-T, and then visualized using Western Pico PLUS HRP substrate (Thermo Fisher) and a G:BOX gel imager (Syngene).
Statistics
Where indicated, the statistical significance of any differences in metabolite titer between conditions was verified using Student’s two-tailed t-test in Microsoft Excel Professional 2013. For yeast experiments, biological replicates are defined as independent cultures inoculated from separate yeast colonies or streaks and cultivated in separate containers. For tobacco experiments, one biological replicate is defined as all infiltrated leaves from a single plant.
Additional software
All figures were prepared using GraphPad Prism 7, ImageJ, PyMOL, and Inkscape.
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this paper.
Data availability
Data supporting the findings of this work are available within the paper and its Supplementary Information files. The datasets generated and analysed during the current study are available from the corresponding author upon reasonable request. Novel genetic sequences identified and characterized in this study are available from the following public databases. 1000Plants (1KP) database63: scaffold-AIOU-2012986-Brugmansia_sanguinea (BsUGT); scaffold-JNVS-2051323-Datura_metel (DmUGT). Medicinal Plant RNA-seq database32: medp_datin_20101112|6354 (DiHDH); medp_datst_20101112|10433 (DsHDH). MSU Medicinal Plant Genomics Resource30: full amino acid sequences and database accession numbers (IDs) for all tested HDH candidates are provided in Supplementary Table 1. Accession numbers for previously reported gene and protein sequences in the GenBank/UniProt databases are provided in Supplementary Table 2. Protein crystal structures used for homology modelling are available from the RCSB Protein Data Bank (PDB) with the following accessions: Arabidopsis thaliana salicylate UDP-glucosyltransferase UGT74F2 with bound UDP, accession 5V2K; P. tremuloides sinapyl alcohol dehydrogenase with bound NADPH, accession 1YQD. Source data are provided with this paper.
Code availability
The custom R script used for identification of HDH candidates via coexpression analysis of A. belladonna RNA sequencing data is available from the Smolke Laboratory GitHub: https://github.com/smolkelab/Oxidoreductase_coexpression_analysis.
References
World Health Organization. WHO Model List of Essential Medicines - 19th List (April 2015).
Grynkiewicz, G. & Gadzikowska, M. Tropane alkaloids as medicinally useful natural products and their synthetic derivatives as new drugs. Pharmacol. Rep. 60, 439–463 (2008).
U.S. Food and Drug Administration. FDA Drug Shortages: Atropine Sulfate Injection (FDA Drug Shortages database, accessed online 14 April 2020); https://www.accessdata.fda.gov/scripts/drugshortages/dsp_ActiveIngredientDetails.cfm?AI=AtropineSulfateInjection&st=c
U.S. Food and Drug Administration. FDA Drug Shortages: Scopolamine Transdermal System (FDA Drug Shortages database, accessed online 14 April 2020); https://www.accessdata.fda.gov/scripts/drugshortages/dsp_ActiveIngredientDetails.cfm?AI=ScopolamineTransdermalSystem&st=r
The Climate Council of Australia. ‘This is Not Normal’: Climate Change and Escalating Bushfire Risk https://www.climatecouncil.org.au/wp-content/uploads/2019/11/CC-nov-Bushfire-briefing-paper.pdf (2019).
Hahn, S. M. FDA Statement: Coronavirus (COVID-19) Supply Chain Update https://www.fda.gov/news-events/press-announcements/coronavirus-covid-19-supply-chain-update (2020).
U.S. Food and Drug Administration. Coronavirus (Covid-19) Update: FDA Takes Further Steps to Help Mitigate Supply Interruptions of Food and Medical Products https://www.fda.gov/news-events/press-announcements/coronavirus-covid-19-update-fda-takes-further-steps-help-mitigate-supply-interruptions-food-and (2020).
Agrawal, G., Ahlawat, H. & Dewhurst, M. Winning Against COVID-19: the Implications for Biopharma https://www.mckinsey.com/industries/pharmaceuticals-and-medical-products/our-insights/winning-against-covid-19-the-implications-for-biopharma?from=groupmessage&isappinstalled=0# (2020).
Kohnen, K. L., Sezgin, S., Spiteller, M., Hagels, H. & Kayser, O. Localization and organization of scopolamine biosynthesis in Duboisia myoporoides R. Br. Plant Cell Physiol. 59, 107–118 (2018).
Ullrich, S. F., Hagels, H. & Kayser, O. Scopolamine: a journey from the field to clinics. Phytochem. Rev. 16, 333–353 (2017).
