Abstract
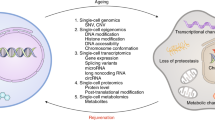
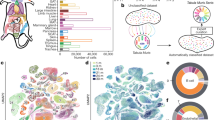
Ageing is the single greatest cause of disease and death worldwide, and understanding the associated processes could vastly improve quality of life. Although major categories of ageing damage have been identified—such as altered intercellular communication, loss of proteostasis and eroded mitochondrial function1—these deleterious processes interact with extraordinary complexity within and between organs, and a comprehensive, whole-organism analysis of ageing dynamics has been lacking. Here we performed bulk RNA sequencing of 17 organs and plasma proteomics at 10 ages across the lifespan of Mus musculus, and integrated these findings with data from the accompanying Tabula Muris Senis2—or ‘Mouse Ageing Cell Atlas’—which follows on from the original Tabula Muris3. We reveal linear and nonlinear shifts in gene expression during ageing, with the associated genes clustered in consistent trajectory groups with coherent biological functions—including extracellular matrix regulation, unfolded protein binding, mitochondrial function, and inflammatory and immune response. Notably, these gene sets show similar expression across tissues, differing only in the amplitude and the age of onset of expression. Widespread activation of immune cells is especially pronounced, and is first detectable in white adipose depots during middle age. Single-cell RNA sequencing confirms the accumulation of T cells and B cells in adipose tissue—including plasma cells that express immunoglobulin J—which also accrue concurrently across diverse organs. Finally, we show how gene expression shifts in distinct tissues are highly correlated with corresponding protein levels in plasma, thus potentially contributing to the ageing of the systemic circulation. Together, these data demonstrate a similar yet asynchronous inter- and intra-organ progression of ageing, providing a foundation from which to track systemic sources of declining health at old age.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
Raw data are available from the Gene Expression Omnibus under accession code GSE132040. Source data are provided with this paper.
References
López-Otín, C., Blasco, M. A., Partridge, L., Serrano, M. & Kroemer, G. The hallmarks of aging. Cell 153, 1194–1217 (2013).
The Tabula Muris Consortium. A single-cell transcriptomic atlas characterizes ageing tissues in the mouse. Nature https://doi.org/10.1038/s41586-020-2496-1 (2020).
The Tabula Muris Consortium. Single-cell transcriptomics of 20 mouse organs creates a Tabula Muris. Nature 562, 367–372 (2018).
Palmer, A. K. & Kirkland, J. L. Aging and adipose tissue: potential interventions for diabetes and regenerative medicine. Exp. Gerontol. 86, 97–105 (2016).
Kohonen, T. The self-organizing map. Proc. IEEE 78, 1464–1480 (1990).
Austad, S. N. & Fischer, K. E. Sex differences in lifespan. Cell Metab. 23, 1022–1033 (2016).
Fuente-Martín, E., Argente-Arizón, P., Ros, P., Argente, J. & Chowen, J. A. Sex differences in adipose tissue. Adipocyte 2, 128–134 (2013).
Zhang, Y. et al. Transcriptional profiling of human liver identifies sex-biased genes associated with polygenic dyslipidemia and coronary artery disease. PLoS ONE 6, e23506 (2011).
Sabolić, I. et al. Gender differences in kidney function. Pflugers Arch. 455, 397–429 (2007).
Hood, S. & Amir, S. The aging clock: circadian rhythms and later life. J. Clin. Invest. 127, 437–446 (2017).
Newman, A. M. et al. Determining cell type abundance and expression from bulk tissues with digital cytometry. Nat. Biotechnol. 37, 773–782 (2019).
Kuk, J. L. et al. Visceral fat is an independent predictor of all-cause mortality in men. Obesity (Silver Spring) 14, 336–341 (2006).
Shaffer, A. L. et al. XBP1, downstream of Blimp-1, expands the secretory apparatus and other organelles, and increases protein synthesis in plasma cell differentiation. Immunity 21, 81–93 (2004).
Vadasz, Z., Haj, T., Kessel, A. & Toubi, E. Age-related autoimmunity. BMC Med. 11, 94 (2013).
