Abstract
Malignant transformation of cells typically involves several genetic lesions, whose combined activity gives rise to cancer1. Here we analyse 1,148 patient-derived B-cell leukaemia (B-ALL) samples, and find that individual mutations do not promote leukaemogenesis unless they converge on one single oncogenic pathway that is characteristic of the differentiation stage of transformed B cells. Mutations that are not aligned with this central oncogenic driver activate divergent pathways and subvert transformation. Oncogenic lesions in B-ALL frequently mimic signalling through cytokine receptors at the pro-B-cell stage (via activation of the signal-transduction protein STAT5)2,3,4 or pre-B-cell receptors in more mature cells (via activation of the protein kinase ERK)5,6,7,8. STAT5- and ERK-activating lesions are found frequently, but occur together in only around 3% of cases (P = 2.2 × 10−16). Single-cell mutation and phospho-protein analyses reveal the segregation of oncogenic STAT5 and ERK activation to competing clones. STAT5 and ERK engage opposing biochemical and transcriptional programs that are orchestrated by the transcription factors MYC and BCL6, respectively. Genetic reactivation of the divergent (suppressed) pathway comes at the expense of the principal oncogenic driver and reverses transformation. Conversely, deletion of divergent pathway components accelerates leukaemogenesis. Thus, persistence of divergent signalling pathways represents a powerful barrier to transformation, while convergence on one principal driver defines a central event in leukaemia initiation. Pharmacological reactivation of suppressed divergent circuits synergizes strongly with inhibition of the principal oncogenic driver. Hence, reactivation of divergent pathways can be leveraged as a previously unrecognized strategy to enhance treatment responses.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
Gel scans are provided in Supplementary Fig. 1. All other data are available from M.M. upon reasonable request. The Gene Expression Omnibus (GEO; https://www.ncbi.nlm.nih.gov/geo/) accession number for the gene-expression profiles of diagnostic samples from 207 patients (COG P9906) is GSE11877 (Supplementary Table 10). ChIP–seq data from human B lymphocytes (GM12878) are from ENCODE. Further information and requests for reagents can directed to and will be fulfilled by M.M. Source data are provided with this paper.
Code availability
Codes for mutual exclusivity analyses are available on GitHub at https://github.com/muschenlab/Stat5-vs-Erkunder the MIT license, which allows for reuse and distribution.
References
Fearon, E. R., Hamilton, S. R. & Vogelstein, B. Clonal analysis of human colorectal tumors. Science 238, 193–197 (1987).
Goetz, C. A., Harmon, I. R., O’Neil, J. J., Burchill, M. A. & Farrar, M. A. STAT5 activation underlies IL7 receptor-dependent B cell development. J. Immunol. 172, 4770–4778 (2004).
Malin, S. et al. Role of STAT5 in controlling cell survival and immunoglobulin gene recombination during pro-B cell development. Nat. Immunol. 11, 171–179 (2010).
Katerndahl, C. D. S. et al. Antagonism of B cell enhancer networks by STAT5 drives leukemia and poor patient survival. Nat. Immunol. 18, 694–704 (2017).
Shaw, A. C., Swat, W., Ferrini, R., Davidson, L. & Alt, F. W. Activated Ras signals developmental progression of recombinase-activating gene (RAG)-deficient pro-B lymphocytes. J. Exp. Med. 189, 123–129 (1999).
Irving, J. et al. Ras pathway mutations are prevalent in relapsed childhood acute lymphoblastic leukemia and confer sensitivity to MEK inhibition. Blood 124, 3420–3430 (2014).
Anderson, L. J. & Longnecker, R. EBV LMP2A provides a surrogate pre-B cell receptor signal through constitutive activation of the ERK/MAPK pathway. J. Gen. Virol. 89, 1563–1568 (2008).
Feldhahn, N. et al. Mimicry of a constitutively active pre-B cell receptor in acute lymphoblastic leukemia cells. J. Exp. Med. 201, 1837–1852 (2005).
Yasuda, T. et al. Erk kinases link pre-B cell receptor signaling to transcriptional events required for early B cell expansion. Immunity 28, 499–508 (2008).
Shojaee, S. et al. PTEN opposes negative selection and enables oncogenic transformation of pre-B cells. Nat. Med. 22, 379–387 (2016).
Martincorena, I. et al. Somatic mutant clones colonize the human esophagus with age. Science 362, 911–917 (2018).
Mandal, M. et al. Ras orchestrates exit from the cell cycle and light-chain recombination during early B cell development. Nat. Immunol. 10, 1110–1117 (2009).