Kohnen-Johannsen, K. L. & Kayser, O. Tropane alkaloids: chemistry, pharmacology, biosynthesis and production. Molecules 24, 1–23 (2019).
American Society of Anesthesiologists (ASA). ASA Urges Federal Government to Take Action on Drug Shortages https://www.asahq.org/advocacy-and-asapac/fda-and-washington-alerts/washington-alerts/2020/04/asa-urges-federal-government-to-take-action-on-drug-shortages (2020).
Bedewitz, M. A., Jones, A. D., D’Auria, J. C. & Barry, C. S. Tropinone synthesis via an atypical polyketide synthase and P450-mediated cyclization. Nat. Commun. 9, 5281 (2018).
Srinivasan, P. & Smolke, C. D. Engineering a microbial biosynthesis platform for de novo production of tropane alkaloids. Nat. Commun. 10, 3634 (2019).
Ping, Y. et al. De novo production of the plant-derived tropine and pseudotropine in yeast. ACS Synth. Biol. 8, 1257–1262 (2019).
Qiu, F. et al. Functional genomics analysis reveals two novel genes required for littorine biosynthesis. New Phytol. 225, 1906–1914 (2019).
Li, R. et al. Functional genomic analysis of alkaloid biosynthesis in Hyoscyamus niger reveals a cytochrome P450 involved in littorine rearrangement. Chem. Biol. 13, 513–520 (2006).
Nasomjai, P. et al. Mechanistic insights into the cytochrome P450-mediated oxidation and rearrangement of littorine in tropane alkaloid biosynthesis. Chem Bio Chem 10, 2382–2393 (2009).
Matsuda, J., Okabe, S., Hashimoto, T. & Yamada, Y. Molecular cloning of hyoscyamine 6 β-hydroxylase, a 2-oxoglutarate-dependent dioxygenase, from cultured roots of Hyoscyamus niger. J. Biol. Chem. 266, 9460–9464 (1991).
Hashimoto, T., Matsuda, J. & Yamada, Y. Two-step epoxidation of hyoscyamine to scopolamine is catalyzed by bifunctional hyoscyamine 6 β-hydroxylase. FEBS Lett. 329, 35–39 (1993).
Wang, Z., Jiang, M., Guo, X., Liu, Z. & He, X. Reconstruction of metabolic module with improved promoter strength increases the productivity of 2-phenylethanol in Saccharomyces cerevisiae. Microb. Cell Fact. 17, 60 (2018).
Zhang, X., Zhang, S., Shi, Y., Shen, F. & Wang, H. A new high phenyl lactic acid-yielding Lactobacillus plantarum IMAU10124 and a comparative analysis of lactate dehydrogenase gene. FEMS Microbiol. Lett. 356, 89–96 (2014).
Sévin, D. C., Fuhrer, T., Zamboni, N. & Sauer, U. Nontargeted in vitro metabolomics for high-throughput identification of novel enzymes in Escherichia coli. Nat. Methods 14, 187–194 (2017).
Qiu, F. et al. A phenylpyruvic acid reductase is required for biosynthesis of tropane alkaloids. Org. Lett. 24, 7807–7810 (2018).
Fujii, T. et al. Novel fungal phenylpyruvate reductase belongs to d-isomer-specific 2-hydroxyacid dehydrogenase family. Biochim. Biophys. Acta. Proteins Proteomics 1814, 1669–1676 (2011).
Zheng, Z. et al. Efficient conversion of phenylpyruvic acid to phenyllactic acid by using whole cells of Bacillus coagulans SDM. PLoS ONE 6, e19030 (2011).
Li, J. F. et al. Directed modification of l-LcLDH1, an l-lactate dehydrogenase from Lactobacillus casei, to improve its specific activity and catalytic efficiency towards phenylpyruvic acid. J. Biotechnol. 281, 193–198 (2018).
Ross, J., Li, Y., Lim, E.-K. & Bowles, D. J. Higher plant glycosyltransferases. Genome Biol. 2, 3004.1–3004.6 (2001).
Wang, H. et al. Engineering Saccharomyces cerevisiae with the deletion of endogenous glucosidases for the production of flavonoid glucosides. Microb. Cell Fact. 15, 134 (2016).
Michigan State University. Medicinal Plant Genomics Resource (accessed online 23 April 23, 2018); http://medicinalplantgenomics.msu.edu
Grabherr, M. G. et al. Full-length transcriptome assembly from RNA-seq data without a reference genome. Nat. Biotechnol. 29, 644–652 (2011).
