Abstract
The mitochondrial electron transport chain (ETC) is necessary for tumour growth1,2,3,4,5,6 and its inhibition has demonstrated anti-tumour efficacy in combination with targeted therapies7,8,9. Furthermore, human brain and lung tumours display robust glucose oxidation by mitochondria10,11. However, it is unclear why a functional ETC is necessary for tumour growth in vivo. ETC function is coupled to the generation of ATP—that is, oxidative phosphorylation and the production of metabolites by the tricarboxylic acid (TCA) cycle. Mitochondrial complexes I and II donate electrons to ubiquinone, resulting in the generation of ubiquinol and the regeneration of the NAD+ and FAD cofactors, and complex III oxidizes ubiquinol back to ubiquinone, which also serves as an electron acceptor for dihydroorotate dehydrogenase (DHODH)—an enzyme necessary for de novo pyrimidine synthesis. Here we show impaired tumour growth in cancer cells that lack mitochondrial complex III. This phenotype was rescued by ectopic expression of Ciona intestinalis alternative oxidase (AOX)12, which also oxidizes ubiquinol to ubiquinone. Loss of mitochondrial complex I, II or DHODH diminished the tumour growth of AOX-expressing cancer cells deficient in mitochondrial complex III, which highlights the necessity of ubiquinone as an electron acceptor for tumour growth. Cancer cells that lack mitochondrial complex III but can regenerate NAD+ by expression of the NADH oxidase from Lactobacillus brevis (LbNOX)13 targeted to the mitochondria or cytosol were still unable to grow tumours. This suggests that regeneration of NAD+ is not sufficient to drive tumour growth in vivo. Collectively, our findings indicate that tumour growth requires the ETC to oxidize ubiquinol, which is essential to drive the oxidative TCA cycle and DHODH activity.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
All data from the manuscript are available from the corresponding author on request. Source data are provided with this paper.
References
DeBerardinis, R. J. & Chandel, N. S. We need to talk about the Warburg effect. Nat. Metab. 2, 127–129 (2020).
Weinberg, F. et al. Mitochondrial metabolism and ROS generation are essential for Kras-mediated tumorigenicity. Proc. Natl Acad. Sci. USA 107, 8788–8793 (2010).
Tan, A. S. et al. Mitochondrial genome acquisition restores respiratory function and tumorigenic potential of cancer cells without mitochondrial DNA. Cell Metab. 21, 81–94 (2015).
Ju, Y. S. et al. Origins and functional consequences of somatic mitochondrial DNA mutations in human cancer. eLife 3, (2014).
Kuntz, E. M. et al. Targeting mitochondrial oxidative phosphorylation eradicates therapy-resistant chronic myeloid leukemia stem cells. Nat. Med. 23, 1234–1240 (2017).
Roth, K. G., Mambetsariev, I., Kulkarni, P. & Salgia, R. The mitochondrion as an emerging therapeutic target in cancer. Trends Mol. Med. 26, 119–134 (2020).
Navarro, P. et al. Targeting tumor mitochondrial metabolism overcomes resistance to antiangiogenics. Cell Rep. 15, 2705–2718 (2016).
Kim, S. H. et al. Phenformin inhibits myeloid-derived suppressor cells and enhances the anti-tumor activity of PD-1 blockade in melanoma. J. Invest. Dermatol. 137, 1740–1748 (2017).
Viale, A. et al. Oncogene ablation-resistant pancreatic cancer cells depend on mitochondrial function. Nature 514, 628–632 (2014).
Maher, E. A. et al. Metabolism of [U-13C]glucose in human brain tumors in vivo. NMR Biomed. 25, 1234–1244 (2012).
Hensley, C. T. et al. Metabolic heterogeneity in human lung tumors. Cell 164, 681–694 (2016).
Hakkaart, G. A., Dassa, E. P., Jacobs, H. T. & Rustin, P. Allotopic expression of a mitochondrial alternative oxidase confers cyanide resistance to human cell respiration. EMBO Rep. 7, 341–345 (2006).