Duy, C. et al. BCL6 is critical for the development of a diverse primary B cell repertoire. J. Exp. Med. 207, 1209–1221 (2010).
Geng, H. et al. Self-enforcing feedback activation between BCL6 and pre-B cell receptor signaling defines a distinct subtype of acute lymphoblastic leukemia. Cancer Cell 27, 409–425 (2015).
Swaminathan, S. et al. Mechanisms of clonal evolution in childhood acute lymphoblastic leukemia. Nat. Immunol. 16, 766–774 (2015).
Walker, S. R. et al. STAT5 outcompetes STAT3 to regulate the expression of the oncogenic transcriptional modulator BCL6. Mol. Cell. Biol. 33, 2879–2890 (2013).
Korotchenko, V. N. et al. In vivo structure-activity relationship studies support allosteric targeting of a dual specificity phosphatase. ChemBioChem 15, 1436–1445 (2014).
Shojaee, S. et al. Erk negative feedback control enables pre-B cell transformation and represents a therapeutic target in acute lymphoblastic leukemia. Cancer Cell 28, 114–128 (2015).
Yang, J. et al. Discovery and characterization of a cell-permeable, small-molecule c-Abl kinase activator that binds to the myristoyl binding site. Chem. Biol. 18, 177–186 (2011).
Heltemes-Harris, L. M. et al. Sleeping Beauty transposon screen identifies signaling modules that cooperate with STAT5 activation to induce B-cell acute lymphoblastic leukemia. Oncogene 35, 3454–3464 (2016).
Porpaczy, E. et al. Aggressive B-cell lymphomas in patients with myelofibrosis receiving JAK1/2 inhibitor therapy. Blood 132, 694–706 (2018).
Xiao, W. et al. Tumor suppression by phospholipase C-beta3 via SHP-1-mediated dephosphorylation of Stat5. Cancer Cell 16, 161–171 (2009).
Fusaki, N. et al. BLNK is associated with the CD72/SHP-1/Grb2 complex in the WEHI231 cell line after membrane IgM cross-linking. Eur. J. Immunol. 30, 1326–1330 (2000).
Imamura, Y. et al. BLNK binds active H-Ras to promote B cell receptor-mediated capping and ERK activation. J. Biol. Chem. 284, 9804–9813 (2009).
Nikolaev, S. I. et al. Frequent cases of RAS-mutated Down syndrome acute lymphoblastic leukaemia lack JAK2 mutations. Nat. Commun. 5, 4654 (2014).
Herold, T. et al. Adults with Philadelphia chromosome-like acute lymphoblastic leukemia frequently have IGH-CRLF2 and JAK2 mutations, persistence of minimal residual disease and poor prognosis. Haematologica 102, 130–138 (2017).
Jerchel, I. S. et al. RAS pathway mutations as a predictive biomarker for treatment adaptation in pediatric B-cell precursor acute lymphoblastic leukemia. Leukemia 32, 931–940 (2018).
Cerami, E. et al. The cBio Cancer Genomics Portal: An opening platform for exploring multidimensional cancer genomics data. Cancer Discov. 2, 401–404 (2012).
Koch, R. et al. Biomarker-driven strategy for MCL1 inhibition in T-cell lymphomas. Blood 133, 566–575 (2019).
Bliss, C. I. The toxicity of poisons applied jointly. Ann. Appl. Biol. 26, 585–615 (1939).
Weibull, W. A statistical distribution function of wide applicability. J. Appl. Mech. 18, 293–297 (1951).
R Development Core Team. A language and environment for statistical computing. R Foundation for Statistical Computing, Vienna (2009).
Acknowledgements
We thank L. Klemm, F. Auer and J. Winchester for help with some of the experiments, and present and former members of the Müschen Laboratory for their support and helpful discussions. Research in the Müschen Laboratory is funded by the National Institutes of Health (NIH) through National Cancer Institute (NCI) R35CA197628, R01CA157644, R01CA213138 and P01CA233412 (to M.M.); the Howard Hughes Medical Institute (HHMI-55108547 to M.M.); the Norman and Sadie Lee Foundation; the Falk Trust through a Falk Medical Research Trust Transformational Award; the Pediatric Cancer Research Foundation (PCRF) and the V Foundation for Cancer Research, the Leukemia and Lymphoma Society through MCL-7000-18 and a Blood Cancer Discoveries Grant BCDG-20327-20 (to M.M.); and the California Institute for Regenerative Medicine (CIRM) through DISC2-10061. D.M.W. is supported by NCI grants R35CA231958 and U54CA217377. M.M. is a Howard Hughes Medical Institute (HHMI) Faculty Scholar. M.M. and S.I. are supported by the Jacki & Bruce Barron Cancer Research Scholars’ Program. M.A.M. is supported by K08 Clinical Investigator Award CA212252-01A1. T.S. is a Lymphoma Research Foundation Grantee. V.K. is supported by Career Development Fellow grant (5491-20) from the Leukemia and Lymphoma Society, and by a Young Investigator Award from Alex’s Lemonade Stand Foundation.