NIH. Medicinal Plant RNA Seq Database (accessed online 12 July 2018); https://medplantrnaseq.org
Bontpart, T., Cheynier, V., Ageorges, A. & Terrier, N. BAHD or SCPL acyltransferase? What a dilemma for acylation in the world of plant phenolic compounds. New Phytol. 208, 695–707 (2015).
Carqueijeiro, I. et al. in Plant Vacuolar Trafficking. Methods and Protocols 33–54 (2018).
Strasser, R. Plant protein glycosylation. Glycobiology 26, 926–939 (2016).
Stehle, F., Stubbs, M. T., Strack, D. & Milkowski, C. Heterologous expression of a serine carboxypeptidase-like acyltransferase and characterization of the kinetic mechanism. FEBS J. 275, 775–787 (2008).
Stack, J. H., Horazdovsky, B. & Emr, S. D. Receptor-mediated protein sorting to the vacuole in yeast: roles for a protein kinase, a lipid kinase and GTP-binding proteins. Annu. Rev. Cell Dev. Biol. 11, 1–33 (1995).
Roberts, C. J., Nothwehr, S. F. & Stevens, T. H. Membrane protein sorting in the yeast secretory pathway: evidence that the vacuole may be the default compartment. J. Cell Biol. 119, 69–83 (1992).
Liu, L., Spurrier, J., Butt, T. R. & Strickler, J. E. Enhanced protein expression in the baculovirus/insect cell system using engineered SUMO fusions. Protein Expr. Purif. 62, 21–28 (2008).
Morita, M. et al. Vacuolar transport of nicotine is mediated by a multidrug and toxic compound extrusion (MATE) transporter in Nicotiana tabacum. Proc. Natl Acad. Sci. USA 106, 2447–2452 (2009).
Shoji, T. et al. Multidrug and toxic compound extrusion-type transporters implicated in vacuolar sequestration of nicotine in tobacco roots. Plant Physiol. 149, 708–718 (2009).
Cardillo, A. B., Perassolo, M., Sartuqui, M., Rodríguez Talou, J. & Giulietti, A. M. Production of tropane alkaloids by biotransformation using recombinant Escherichia coli whole cells. Biochem. Eng. J. 125, 180–189 (2017).
Lesuisse, E., Crichton, R. R. & Labbe, P. Iron-reductases in the yeast Saccharomyces cerevisiae. Protein Struct. Mol. 1038, 253–259 (1990).
Hu, Y., Zhu, Z., Nielsen, J. & Siewers, V. Heterologous transporter expression for improved fatty alcohol secretion in yeast. Metab. Eng. 45, 51–58 (2018).
Dastmalchi, M. et al. Purine permease-type benzylisoquinoline alkaloid transporters in opium poppy. Plant Physiol. 181, 916–933 (2019).
Ro, D. K. et al. Induction of multiple pleiotropic drug resistance genes in yeast engineered to produce an increased level of anti-malarial drug precursor, artemisinic acid. BMC Biotechnol. 8, 83 (2008).
Brown, S., Clastre, M., Courdavault, V. & O’Connor, S. E. De novo production of the plant-derived alkaloid strictosidine in yeast. Proc. Natl Acad. Sci. USA 112, 3205–3210 (2015).
Galanie, S., Thodey, K., Trenchard, I. J., Filsinger Interrante, M. & Smolke, C. D. Complete biosynthesis of opioids in yeast. Science 349, 1095–1100 (2015).
Cravens, A., Payne, J. & Smolke, C. D. Synthetic biology strategies for microbial biosynthesis of plant natural products. Nat. Commun. 10, 2142 (2019).
Redford, K. H., Brooks, T. M., Macfarlane, N. B. W. & Adams, J. S. Genetic Frontiers for Conservation: an Assessment of Synthetic Biology and Biodiversity Conservation https://portals.iucn.org/library/node/48408 (2019).
Entian, K. D. & Kötter, P. 25 yeast genetic strain and plasmid collections. Methods Microbiol. 36, 629–666 (2007).
Kwiatkowski, T. J., Jr, Zoghbi, H. Y., Ledbetter, S. A., Ellison, K. A. & Chinault, A. C. Rapid identification of yeast artificial chromosome clones by matrix pooling and crude lysate PCR. Nucleic Acids Res. 18, 7191–7192 (1990).
