Abstract
Mutations in PLP1, the gene that encodes proteolipid protein (PLP), result in failure of myelination and neurological dysfunction in the X-chromosome-linked leukodystrophy Pelizaeus–Merzbacher disease (PMD)1,2. Most PLP1 mutations, including point mutations and supernumerary copy variants, lead to severe and fatal disease. Patients who lack PLP1 expression, and Plp1-null mice, can display comparatively mild phenotypes, suggesting that PLP1 suppression might provide a general therapeutic strategy for PMD1,3,4,5. Here we show, using CRISPR–Cas9 to suppress Plp1 expression in the jimpy (Plp1jp) point-mutation mouse model of severe PMD, increased myelination and restored nerve conduction velocity, motor function and lifespan of the mice to wild-type levels. To evaluate the translational potential of this strategy, we identified antisense oligonucleotides that stably decrease the levels of Plp1 mRNA and PLP protein throughout the neuraxis in vivo. Administration of a single dose of Plp1-targeting antisense oligonucleotides in postnatal jimpy mice fully restored oligodendrocyte numbers, increased myelination, improved motor performance, normalized respiratory function and extended lifespan up to an eight-month end point. These results suggest that PLP1 suppression could be developed as a treatment for PMD in humans. More broadly, we demonstrate that oligonucleotide-based therapeutic agents can be delivered to oligodendrocytes in vivo to modulate neurological function and lifespan, establishing a new pharmaceutical modality for myelin disorders.
This is a preview of subscription content, access via your institution
Access options




Similar content being viewed by others
Data availability
All data generated or analysed during this study are included in this article and its Supplementary Information. Animals and iPS cell lines are available from P.J.T. upon request. Source data are provided with this paper.
References
Inoue, K. Pelizaeus–Merzbacher Disease: molecular and cellular pathologies and associated phenotypes. Adv. Exp. Med. Biol. 1190, 201–216 (2019).
Wolf, N. I., van Spaendonk, R. M. L., Hobson, G. M. & Kamholz, J. in Gene Reviews (eds Adam, M. P. et al.) https://www.ncbi.nlm.nih.gov/books/NBK1182/ (1993).
Garbern, J. Y. et al. Patients lacking the major CNS myelin protein, proteolipid protein 1, develop length-dependent axonal degeneration in the absence of demyelination and inflammation. Brain 125, 551–561 (2002).
Griffiths, I. et al. Axonal swellings and degeneration in mice lacking the major proteolipid of myelin. Science 280, 1610–1613 (1998).
Klugmann, M. et al. Assembly of CNS myelin in the absence of proteolipid protein. Neuron 18, 59–70 (1997).
Goldman, S. A., Nedergaard, M. & Windrem, M. S. Glial progenitor cell-based treatment and modeling of neurological disease. Science 338, 491–495 (2012).
Gupta, N. et al. Neural stem cell engraftment and myelination in the human brain. Sci. Transl. Med. 4, 155ra137 (2012).
Saher, G. et al. Therapy of Pelizaeus–Merzbacher disease in mice by feeding a cholesterol-enriched diet. Nat. Med. 18, 1130–1135 (2012).
Wishnew, J. et al. Umbilical cord blood transplantation to treat Pelizaeus–Merzbacher disease in 2 young boys. Pediatrics 134, e1451–e1457 (2014).
Tantzer, S., Sperle, K., Kenaley, K., Taube, J. & Hobson, G. M. Morpholino antisense oligomers as a potential therapeutic option for the correction of alternative splicing in PMD, SPG2, and HEMS. Mol. Ther. Nucleic Acids 12, 420–432 (2018).
Li, H. et al. Gene suppressing therapy for Pelizaeus–Merzbacher disease using artificial microRNA. JCI Insight 4, 125052 (2019).
Nobuta, H. et al. Oligodendrocyte death in Pelizaeus–Merzbacher disease is rescued by iron chelation. Cell Stem Cell 25, 531–541 (2019).
Gruenenfelder, F. I. et al. Neural stem cells restore myelin in a demyelinating model of Pelizaeus–Merzbacher disease. Brain 143, 1383–1399 (2020).
Elitt, M. S. et al. Chemical screening identifies enhancers of mutant oligodendrocyte survival and unmasks a distinct pathological phase in Pelizaeus–Merzbacher Disease. Stem Cell Rep. 11, 711–726 (2018).