Author information
Authors and Affiliations
Contributions
L.N.C. designed and performed experiments and contributed to all aspects of the study, in particular western blotting, single-cell phosphoprotein analyses, single-cell mutation analyses, CRISPR-mediated gene deletion, flow-cytometry analysis, growth competition assays, cell sorting, colony-forming assays, viable cell counts, determination of drug responses, in vitro combination treatment assays, in vivo transplantation experiments, and data analysis. M.A.M. performed in vivo ponatinib treatment of patient-derived xenograft models of Ph+ ALL cases in NSG mice, and subsequent targeted sequencing of leukaemia cells harvested from sacrificed mice; M.A.M. also analysed data and provided PDXs. M.E.R. designed figures, performed mutational exclusivity analyses and biostatistical analyses. R.C. performed western blotting, flow cytometry and data analysis. T.S. performed single-cell mutation analysis, flow-cytometry analysis and data analysis and provided expertise in single-cell western blotting. J.W.L. and G.X. performed flow-cytometry analysis and analysed data. K.N.C. performed in vivo transplantation experiments, flow-cytometry analysis and data analysis. K.K. generated the B-cell-specific Myc–eGFP × Bcl6–mCherry double reporter knock-in mouse model, performed flow-cytometry analysis and generated Contour plots for visualization of data obtained from single-cell phosphoprotein analysis. V.K. performed annexin V staining and senescence β-galactosidase staining and analysed data. M.A.A. and E.A. performed in vivo transplantation experiments and data analysis. G.D. performed growth competition assays and data analysis. C. Hurtz, S.S., C. Hong and Z.C. performed in vitro experiments and analysed data. P.P. performed analysis of TARGET data. C.W.C. and J.C. provided expertise for gene-editing experiments. M.H. provided expertise in bioinformatics analysis. A.V. provided the small-molecule inhibitor BCI-215 and D.M.W. provided PDXs. M.A.N. and A.P.W. provided expertise in surface proteomics in leukaemia biology. S.I. and O.L. provided expertise in leukaemia biology. H.G. performed biostatistical analysis and modeling synergy of in vivo transplantation experiments. M.M. secured funding, developed the ‘divergent pathway’ concept, provided mentorship and wrote the manuscript with the input of all co-authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature thanks Michael Reth, Oscar Rueda, Veronika Sexl and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Segregation of STAT5- and ERK-activating mutations in human ALL and AML.
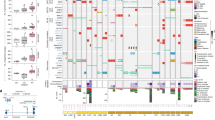
a–e, We studied STAT5- and ERK-pathway mutations in 1,148 patient-derived B-ALL samples (Supplementary Table 1). The null hypothesis is that STAT5- and ERK-pathway mutations occur independently of each other. The expected co-occurrence of mutations in both pathways under the null hypothesis was 121. a, The observed co-occurrence of the two mutations was 35, significantly lower than expected (odds ratio (OR) = 0.13; P = 2.2 × 10−16; Fisher’s exact test). b, To analyse gene–gene co-occurrence in a more unsupervised manner, we ran a Fisher’s test for each gene pair and plotted the results as a co-occurrence network. The pathway assignment for each gene is indicated by the node colour (white, STAT5; grey, ERK), and the mutation frequency by the node size. The direction of the Fisher’s result is indicated by the line colour (green, positive/greater than expected; red, negative/lower than expected); the line width represents the strength of association (–log10 P-value × |logOR|). c, Volcano plot of gene–gene co-occurrence results. Each point represents a gene pair, coloured by the pathway assignment for the pair (green, both STAT5; red, both ERK; grey, interpathway); selected gene pairs are labelled. The overall difference in co-occurrence between pathways was tested by two complementary methods, as follows. d, First, a Fisher’s test was run over 10,000 permutations, with random shuffling of gene-pathway assignments on each iteration, to generate a null distribution for the hypothesis that the pathway does not affect co-occurrence, which we compared against the observed value from ALL patient data. e, Second, overall shifts in the distribution of logORs between pathways for Fisher’s results on individual gene pairs were tested by Welch’s two-sample t-test (left-hand part of panel; two-sided, P = 0.001) and one-way analysis of variance (ANOVA; right-hand part of panel; P = 0.0004) with Tukey’s HSD post-test (P = 0.0003 and P = 0.16). Low-frequency, non-significant gene pairs were excluded to avoid extreme odds ratios biasing results. f–j, The same analyses were carried out on 916 patient-derived AML samples (Supplementary Table 2). The expected co-occurrence of mutations in both pathways under the null hypothesis was 34; the observed co-occurrence was 24, significantly lower than expected (f; OR = 0.59, P = 0.033, Fisher’s exact test). j, Left, Welch’s two-sample t-test (two-sided, P = 0.82); right, one-way ANOVA (P = 0.57).