Titov, D. V. et al. Complementation of mitochondrial electron transport chain by manipulation of the NAD+/NADH ratio. Science 352, 231–235 (2016).
Martínez-Reyes, I. et al. TCA cycle and mitochondrial membrane potential are necessary for diverse biological functions. Mol. Cell 61, 199–209 (2016).
Garcia-Bermudez, J. et al. Aspartate is a limiting metabolite for cancer cell proliferation under hypoxia and in tumours. Nat. Cell Biol. 20, 775–781 (2018).
Sullivan, L. B. et al. Supporting aspartate biosynthesis is an essential function of respiration in proliferating cells. Cell 162, 552–563 (2015).
Sullivan, L. B. et al. Aspartate is an endogenous metabolic limitation for tumour growth. Nat. Cell Biol. 20, 782–788 (2018).
Birsoy, K. et al. An essential role of the mitochondrial electron transport chain in cell proliferation is to enable aspartate synthesis. Cell 162, 540–551 (2015).
Bajzikova, M. et al. Reactivation of dihydroorotate dehydrogenase-driven pyrimidine biosynthesis restores tumor growth of respiration-deficient cancer cells. Cell Metab. 29, 399–416 (2019).
Mullen, A. R. et al. Reductive carboxylation supports growth in tumour cells with defective mitochondria. Nature 481, 385–388 (2011).
Lussey-Lepoutre, C. et al. Loss of succinate dehydrogenase activity results in dependency on pyruvate carboxylation for cellular anabolism. Nat. Commun. 6, 8784 (2015).
Cardaci, S. et al. Pyruvate carboxylation enables growth of SDH-deficient cells by supporting aspartate biosynthesis. Nat. Cell Biol. 17, 1317–1326 (2015).
DeBerardinis, R. J. & Chandel, N. S. Fundamentals of cancer metabolism. Sci. Adv. 2, e1600200 (2016).
Guo, J. Y. et al. Activated Ras requires autophagy to maintain oxidative metabolism and tumorigenesis. Genes Dev. 25, 460–470 (2011).
Romero, R. et al. Keap1 loss promotes Kras-driven lung cancer and results in dependence on glutaminolysis. Nat. Med. 23, 1362–1368 (2017).
Momcilovic, M. et al. In vivo imaging of mitochondrial membrane potential in non-small-cell lung cancer. Nature 575, 380–384 (2019).
Molina, J. R. et al. An inhibitor of oxidative phosphorylation exploits cancer vulnerability. Nat. Med. 24, 1036–1046 (2018).
Alistar, A. et al. Safety and tolerability of the first-in-class agent CPI-613 in combination with modified FOLFIRINOX in patients with metastatic pancreatic cancer: a single-centre, open-label, dose-escalation, phase 1 trial. Lancet Oncol. 18, 770–778 (2017).
DuPage, M., Dooley, A. L. & Jacks, T. Conditional mouse lung cancer models using adenoviral or lentiviral delivery of Cre recombinase. Nat. Protoc. 4, 1064–1072 (2009).
Chong, J. et al. MetaboAnalyst 4.0: towards more transparent and integrative metabolomics analysis. Nucleic Acids Res. 46 (W1), W486–W494 (2018).
