Abstract
At the end of mitosis, eukaryotic cells must segregate the two copies of their replicated genome into two new nuclear compartments1. They do this either by first dismantling and later reassembling the nuclear envelope in an ‘open mitosis’ or by reshaping an intact nucleus and then dividing it into two in a ‘closed mitosis’2,3. Mitosis has been studied in a wide variety of eukaryotes for more than a century4, but how the double membrane of the nuclear envelope is split into two at the end of a closed mitosis without compromising the impermeability of the nuclear compartment remains unknown5. Here, using the fission yeast Schizosaccharomyces pombe (a classical model for closed mitosis5), genetics, live-cell imaging and electron tomography, we show that nuclear fission is achieved via local disassembly of nuclear pores within the narrow bridge that links segregating daughter nuclei. In doing so, we identify the protein Les1, which is localized to the inner nuclear envelope and restricts the process of local nuclear envelope breakdown to the bridge midzone to prevent the leakage of material from daughter nuclei. The mechanism of local nuclear envelope breakdown in a closed mitosis therefore closely mirrors nuclear envelope breakdown in open mitosis3, revealing an unexpectedly high conservation of nuclear remodelling mechanisms across diverse eukaryotes.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
Bulk microscopy time-series data, comprising more than 50 files with an average size over 1 GB, are available upon request. The S. pombe strains generated for and used in this study (Extended Data Table 1) are available upon request. Source data are provided with this paper.
Code availability
All custom software designed for and used in this study is freely available on GitHub in a public repository at https://github.com/superresolusian/local-NEB. The use of this code is governed by an MIT license.
References
LaJoie, D. & Ullman, K. S. Coordinated events of nuclear assembly. Curr. Opin. Cell Biol. 46, 39–45 (2017).
Sazer, S., Lynch, M. & Needleman, D. Deciphering the evolutionary history of open and closed mitosis. Curr. Biol. 24, R1099–R1103 (2014).
Ungricht, R. & Kutay, U. Mechanisms and functions of nuclear envelope remodelling. Nat. Rev. Mol. Cell Biol. 18, 229–245 (2017).
Yanagida, M. The role of model organisms in the history of mitosis research. Cold Spring Harb. Perspect. Biol. 6, a015768 (2014).
Zhang, D. & Oliferenko, S. Remodeling the nuclear membrane during closed mitosis. Curr. Opin. Cell Biol. 25, 142–148 (2013).
Walczak, C. E., Cai, S. & Khodjakov, A. Mechanisms of chromosome behaviour during mitosis. Nat. Rev. Mol. Cell Biol. 11, 91–102 (2010).
Tamm, T. et al. Brr6 drives the Schizosaccharomyces pombe spindle pole body nuclear envelope insertion/extrusion cycle. J. Cell Biol. 195, 467–484 (2011).
Boettcher, B. & Barral, Y. The cell biology of open and closed mitosis. Nucleus 4, 160–165 (2013).
Harel, A. et al. Persistence of major nuclear envelope antigens in an envelope-like structure during mitosis in Drosophila melanogaster embryos. J. Cell Sci. 94, 463–470 (1989).
Lénárt, P. et al. Nuclear envelope breakdown in starfish oocytes proceeds by partial NPC disassembly followed by a rapidly spreading fenestration of nuclear membranes. J. Cell Biol. 160, 1055–1068 (2003).
Gu, Y., Yam, C. & Oliferenko, S. Divergence of mitotic strategies in fission yeasts. Nucleus 3, 220–225 (2012).
Ding, R., McDonald, K. L. & McIntosh, J. R. Three-dimensional reconstruction and analysis of mitotic spindles from the yeast, Schizosaccharomyces pombe. J. Cell Biol. 120, 141–151 (1993).
Ward, J. J., Roque, H., Antony, C. & Nédélec, F. Mechanical design principles of a mitotic spindle. eLife 3, e03398 (2014).