Conboy, I. M. et al. Rejuvenation of aged progenitor cells by exposure to a young systemic environment. Nature 433, 760–764 (2005).
Villeda, S. A. et al. Young blood reverses age-related impairments in cognitive function and synaptic plasticity in mice. Nat. Med. 20, 659–663 (2014).
Castellano, J. M. Blood-based therapies to combat aging. Gerontology 65, 84–89 (2019).
Lehallier, B. et al. Undulating changes in human plasma proteome profiles across the lifespan. Nat. Med. 25, 1843–1850 (2019).
Yousef, H. et al. Aged blood impairs hippocampal neural precursor activity and activates microglia via brain endothelial cell VCAM1. Nat. Med. 25, 988–1000 (2019).
Graja, A. et al. Loss of periostin occurs in aging adipose tissue of mice and its genetic ablation impairs adipose tissue lipid metabolism. Aging Cell 17, e12810 (2018).
Gan, K. J. & Südhof, T. C. Specific factors in blood from young but not old mice directly promote synapse formation and NMDA-receptor recruitment. Proc. Natl Acad. Sci. USA 116, 12524–12533 (2019).
Mahmoudi, S., Xu, L. & Brunet, A. Turning back time with emerging rejuvenation strategies. Nat. Cell Biol. 21, 32–43 (2019).
Picelli, S. et al. Full-length RNA-seq from single cells using Smart-seq2. Nat. Protoc. 9, 171–181 (2014).
Picelli, S. et al. Tn5 transposase and tagmentation procedures for massively scaled sequencing projects. Genome Res. 24, 2033–2040 (2014).
Hennig, B. P. et al. Large-scale low-cost NGS library preparation using a robust Tn5 purification and tagmentation protocol. G3 (Bethesda) 8, 79–89 (2018).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Alexa, A. & Rahnenfuhrer, J. topGO: enrichment analysis for gene ontology (Bioconductor, 2016).
Yu, G., Wang, L. G., Han, Y. & He, Q. Y. clusterProfiler: an R package for comparing biological themes among gene clusters. OMICS 16, 284–287 (2012).
Carlson, M. org.Mm.eg.db: Genome wide annotation for Mouse (Bioconductor, 2017).
Butler, A., Hoffman, P., Smibert, P., Papalexi, E. & Satija, R. Integrating single-cell transcriptomic data across different conditions, technologies, and species. Nat. Biotechnol. 36, 411–420 (2018).
Hafemeister, C. & Satija, R. Normalization and variance stabilization of single-cell RNA-seq data using regularized negative binomial regression. Genome Biol. 20, 296 (2019).
Zheng, G. X. Y. et al. Massively parallel digital transcriptional profiling of single cells. Nat. Commun. 8, 14049 (2017).
Fox, J. & Weisberg, S. An R Companion to Applied Regression (Sage, 2011).
Benjamini, Y. & Hochberg, Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J. R. Stat. Soc. B 57, 289–300 (1995).
Szklarczyk, D. et al. STRING v11: protein-protein association networks with increased coverage, supporting functional discovery in genome-wide experimental datasets. Nucleic Acids Res. 47, D607–D613 (2019).
Wehrens, R. & Lutgarde, M. C. Buydens. “Self-and super-organizing maps in R: the Kohonen package. J. Stat. Softw. 21, 1–19 (2007).
Paulson, J. N. et al. Tissue-aware RNA-seq processing and normalization for heterogeneous and sparse data. BMC Bioinformatics 18, 437 (2017).
Acknowledgements
We thank members of the Wyss-Coray laboratory for volunteering help during organ collection and processing: E. Berber, K. Brewer, B. Chang, J. Marschallinger, L. Bonanno, J. Sun, M. F. Lugo-Fagundo, A. Yang and M. W. McNerny. We also thank the members of the Wyss-Coray laboratory and the Chan-Zuckerberg Biohub for feedback and support, and H. Zhang and K. Dickey for laboratory management. This work was funded by the Department of Veterans Affairs (BX004599 to T.W.-C.), the National Institute on Aging (R01-AG045034 and DP1-AG053015 to T.W.-C.), the NOMIS Foundation (T.W.-C.), The Glenn Foundation for Medical Research (T.W.-C.) and the Wu Tsai Neurosciences Institute (T.W.-C.).