Acknowledgements
This work was supported by the NIH (5R35CA197532) to N.S.C.; a postdoctoral fellowship by Ramon Areces Foundation of Spain to I.M.R.; NCI (T32 CA009560) and NIH (2T32HL076139-16) to H. Kong; NIH (5 T32 CA 9560-33) and Northwestern University Pulmonary department’s Cugell fellowship to K.V.; NIH (T32CA09560 and T32HL076139-13) to G.S.M., and NIH (T32 T32HL076139) to S.E.W. E.M.S. is a Cancer Research Institute Irvington Fellow supported by the Cancer Research Institute. We thank Robert H. Lurie Cancer Center Flow Cytometry facility supported by NCI CCSG P30 CA060553 for their invaluable assistance. Imaging work was performed at the Northwestern University Center for Advanced Microscopy generously supported by NCI CCSG P30 CA060553 awarded to the Robert H Lurie Comprehensive Cancer Center. We thank T. Papagiannakopoulos and S. Leboeuf for providing KP cells. We thank C. Moraes for CYTB-Δ cells. Original LbNOX constructs were generated by V. Mootha’s laboratory and acquired from Addgene. We thank E. Dufour and H. Jacobs for original NDI1 and AOX constructs. We thank P. Ntziachristos for the MIGR1-Notch1ΔE-GFP vector. We thank the members of the Chandel laboratory for discussions.
Author information
Authors and Affiliations
Contributions
I.M.-R. and N.S.C. conceptualized the study, interpreted the data, and wrote the manuscript with the input of co-authors. I.M.-R., L.R.C., H. Kong, M.W., H. Kihshen, G.S.M., C.R.R. and K.V. carried out most of the experiments in the paper. I.M.-R., L.R.C. and H. Kong performed leukaemia experiments. I.M.-R., L.R.C., H. Kong and K.V. performed lung tumour experiments. I.M.-R., L.R.C., C.R.R, G.S.M. and S.E.W. carried out xenograft experiments. I.M.-R., M.W. and H. Kihshen performed experiments to analyse the OCR in intact and permeabilized cells. I.M.-R., C.R.R. and M.W performed experiments to assess proliferation. I.M.-R. and M.W. performed western blot experiments. I.M.-R., L.R.C., G.S.M. and P.G. conducted and analysed metabolomics and carbon flux experiments. R.P., G.R.S.B., E.M.S. and S.E.W. provided technical expertise with mouse experiments.
Corresponding author
Ethics declarations
Competing interests
N.S.C. is a scientific advisory board member of Rafael Pharmaceuticals.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Metabolite changes in complex III deficient cells in the presence or absence of pyruvate and uridine.
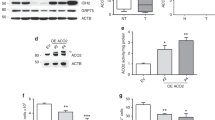
a, b, Schematic representation of the ETC in 143B-CYTB-WT (a) and 143B-CYTB-Δ cells (b). c, Coupled OCR of 143B-CYTB-WT and 143B-CYTB-Δ cells (n = 5 biologically independent experiments). d, 143B-CYTB-WT and 143B-CYTB-Δ cells were grown in the presence or absence of methyl pyruvate and/or uridine and cell number was assessed after 72 h (n = 5 biologically independent experiments). e, Intracellular aspartate levels in the presence of methyl pyruvate and uridine in 143B-CYTB-WT and 143B-CYTB-Δ cells (n = 4 biologically independent experiments). f, Intracellular aspartate levels in the absence of methyl pyruvate and uridine in 143B-CYTB-WT and 143B-CYTB-Δ cells (n = 5 biologically independent experiments). g, Intracellular NAD+/NADH ratio in the absence of methyl pyruvate and uridine of 143B-CYTB-WT and 143B-CYTB-Δ cells (n = 5 biologically independent experiments). h, The heat map displays the relative abundance of significantly changed metabolites in 143B-CYTB-WT, 143B-CYTB-Δ cells and in 143B-CYTB-Δ cells expressing either GFP or AOX in the absence of methyl pyruvate and uridine. A red–blue colour scale depicts the abundance of the metabolites (red: high, blue: low) (n = 5 biologically independent experiments). i, The heat map displays the relative abundance of significantly changed metabolites in 143B-CYTB-WT and 143B-CYTB-Δ cells in the presence of methyl pyruvate and uridine (n = 4 biologically independent experiments). j, Tumour mass of xenografts from 143B-CYTB-WT and 143B-CYTB-Δ cells (n = 10 mice per group from two independent cohorts). k, Coupled OCR of KP-NT and KP-QPC_KO cells (n = 10 technical replicates from two independent experiments). Data are mean ± s.e.m. (c–g, j) or mean ± s.d. (k). *P < 0.05, **P < 0.01, two-tailed t-tests (c, g, j), two-way ANOVA (d) with a Bonferroni test for multiple comparisons or one-way ANOVA (k) with a Bonferroni test for multiple comparisons (exact P values are in the Source Data). Metabolites levels were analysed with multiple one-way ANOVA using an FDR of 0.1 and Fisher’s least significant difference test post hoc analyses Q = 10%. For two-group heat maps, t-tests with an FDR cut-off value of 0.1 were used to identify significantly changed metabolites. Each row was analysed individually. (*Q < 0.1; exact Q values are in the Source Data).