Loiodice, I. et al. Quantifying tubulin concentration and microtubule number throughout the fission yeast cell cycle. Biomolecules 9, 86 (2019).
Zhang, D., Vjestica, A. & Oliferenko, S. The cortical ER network limits the permissive zone for actomyosin ring assembly. Curr. Biol. 20, 1029–1034 (2010).
Frost, A. et al. Functional repurposing revealed by comparing S. pombe and S. cerevisiae genetic interactions. Cell 149, 1339–1352 (2012).
Krüger, L. K., Sanchez, J.-L., Paoletti, A. & Tran, P. T. Kinesin-6 regulates cell-size-dependent spindle elongation velocity to keep mitosis duration constant in fission yeast. eLife 8, e42182 (2019).
Flor-Parra, I. et al. Importin α and vNEBD control meiotic spindle disassembly in fission yeast. Cell Rep. 23, 933–941 (2018).
Lucena, R. et al. Nucleocytoplasmic transport in the midzone membrane domain controls yeast mitotic spindle disassembly. J. Cell Biol. 209, 387–402 (2015).
Gatta, A. T. & Carlton, J. G. The ESCRT-machinery: closing holes and expanding roles. Curr. Opin. Cell Biol. 59, 121–132 (2019).
Olmos, Y., Hodgson, L., Mantell, J., Verkade, P. & Carlton, J. G. ESCRT-III controls nuclear envelope reformation. Nature 522, 236–239 (2015).
Vietri, M. et al. Spastin and ESCRT-III coordinate mitotic spindle disassembly and nuclear envelope sealing. Nature 522, 231–235 (2015).
Pieper, G. H., Sprenger, S., Teis, D. & Oliferenko, S. ESCRT-III/Vps4 controls heterochromatin–nuclear envelope attachments. Dev. Cell 53, 27–41.e6 (2020).
von Appen, A. et al. LEM2 phase separation promotes ESCRT-mediated nuclear envelope reformation. Nature 582, 115–118 (2020).
Gu, M. et al. LEM2 recruits CHMP7 for ESCRT-mediated nuclear envelope closure in fission yeast and human cells. Proc. Natl Acad. Sci. USA 114, E2166–E2175 (2017).
Thaller, D. J. et al. An ESCRT–LEM protein surveillance system is poised to directly monitor the nuclear envelope and nuclear transport system. eLife 8, e45284 (2019).
Yam, C., He, Y., Zhang, D., Chiam, K.-H. & Oliferenko, S. Divergent strategies for controlling the nuclear membrane satisfy geometric constraints during nuclear division. Curr. Biol. 21, 1314–1319 (2011).
McIntosh, J. R., Roos, U. P., Neighbors, B. & McDonald, K. L. Architecture of the microtubule component of mitotic spindles from Dictyostelium discoideum. J. Cell Sci. 75, 93–129 (1985).
Yang, J. et al. The I-TASSER suite: protein structure and function prediction. Nat. Methods 12, 7–8 (2015).
Roy, A., Kucukural, A. & Zhang, Y. I-TASSER: a unified platform for automated protein structure and function prediction. Nat. Protoc. 5, 725–738 (2010).
Eddy, S. R. Accelerated profile HMM searches. PLoS Comput. Biol. 7, e1002195 (2011).
Katoh, K., Misawa, K., Kuma, K. & Miyata, T. MAFFT: a novel method for rapid multiple sequence alignment based on fast Fourier transform. Nucleic Acids Res. 30, 3059–3066 (2002).
Katoh, K., Rozewicki, J. & Yamada, K. D. MAFFT online service: multiple sequence alignment, interactive sequence choice and visualization. Brief. Bioinform. 20, 1160–1166 (2019).
Capella-Gutiérrez, S., Silla-Martínez, J. M. & Gabaldón, T. trimAl: a tool for automated alignment trimming in large-scale phylogenetic analyses. Bioinformatics 25, 1972–1973 (2009).