Alberti, S., Gitler, A. D. & Lindquist, S. A suite of Gateway cloning vectors for high-throughput genetic analysis in Saccharomyces cerevisiae. Yeast 24, 913–919 (2007).
Sainsbury, F., Thuenemann, E. C. & Lomonossoff, G. P. pEAQ: versatile expression vectors for easy and quick transient expression of heterologous proteins in plants. Plant Biotechnol. J. 7, 682–693 (2009).
Ryan, O. W. et al. Selection of chromosomal DNA libraries using a multiplex CRISPR system. eLife 3, 1–15 (2014).
Thodey, K., Galanie, S. & Smolke, C. D. A microbial biomanufacturing platform for natural and semisynthetic opioids. Nat. Chem. Biol. 10, 837–844 (2014).
Allen, F., Pon, A., Wilson, M., Greiner, R. & Wishart, D. CFM-ID: a web server for annotation, spectrum prediction and metabolite identification from tandem mass spectra. Nucleic Acids Res. 42, W94–W99 (2014).
Guijas, C. et al. METLIN: a technology platform for identifying knowns and unknowns. Anal. Chem. 90, 3156–3164 (2018).
Amberg, D. C., Burke, D. J. & Strathern, J. N. Yeast vital stains: visualizing vacuoles and endocytic compartments with FM4-64. Cold Spring Harb. Protoc. 2006, pdb.prot4165 (2006).
Wallace, W., Schaefer, L. H. & Swedlow, J. R. A workingperson’s guide to deconvolution in light microscopy. Biotechniques 31, 1076–1078, 1080, 1082 passim (2001).
Haas, B. J. et al. De novo transcript sequence reconstruction from RNA-seq using the Trinity platform for reference generation and analysis. Nat. Protoc. 8, 1494–1512 (2013).
Bryant, D. M. et al. A tissue-mapped axolotl de novo transcriptome enables identification of limb regeneration factors. Cell Rep. 18, 762–776 (2017).
Matasci, N. et al. Data access for the 1,000 Plants (1KP) project. Gigascience 3, 17 (2014).
Källberg, M. et al. Template-based protein structure modeling using the RaptorX web server. Nat. Protoc. 7, 1511–1522 (2012).
Bomati, E. K. & Noel, J. P. Structural and kinetic basis for substrate selectivity in Populus tremuloides sinapyl alcohol dehydrogenase. Plant Cell 17, 1598–1611 (2005).
Weigel, D. & Glazebrook, J. Transformation of agrobacterium using the freeze-thaw method. Cold Spring Harb. Protoc. 2006, pdb.prot4666–pdb.prot4666 (2006).
Kushnirov, V. V. Rapid and reliable protein extraction from yeast. Yeast 16, 857–860 (2000).
Acknowledgements
We thank A. Cravens for the yeast multiplex CRISPR–Cas9/sgRNA plasmids (pCS3410, 3411, 3414 and 3700-3703); T. Lozanoski (Jarosz laboratory) for a plasmid containing yeast codon-optimized mVenus; J. Payne for assistance with the generation of DFT energy-optimized ligand structures for docking studies; P. Dykstra for assistance with fluorescence microscopy and the Stanford Cell Sciences Imaging Facility for access to microscopy equipment and training; R. Nett for suggestions regarding de novo transcriptome assembly using Trinity; W. Cody and the Sattely laboratory for providing N. benthamiana plants, Agrobacterium strains, a pEAQ-HT binary vector backbone plasmid, and suggestions for tobacco agroinfiltration experiments; D. Endy and J. Payne for discussions and feedback in the preparation of this manuscript. This work was supported by the National Institutes of Health and the Natural Sciences and Engineering Research Council of Canada (doctoral postgraduate scholarship to P.S.).
Author information
Authors and Affiliations
Contributions
P.S. and C.D.S. conceived of the project, designed the experiments, analysed the results, and wrote the manuscript. P.S. performed the experiments.
Corresponding author
Ethics declarations
Competing interests
P.S. and C.D.S. are inventors on a pending patent application. C.D.S. is a founder and CEO of Antheia, Inc.
Additional information
Peer review information Nature thanks Jose Avalos, Oliver Kayser and the other anonymous reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Design of genomic integrations for pathway construction in yeast.
Block arrows represent gene expression cassettes with unique promoter and terminator for each locus. Genus and species sources for heterologous genes are indicated by two letters preceding the gene symbol. Superscript annotations on gene symbols indicate N- or C-terminal modifications; dash (-) indicates fusion protein. See Supplementary Tables 1–2 for gene sources and Supplementary Table 3 for yeast strain genotypes.