Author information
Authors and Affiliations
Consortia
Contributions
N.S., B.L., S.R.Q. and T.W.-C. conceptualized the study. N.S., O.H., B.L., R.P. and T.W.-C. conceptualized the analysis. O.H., B.L. and R.P., with contributions from A.K. and T.F., conducted the transcriptomic analyses. N.S., S.E.L., D.P.L., M.E.Z., H.Z. and D.B. collected samples and extracted RNA. S.H., A.Z. and W.T. conducted cDNA and library preparation. P.M.L. created the Shiny web interface. R.S. and M.T. performed sequencing and library quality control. S.H., A.O.P., J.T.W. and A.M. processed raw sequencing data. The Tabula Muris Consortium generated the single-cell sequencing database. N.S. and K.C. performed and analysed FACS and RNAscope experiments. N.S., B.L., O.H., T.W.-C. and S.R.Q. wrote and edited the manuscript. T.W.-C., S.R.Q., S.D., N.F.N. and J.K. supervised the work. A full list of author contributions for The Tabula Muris Consortium can be found in the Supplementary Information.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature thanks Fan Zhang and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Extended data figures and tables
Extended Data Fig. 1 Gene expression variance analysis.
a, Visualization of the principal variance component analysis, displaying the gene expression variance explained by residuals (that is, biological and technical noise) or experimental factors such as tissue, age, sex and respective combinations. n = 904 total samples. b, c, t-SNE visualization of all samples, based on the first six principal components coloured by age (b) and sex (c). d, Hierarchical clustering of all samples using Ward’s algorithm. Samples are annotated by tissue, sex and age. Highlighted are samples clustering by sex in selected tissues. Non-specific clustering of samples derived from white adipose tissues is further highlighted.
Extended Data Fig. 2 Validation of differential gene expression analysis.
a, Heat map displaying the number of DEGs per tissue for pairwise analysis on adjacent time points. b, Heat map displaying the number of DEGs per tissue for pairwise comparisons, referenced to tissues from 1-month-old mice. c, Heat map displaying the number of DEGs per tissue for pairwise comparisons, referenced to 6-month tissues. d, Box plot displaying the number of DEGs per tissue (n = 17 tissues) for pairwise comparisons with a 3-month reference. In box plots, the centre line denotes the mean, box boundaries denote the first and third quartiles. Outliers show tissues undergoing exceptionally strong expression shifts at a given age. e, Enrichment for functional categories in the top 100 genes that are differentially expressed in the most tissues (ranked using pairwise comparisons with a 3-month reference). Pathway enrichment with GO, Reactome and KEGG databases. Enrichment was tested using Fisher’s exact test (GO) and the hypergeometric test (Reactome and KEGG). To estimate the contribution of each tissue, we used the number of genes per pathway in the top 100 DEGs and estimated the percentage of significant genes per tissue. q-values were estimated using the Benjamini–Hochberg procedure for each database separately, and for GO classes (molecular function, cellular component, biological process) independently. Values of n are as in d. f, Cumulative sum of DEGs per tissue in the ranked top 100 genes. g, Number of DEGs per tissue in the top 100 genes. n = 54 (MAT), 52 (kidney), 52 (GAT), 54 (spleen), 50 (liver), 54 (lung), 50 (intestine), 55 (SCAT), 51 (skin), 53 (BAT), 52 (heart), 52 (muscle), 53 (brain), 52 (WBC), 54 (bone), 51 (marrow), 46 (pancreas). q-values are as in e. h, STRING analysis of the top 30 genes in Fig. 1g.
Extended Data Fig. 3 SOMs of gene correlation with age and sex.
a, b, SOMs were generated from transcriptome-wide gene expression correlation (Spearman’s rank correlation coefficient) of each gene (n = 12,462 genes) with age (a) and sex (b). Genes with similar correlation are mapped to the same cell, and cells are grouped by similarity. The SOM cell layout is common across organs, with the average across all organs at the bottom.