Nguyen, L.-T., Schmidt, H. A., von Haeseler, A. & Minh, B. Q. IQ-TREE: a fast and effective stochastic algorithm for estimating maximum-likelihood phylogenies. Mol. Biol. Evol. 32, 268–274 (2015).
Kalyaanamoorthy, S., Minh, B. Q., Wong, T. K. F., von Haeseler, A. & Jermiin, L. S. ModelFinder: fast model selection for accurate phylogenetic estimates. Nat. Methods 14, 587–589 (2017).
Finn, R. D. et al. Pfam: the protein families database. Nucleic Acids Res. 42, D222–D230 (2014).
El-Gebali, S. et al. The Pfam protein families database in 2019. Nucleic Acids Res. 47 (D1), D427–D432 (2019).
Robert, X. & Gouet, P. Deciphering key features in protein structures with the new ENDscript server. Nucleic Acids Res. 42, W320–W324 (2014).
Forsburg, S. L. & Rhind, N. Basic methods for fission yeast. Yeast 23, 173–183 (2006).
Moreno, S., Klar, A. & Nurse, P. Molecular genetic analysis of fission yeast Schizosaccharomyces pombe. Methods Enzymol. 194, 795–823 (1991).
Bähler, J. et al. Heterologous modules for efficient and versatile PCR-based gene targeting in Schizosaccharomyces pombe. Yeast 14, 943–951 (1998).
Murray, J. M., Watson, A. T. & Carr, A. M. Transformation of Schizosaccharomyces pombe: lithium acetate/ dimethyl sulfoxide procedure. Cold Spring Harb. Protoc. 2016, pdb.prot090969 (2016).
Zhang, M. et al. Rational design of true monomeric and bright photoactivatable fluorescent proteins. Nat. Methods 9, 727–729 (2012).
Escorcia, W. & Forsburg, S. L. Tetrad dissection in fission yeast. Methods Mol. Biol. 1721, 179–187 (2018).
Huff, J. et al. The new 2D superresolution mode for ZEISS Airyscan. Nat. Methods 14, 1223 (2017).
Schindelin, J. et al. Fiji: an open-source platform for biological-image analysis. Nat. Methods 9, 676–682 (2012).
Schneider, C. A., Rasband, W. S. & Eliceiri, K. W. NIH Image to ImageJ: 25 years of image analysis. Nat. Methods 9, 671–675 (2012).
Gustafsson, N. et al. Fast live-cell conventional fluorophore nanoscopy with ImageJ through super-resolution radial fluctuations. Nat. Commun. 7, 12471 (2016).
Lehtinen, J. et al. Noise2Noise: learning image restoration without clean data. In 35th Int. Conf. Mach. Learn. 7, 4620–4631 (2018).
Weigert, M. et al. Content-aware image restoration: pushing the limits of fluorescence microscopy. Nat. Methods 15, 1090–1097 (2018).
Ballard, D. H. Generalizing the Hough transform to detect arbitrary shapes. Pattern Recognit. 13, 111–122 (1981).
Asakawa, H. et al. Asymmetrical localization of Nup107–160 subcomplex components within the nuclear pore complex in fission yeast. PLoS Genet. 15, e1008061 (2019).
Ho, J., Tumkaya, T., Aryal, S., Choi, H. & Claridge-Chang, A. Moving beyond P values: data analysis with estimation graphics. Nat. Methods 16, 565–566 (2019).
Kukulski, W. et al. Precise, correlated fluorescence microscopy and electron tomography of lowicryl sections using fluorescent fiducial markers. Methods Cell Biol. 111, 235–257 (2012).
Bharat, T. A. M., Hoffmann, P. C. & Kukulski, W. Correlative microscopy of vitreous sections provides insights into BAR-domain organization in situ. Structure 26, 879–886 (2018).
Hohmann-Marriott, M. F. et al. Nanoscale 3D cellular imaging by axial scanning transmission electron tomography. Nat. Methods 6, 729–731 (2009).