Extended Data Fig. 2 Substrate specificity and structure-guided active site engineering of UGT84A27 in engineered yeast.
a, Phenylpropanoids tested as glucose (Glc) acceptors for UGT84A27 in engineered yeast. Top, (d)-3-phenyllactic acid (PLA); middle, trans-cinnamic acid (CA); bottom, trans-ferulic acid (FA). b, Heat map of the percentage conversion of fed phenylpropanoids to glucosides by yeast engineered for UGT84A27 expression. UGT84A27 orthologues or a BFP negative control were expressed from low-copy plasmids in CSY1251. Transformed cells were cultured in selective medium supplemented with 500 μM PLA, CA or FA for 72 h before LC–MS/MS analysis of culture supernatant. Data represent the mean of n = 3 biologically independent samples ± s.d. c, Representative LC–MS/MS traces showing conversion of PLA, CA and FA to cognate glucosides by AbUGT in CSY1251 cultured as in b for 120 h to enable more complete glucosylation. For PLA, acid and glucoside were distinguished by different NH4+ adduct parent masses as well as different retention times. For CA and FA, rapid fragmentation necessitated detection of the glucosides based on the lower-retention peaks produced by their phenylpropanoid fragments. d, Homology model of AbUGT84A27 constructed based on the crystal structure of Arabidopsis thaliana salicylate UDP-glucosyltransferase UGT74F2 with bound UDP (PDB: 5V2K). PLA (orange) is shown in the preferred binding pose with UDP-glucose (pink) based on docking simulations. e, Zoomed view of AbUGT active site with docked d-PLA and UDP-glucose. Potential mutations identified to improve PLA selectivity (F130Y, L205F, I292Q) are shown; dashed lines indicate putative polar/hydrogen bond interactions. f, Heat map of the percentage conversion of fed phenylpropanoids to glucosides by yeast engineered for expression of AbUGT mutants. AbUGT wild-type, active site mutants, or a BFP negative control were expressed from low-copy plasmids in CSY1251. Transformed cells were cultured in selective media supplemented with 500 μM PLA, CA or FA for 72 h before LC–MS/MS analysis of culture supernatant. Data represent the mean of n = 3 biologically independent samples ± s.d.
Extended Data Fig. 3 Coexpression analysis, active site mutagenesis, and orthologue identification for AbHDH.
a, Heat map of tissue-specific expression profiles for HDH gene candidates identified from the A. belladonna transcriptome. Transcript expression is scaled by row using a normal distribution. Dendrogram indicates hierarchical clustering of candidates by tissue-specific expression profile. Colour scheme for gene IDs: purple, known TA pathway genes; blue, putative HDH candidates with complete open reading frame sequences; black, putative HDH candidates with incomplete open reading frame sequences. Gene abbreviations (vertical axis): CPA, N-carbamoylputrescine amidase. Tissue abbreviations (horizontal axis): F, flower; MS, mature seed; PTR, primary tap root; SS, sterile seedling; CA, callus; SR, secondary root; S, stem; RF, ripe fruit; GF, green fruit; L, leaf; FB, flower bud. b, Wild-type (WT) AbHDH, active site mutants, or a negative control (BFP) were expressed from low-copy plasmids in CSY1292. Transformed strains were cultured in selective media with 1 mM littorine at 25 °C for 72 h before quantification of scopolamine production by LC–MS/MS analysis of culture supernatant. Data indicate the mean of n = 3 biologically independent samples (open circles) and error bars show s.d. *P < 0.05, **P < 0.01, ***P < 0.001, Student’s two-tailed t-test. Statistical significance is shown relative to wild type. C52A, P = 1.68 × 10−5; C52S, P = 2.06 × 10−5; S54A, P = 8.19 × 10−4; S54C, P = 5.27 × 10−6; H74A, P = 3.40 × 10−6; H74F, P = 2.78 × 10−5; C168A, P = 1.75 × 10−5; C168S, P = 2.39 × 10−5. c, Phylogenetic tree of the three identified HDH orthologues (AbHDH, DiHDH, DsHDH) together with closest protein hits in the UniProt/SwissProt database. Phylogenetic relationships were derived via bootstrap neighbour-joining with n = 1,000 trials in ClustalX2 and the resulting tree was visualized with FigTree software. 8HGDH, 8-hydroxygeraniol dehydrogenase; ADH, alcohol dehydrogenase; CADH, cinnamyl alcohol dehydrogenase; DPAS, dehydroprecondylocarpine acetate synthase; GDH, geraniol dehydrogenase; GS, geissoschizine synthase; MTDH, mannitol dehydrogenase; REDX, unspecified redox protein. d, HDH orthologues (AbHDH, DiHDH and DsHDH) were co-expressed with either a BFP negative control (‘−’) or an additional copy of DsH6H (‘+’) from low-copy plasmids in CSY1292. Transformed cells were cultured in selective medium supplemented with 1 mM littorine for 72 h before LC–MS/MS analysis of culture supernatant. Data represent the mean of n = 3 biologically independent samples (open circles) and error bars show s.d. *P < 0.05, **P < 0.01, ***P < 0.001, Student’s two-tailed t-test. AbHDH + DsH6H versus AbHDH only, scopolamine, P = 4.68 × 10−5; DiHDH versus AbHDH, hyoscyamine aldehyde, P = 0.0141; DsHDH versus AbHDH, hyoscyamine aldehyde, P = 0.0372.