Southwood, C. M., Garbern, J., Jiang, W. & Gow, A. The unfolded protein response modulates disease severity in Pelizaeus–Merzbacher disease. Neuron 36, 585–596 (2002).
Jinek, M. et al. A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science 337, 816–821 (2012).
Cong, L. et al. Multiplex genome engineering using CRISPR/Cas systems. Science 339, 819–823 (2013).
Tatar, C. L. et al. Increased Plp1 gene expression leads to massive microglial cell activation and inflammation throughout the brain. ASN Neuro 2, e00043 (2010).
Uschkureit, T., Spörkel, O., Büssow, H. & Stoffel, W. Rumpshaker-like proteolipid protein (PLP) ratio in a mouse model with unperturbed structural and functional integrity of the myelin sheath and axons in the central nervous system. Glia 35, 63–71 (2001).
Finkel, R. S. et al. Nusinersen versus sham control in infantile-onset spinal muscular atrophy. N. Engl. J. Med. 377, 1723–1732 (2017).
Kordasiewicz, H. B. et al. Sustained therapeutic reversal of Huntington’s disease by transient repression of huntingtin synthesis. Neuron 74, 1031–1044 (2012).
Mazur, C. et al. Brain pharmacology of intrathecal antisense oligonucleotides revealed through multimodal imaging. JCI Insight 4, 129240 (2019).
Wu, Q. et al. Elevated levels of the chemokine GRO-1 correlate with elevated oligodendrocyte progenitor proliferation in the jimpy mutant. J. Neurosci. 20, 2609–2617 (2000).
Miller, M. J. et al. Proteolipid protein gene mutation induces altered ventilatory response to hypoxia in the myelin-deficient rat. J. Neurosci. 23, 2265–2273 (2003).
Ueda, A. et al. Pelizaeus–Merzbacher disease can be a differential diagnosis in males presenting with severe neonatal respiratory distress and hypotonia. Hum. Genome Var. 5, 18013 (2018).
Osorio, M. J. et al. Concise Review: stem cell-based treatment of Pelizaeus–Merzbacher disease. Stem Cells 35, 311–315 (2017).
Renier, W. O. et al. Connatal Pelizaeus-Merzbacher disease with congenital stridor in two maternal cousins. Acta Neuropathol. 54, 11–17 (1981).
Lee, Y. et al. Oligodendroglia metabolically support axons and contribute to neurodegeneration. Nature 487, 443–448 (2012).
Fünfschilling, U. et al. Glycolytic oligodendrocytes maintain myelin and long-term axonal integrity. Nature 485, 517–521 (2012).
Freeman, S. A. et al. Acceleration of conduction velocity linked to clustering of nodal components precedes myelination. Proc. Natl Acad. Sci. USA 112, E321–E328 (2015).
Nave, K. A., Lai, C., Bloom, F. E. & Milner, R. J. Jimpy mutant mouse: a 74-base deletion in the mRNA for myelin proteolipid protein and evidence for a primary defect in RNA splicing. Proc. Natl Acad. Sci. USA 83, 9264–9268 (1986).
Hsu, P. D. et al. DNA targeting specificity of RNA-guided Cas9 nucleases. Nat. Biotechnol. 31, 827–832 (2013).
Nakagata, N., Okamoto, M., Ueda, O. & Suzuki, H. Positive effect of partial zona-pellucida dissection on the in vitro fertilizing capacity of cryopreserved C57BL/6J transgenic mouse spermatozoa of low motility. Biol. Reprod. 57, 1050–1055 (1997).
Schmid-Burgk, J. L. et al. OutKnocker: a web tool for rapid and simple genotyping of designer nuclease edited cell lines. Genome Res. 24, 1719–1723 (2014).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Stemmer, M., Thumberger, T., Del Sol Keyer, M., Wittbrodt, J. & Mateo, J. L. CCTop: an intuitive, flexible and reliable CRISPR/Cas9 target prediction tool. PLoS ONE 10, e0124633 (2015).
Bae, S., Park, J. & Kim, J. S. Cas-OFFinder: a fast and versatile algorithm that searches for potential off-target sites of Cas9 RNA-guided endonucleases. Bioinformatics 30, 1473–1475 (2014).
