Abstract
In metazoans, the secreted proteome participates in intercellular signalling and innate immunity, and builds the extracellular matrix scaffold around cells. Compared with the relatively constant intracellular environment, conditions for proteins in the extracellular space are harsher, and low concentrations of ATP prevent the activity of intracellular components of the protein quality-control machinery. Until now, only a few bona fide extracellular chaperones and proteases have been shown to limit the aggregation of extracellular proteins1,2,3,4,5. Here we performed a systematic analysis of the extracellular proteostasis network in Caenorhabditis elegans with an RNA interference screen that targets genes that encode the secreted proteome. We discovered 57 regulators of extracellular protein aggregation, including several proteins related to innate immunity. Because intracellular proteostasis is upregulated in response to pathogens6,7,8,9, we investigated whether pathogens also stimulate extracellular proteostasis. Using a pore-forming toxin to mimic a pathogenic attack, we found that C. elegans responded by increasing the expression of components of extracellular proteostasis and by limiting aggregation of extracellular proteins. The activation of extracellular proteostasis was dependent on stress-activated MAP kinase signalling. Notably, the overexpression of components of extracellular proteostasis delayed ageing and rendered worms resistant to intoxication. We propose that enhanced extracellular proteostasis contributes to systemic host defence by maintaining a functional secreted proteome and avoiding proteotoxicity.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Data availability
All relevant data are available and/or included with the manuscript or its Supplementary Information. RNA-sequencing data have been uploaded to the European Nucleotide Archive under the study accession PRJEB36386. Source data are provided with this paper.
Code availability
The source code for the bioinformatics analysis of homologues is available at https://github.com/Ashafix/C_Elegans_Homologs.
References
Hoshino, A. et al. A novel function for proSAAS as an amyloid anti-aggregant in Alzheimer’s disease. J. Neurochem. 128, 419–430 (2014).
Helwig, M. et al. The neuroendocrine protein 7B2 suppresses the aggregation of neurodegenerative disease-related proteins. J. Biol. Chem. 288, 1114–1124 (2013).
Wyatt, A. R., Yerbury, J. J., Ecroyd, H. & Wilson, M. R. Extracellular chaperones and proteostasis. Annu. Rev. Biochem. 82, 295–322 (2013).
Genereux, J. C. et al. Unfolded protein response-induced ERdj3 secretion links ER stress to extracellular proteostasis. EMBO J. 34, 4–19 (2015).
Tucker, H. M. et al. The plasmin system is induced by and degrades amyloid-beta aggregates. J. Neurosci. 20, 3937–3946 (2000).
Ermolaeva, M. A. et al. DNA damage in germ cells induces an innate immune response that triggers systemic stress resistance. Nature 501, 416–420 (2013).
Pellegrino, M. W. et al. Mitochondrial UPR-regulated innate immunity provides resistance to pathogen infection. Nature 516, 414–417 (2014).
Reddy, K. C. et al. An intracellular pathogen response pathway promotes proteostasis in C. elegans. Curr. Biol. 27, 3544–3553 (2017).
Richardson, C. E., Kooistra, T. & Kim, D. H. An essential role for XBP-1 in host protection against immune activation in C. elegans. Nature 463, 1092–1095 (2010).
Klaips, C. L., Jayaraj, G. G. & Hartl, F. U. Pathways of cellular proteostasis in aging and disease. J. Cell Biol. 217, 51–63 (2018).
Altun, Z. F. & Hall, D. H. Pericellular Structures https://www.wormatlas.org/hermaphrodite/pericellular/Periframeset.html (2009).
Fares, H. & Greenwald, I. Genetic analysis of endocytosis in Caenorhabditis elegans: coelomocyte uptake defective mutants. Genetics 159, 133–145 (2001).
David, D. C. et al. Widespread protein aggregation as an inherent part of aging in C. elegans. PLoS Biol. 8, e1000450 (2010).
Ruskamo, S. et al. Molecular mechanisms of Charcot-Marie-Tooth neuropathy linked to mutations in human myelin protein P2. Sci. Rep. 7, 6510 (2017).