Mastronarde, D. N. Automated electron microscope tomography using robust prediction of specimen movements. J. Struct. Biol. 152, 36–51 (2005).
Kremer, J. R., Mastronarde, D. N. & McIntosh, J. R. Computer visualization of three-dimensional image data using IMOD. J. Struct. Biol. 116, 71–76 (1996).
Acknowledgements
We thank M. Balasubramanian, S. Oliferenko, S. Hauf, K. Gould, J. Bahler, P. Nurse and their laboratories for sharing S. pombe strains, plasmids, expertise and S. pombe protocols; J. O. Patterson for the gift of pFA6a-mNeonGreen plasmids; the LMB electron microscopy facility for electron microscopy support; T.-O. Buchholz for advice on the Noise2Noise implementation.; and D. Albrecht, I. Raote, A. Chaigne, P. Pereira, C. Jacobs and members of the Baum laboratory, in particular G. Paci, G. Cazzagon and H. Matthews, for their feedback on this manuscript. G.D. was funded by a European Union Marie Sklodowska-Curie Individual Fellowship (704281-CCDSA) and the Wellcome Trust (203276/Z/16/Z). S. Culley and R.H. were supported by the UK Biotechnology and Biological Sciences Research Council (BB/R021805/1; BB/S507532/1), the UK Medical Research Council (MR/K015826/1) and the Wellcome Trust (203276/Z/16/Z). S. Curran was supported by the Francis Crick Institute, which receives its core funding from Cancer Research UK (FC001121), the UK Medical Research Council (FC001121) and the Wellcome Trust (FC001121). W.K. was funded by the Medical Research Council (MC_UP_1201/8). B.B. was supported by UCL’s Institute for the Physics of Living Systems, the MRC-LMCB, the Wellcome Trust (203276/Z/16/Z) and Cancer Research UK (C1529/A28276).
Author information
Authors and Affiliations
Contributions
G.D. co-conceived the project, designed and implemented all the experiments, generated strains and reagents, acquired, analysed and interpreted the data (with the exception of the electron microscopy data shown in parts of Figs. 2, 4, Extended Data Figs. 3, 7), and led the drafting of the paper. S. Culley and U.S. created new software used in the work, and analysed and assisted in the interpretation of data. S. Curran advised on experimental design, helped to generate strains and reagents, and provided protocols and training. R.H. advised on the creation of new software used for the project. W.K. performed all electron tomography and analysis of electron tomograms (parts of Figs 2, 4, Extended Data Figs. 3, 7). B.B. co-conceived and supervised the project, provided advice on experimental design, implementation and analysis and co-drafted the manuscript. All authors provided input during the manuscript drafting stage.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature thanks Brian Burke and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 ER topology, Les1 structure and phylogeny.
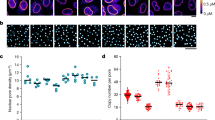
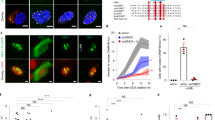
a, Airyscan reconstructions of cells expressing the mCherry-AHDL synthetic construct. Orange arrows indicate tubules linking the outer nuclear envelope to the cortical ER. Purple arrows indicate the displacement of the cortical ER by the division ring (not shown). The blue bar indicates the central ER plate formed during late anaphase. Representative of more than 20 cells across 3 technical repeats. Scale bar. 2 μm. b, Genome-wide genetic interaction scores (E-MAP) with Les1/SPAC23C4.05c. Data from ref. 16. c, Schematic of Les1 sequence with positions of conserved motifs indicated. d, Phylogenetic tree of Les1 homologues across Schizosaccharomyces species with single S. cerevisiae homologue MSC1 grouping separately, indicating a duplication in the lineage leading up to Schizosaccharomyces. Bootstrap values indicated at nodes. See Methods for details of tree construction. e, Alignment of Les1 homologues with key conserved motifs highlighted. Colour-coded by % similarity. See Methods for details.