Extended Data Fig. 2 Mitochondrial complex III is required for T-ALL growth in vivo.
a, Schematic representation of the T-ALL experiments. b, c, Percentage of GFP+ T-ALL cells from the spleen (b) or bone marrow (c) of QPC-WT and QPC-KO recipients (WT: n = 7; KO: n = 5 mice). d, e, The absolute number of GFP+ T-ALL cells from the spleen (d) or bone marrow (e) of QPC-WT and QPC-KO recipients (WT: n = 7; KO: n = 5 mice). f, Weight of spleens from QPC-WT and QPC-KO recipients (WT: n = 6; KO: n = 4 mice). g, Survival of mice injected with QPC-WT or QPC-KO T-ALL cells (WT: n = 7; KO: n = 4 mice). Data are mean ± s.e.m. from three independent experiments. *P < 0.05, **P < 0.01, two-tailed t-tests with a Welch’s correction (exact P values are in the Source Data). Survival curves were compared using the log-rank test (P < 0.0001). An example of the gating strategy is provided in Supplementary Fig. 7.
Extended Data Fig. 3 Complex III-deficient cells are auxotrophic for uridine.
a, Schematic representation of the ETC in AOX expressing 143B-CYTB-Δ cells. b, Coupled OCR of 143B-CYTB-Δ-GFP and 143B-CYTB-Δ-AOX cells (n = 5 biologically independent experiments). c, Tumour mass of xenografts from 143B-CYTB-Δ-GFP and 143B-CYTB-Δ-AOX cells (n = 9 mice per group from two independent cohorts). d, Coupled OCR of KP-QPC_KO-GFP and KP-QPC_KO-AOX cells (n = 7 replicates from one representative of five biologically independent experiments). e, 143B-CYTB-WT treated or untreated with piericidin A (0.5 μM) or antimycin A (0.5 μM) were grown in the presence or absence of methyl pyruvate and/or uridine and cell number was assessed after 72 h (n = 4 biologically independent experiments). f, The dihydroorotate-to-orotate ratio was assessed in 143B-CYTB-WT treated or untreated with piericidin A (0.5 μM) or antimycin A (0.5 μM) (n = 6 biologically independent experiments). Data are mean ± s.e.m. (b, c, e, f) or mean ± s.d. (d). *P < 0.05, **P < 0.01, two-tailed t-tests (b–d), two-way ANOVA (e) with a Bonferroni test for multiple comparisons or one-way ANOVA (f) with a Bonferroni test for multiple comparisons (exact P values are in the Source Data).
Extended Data Fig. 4 De novo pyrimidine synthesis is necessesary for tumour growth.