Extended Data Fig. 4 Sex-specific expression changes across organs.
a, Smoothed line plot displaying the number of DEGs between female and male mice at each age. Positive (negative) values represent upregulated (downregulated) genes. Grey lines represent all other tissues. b, Heat map representation of a. c, Expression of Apoe mRNA in GAT and Axin2 mRNA in spleen. The black line indicates LOESS regression. n = 45 (GAT) and n = 47 (spleen) independent samples. d, Venn diagrams depicting the overlap of DEGs in female and male mice detected at 3 months and 18 months of age in GAT, SCAT, liver and kidney. One-sided Fisher’s exact test, ***P < 0.0001. e–h, Top 10 GO terms enriched among the DEGs between female and male mice at 18 months of age in GAT (e), SCAT (f), liver (g) and kidney (h). Data are mean ± s.e.m. n = 2 (female) and n = 4 (male) independent mice for each organ. q values were estimated using the Benjamini–Hochberg procedure for each database separately, and for GO classes (molecular function, cellular component, biological process) independently.
Extended Data Fig. 5 Organ-specific gene expression dynamics.
For each of the 17 organs (rows), the average trajectory of the 15,000 most highly expressed genes is represented in column 1. Five clusters were used (columns 2–6) for further analysis. Average trajectories for each cluster ± s.d. are represented.
Extended Data Fig. 6 Pathway enrichment analysis of organ-specific clusters.
Clusters from Extended Data Fig. 5 show enrichment for genes in functional categories. Pathway enrichment was tested using GO, Reactome and KEGG databases. Enrichment was tested using Fisher’s exact test (GO) and the hypergeometric test (Reactome and KEGG). The top five pathways for each cluster are shown. q values were estimated with the Benjamini–Hochberg procedure for each database separately, and for GO classes (molecular function, cellular component, biological process) independently. Sample size per cluster or tissue is indicated in Extended Data Fig. 5.
Extended Data Fig. 7 Analysis of cytokines and transcription factors.
a, Age-related changes in inflammatory cytokine and chemokines (cytokine-mediated signalling pathways GO:0019221; n = 501 genes), and transcription factors (TRANSFAC database; n = 334 genes). Thicker lines outlined with white represent the average trajectory for each cluster ± s.d. b, c, Spearman correlation coefficient for ageing genes in a.
Extended Data Fig. 8 Integration of bulk and single-cell transcriptomic data.
a, b, Representative GO terms enriched among the genes with highly disperse (a) and cell-specific (b) expression patterns. n = 1,108 cells. q values were estimated using the Benjamini–Hochberg procedure for each database separately, and for GO classes (molecular function, cellular component, biological process) independently. c, Expression of Aco2 mRNA in the kidney. Black line shows LOESS regression; ρ, Spearman’s rank correlation coefficient. n = 52 independent samples. Data are mean ± s.e.m. d, e, t-SNE visualization of scRNA-seq data (FACS) from the kidney, coloured by expression of Aco2 (d) and Cs (e). n = 1,108 cells. f, Violin plot representing expression of Aco1 and Aco2 across all profiled cell types in the kidney. Points indicate cell-wise expression levels and the violin indicates average distribution split by age. Significance was assessed using a Student’s t-test. n = 325 cells (3 months) and 783 cells (24 months). g, Spearman’s rank correlation for cell type fractions significantly (P < 0.05) changing with age, based on deconvolution with FACS or droplet scRNA-seq expression signatures. n = 38 (FACS, BAT), n = 37 (droplet, GAT), n = 37 (FACS, GAT), n = 34 (droplet, kidney), n = 35 (FACS, kidney), n = 35 (droplet, liver), n = 35 (FACS, liver), n = 37 (droplet, lung), n = 37 (FACS, lung), n = 38 (droplet, marrow), n = 36 (FACS, marrow), n = 38 (droplet, MAT), n = 39 (FACS, MAT), n = 34 (droplet, pancreas), n = 32 (FACS, pancreas), n = 37 (droplet, SCAT), n = 38 (FACS, SCAT), n = 35 (droplet, skin), n = 33 (FACS, skin), n = 36 (droplet, spleen), n = 37 (FACS, spleen) independent samples. h, Pairwise comparisons cell fractions between scRNA-seq (FACS), scRNA-seq (droplet), FACS-based bulk RNA-seq deconvolution and droplet-based bulk RNA-seq deconvolution. Each point represents an individual cell type in an individual tissue type.