Extended Data Fig. 2 Image analysis workflow and its application to NE protein localization and spindle breakdown timings.
a, Schematic demonstrating pipeline for detecting and measuring nuclei and bridges in time-lapse data. Individual steps are described fully in the Methods. Representative images shown here are of Les1-mNeonGreen and Atb2-mCherry. b, The delay between reaching maximum spindle length (spindle max.) and spindle breakage, in seconds. Spindle breakage always follows maximal extension. Sample size (n) = 39 individual cells drawn from 3 biological repeats of the entire experiment. Central line represents the mean and the error bars represent the standard deviation of the population. c, Images of two-colour strains used for relative localization of nuclear envelope and NPC components and nucleus radii (r) as measured using the circular Hough transform (Methods), representative of more than 130 individual cells each drawn from 2 biological repeats. d, e, The vertical displacement (relative to the plane of the nuclear envelope) of various nuclear pore complex components (e) (schematized in d; Nup60, in the nuclear ring; Alm1 and Nup211 in the basket; transmembrane Nup Pom34) and nuclear envelope (NE) membrane proteins (Cta4) relative to Cut11. These measurements correspond to the differences in r as shown in c. Nup60-mEOS was used as an internal control, with the values representing the displacement of photo-converted Nup60-mEOS (red channel) relative to un-converted Nup60-mEOS (green channel). The dotted lines represent an estimate of the thickness of the nuclear envelope (see Methods for details). n = 393 (Cta4), 455 (Pom34), 266 (Les1), 185 (Nup60), 280 (Nup211), 218 (Alm1), and 135 (Nup60-mEOS) individual cells, each drawn from across 2 biological repeats of the entire experiment. The measure of centre (central line) shows the mean of each population and the error bars represent the standard deviation.
Extended Data Fig. 3 Electron tomography of mid- and early-stage bridges with correlative fluorescence images.
a, Fluorescence images of resin sections through cells expressing Les1-mNeonGreen (green) and Atb2-mCherry (magenta), corresponding to cells that were imaged by electron tomography and are shown in Fig. 2b (left image), Fig. 2c top panel (middle image) and Fig. 2c lower panel (right image). Images have been rotated to match approximately the orientation of electron tomograms. Scale bars, 1 μm. Representative of 1 (Fig. 2b) and 5 technical repeats (Fig. 2c), respectively. b, Fluorescence image of resin section through a cell expressing Les1-mNeonGreen (green) and Atb2-mCherry (magenta), corresponding to cell imaged by electron tomography shown to the right, indicated by white region. Scale bar, 1 μm. The right two panels are virtual slices through electron tomograms of the cell shown to the left. The approximate overlap in field of view of the tomograms is indicated by dashed lines. No NPCs are visible in the stalk part shown in top image. NPCs (indicated by blue arrowheads) are constrained to the midzone (bottom image). Scale bars, 100 nm. Representative of 2 technical repeats. c, Early stage bridge of a dividing cell; same electron tomogram and cell as shown in Fig. 2b. Top panel is the segmentation model; white area corresponds to the panels below. Inset slice 1 is a virtual slice through the electron tomogram showing the centres of microtubules, indicated by white arrowheads. Inset slice 2 is 18 nm apart in z and shows a nuclear pore, indicated by blue arrowheads. The NPC thus projects less than 18 nm into the nucleoplasm before encountering the microtubules. 1 technical repeat. Scale bars, 50 nm.