a, Schematic representation of the ETC in 143B-CYTB-Δ–DHODH_KO-AOX cells. b, Western blot analysis of DHODH in 143B-CYTB-Δ non-targeting (NT) and 143B-CYTB-Δ-DHODH_KO cells. Tubulin was used as a loading control. Data are representative of two independent experiments. c, 143B-CYTB-Δ-NT or 143B-CYTB-Δ-DHODH-KOs expressing GFP or AOX were grown in the presence or absence of uridine and cell number was assessed after 72 h (n = 5 biologically independent experiments). d, e, Average tumour volume (d) and tumour mass (e) of xenografts from 143B-CYTB-Δ-NT-AOX and 143B-CYTB-Δ-DHODH_KO2-AOX cells (n = 10 mice per group from two independent cohorts). f, Western blot analysis of DHODH protein levels in 143B-CYTB-Δ-NT, 143B-CYTB-Δ-DHODH_KO2-AOX-RFP and 143B-CYTB-Δ-DHODH_KO2-AOX-cDNA DHODH cells. Data are representative of three independent experiments. g, 143B-CYTB-Δ-DHODH_KO2-AOX-RFP and 143B-CYTB-Δ-DHODH_KO2-AOX-cDNA DHODH cells were grown in the presence or absence of uridine and cell number was assessed after 72 h (n = 5 biologically independent experiments). h, i, Average tumour volume (h) and tumour mass (i) of xenografts from 143B-CYTB-Δ-DHODH_KO2-AOX-RFP and 143B-CYTB-Δ-DHODH_KO2-AOX-cDNA DHODH cells (n = 9 mice per group from two independent cohorts). Data are mean ± s.e.m. (c–e, g–i) *P < 0.05, **P < 0.01, two-tailed t-tests (e, i) or two-way ANOVA (c, d, g, h) with a Bonferroni test for multiple comparisons (exact P values are in the Source Data). For gel source data, see Supplementary Fig. 4.
Extended Data Fig. 5 Restoration of complex I by ectopic expression of NDUFS2 cDNA rescues tumour growth.
a, Schematic representation of the ETC in complex I-deficient 143B-CYTB-Δ-NDUFS2_KO-AOX cells. b, Tumour mass of xenografts from 143B-CYTB-Δ-NT-AOX and 143B-CYTB-Δ-NDUFS2_KO1-AOX cells (n = 10 mice per group from two independent cohorts). c, Western blot analysis of NDUFS2 protein levels in 143B-CYTB-Δ-NT cells, and in AOX-expressing 143B-CYTB-Δ-NDUFS2_KO1 clone transduced with either RFP or human NDUFS2 cDNA. GAPDH was used as a loading control. Data representative of two independent experiments. d, Basal OCR of AOX expressing 143B-CYTB-Δ-NDUFS2_KO1 cells transduced with either RFP or human NDUFS2 cDNA. e,143B-CYTB-Δ-NDUFS2_KO1-AOX-RFP and 143B-CYTB-Δ-NDUFS2_KO1-AOX-cDNA NDUFS2 cells were grown in the presence or absence of methyl pyruvate and cell number was assessed after 72 h (n = 5 biologically independent experiments). f, g, Average tumour volume (f) and tumour mass (g) of xenografts from AOX-expressing 143B-CYTB-Δ-NDUFS2_KO1 cells transduced with either RFP or human NDUFS2 cDNA (n = 9 mice per group from two independent cohorts). Data are mean ± s.e.m. (b, d–g). *P < 0.05, **P < 0.01, two-tailed t-tests (b, d, g) or two-way ANOVA (e, f) with a Bonferroni test for multiple comparisons (exact P values are in the Source Data). For gel source data, see Supplementary Fig. 2.
Extended Data Fig. 6 NDI1 expression in complex I-deficient cells rescues electron transfer but not ATP production.