Extended Data Fig. 2 Single-cell phosphoprotein analyses of patient-derived B-ALL samples reveal segregation of STAT5 and ERK phosphorylation to competing clones.
a–e, For single-cell phosphoprotein analyses, scWest chips were used to capture individual cells and perform size-based protein separation. Each chip includes 16 arrays of 400-well blocks (6,400 wells) on a polyacrylamide gel. Single-cell suspensions of patient-derived B-ALL cells were loaded onto the scWest chip and inserted into Milo (ProteinSimple) for cell lysis, size-based protein separation and UV capture to immobilize protein bands. a, A representative scanned image of the scWest chip probed with histone H3 antibodies to confirm cell occupancy, followed by fluorescent secondary antibodies (n = 16 independent biological samples). b, c, Chips were probed with anti-STAT5-pY694 and anti-ERK-pT202/Y204 antibodies followed by fluorescent secondary antibodies for simultaneous detection of both phosphoproteins. At the left are representative images of signals observed for STAT5-pY694 and ERK-pT202/Y204. At the right, fluorescence intensity was plotted against distance from the well centre (peak location, in μm). In addition to histone H3, chips were stained for DNA using TOTO-1 dye to verify cell occupancy. Each signal was inspected to confirm that it was associated with a peak located at the correct distance from the well centre (n = 16 independent biological samples, each in triplicate). Chips were scanned using a microarray scanner, and peak identification was performed using Scout Software (ProteinSimple). For single-cell western analyses, optimal cell loading is achieved when 2% or fewer wells contain multiple cells (https://www.proteinsimple.com/milo.html). d, e, Single-cell phosphoprotein analyses for STAT5-pY694 and ERK-pT202/Y204 were performed for patient-derived B-ALL samples (six independent biological samples, each in triplicate) with anti-STAT5-pY694 and anti-ERK-pT202/Y204 antibodies for simultaneous detection of both STAT5 and ERK phosphorylation. scWest chips were then probed for histone H3 and TOTO-1 (DNA stain) to verify cell occupancy. Scout Software (ProteinSimple) was used for peak identification and data analysis. Each data point was inspected to confirm that the signal detected was associated with a peak located at the correct peak location (distance from the well centre). d, Heatmaps illustrating cells that express STAT5-pY694 (green) and/or ERK-pT202/Y204 (red). Sample names and genetic characteristics are shown at the top of each heatmap. e, Tables summarizing the number of cells expressing neither STAT5-pY694 nor ERK-pT202/Y204 and the number of cells expressing either STAT5-pY694 or ERK-pT202/Y204, or in rare cases, both. For single-cell western analyses, optimal cell loading is achieved when 2% or fewer wells contain multiple cells.
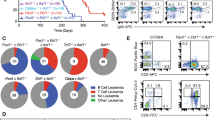
Extended Data Fig. 3 Phenotypic characterization of B-ALL cells driven by oncogenic STAT5 or ERK.
a–g, Eight different B-ALL PDX samples (Supplementary Table 5) were analysed by flow cytometry for surface expression of CRLF2 (a), VpreB (b), CD20 (c), IgM (d), Ig-κλ (e), CD34 (f) and IL7R (g). Normal CD19+ B-cell precursors from a normal human bone-marrow (BM) sample and mature B cells from a normal human peripheral blood (PB) sample, as well a mature B-cell lymphoma cell line (DLBCL), were used as references. Shown are representative FACS plots from three independent experiments.