Extended Data Fig. 4 Screening H6H orthologues from TA-producing Solanaceae in yeast.
H6H orthologues from Anisodus acutangulus (AaH6H), Brugmansia arborea (BaH6H), Datura metel (DmH6H), Datura stramonium (DsH6H), or a negative control (BFP) were expressed from low-copy plasmids in CSY1251. Transformed cells were cultured in selective media supplemented with 1 mM hyoscyamine for 72 h before LC–MS/MS analysis of culture supernatant. Data represent the mean of n = 3 biologically independent samples (open circles) and error bars show s.d. *P < 0.05, **P < 0.01, ***P < 0.001, Student’s two-tailed t-test. AaH6H versus control, P = 0.00146; BaH6H versus control, P = 0.00486; DmH6H versus AaH6H, P = 0.00739; DsH6H versus DmH6H, 6.67 × 10−6.
Extended Data Fig. 5 Phylogenetic analysis of HDH.
a, Sequence alignment of AbHDH, DiHDH and DsHDH (green) with selected BLASTp hits from the cinnamyl alcohol dehydrogenase (CAD)-like (red) and sinapyl alcohol dehydrogenase (SAD)-like (blue) families. Catalytic Zn2+ residues are highlighted in bold (black); structural Zn2+-binding residues are highlighted in pink; and key conserved residues which differentiate CAD- and SAD-like ADHs are highlighted in red/blue/green. b, Zoomed surface view of AbHDH substrate-binding pocket with bound NADPH (orange) and hyoscyamine aldehyde (green) based on docking simulations. c, Zoomed inverse surface (cavity) view of AbHDH active site pocket with bound NADPH (orange) and hyoscyamine aldehyde (green) based on docking simulations as in b. Residues W63, H74 and M124 form a tight binding pocket for the aryl moiety of hyoscyamine aldehyde, leaving the tropine moiety to extend out of the active site.
Extended Data Fig. 6 Analysis of AbLS localization, N-glycosylation, and proteolytic processing patterns in yeast and tobacco.
a, Illustration of the canonical plant ER-to-vacuole trafficking and maturation pathway for SCPL acyltransferases (SCPL-ATs), with A. belladonna littorine synthase (AbLS) shown as example. Circled numbers indicate major steps in SCPL-AT expression and activity, including maturation in the (1) ER lumen and (2) Golgi, (3) trafficking to the vacuole, and vacuolar (4) substrate import and (5) product export. b, Additional fields of view (see Fig. 3a) of yeast epifluorescence microscopy showing N-terminal GFP-tagged AbLS (GFP–AbLS), the vacuolar membrane stain FM4-64, and brightfield merged images. Microscopy was performed on CSY1294 expressing GFP–AbLS from a low-copy plasmid. 2D deconvolution analysis was performed as noted in the Methods. Scale bar, 5 μm. Images are representative of two independent experiments. c, Western blot of wild-type AbLS expressed in tobacco and treated with deglycosylases. C-terminal HA-tagged AbLS was transiently expressed in N. benthamiana leaves via agroinfiltration. Crude leaf extracts were either untreated (lane 1: ‘–’), or treated with peptide N-glycosidase F (PNGase F; lane 2: ‘N’) or O-glycosidase (lane 3: ‘O’) to remove N- or O-linked glycosylation, respectively. Lane ‘L’, Bio-Rad Precision Plus Dual Colour protein ladder. d, e, Western blot of AbLS glycosylation site mutants expressed in yeast and tobacco. C-terminal HA-tagged wild-type AbLS, single glycosylation site point mutants (N to Q), or a quadruple mutant were expressed transiently via agroinfiltration in N. benthamiana (‘Nb’) (d) or from low-copy plasmids in CSY1294 (‘yeast’) (e). For d and e, corresponding