a, Schematic representation of the ETC in complex I deficient 143B-CYTB-Δ-NDUFS2_KO-AOX cells expressing NDI1. b, 143B-CYTB-Δ-NDUFS2_KO1-AOX-RFP and 143B-CYTB-Δ-NDUFS2_KO1-AOX-NDI1 cells were grown in the presence or absence of methyl pyruvate and cell number was assessed after 72 h (n = 6 biologically independent experiments). c, The heat map displays the relative abundance of significantly changed metabolites in 143B-CYTB-Δ–NDUFS2_KO1-AOX cells expressing RFP, NDI1 or LbNOX in either mitochondria or cytosol (n = 4 biologically independent experiments). A red–blue colour scale depicts the abundance of the metabolites (red: high, blue: low). Metabolites levels were analysed with multiple one-way ANOVA using an FDR of 0.1 and Fisher’s least significant difference test post hoc analyses Q = 10%. Each row was analysed individually. (*Q < 0.1; exact Q values are in the Source Data.) d, 143B-CYTB-Δ-NT-AOX, 143B-CYTB-Δ-NDUFS2_KO1-AOX-RFP and 143B-CYTB-Δ-NDUFS2_KO1-AOX-NDI1 cells were grown in media containing 10 mM glucose or 10 mM galactose for 48 h and assessed for cell death (n = 4 biologically independent experiments). e, Tumour mass of xenografts from 143B-CYTB-Δ-NDUFS2_KO1 cells expressing AOX and either RFP or NDI1 (n = 10 mice per group from two independent cohorts). Data are mean ± s.e.m. (b, d, e). *P < 0.05, **P < 0.01, two-tailed t-tests (e) or two-way ANOVA (b, d) with a Bonferroni test for multiple comparisons (exact P values are in the Source Data).
Extended Data Fig. 7 LbNOX expression in mitochondria or cytosol promotes major changes in the metabolome of complex III-deficient cells.
a, Schematic representation of the ETC in 143B-CYTB-Δ cells expressing LbNOX in mitochondria. b, Intracellular NAD+/NADH ratio in 143B-CYTB-Δ-RFP, 143B-CYTB-Δ-LbNOX-Mito and 143B-CYTB-Δ-LbNOX-Cyto cells in the absence of methyl pyruvate (n = 5 biologically independent experiments). c–e, Intracellular aspartate (c), succinate (d) and α-ketoglutarate levels (e) in 143B-CYTB-Δ-RFP, 143B-CYTB-Δ-LbNOX-Mito and 143B-CYTB-Δ-LbNOX-Cyto cells in the absence of methyl pyruvate (n = 5 biologically independent experiments). f, The heat map displays the relative abundance of significantly changed metabolites in 143B-CYTB-Δ-RFP, 143B-CYTB-Δ-LbNOX-Mito and 143B-CYTB-Δ-LbNOX-Cyto cells in the absence of methyl pyruvate (n = 5 biologically independent experiments). A red–blue colour scale depicts the abundance of the metabolites (red: high, blue: low). g, Tumour mass of xenografts from 143B-CYTB-Δ-RFP, 143B-CYTB-Δ-LbNOX-Mito and 143B-CYTB-Δ-LbNOX-Cyto cells (n = 9 mice per group from two independent cohorts). Data are mean ± s.e.m. (b–e, g). *P < 0.05, **P < 0.01, one-way ANOVA (b, g) with a Bonferroni test for multiple comparisons (exact P values are in the Source Data). Metabolites levels (c–f) were analysed with multiple one-way ANOVA using an FDR of 0.1 and Fisher’s least significant difference test post hoc analyses Q = 10%. Each row was analysed individually (*Q < 0.1; exact Q values in Source Data).
Extended Data Fig. 8 LbNOX expression in mitochondria or cytosol promotes major changes in the metabolome of complex I deficient cells.