Extended Data Fig. 4 Phenotypic differences between B-ALL cells driven by oncogenic STAT5 or ERK reflect developmental rewiring during early B-cell development.
a, Mouse B-cell precursors, STAT5-driven B-ALL cells, BCR–ABL1-driven B-ALL cells and NRASG12D-driven B-ALL cells were analysed by flow cytometry for surface expression of CD43, IL7R, VpreB, IgM, Ig-κ, IgD and CD21. Normal CD19+ B-cell precursors from bone-marrow (BM) cells and CD19+ spleen B cells from wild-type mice were used for reference. Shown are representative FACS plots from three independent biological replicates. b, For STAT5-driven B-ALL cases (n = 26) and ERK-driven B-ALL cases (n = 67), mRNA levels of IGLV, IGHM, IKZF1 and BCL6 were analysed in microarray results and associated with clinical outcome (COG P9906) at the time of diagnosis. Patients in each group were segregated into two subgroups on the basis of higher- versus lower-than-median mRNA levels for each of these four genes. Overall survival and relapse-free survival for the Kaplan–Meier plots are shown. Mantel–Cox log-rank tests (two-sided) were used to determine statistical significance. y-axes indicate time (years).
Extended Data Fig. 5 Concurrent activation of STAT5 and ERK signalling induces B-cell senescence and cell death.
a, Levels of NRASG12D, STAT5-pY694, STAT5, STAT3-pS727, STAT3, ERK1/2-pT202/Y204 and ERK1/2 upon doxycycline-induced expression of NRASG12D or STAT5ACA (n = 3 independent experiments). b, c, IL7-dependent mouse pro-B cells (b) or pre-B cells (c) were transduced with empty vector (EV), BCR–ABL1–GFP or LMP2A–GFP. Changes in proportions of GFP+ cells were monitored by flow cytometry (n = 3 independent experiments; means ± s.d.). d, e, FACS analysis of annexin V/7AAD (left) and senescence β-galactosidase staining (centre) were performed with pro-B cells (d) and pre-B cells (e) expressing EV, LMP2A or BCR–ABL1 as indicated. Right, levels of p16 and p21 were examined (n = 3 independent experiments). Shown are representative FACS plots and images from three independent experiments. P-values were determined by two-tailed t-test. β-Galactosidase staining for senescent cells is quantified as the mean percentage of cells positive for staining ± s.d. f, As in normal B-cell development, B-ALL subtypes can be traced to specific differentiation stages with distinct requirements for survival and proliferation signals. For instance, pro-B cells depend on cytokine-receptor signalling and activation of STAT5 but not ERK. Conversely, pre-B cells depend on pre-BCR signalling and activation of ERK but not STAT5. STAT5-driven B-ALL cells (left) depend on oncogenic mimics of cytokine-receptor signalling and resemble pro-B cells that depend on STAT5 activation downstream of cytokine receptors. Mimicking pre-BCR signalling at the pro-B-cell to pre-B-cell transition (right), oncogenic RAS signalling suppresses STAT5 signalling and induces de novo expression of BCL6. For gel source data, see Supplementary Fig. 1.
Extended Data Fig. 6 Pharmacological inhibition of divergent pathways facilitates B-leukaemogenesis.
a, IL7-dependent pro-B cells were first retrovirally transduced with EV–GFP (left), BCR–ABL1–GFP (second from left), EV–Orange (second from right), or NRASG12D–Orange (right). One week later, GFP+ B-ALL cells were transduced with NRASG12D–Orange for concurrent activation of ERK, and Orange+ B-ALL cells were transduced with BCR–ABL1–GFP for concurrent activation of STAT5. The ability of oncogenic STAT5 (GFP+) and ERK (Orange+) signalling to contribute to the dominant clone was monitored by flow cytometry over time. Flow cytometry was performed to monitor the proportions of GFP+, Orange+ and double-positive cells at various time points following transductions. Shown are representative FACS plots from three independent biological replicates. BA–GFP, BCR–ABL1–GFP. b, Cells from a were sorted for double-positive (GFP+ and Orange+) populations, and 10, 000 cells were seeded in methylcellulose for colony-formation assays (10 days; n = 3 independent biological replicates; means ± s.d.). P = 0.003 (left) and P = 0.007 (right) (two-tailed t-test). c–e, Patient-derived BCR–ABL1 B-ALL cells (MXP2) expressing Tet-On NRASG12D and patient-derived KRASG12V B-ALL cells (LAX7R) expressing Tet-On BCR–ABL1 were induced with doxycycline. Viable cell counts (c) were measured upon induction with doxycycline. Annexin V/7AAD and senescence β-galactosidase (d, e) staining were also performed, and levels of p16 and p21 were assessed (n = 3 independent experiments). Shown are representative FACS plots and images. P-values were determined by two-tailed t-tests. Quantification for senescence β-galactosidase staining: mean percentage of cells positive for staining ± s.d. f, Murine wild-type BCR–ABL1 B-ALL cells were primed with vehicle control, DPH (1 μM), trametinib (1 nM) or both for 10 days before colony-forming assays were carried out (n = 3 independent biological replicates; means ± s.d.) g, Mouse wild-type NRASG12D B-ALL cells cultured in the presence of IL7 were primed with vehicle control, BCI-215 (50 nM), ruxolitinib (10 nM), or both for 10 days before colony-forming assays were carried out (n = 3 independent biological replicates; means ± s.d.). P-values were determined by two-tailed t-test (f, g). For gel source data, see Supplementary Fig. 1.