Extended Data Fig. 4 Dynamics of individual nucleoporins.
a, Averaged normalized intensity traces for Alm1-mNeonGreen (between 9 cells at t = -300 to 26 cells at t = 0) aligned by spindle max (t = 0). The central line represents the mean of the population; the shaded area depicts the standard deviation. Images on right are confocal maximum intensity projections of dividing cells expressing Alm1-mNeonGreen and Cut11-mCherry, representative of more than 20 cells across 3 biological repeats. Scale bar, 2 μm. b, Kymograph of intensities averaged across a single confocal plane of a dividing cell expressing Cut11-mCherry and Alm1-mNeonGreen and imaged at 3 frames per second followed by denoising (Methods). Representative of more than 10 cells across 2 technical repeats. c, Confocal maximum intensity projections of dividing cells expressing Nup211-mNeonGreen and Cut11-mCherry. Representative of more than 30 cells across 3 biological repeats. Scale bar, 2 μm. d, Schematic indicating effect of deleting Nem1 on nuclear envelope surface area and tension. e, Confocal maximum intensity projections of nem1∆ cells expressing Nup211-mNeonGreen and Cut11-mCherry. Representative of more than 30 cells across 2 biological repeats. Scale bar, 2 μm. The basket Nup is able to enter the nuclear bridge in this condition. f, Confocal maximum intensity projections of dividing cells expressing various NPC subcomplex components tagged with mNeonGreen at their C termini, co-expressed in each case with Cut11-mCherry. Each set of panels is representative of >20 cells across 2 biological repeats. Scale bars, 2 μm.
Extended Data Fig. 5 Relative localization of Les1 and nucleoporins.
a, Averaged line traces (darker lines = mean, lighter bands = standard deviation) of Les1 and Cut11 intensities along the bridge for 16 cells aligned at bridge length 3 μm. b, Time from bridge formation to maximum spindle length measured for strains pooled to generate Fig. 3e (GD173, GD250) and Extended Data 7d (GD176, GD253). Numbers in brackets (n) indicate number of cells in each population, with bars representing mean and standard deviation. The ANOVA F statistic and P value are listed above each plot. The line and pairwise P value within each plot refer to the comparison between pooled strains. c, Single Airyscan reconstructions of cells expressing Cut11 tagged at the endogenous locus with mNeonGreen and a synthetic mCherry-AHDL construct. Magenta arrow highlights stray nuclear pore cluster accompanied by widening of the nuclear envelope. Representative of more than 10 cells across 2 technical replicates. Scale bar, 2 μm. d, Maximum intensity projections of confocal images acquired every 60 s of cells expressing Les1-mNeonGreen and Cut11-mCherry. Bridge formation is at t = 0. Green arrows mark boundaries of Les1 stalks, magenta arrow indicates stray pore cluster. Representative of more than 50 cells across 6 biological repeats. Scale bar, 2 μm. e, SRRF reconstruction from single confocal planes of a cell expressing Les1-mNeonGreen and Cut11-mCherry. Magenta arrow indicates stray nuclear pore cluster, with examples marked upon the line scans along the bridges of 3 illustrative cells in f, Representative of more than 20 cells across 4 biological repeats. g, Confocal maximum intensity projections of cells expressing Cut11-GFP and Atb2-mCherry and acquired at 10 s intervals. Scale bars, 2 μm. Representative of more than 20 cells across 2 biological repeats. h, Relative NPC decay rates, calculated for a strain expressing both Nup60 and Cut11 (GD173; between 11 cells at t = -300 and 34 cells at t = 0), showing similar relative rates to a cross-strain comparison, as in b or Fig. 3e. Line shows mean of the population. i, Kymograph generated using 10 fps single plane imaging of a strain expressing Nup60 and Cut11 tagged at the endogenous loci with mNeonGreen and mCherry respectively. Blue (Nup60) and magenta (Cut11) arrows represent the staggered decay of individual clusters of nuclear pores. Representative of more than 10 cells drawn from 2 technical replicates.