Haeussler, M. et al. Evaluation of off-target and on-target scoring algorithms and integration into the guide RNA selection tool CRISPOR. Genome Biol. 17, 148 (2016).
Robinson, J. T. et al. Integrative genomics viewer. Nat. Biotechnol. 29, 24–26 (2011).
Wiśniewski, J. R., Zougman, A., Nagaraj, N. & Mann, M. Universal sample preparation method for proteome analysis. Nat. Methods 6, 359–362 (2009).
Tran, N. H. et al. Deep learning enables de novo peptide sequencing from data-independent-acquisition mass spectrometry. Nat. Methods 16, 63–66 (2019).
Tran, N. H., Zhang, X., Xin, L., Shan, B. & Li, M. De novo peptide sequencing by deep learning. Proc. Natl Acad. Sci. USA 114, 8247–8252 (2017).
Najm, F. J. et al. Rapid and robust generation of functional oligodendrocyte progenitor cells from epiblast stem cells. Nat. Methods 8, 957–962 (2011).
Lager, A. M. et al. Rapid functional genetics of the oligodendrocyte lineage using pluripotent stem cells. Nat. Commun. 9, 3708 (2018).
Hagemann, T. L. et al. Antisense suppression of glial fibrillary acidic protein as a treatment for Alexander disease. Ann. Neurol. 83, 27–39 (2018).
Langmead, B., Trapnell, C., Pop, M. & Salzberg, S. L. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol. 10, R25 (2009).
Glascock, J. J. et al. Delivery of therapeutic agents through intracerebroventricular (ICV) and intravenous (IV) injection in mice. J. Vis. Exp. 56, 2968 (2011).
Acknowledgements
This research was supported, in part, by grants from the National Institutes of Health R01NS093357 (P.J.T.), T32GM007250 (M.S.E, Z.S.N. and K.C.A), F30HD084167 (Z.S.N.), F30HD096784 (K.C.A.) and T32NS077888 (K.C.A.); the New York Stem Cell Foundation (P.J.T); the European Leukodystrophy Association (P.J.T.); and philanthropic contributions from the Research Institute for Children’s Health and the Geller, Goodman, Fakhouri, Long, Matreyek, Peterson and Weidenthal families. Additional support was provided by the Genomics, Small Molecule Drug Development, Transgenic and Rodent Behavioral core facilities of the Case Western Reserve University (CWRU) Comprehensive Cancer Center (P30CA043703), the Data Analytics Core of the Department of Population and Quantitative Health Sciences at CWRU, the CWRU Light Microscopy Imaging Center (S10OD016164), the electron microscopy division of the Cleveland Clinic Lerner Research Institute Imaging Core, and the University of Chicago Genomics Facility. We thank L. Landmesser, P. MacFarlane, R. Miller, P. Scacheri, T. Wynshaw-Boris, B. Clayton, S. Edelheit, A. Miron, H. Arakawa, P. Philippidou, A. Vagnozzi, L. Hu, E. Cohn, M. Scavuzzo, C. Allan and J. Cregg for technical assistance, discussion and review of the manuscript.
Author information
Authors and Affiliations
Contributions
M.S.E. and P.J.T. conceived and managed the overall study. H.E.S and M.S.E. maintained the animal colonies and tracked survival. M.S.E. captured video recordings. L.B. and M.S.E. designed and tested sgRNAs. S.H. performed data analysis for CRISPR off-target assessments. D.F.L., R.A.C. and W.J. performed zygote electroporation and oviduct transfers. H.E.S., B.S.N., K.C.A. and L.B. performed western blot experiments and protein quantification. D.M.S. and H.E.S. performed mass spectrometry sample preparation and analysis. B.E.P., L.B., B.S.N. and K.C.A. performed RT–qPCR. M.M., B.S.N, L.B., H.E.S., A.S.G. and M.S.E. generated and quantified the immunohistochemistry data. Y.M.-H. performed optic nerve electrophysiology studies and analysed the data. Y.M.-H., M.H., M.S.E. and H.E.S. processed samples for electron microscopy. Y.M.-H. analysed and quantified electron microscopy images. M.S.E., K.C.A., B.S.N. and L.B. performed rotarod and open-field experiments. M.S.E., B.S.N., K.C.A. and H.E.O. generated and characterized iPS cells and OPCs in vitro. H.T.Z. and A.S. generated Hdac2-targeting ASO data. B.E.P. and F.R. designed and characterized Plp1-targeting ASOs, tested tolerability in adult mice, recommended the use of ASOs, and contributed to the study design and interpretation of results in the ASO-treated disease model. M.S.E. performed ASO injections in jimpy mice. Y.M.-H. and B.S.N. performed all respiratory evaluations and analysed the data. Z.S.N. contributed key components to experimental design, data analysis and manuscript composition. M.S.E. and L.B. performed statistical analyses. M.S.E., M.M., Y.M.-H., L.B. and P.J.T. assembled figures. M.S.E. and P.J.T. wrote the manuscript with input from all authors.