Extended Data Fig. 7 Central role of PTPN6 in enabling oncogenic ERK signalling.
a, Levels of NRAS, PTPN6-pY564, PTPN6, ERK1/2-pT202/Y204 and ERK1/2 were assessed by western blotting following doxycycline-induced expression of NRASG12D in mouse B-cell precursors (n = 3 independent experiments). b, ChIP–seq analyses of human B lymphocytes GM12878 (ENCODE) revealed binding of ERK-dependent transcription factors ELK1, CREB1, c-JUN and JUNB to the PTPN6 locus. c, Patient-derived B-ALL cells (PDX2) were treated with BCI-215 (1 μmol l−1) for various times. Levels of STAT5-pY694, STAT5, ERK1/2-pT202/pY204 and ERK1/2 were then measured by western blotting (n = 3 independent experiments). d, Patient-derived B-ALL cells (PDX2) were treated with BCI-215 (1 μmol l−1) for various times, and levels of STAT5-pY694, STAT5, PTPN6-pY564 and PTPN6 were measured by western blotting (n = 3 independent experiments). e, Ptpn6fl/fl B-cell precursors were transduced with 4-OHT-inducible Cre or EV and then induced with 4-OHT for various times and levels of STAT5-pY694, STAT5, ERK1/2-pT202/Y204, ERK1/2 and PTPN6 were measured (n = 3 independent experiments). f, Ptpn6fl/fl B-cell precursors expressing NRASG12D were transduced with GFP-tagged, 4-OHT-inducible Cre or EV. Following induction with 4-OHT, enrichment or depletion of GFP+ cells was monitored by flow cytometry. Shown are average relative changes (means ± s.d.) of GFP+ cells following induction (n = 6 independent biological replicates). g, Quantification (n = 3 independent experiments; means ± s.d.) and representative images from serial replating assays of pre-B cells transformed with NRASG12D following Cre-mediated deletion of Ptpn6. We seeded 10, 000 cells in semisolid methylcellulose and monitored colony formation for 14 days. P-values were determined by two tailed t-test. For gel source data, see Supplementary Fig. 1.
Extended Data Fig. 8 BLNK enables oncogenic ERK signalling at the expense of STAT5–MYC.
a, Blnk+/+ and Blnk−/− mouse B-cell precursors expressing doxycycline-inducible Ig-HC were treated with doxycycline for 48 h. Levels of ERK1/2-pT202/pY204, ERK1/2, PTPN6-pY564, PTPN6, STAT5-pY694, STAT5, MYC, BCL6 and BLNK were measured by western blotting (n = 3 independent experiments). b, Blnk+/+ and Blnk−/− B-cell precursors expressing doxycycline-inducible NRASG12D were treated with doxycycline for 48 h. Levels of NRAS, ERK1/2-pT202/pY204, ERK1/2, PTPN6-pY564, PTPN6, STAT5-pY694, STAT5, MYC, BCL6 and BLNK were measured by western blotting (n = 3 independent experiments). c–f, Viable cell counts (c, d) and viability changes (e, f) were measured upon doxycycline-induced expression of STAT5ACA or of NRASG12D in Blnk+/+ and Blnk−/− B-cell precursors at various time points (n = 3 independent experiments; means ± s.d.). g, h, The colony-forming ability of Blnk+/+ and Blnk−/− B-cell precursors was assessed upon doxycycline-induced expression of STAT5ACA (g) or NRASG12D (h). We plated 10,000 cells; shown are mean ± s.d. values of three independent experiments and representative images. P-values were determined by two tailed t-test. For gel source data, see Supplementary Fig. 1.