Extended Data Fig. 6 Response of nuclear division dynamics to acute perturbations.
a, Confocal maximum intensity projections of nem1∆ cells expressing Les1-mNeonGreen and Cut11-mCherry. Note the absence of Les1 stalk formation. Schematic indicates opposing effects of nem1∆ and Cerulenin treatment. Representative of more than 30 cells drawn from 2 biological repeats. Scale bar, 2 μm. b, Two examples of nuclei attempting to divide in the presence of 10 μM Cerulenin. Note Les1 stalk formation along aberrant bridge-like projections. Magenta arrows indicate NPC clusters. Representative of more than 30 cells drawn from 3 biological repeats. c, Dividing cells expressing the Ark1-as3 analogue-sensitive allele, either co-expressing Les1-mNeonGreen and mCherry-Atb2 or Les1-mNeonGreen and Cut11-mCherry and treated with 5 nM 1NM-PP1 and 5 μM latrunculin A. In both cases, representative of more than 30 cells across 2 biological repeats. Scale bars, 2 μm. d, iSIM image of imp1∆ cells expressing Les1-mNeonGreen and Cut11-mCherry. Arrows highlight isolated NPC clusters in Les1-depleted regions. Representative of more than 40 cells across 2 biological replicates. Scale bar, 2 μm. e, Time-lapse confocal images of cells expressing Cut11-GFP, Cdc15-mNeonGreen, and mCherry-AHDL acquired at 60 s intervals with frames displayed at 3-min intervals. Representative of more than 50 cells across 3 biological replicates. Scale bars, 4 μm. f, Treatment with latrunculin A depolymerizes the actin ring (marked by Cdc15) but has a minimal effect on the time of nuclear division, as marked by the time from bridge formation to complete NPC signal loss. Numbers above and below the horizontal bar represent the difference in means with 95% confidence interval and the two-sided Mann–Whitney P value. n = 180 (untreated) and 146 (Lat A) individual cells in each population, in each case pooled from 3 biological replicates. Overlaid on individual data points, the upper and lower extent of the boxes span the inter-quartile range and the central bar denotes the median. g, Cytokinetic ring constriction only begins after nuclear division completes, and it takes almost 30 min for the ring to constrict and septation to complete. n = 222 (NPC loss to constriction start), 103 (constriction start to end) and 111 (constriction end to septation) individual cells in each population, in each case pooled from 3 biological replicates. Overlaid on individual data points, the upper and lower extent of the boxes span the inter-quartile range and the central bar denotes the median.
Extended Data Fig. 7 Nucleoporin localization and dynamics in les1Δ cells.
a, Fluorescence image of resin section through a les1Δ cell expressing Cut11-GFP (green) and a synthetic mCherry-AHDL construct (magenta), corresponding to the cell that was imaged by electron tomography shown in Fig. 4c. Representative of 2 technical repeats. Scale bar, 1 μm. b, Inter-pore distances for tomograms shown in Extended Data Fig. 2b (WT Early stage), 3b (WT Mid-stage), and tomogram not shown (les1∆ Mid-stage). See Methods for details on measurement. n = 26 (WT mid-stage), 18 (WT early stage) and 18 (les1∆ mid-stage) individual nuclear pores in each dataset. The central lines represent the mean of the population, with the error bars representing standard deviation. c, Bridge intensity at bridge formation as an indirect readout of nucleoporin copy number in individual cells, for wild-type (WT) and les1∆ cells. n = 19 (WT Cut11), 27 (les1∆ Cut11), 44 (WT Nup60) and 102 (les1∆ Nup60) individual cells pooled from a minimum of 2 biological replicates. Bars overlaid on top of individual data points represent the mean (central line) and standard deviation (error bars). P values derive from a two-tailed unpaired t-test. See Methods for details. d, Decay curves for NLS-GFP (orange, from 28 cells at t = -300 to 39 cells at t = 0), Nup60 (blue, from 10 cells at t = -300 to 15 cells at t = 0) and Cut11 (magenta, from 10 cells at t = -300 to 15 cells at t = 0) in a les1∆ background, drawn from a minimum of 2 biological replicates. Each trace