Corresponding author
Ethics declarations
Competing interests
P.J.T. and M.S.E. are listed as inventors on pending patent claims (PCT/US2017/064870) filed by CWRU covering methods of PLP1 suppression. P.J.T. is a co-founder and consultant for Convelo Therapeutics, which has licensed patents unrelated to the current study from CWRU inventors (P.J.T., M.S.E., Z.S.N. and M.M.). P.J.T. and CWRU retain equity in Convelo Therapeutics. P.J.T. is a consultant and on the Scientific Advisory Board of Cell Line Genetics, which performed karyotyping in this study. P.J.T. is Chair of the Scientific Advisory Board (volunteer position) for the Pelizaeus-Merzbacher Disease Foundation. B.E.P., H.T.Z., A.S. and F.R. are employees of Ionis Pharmaceuticals. No other authors declare competing interests.
Additional information
Peer review information Nature thanks Evan Goldstein and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 CRISPR nuclease induction of Plp1 frameshift mutations in jimpy with high accuracy.
a, Annotated Sanger sequencing traces of wild-type, jimpy, and CR-impy mice showing the complex, frameshift in Plp1 exon 3 from dual cutting of CRISPR/spCas9 sgRNAs in CR-impy mice as well as the jimpy point mutation in intron 4. sgRNA 3 and 7 sequences outlined by black boxes with the predicted double strand break site shown a black arrow. b, Table showing the top predicted on- and off-target sites for sgRNAs 3 and 7. CRISPR-induced indels were detected by whole genome sequencing of the CR-impy founder and three independent CR-impy F2 generation males, and consisted of an on-target 80bp complex deletion (CR-impy deletion) in exon 3 of Plp1 (green), an off-target 1 bp insertion in chromosome 6 (red), and an off-target 1 bp insertion in chromosome 11 (yellow). c–e, Integrative Genomics Viewer browser images showing aligned reads for the CR-impy founder, the jimpy control, and three CR-impy F2 males along with the detected indels at the on-target locus at exon 3 of Plp1 on chromosome X (c), and off-targets on chromosome 6 (d) and chromosome 11 (e) depicted by the dashed green, red, and yellow boxes, respectively. sgRNA 3 or sgRNA 7 targeted sequences are depicted by black bars.
Extended Data Fig. 2 CRISPR-mediated suppression of Plp1 in jimpy mice increases Mbp expression across multiple CNS regions.
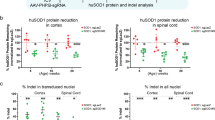
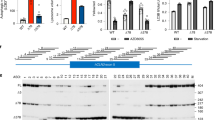
a, RT–qPCR data showing the levels of Plp1 transcript at 6 months (n = 3 mice). b, western blot data demonstrating the levels of MBP protein at 3 weeks (n = 3 mice). c, RT–qPCR data showing the levels of Mbp transcript at 6 months (n = 3 mice). d, western blot data demonstrating the levels of MBP protein at 6 months (n = 3 mice). Individual data points represent the mean value of 4 technical replicates for each biological replicate (a, c) or independent biological replicates (b, d). Biological replicates (individual mice) indicated by open circles. Graph bars indicate mean ± standard deviation. p-values calculated using one-way ANOVA with Tukey correction at 3 weeks or two-way, an unpaired two-sided t-test at later time points. p-values stated for P < 0.1, otherwise not significant (n.s). See Supplementary Data 2 for full western blot images for all samples.