Extended Data Fig. 9 Divergent drug responses in a STAT5- and ERK-driven pair of primary and relapse B-ALLs.
a, c–d, LAX7 (at diagnosis; STAT5-driven, IL7RSI246S) and LAX7R (relapsed; ERK-driven, KRASG12V) cells were transfected with Cas9/RNPs carrying NT or BLNK guide RNAs and then mixed with GFP+ LAX7 and GFP+ LAX7R competitor cells, respectively. a, d, Enrichment or depletion of GFP+ cells was monitored by flow cytometry (n = 3 independent experiments). Data are means ± s.d. (note that the direction of the y-axis in a is reversed, starting from a fold change of 2.5 at the bottom). b, Levels of ERK1/2-pT202/Y204, ERK1/2, STAT5-pY694 and STAT5 in patient-derived B-ALL cells (IL7RSI246S LAX7, at diagnosis; KRASG12V LAX7R, relapsed) were examined by western blotting (n = 3 independent experiments, shown are two of the technical replicates from an independent experiment). c, Efficiency of CRISPR/Cas9-mediated deletion of BLNK was assessed by western blot (crRNAs, CRISPR RNAs). e, Western blotting analyses were performed to assess levels of STAT5-pY694, STAT5, STAT3-pS727, STAT3, ERK1/2-pT202/Y204 and ERK1/2 upon overnight treatment of LAX7 cells with ruxolitinib (left) or of LAX7R cells with trametinib (right); n = 3 independent experiments. f, Patient-derived B-ALL cells (LAX7 and LAX7R) were treated with increasing concentrations of ruxolitinib or trametinib for 72 h. Relative viability (n = 3; mean ± s.d.) was measured. The x-axes indicate the concentrations of ruxolitinib or trametinib used. For gel source data, see Supplementary Fig. 1.
Extended Data Fig. 10 Pharmacological reactivation of suppressed divergent pathways as a therapeutic strategy in B-ALL.
a, h, Patient-derived B-ALL cells (STAT5-driven) LAX7 (a) and JFK125R (h) were treated overnight with vehicle control (DMSO), BCI-215 (1 μM), ruxolitinib (500 nM), or a combination of both. Western blotting was performed to measure levels of STAT5-pY694, STAT5, ERK1/2-pT202/pY204 and ERK1/2 (n = 3 independent experiments). b, Patient-derived B-ALL cells (ERK-driven, LAX7R) were treated overnight with vehicle control, DPH (1 μM), trametinib (500 nM), or a combination of both. Levels of STAT5-pY694, STAT5, ERK1/2-pT202/Y204 and ERK1/2 were assessed (n = 3 independent experiment). c, i, Patient-derived B-ALL cells (LAX7 and LAX7R (c) and JFK125R (i)) were treated with increasing concentrations of BCI-215, ruxolitinib or both, or DPH, trametinib, or both for three days. Percentage growth inhibition at each concentration is shown as heatmaps (n = 3 independent experiments). Combination indices (Cis) were calculated to determine synergy for treatment combinations. DPH and trametinib concentrations used for LAX7, LAX7R and JFK125R (in μM): DPH, 0, 0.16, 0.31, 0.63,1.3, 2.5, 5; trametinib, 0, 0.032, 0.063, 0.13, 0.25, 0.5, 1. Ruxolitinib and BCI-215 concentrations used for LAX7 (in μM): ruxolitinib, 0, 0.125, 0.25, 0.5, 1, 2; BCI-215, 0, 0.04, 0.08, 0.16, 0.31, 0.63. Ruxolitinib and BCI-215 concentrations used for LAX7R and JFK125R (in μM): ruxolitinib, 0, 0.063, 0.125, 0.25, 0.5, 1, 2; BCI-215, 0, 0.02, 0.04, 0.08, 0.16, 0.31, 0.63. d, j, Single-cell phosphoprotein analyses for STAT5-pY694 and ERK-pT202/Y204 were performed for patient-derived B-ALL cells LAX7 and LAX7R (d) and JFK125R (j) before in vivo treatment (n = 3 independent experiments). e, k, Patient-derived LAX7R (e) or JFK125R (k) B-ALL cells were injected into sublethally irradiated (2 Gy) NSG mice. Recipient mice injected with LAX7R were treated six times a week for four weeks with 2 mg kg−1 DPH, 0.5 mg kg−1 trametinib or both (e). Recipient mice injected with JFK125R were treated five times a week for four weeks with 2 mg kg−1 BCI-215, 30 mg kg−1 ruxolitinib or both (k). Mice were killed when they showed signs of overt leukaemia (hunched back, weight loss and inability to move). The y-axes indicate overall survival (%).Survival curves are shown (n = 6 per group). To assess additive versus synergistic activity of single versus combination treatments in vivo, we adapted the Bliss independence model for survival analysis. With this approach, treatments are Bliss ‘independent’ if the fraction of cells surviving combination therapy equals the product of fractions that survive the individual treatments. A Weibull distribution, {exp[−(t/β)α]}, is fitted to survival data for each condition, and distributions of survival benefits (treated survival time minus untreated survival time) are computed for each treatment. Survival benefits of drugs 1 and 2 are summed and added to the untreated survival distribution to compose a ‘sum of benefits’ survival distribution (see Methods). f, g, l, m, Although combination treatments prolonged overall survival, transplant-recipient mice ultimately developed overt leukaemia. B-ALLs that developed in mice bearing LAX7R (after treatment with DPH plus trametinib; e) and JFK125R (after treatment with BCI-215 plus ruxolitinib; k) were isolated from bone marrow, suspended in cell culture medium and treated with increasing concentrations of BCI-215, ruxolitinib or both, or DPH, trametinib, or both. f, l, Percentage growth inhibition at each concentration is shown as heatmaps (n = 3 independent experiments). Combination indices were calculated to determine synergy of treatment combinations. g, m, To elucidate the clonal composition of LAX7R B-ALL post-treatment (e) and that of JFK125R post-treatment (k), we performed single-cell phosphoprotein analyses for STAT5-pY694 and ERK-pT202/Y204 (n = 3 independent experiments). For gel source data, see Supplementary Fig. 1.