Extended Data Fig. 9 Identifying Igjhigh B cells with FACS and droplet scRNA-seq.
a, t-SNE visualization of all Cd79a-expressing cells present in the Tabula Muris Senis FACS dataset (17 tissues). Coloured clusters as identified with the Seurat software toolkit. Igjhigh B cell cluster 11 is highlighted. n = 10,867 cells. b, t-SNE in a coloured by the Igjhigh B cell markers Igj, Xbp1 and Derl3. c, GO terms enriched among the top 300 marker genes of Igjhigh (n = 129 cells) versus Igjlow(n = 10,738 cells) (FACS). q values were estimated using the Benjamini–Hochberg procedure for each database separately, and for GO classes (molecular function, cellular component, biological process) independently. d, Distribution of Igjhigh as percentages of Cd79a-expressing cells per tissue. e, Percentage of Igjhigh B cells of all Cd79a-expressing cells across all tissues. n = 5 (3 months) and n = 4 (24 months) independent mice. Significance was assessed using a Student’s t-test. Data are mean ± s.e.m. f, t-SNE visualization of all Cd79a-expressing cells present in the Tabula Muris Senis droplet dataset (17 tissues). Coloured clusters are as identified with the Seurat software toolkit. IgJhigh B cell cluster 5 is highlighted. n = 23,796 cells. g, t-SNE in f coloured by the B cell marker Cd79a and Igjhigh B cell marker Derl3. h, Percentage of Igjhigh B cells of all Cd79a-expressing cells across all tissues. i, Heat map of the z-transformed Igj expression trajectories across bone (n = 54), marrow (n = 51), spleen (n = 54), liver (n = 50), GAT (n = 52), kidney (n = 52), heart (n = 52) and muscle (n = 52). j, Change in Igj mRNA expression in human visceral fat (aged 20–29, n = 25; aged 50–59, n = 124; aged 70–79, n = 12) and subcutaneous fat (aged 20–29, n = 32; aged 50–59, n = 149; aged 70–79, n = 13) (data from GTEx consortium). In the box plot, the median is represented by the centre line and the box boundaries represent the first and third quartiles. k, Number of Igjhigh B cells with a successfully assembled B cell receptor locus, split by mouse and immunoglobulin class. l, Clonally amplified Igjhigh B cells as detected in mouse 1 and mouse 3, grouped by tissue of origin (colour) and immunoglobulin class (shape).
Extended Data Fig. 10 STRING analysis of top correlating plasma proteins.
a, The top seven plasma proteins correlated with gene expression in muscle, coloured by pathway. b, The top 25 plasma proteins correlated with gene expression in any organ.
Supplementary information
Supplementary Information
This file contains a list of consortium authors.
Supplementary Figure
Supplementary Figure 1. Example gating strategy for detecting plasma cells. Gating strategy for detecting plasma cells as in Figure 3j.
Supplementary Tables
This file contains Supplementary Tables 1-15.
Source data
Rights and permissions
About this article
Cite this article
Schaum, N., Lehallier, B., Hahn, O. et al. Ageing hallmarks exhibit organ-specific temporal signatures. Nature 583, 596–602 (2020). https://doi.org/10.1038/s41586-020-2499-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-020-2499-y
This article is cited by
-
Proteomics of mouse brain endothelium uncovers dysregulation of vesicular transport pathways during aging
Nature Aging (2024)
-
Nonlinear DNA methylation trajectories in aging male mice
Nature Communications (2024)
-
Characterizing expression changes in noncoding RNAs during aging and heterochronic parabiosis across mouse tissues
Nature Biotechnology (2024)
-
Inhibition of S6K lowers age-related inflammation and increases lifespan through the endolysosomal system
Nature Aging (2024)
-
Tissue-specific profiling of age-dependent miRNAomic changes in Caenorhabditis elegans
Nature Communications (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.