a, Schematic representation of the ETC in 143B-CYTB-Δ-NDUFS2_KO-AOX cells expressing LbNOX in mitochondria. b, 143B-CYTB-Δ-NDUFS2_KO1-AOX-LbNOX-Mito and 143B-CYTB-Δ-NDUFS2_KO1-AOX-LbNOX-Cyto were grown in the presence or absence of methyl pyruvate and cell number was assessed after 72 h (n = 5 biologically independent experiments). c, Intracellular NAD+/NADH ratio of 143B-CYTB-Δ-NDUFS2_KO1-AOX-RFP, 143B-CYTB-Δ-NDUFS2_KO1-AOX-LbNOX-Mito and 143B-CYTB-Δ-NDUFS2_KO1-AOX-LbNOX-Cyto cells in the absence of methyl pyruvate and uridine (n = 4 biologically independent experiments). d, Intracellular aspartate levels of 143B-CYTB-Δ-NDUFS2_KO1-AOX-RFP, 143B-CYTB-Δ-NDUFS2_KO1-AOX-LbNOX-Mito and 143B-CYTB-Δ-NDUFS2_KO1-AOX-LbNOX-Cyto cells in the absence of methyl pyruvate and uridine (n = 4 biologically independent experiments). e, 143B-CYTB-Δ-NDUFS2_KO1-AOX-LbNOX-Mito and 143B-CYTB-Δ-NDUFS2_KO1-AOX-LbNOX-Cyto cells were grown in medium containing 10 mM glucose or 10 mM galactose for 48 h and assessed for cell death (n = 4 biologically independent experiments). f, Tumour mass of xenografts from 143B-CYTB-Δ-NDUFS2_KO1-AOX cells expressing LbNOX in either mitochondria or cytosol (n = 10 mice per group from two independent cohorts). g, Western blot analysis (data representative of two independent experiments) of LbNOX expression in xenograft tumours from 143B-CYTB-Δ-NDUFS2_KO1-AOX-RFP, 143B-CYTB-Δ-NDUFS2_KO1-AOX-LbNOX-Mito and 143B-CYTB-Δ-NDUFS2_KO1-AOX-LbNOX-Cyto cells. Tubulin was used as a loading control. Data are mean ± s.e.m. (b–f). *P < 0.05, **P < 0.01, two-tailed t-tests (f), one-way ANOVA (c) with a Bonferroni test for multiple comparisons or a two-way ANOVA (b, e) with a Bonferroni test for multiple comparisons (exact P values are in the Source Data). Metabolites levels (d) were analysed with multiple one-way ANOVA using an FDR of 0.1 and Fisher’s least significant difference test post hoc analyses Q = 10%. Each row was analysed individually. (*Q < 0.1; exact Q values in Source Data.) For gel source data, see Supplementary Fig. 5.
Extended Data Fig. 9 Complex I-deficient cells expressing LbNOX in the cytosol perform glutamine reductive carboxylation.
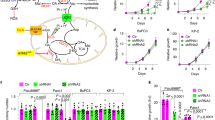
a, Schematic representation for oxidative and reductive glutamine metabolism. Metabolism of [U-13C]glutamine generates fully labelled α-ketoglutarate. Oxidation of α-ketoglutarate in the TCA cycle produces metabolites with four 13C-carbons (m + 4), while reduction of α-ketoglutarate through the reductive carboxylation pathway produces citrate with five 13C-carbons (m + 5). Further reductive metabolism of the m + 5 citrate yields metabolites with three 13C-carbons (m + 3). b–h, 143B-CYTB-Δ-NDUFS2_KO1-AOX-RFP, 143B-CYTB-Δ-NDUFS2_KO1-AOX-LbNOX-Mito and 143B-CYTB-Δ-NDUFS2_KO1-AOX-LbNOX-Cyto cells were labelled for 6 h with [U-13C]glutamine in the presence (b–d) or absence (e–h) of methyl pyruvate, and the percentage of labelled metabolite pools was examined. m + 5 and m + 3 pools result from glutamine flow through reductive metabolism. m + 4 pools result from glutamine flow through oxidative metabolism. Data are mean ± s.e.m. of four biologically independent experiments.