Supplementary information
Supplementary Figure 1
The original source images for all data obtained by electrophoretic separation with molecular weight markers and controls. Blots are labeled according to the corresponding Figure panel within the main or Extended Data Figures.
Supplementary Table 1| Identification of STAT5- and ERK-activating lesions in 1148 B-ALL patient samples.
1148 B-ALL samples were studied for STAT5-activating (CRLF2, IL3RA, IL7R, PDGFRA, PDGFRB, EPOR, ABL1, JAK1, JAK2, JAK3, TYK2 and STAT5A) and ERK-activating (NRAS, KRAS, PTPN11, NF1, RAF1 and FLT3) mutations. The samples originate from six different sources, including the St. Jude Children's Research Hospital Pecan Data Portal (https://pecan.stjude.cloud/proteinpaint/heatmap/HM,BALL), published studies (Nikolaev et al., 2014, Herold et al., 2017, Jerchel et al., 2018) as well as samples analyzed in the Müschen and Weinstock laboratories. Null hypothesis for this analysis was that STAT5- and ERK-pathway mutations occur independently of each other. The expected co-occurrence of the two mutations under the null hypothesis was 121 in 1148 B-ALL samples. The observed co-occurrence of mutations in the two pathways was 35, significantly lower than the expected (odds ratio 0.13, P=2.2e-16, Fisher’s exact test, two-sided).
Supplementary Table 2| Identification of STAT5- and ERK-activating lesions in 916 AML patient samples.
916 AML samples were studied for STAT5-activating (CALR, CBL, JAK1, JAK2, JAK3, KIT, MPL, PDGFRA, PDGFRB and STAT5A) and ERK-activating (RAF1, FLT3, KRAS, NF1, NRAS and PTPN11) mutations. The samples originate from the Catalogue of Somatic Mutations in Cancer (COSMIC; date of access: 2019-07-27). Null hypothesis for this analysis was that STAT5- and ERK-pathway mutations occur independently of each other. The expected co-occurrence of the two mutations under the null hypothesis was 34 in 916 AML samples. The observed co-occurrence of mutations in the two pathways was 24, significantly lower than the expected (odds ratio is 0.59, P=0.033, Fisher’s exact test, two-sided).
Supplementary Tables 3-11
This file contains a description of Supplementary Tables 1 and 2 (provided as separate Excel files), and Supplementary Tables 3-11.
Rights and permissions
About this article
Cite this article
Chan, L.N., Murakami, M.A., Robinson, M.E. et al. Signalling input from divergent pathways subverts B cell transformation. Nature 583, 845–851 (2020). https://doi.org/10.1038/s41586-020-2513-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-020-2513-4
This article is cited by
-
The dopamine transporter antagonist vanoxerine inhibits G9a and suppresses cancer stem cell functions in colon tumors
Nature Cancer (2024)
-
DUSP6 mediates resistance to JAK2 inhibition and drives leukemic progression
Nature Cancer (2022)
-
CCND3 is indispensable for the maintenance of B-cell acute lymphoblastic leukemia
Oncogenesis (2022)
-
Nuclear corepressors NCOR1/NCOR2 regulate B cell development, maintain genomic integrity and prevent transformation
Nature Immunology (2022)
-
Dual-specificity phosphatases: therapeutic targets in cancer therapy resistance
Journal of Cancer Research and Clinical Oncology (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.