Extended Data Fig. 3 CRISPR-mediated suppression of Plp1 in jimpy mice reduces markers of activated microglia and astrocytes.
a, Immunohistochemical images of whole-brain sagittal sections showing Iba1+ microglia (red) and DAPI+ nuclei (blue) across genotypes. Scale bar, 2mm. b, Immunohistochemical images of whole-brain sagittal sections showing GFAP+ astrocytes (red) and DAPI+ nuclei (blue) staining across genotypes. Scale bar, 2mm. c, d, Normalized mean signal intensity of (c) Iba1+ microglia and (d) GFAP+ astrocytes across genotypes and CNS regions (n = 3 mice). Biological replicates (individual mice) indicated by open circles. Graph bars indicate mean ± standard deviation. p-values calculated using one-way ANOVA with Tukey correction. p-values stated for P < 0.1, otherwise not significant (n.s). See Supplementary Data 3-5 for representative source images of Iba-1 and GFAP staining.
Extended Data Fig. 4 Plp1 suppression in jimpy OPCs rescues survival of differentiating oligodendrocytes in vitro.
a, Phase and immunocytochemistry images of Oct4+ and Nanog+ iPS cells, along with DAPI+ nuclei and b, normal karyotype of a CR-impy iPS cell line used to generate OPCs. Scale bar, 50μm. c, Immunocytochemistry images showing Olig2+ and Sox10+ cells in OPC cultures, along with DAPI+ nuclei, derived from iPS cells. Scale bar, 100μm. d, Percentage of Sox10+ and Olig2+ cells in OPC cultures. e, Immunocytochemistry images of MBP+ and PLP+ oligodendrocytes. f, g, Quantification of (f) MBP+ oligodendrocytes and (g) total cell number (DAPI+ nuclei) from iPS-cell-derived OPCs differentiated in vitro for 3 days. Scale bar, 50μm. Technical replicates (individual wells) for a single cell line per genotype indicated by black circles. Graph bars indicate mean ± standard deviation.
Extended Data Fig. 5 Plp1-targeted ASOs do not suppress off-target transcripts or activate glial cells.
a, b, RT–qPCR data showing the level of (a) Plp1 transcript levels or (b) expression levels of off-target transcripts (up to 3 base mismatches) in the spinal cord for Plp1-tageting ASOs, including Xylt1 (off-target for ASO Plp1.a), Scfd1, or Tpk1 (off-targets for ASO Plp1.b), 2 weeks post-injection of Plp1-targeting ASOs (30μg, 100μg, and 300μg doses) or PBS control in 8 week old adult wild-type (wt) mice (n = 3 mice). c, d, RT–qPCR data showing Plp1 transcript levels or tolerability by expression levels of Gfap, Aif1, and Cd68 transcripts in the cerebral cortex and spinal cord, 8 weeks post-injection with the indicated ASOs (300μg dose) or PBS control in 8 week old wild-type mice (n = 3 mice). e–h, Immunohistochemistry images with haematoxylin counterstain showing Iba1+ or GFAP+ astrocytes in e, Cortical layers I-IV (Iba1), (f) cortical layers I-III (GFAP), (g) spinal cord dorsal horn grey/white matter intersection (Iba1), and (h) spinal cord (GFAP), 8 weeks post-injection with the indicated ASOs (300μg dose) or PBS control in 8 week old wild-type mice. Scale bar, 500μm. Biological replicates (individual mice) indicated by open circles, representing the mean value of 3 technical replicates. Graph bars indicate mean ± standard deviation. p-values calculated using one-way ANOVA with Dunnett’s correction for multiple comparisons. p-values stated for P < 0.1, otherwise not significant (n.s).
Extended Data Fig. 6 Plp1-targeted ASOs distribute widely throughout the CNS after ICV injection in postnatal mice.
a, b, Immunohistochemical images of brain sagittal sections showing ASO+ staining and DAPI+ nuclei (blue) of WT + ASOPlp1.a, WT + ASOPlp1.b and WT uninjected (a) or jp + ASOPlp1.a, jp + ASOPlp1.b and jimpy uninjected mice (b), 3 weeks post-ASO injection (30 μg dose at birth). Scale bar, 50 μm.
Extended Data Fig. 7 Plp1-targeting ASOs increase Mbp expression and rescue oligodendrocyte numbers in jimpy mice.