yeast- and tobacco-expressed controls are included for comparison. Lane ‘L’, Bio-Rad Precision Plus Dual Colour protein ladder. f, Western blot of untagged and HA-tagged wild-type AbLS expressed in tobacco. Untagged (lane 1: ‘–’), N-terminal HA-tagged (lane 2: ‘N’), or C-terminal HA-tagged (lane 3: ‘C’) AbLS was transiently expressed in N. benthamiana leaves via agroinfiltration. Lane ‘L’, Bio-Rad Precision Plus Dual Colour protein ladder. g, h, Analysis of proteolytic cleavage patterns for AbLS split controls and putative propeptide-swapped variants (g) in yeast via western blot (h). C-terminal HA-tagged AbLS variants were expressed from low-copy plasmids in CSY1294 (lanes 1–6); HA-tagged wild-type AbLS expressed in Nicotiana benthamiana (Nb) is shown as an additional control (lane 7). Lane symbols: L, protein molecular mass ladder; WT, wild-type AbLS; SPL, AbLS split at putative propeptide with signal peptides on both fragments; SPL-T, AbLS split at putative propeptide without signal peptides on either fragment; GS, AbLS variant with wild-type propeptide swapped for flexible Gly-Ser linker; SCT, AbLS variant with wild-type propeptide swapped for AtSCT propeptide sequence; CUT, AbLS variant with wild-type propeptide swapped for synthetic poly-arginine site recognized and cleaved by Kex2p protease. For blots in c–h, sample preparation, electrophoresis and protein transfer steps were performed under denaturing and disulfide-reducing (c–e, h) or non-reducing (f) conditions; AbLS detection was performed using a chimeric rabbit IgGκ anti-HA HRP-conjugated antibody (Methods). Blots are representative of three (c) or two (d–f, h) independent experiments. For gel source data, see Supplementary Fig. 1.
Extended Data Fig. 7 Analysis of putative endoproteolytic propeptide removal in AbLS.
a, Sequence alignment of AbLS with characterized serine carboxypeptidases and SCPL acyltransferases known to possess (AtSCT, AsSCPL1, TaCBP2) or lack (AtSMT, yPRC1) proteolytically-removed internal propeptide linkers (red). Putative N-terminal signal peptides are indicated in bold; disulfide bonds are indicated in blue. AtSCT, Arabidopsis thaliana sinapoylglucose:choline sinapoyltransferase; AtSMT, A. thaliana sinapoylglucose:malate sinapoyltransferase; AbLS, Atropa belladonna littorine synthase; AsSCPL1, Avena strigosa avenacin synthase; TaCBP2, Triticum aestivum carboxypeptidase 2; yPRC1, yeast carboxypeptidase Y. b, d, Crystal structure of TaCBP2 (PDB: 1WHT) in cartoon (b) and surface (d) representation showing disulfide bonds (yellow) and internal propeptide removal sites. c, e, Homology model of AbLS based on the crystal structure of TaCBP2 in cartoon (c) and surface (e) representation showing N-terminal signal peptide (red, bottom right in c), disulfide bonds (yellow), and putative internal propeptide (red, top/middle), which appears to block active site access. Note that surface views in d and e are rotated 90° towards the viewer.
Extended Data Fig. 8 Fluorescence microscopy of tobacco alkaloid transporters expressed in CSY1296 for alleviation of vacuolar TA transport limitations.
a, b, C-terminal GFP fusions of NtJAT1 (a) and NtMATE2 (b) were expressed from low-copy plasmids in CSY1296. Strains were cultured, imaged, and analysed via widefield epifluorescence microscopy and 2D deconvolution analysis as described in the Methods. Images are representative of two independent experiments. Scale bar, 5 μm.