was normalized by division by maximum bridge signal for that cell before averaging. The plot on the left represents the population averages for each marker in a les1∆ background, and the dotted orange line indicates exponential fit to the NLS-GFP average. The 3 subsequent plots show the averages (darker line) and standard deviation (shaded area) for each marker (NLS-GFP, orange; Nup60, blue; Cut11, magenta) in a les1∆ background, overlaid on the wild-type equivalents in grey. e, les1∆ cells expressing Alm1-mNeonGreen and Cut11-mCherry. Arrow indicates a small cluster of Alm1 that enters the bridge but then disappears. Representative of more than 20 cells from 2 biological repeats. Scale bar, 2 μm. f. SRRF reconstructions of confocal slices at 28 s intervals of les1∆ cells expressing Nup60-mNeonGreen and Cut11-mCherry tagged at the endogenous loci, aligned relative to spindle max (t = 0). Representative of more than 10 cells across 2 technical replicates. Magenta arrow indicates breakpoint. Scale bar, 2 μm. On the right, the cumulative distribution (42 cells from 2 strains across 3 biological replicates) of breakpoint locations relative to the midzone in les1∆ cells. The shaded grey area represents the mean +/− standard deviation of breakpoint locations in wild-type cells (31 cells from 2 biological replicates), with the cumulative distribution as a grey line.
Extended Data Fig. 8 Les1 truncations and genetic interactions.
a, Schematic indicating Les1 truncation constructs, with numbers representing amino acid positions starting from 1 at the N terminus. Lower panels, cells expressing Les1 (1-291)-mNeonGreen, replacing Les1 at the endogenous locus, as well as Cut11-mCherry. See Methods for details on the truncation constructs (all at endogenous locus, replacing endogenous copy). Note the absence of detectable stalks and the delocalization of Cut11 in the bridge. Representative of more than 10 cells across 2 biological repeats. Scale bar, 5 μm. b, Confocal maximum intensity projections of cells expressing Cmp7-mNeonGreen and Les1-mScarlet. Arrows indicate Cmp7 foci at the tips of retracting stalks. Representative of more than 20 cells across 4 biological repeats. Scale bar, 2 μm. c, Single iSIM slice of a dividing cell expressing Cmp7-mNeonGreen and Les1-mScarlet, representative of more than 5 cells drawn from 2 technical repeats. Scale bar, 2 μm. d, Tetrad dissection assay for les1∆ crossed with either lem2∆ or cmp7∆, showing colonies grown from individual spores. See Methods for details. e, Single image of double deletion strains, derived from the tetrad assay clones shown in d, expressing a synthetic NLS-GFP construct. Scale bar, 10 μm. Representative of more than 200 cells drawn from 2 biological repeats. f, Averaged line traces (darker lines = mean, lighter bands = standard deviation) of Nup60-mCherry intensities in les1∆ cells (grey: control; blue: treated with 10 μM Cerulenin) at bridge length 3 μm (n = 45 cells for cerulenin-treated; n = 27 cells for control). g, Confocal maximum intensity projections of les1∆ lem2∆ cells expressing Cut11-mCherry, either in the presence (lower panel) or absence (upper panel) of 5 μM latrunculin A. Representative of more than 20 cells across 3 technical repeats. Scale bar, 2 μm.
Supplementary information
Rights and permissions
About this article
Cite this article
Dey, G., Culley, S., Curran, S. et al. Closed mitosis requires local disassembly of the nuclear envelope. Nature 585, 119–123 (2020). https://doi.org/10.1038/s41586-020-2648-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-020-2648-3
This article is cited by
-
A diffusion barrier limits nuclear leaks
Nature Cell Biology (2023)
-
High-fidelity 3D live-cell nanoscopy through data-driven enhanced super-resolution radial fluctuation
Nature Methods (2023)
-
Expansion-enhanced super-resolution radial fluctuations enable nanoscale molecular profiling of pathology specimens
Nature Nanotechnology (2023)
-
Forever young: the key to rejuvenation during gametogenesis
Current Genetics (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.