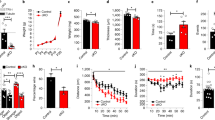
a, Western blot data showing the level of MBP protein (n = 3 mice). b, RT–qPCR data showing the level of Mbp transcript (n = 3 mice). c, Western blot data showing the level of MBP (n = 3 mice). d, Immunohistochemistry images with haematoxylin counterstain of whole brain sagittal sections showing MBP+ myelin. Scale bar, 1mm. e, Quantification of cleaved-caspase 3+ apoptotic cells (n = 3 mice). f, Quantification of CC1+/Olig2+ oligodendrocytes (n = 4 mice). g, Quantification of the number of Olig2+ glial lineage cells (n = 4 mice). h, Quantification of the number of PDGFRα+/Olig2+ OPCs (n = 4 mice). All data collected at 3 weeks post-ASO injection (30μg dose at birth). Individual data points represent the mean value of 4 technical replicates for each biological replicate (individual mice) (b) or independent biological replicates (individual mice) (a, c–h), indicated by open circles. Graph bars indicate mean ± standard deviation. p-values calculated using one-way ANOVA with Dunnett’s correction for multiple comparisons. p-values stated for P < 0.1, otherwise not significant (n.s). See Supplementary Data 4 for full western blot images for all samples.
Extended Data Fig. 8 Plp1-targeted ASOs induce sustained myelination throughout the neuraxis in jimpy mice.
a, b, Electron micrograph images showing myelination of WT + ASOctr or jp + ASOPlp1.b at 2 months (a) and 8 months (b). For a, scale bar, 0.5 μm. In b, the bottom panel is a higher magnification of red boxed area in the top panel. Scale bars, 5 μm (top) and 0.5 μm (bottom).
Supplementary information
Supplementary Information
This merged pdf file contains Supplementary Tables 1-4, Supplementary Data 1-14, which provide metadata for in vivo studies, uncropped scans of immunoblots, and source immunohistochemical images for cell number quantifications, and supplementary references.
41586_2020_2494_MOESM3_ESM.mp4
Video 1 Plp1 suppression in jimpy mice rescues neurological phenotypes at 3 weeks of age. Video comparison of wild-type, jimpy, and CR-impy mice at 3 weeks of age. Representative videos of n=25, 20, 12 wild-type, CR-impy, and jimpy mice, respectively.
41586_2020_2494_MOESM4_ESM.mp4
Video 2 Plp1 suppression in jimpy mice shows sustained rescue of neurological phenotypes at 18 months of age. Video comparison of wild-type and CR-impy mice at 18 months of age (study endpoint). Representative videos of n=4, 5 wild-type and CR-impy mice, respectively.
Video 3
Postnatal delivery of Plp1-targeted ASOs to jimpy mice rescues neurological phenotypes at 3 weeks of age. Video comparison of wild-type and jimpy mice injected with control and Plp1-targeting ASOs at 3 weeks of age. Representative videos of n = 5 (uninjected wild-type), 5 (jimpy + ASOPlp1.a), and 8 (jimpy + ASOPlp1.b) mice.
Video 4
Postnatal delivery of Plp1-targeted ASOs to jimpy mice shows sustained rescue of neurological phenotypes at 6 months of age. Video comparison of wild-type and jimpy mice injected with control and Plp1-targeting ASOs at 6 months of age. Representative videos of n = 5 (uninjected wild-type), 5 (jimpy + ASOPlp1.a), and 7 (jimpy + ASOPlp1.b) mice.
Rights and permissions
About this article
Cite this article
Elitt, M.S., Barbar, L., Shick, H.E. et al. Suppression of proteolipid protein rescues Pelizaeus–Merzbacher disease. Nature 585, 397–403 (2020). https://doi.org/10.1038/s41586-020-2494-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-020-2494-3
This article is cited by
-
Pervasive environmental chemicals impair oligodendrocyte development
Nature Neuroscience (2024)
-
Oligodendrocyte differentiation alters tRNA modifications and codon optimality-mediated mRNA decay
Nature Communications (2022)
-
The Oligodendrocyte Transcription Factor 2 OLIG2 regulates transcriptional repression during myelinogenesis in rodents
Nature Communications (2022)
-
Hypomyelinating leukodystrophies — unravelling myelin biology
Nature Reviews Neurology (2021)
-
Oligodendrocytes depend on MCL-1 to prevent spontaneous apoptosis and white matter degeneration
Cell Death & Disease (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.