Extended Data Fig. 10 Complex II is necessary for tumour growth.
a, Schematic representation of the ETC in complex II deficient 143B-CYTB-Δ cells expressing AOX. b, Western blot analysis of SDHA in 143B-CYTB-Δ non-targeting and 143B-CYTB-Δ-SDHA_KO cells. Tubulin was used as a loading control. Data representative of two independent experiments. c, Complex II-driven OCR of permeabilized 143B-CYTB-Δ-NT-AOX and 143B-CYTB-Δ-SDHA_KO2-AOX cells. Piericidin A (1 μM) and antimycin A (1 μM) were used to inhibit complex I and III, respectively. SHAM (2 mM) was used to inhibit AOX activity (n = 4 biologically independent experiments). d, 143B-CYTB-Δ-SDHA-KOs expressing GFP or AOX were grown in the presence or absence of methyl pyruvate and cell number was assessed after 72 h (n = 5 biologically independent experiments). e, f, Average tumour volume (e) and tumour mass (f) of xenografts from 143B-CYTB-Δ-NT-AOX and 143B-CYTB-Δ-SDHA_KO2-AOX cells (n = 8 mice per group from two independent cohorts). Data are mean ± s.e.m. (c–f). *P < 0.05; **P < 0.01, two-tailed t-tests (f) or two-way ANOVA (d, e) with a Bonferroni test for multiple comparisons (exact P value are in the Source Data). For gel source data, see Supplementary Fig. 6.
Extended Data Fig. 11 Restoration of complex II by ectopic expression of SDHA cDNA rescues tumour growth.
a, Western blot analysis of SDHA protein levels in 143B-CYTB-Δ-NT, 143B-CYTB-Δ-SDHA_KO2-AOX-RFP and 143B-CYTB-Δ-SDHA_KO2-AOX-cDNA SDHA cells. Data representative of three independent experiments. b, Complex II-driven OCR of permeabilized 143B-CYTB-Δ-SDHA_KO2-AOX-RFP and 143B-CYTB-Δ-SDHA_KO2-AOX-cDNA SDHA cells. Succinate and ADP were provided as substrates. Piericidin A (1 μM) and antimycin A (1 μM) were used to inhibit complex I and III respectively. SHAM (2 mM) was used to inhibit AOX activity (n = 4 biologically independent experiments). c, 143B-CYTB-Δ-SDHA_KO2-AOX-RFP and 143B-CYTB-Δ-SDHA_KO2-AOX-cDNA SDHA cells were grown in the presence or absence of methyl pyruvate and cell number was assessed after 72 h (n = 5 biologically independent experiments). d, e, Average tumour volume (d) and tumour mass (e) of xenografts from 143B-CYTB-Δ-SDHA_KO2-AOX-RFP and 143B-CYTB-Δ-SDHA_KO2-AOX-cDNA SDHA cells (n = 8 mice per group from two independent cohorts). Data are mean ± s.e.m. (b–e). *P < 0.05, **P < 0.01, two-tailed t-tests (e) or two-way ANOVA (c, d) with a Bonferroni test for multiple comparisons (exact P values are in the Source Data). For gel source data, see Supplementary Fig. 6.
Supplementary information
Supplementary Information
This file contains Supplementary Figures 1–6 (source data for western blots), Supplementary Figure 7 (the gating strategy for T-ALL experiments) and Supplementary Table 1 (the sequences of the sgRNAs used in this study).
Source data
Rights and permissions
About this article
Cite this article
Martínez-Reyes, I., Cardona, L.R., Kong, H. et al. Mitochondrial ubiquinol oxidation is necessary for tumour growth. Nature 585, 288–292 (2020). https://doi.org/10.1038/s41586-020-2475-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-020-2475-6
This article is cited by
-
Manipulating mitochondrial electron flow: a novel approach to enhance tumor immunogenicity
Molecular Biomedicine (2024)
-
Selenium reduction of ubiquinone via SQOR suppresses ferroptosis
Nature Metabolism (2024)
-
CIP2A induces PKM2 tetramer formation and oxidative phosphorylation in non-small cell lung cancer
Cell Discovery (2024)
-
A break in mitochondrial endosymbiosis as a basis for inflammatory diseases
Nature (2024)
-
Energy metabolism as the hub of advanced non-small cell lung cancer management: a comprehensive view in the framework of predictive, preventive, and personalized medicine
EPMA Journal (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.