Extended Data Fig. 9 Effect of extra gene copies on accumulation of TA pathway intermediates and products in scopolamine-producing strain CSY1296.
a–f, An additional copy of each biosynthetic enzyme between tropine and scopolamine was expressed from the following low-copy plasmids in strain CSY1296: WfPPR, pCS4436 (a); AbUGT, pCS4440 (b); DsRed–AbLS, pCS4526 (c); AbCYP80F1, pCS4438 (d); DsHDH, pCS4478 (e); DsH6H, pCS4439 (f); or a BFP control (pCS4208, pCS4212, or pCS4213). Transformed strains were cultured in appropriate selective media at 25 °C for 96 h before quantification of metabolites in the growth medium by LC–MS/MS analysis of culture supernatant. Note that littorine accumulation was not observed for any samples. Data indicate the mean of n = 3 biologically independent samples (open circles) and error bars denote s.d. Metabolite titres are shown relative to the BFP control with the same auxotrophic marker as the biosynthetic gene: WfPPR (a), DsRed–AbLS (c), AbCYP80F1 (d), and DsH6H (f) relative to the pCS4213 control (LEU2); AbUGT (b) relative to the pCS4208 control (URA3); and DsHDH (e) relative to the pCS4212 control (TRP1). *P < 0.05, **P < 0.01, ***P < 0.001, Student’s two-tailed t-test. Statistical significance is shown relative to corresponding controls. (a) PLA, P = 0.00950; PLA glucoside, P = 3.36 × 10−4; hyoscyamine, P = 0.00221; (b) tropine, P = 0.00500; PLA glucoside, P = 3.59 × 10−4; hyoscyamine, P = 0.0192; scopolamine, P = 6.90 × 10−4; (c) PLA glucoside, P = 0.00544; hyoscyamine, P = 0.0165; scopolamine, P = 3.43 × 10−4; (d) PLA, P = 0.0487; hyoscyamine, P = 0.0154; scopolamine, P = 0.0159; (f) PLA, P = 0.0153; PLA glucoside, P = 0.0453; hyoscyamine, P = 0.00958; scopolamine, P = 0.00816.
Extended Data Fig. 10 Time courses of de novo TA and precursor production in pseudo-fed-batch cultures of CSY1297 and CSY1298.
a–e, TA-producing strains CSY1297 and CSY1298 (with pCS4213) were respectively cultured in non-selective or selective (leucine dropout) media with 50 mM 2-OG and 15 mg l−1 Fe2+ under pseudo-fed-batch conditions at 25 °C for 120 h. Culture supernatants were sampled for PLA (a), tropine (b), hyoscyamine (c) and scopolamine (d) titres by LC–MS/MS analysis and optical density at 600 nm (OD600) (e) every 24 h. Cultures were supplemented with additional dextrose, glycerol and amino acids to final concentrations of 2%, 2% and 1× at 72 h (vertical dotted line). Data indicate the mean of n = 3 biologically independent samples and error bars denote s.d. Note that no littorine accumulation was observed in any samples. f–i, LC–MS/MS verification of de novo medicinal TA production in CSY1297 and CSY1298. Panels show LC–MS/MS chromatograms of hyoscyamine (f, g) and scopolamine (h, i) detected in CSY1297 and CSY1298 cultures grown in non-selective medium or selective medium, respectively (samples from experiment in c and d, 120 h), and of authentic 100 nM chemical standards. Primary MRM transitions used for detection and quantification are shown in f and h; additional characteristic transitions are shown in g and i (‘**’ indicates any detected fragment mass in product ion scan). Traces are representative of n = 3 biologically independent samples.
Supplementary information
Supplementary Information
Supplementary Notes 1 to 12 provide additional results and discussion omitted in the main text. Supplementary Tables 1 to 5 provide protein sequences, accession numbers, yeast strain genotypes, LC-MS/MS parameters, and statistical tests relevant to this study. Captions for Supplementary Data 1 and 2 are also included.
Supplementary Figure
Supplementary Figure 1. Original source images for all Western blots analyzed in this study.
Supplementary Data
Supplementary Data 1. Oligonucleotide and primer sequences used for plasmid and strain construction.
Supplementary Data
Supplementary Data 2. Plasmids used in this study.
Rights and permissions
About this article
Cite this article
Srinivasan, P., Smolke, C.D. Biosynthesis of medicinal tropane alkaloids in yeast. Nature 585, 614–619 (2020). https://doi.org/10.1038/s41586-020-2650-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-020-2650-9
This article is cited by
-
Engineered yeast brews plant hormones
Nature Synthesis (2024)
-
Biosensor and machine learning-aided engineering of an amaryllidaceae enzyme
Nature Communications (2024)
-
Convenient synthesis and delivery of a megabase-scale designer accessory chromosome empower biosynthetic capacity
Cell Research (2024)
-
The forefront of chemical engineering research
Nature Chemical Engineering (2024)
-
Reconstitution of early paclitaxel biosynthetic network
Nature Communications (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.