Evans, E. A., Kawli, T. & Tan, M. W. Pseudomonas aeruginosa suppresses host immunity by activating the DAF-2 insulin-like signaling pathway in Caenorhabditis elegans. PLoS Pathog. 4, e1000175 (2008).
Bischof, L. J. et al. Activation of the unfolded protein response is required for defenses against bacterial pore-forming toxin in vivo. PLoS Pathog. 4, e1000176 (2008).
Wei, J. Z. et al. Bacillus thuringiensis crystal proteins that target nematodes. Proc. Natl Acad. Sci. USA 100, 2760–2765 (2003).
Kao, C. Y. et al. Global functional analyses of cellular responses to pore-forming toxins. PLoS Pathog. 7, e1001314 (2011).
Mizuno, T. et al. The Caenorhabditis elegans MAPK phosphatase VHP-1 mediates a novel JNK-like signaling pathway in stress response. EMBO J. 23, 2226–2234 (2004).
Kim, D. H. et al. Integration of Caenorhabditis elegans MAPK pathways mediating immunity and stress resistance by MEK-1 MAPK kinase and VHP-1 MAPK phosphatase. Proc. Natl Acad. Sci. USA 101, 10990–10994 (2004).
Huffman, D. L. et al. Mitogen-activated protein kinase pathways defend against bacterial pore-forming toxins. Proc. Natl Acad. Sci. USA 101, 10995–11000 (2004).
Melo, J. A. & Ruvkun, G. Inactivation of conserved C. elegans genes engages pathogen- and xenobiotic-associated defenses. Cell 149, 452–466 (2012).
Taylor, R. C. & Dillin, A. XBP-1 is a cell-nonautonomous regulator of stress resistance and longevity. Cell 153, 1435–1447 (2013).
Wan, L. et al. Bacillus thuringiensis targets the host intestinal epithelial junctions for successful infection of Caenorhabditis elegans. Environ. Microbiol. 21, 1086–1098 (2019).
Khoonsari, P. E. et al. Analysis of the cerebrospinal fluid proteome in Alzheimer’s disease. PLoS One 11, e0150672 (2016).
Lambert, J. C. et al. Genome-wide association study identifies variants at CLU and CR1 associated with Alzheimer’s disease. Nat. Genet. 41, 1094–1099 (2009).
Harold, D. et al. Genome-wide association study identifies variants at CLU and PICALM associated with Alzheimer’s disease. Nat. Genet. 41, 1088–1093 (2009).
Yeh, F. L., Wang, Y., Tom, I., Gonzalez, L. C. & Sheng, M. TREM2 binds to apolipoproteins, including APOE and CLU/APOJ, and thereby facilitates uptake of amyloid-beta by microglia. Neuron 91, 328–340 (2016).
Jonsson, T. et al. Variant of TREM2 associated with the risk of Alzheimer’s disease. N. Engl. J. Med. 368, 107–116 (2013).
Guerreiro, R. et al. TREM2 variants in Alzheimer’s disease. N. Engl. J. Med. 368, 117–127 (2013).
Petersen, T. N., Brunak, S., von Heijne, G. & Nielsen, H. SignalP 4.0: discriminating signal peptides from transmembrane regions. Nat. Methods 8, 785–786 (2011).
Krogh, A., Larsson, B., von Heijne, G. & Sonnhammer, E. L. Predicting transmembrane protein topology with a hidden Markov model: application to complete genomes. J. Mol. Biol. 305, 567–580 (2001).
Hansen, M., Hsu, A. L., Dillin, A. & Kenyon, C. New genes tied to endocrine, metabolic, and dietary regulation of lifespan from a Caenorhabditis elegans genomic RNAi screen. PLoS Genet. 1, 119–128 (2005).
Groh, N. et al. Methods to study changes in inherent protein aggregation with age in Caenorhabditis elegans. J. Vis. Exp. 129, 56464 (2017).
Shen, X., Ellis, R. E., Sakaki, K. & Kaufman, R. J. Genetic interactions due to constitutive and inducible gene regulation mediated by the unfolded protein response in C. elegans. PLoS Genet. 1, e37 (2005).
Huang, C. et al. Intrinsically aggregation-prone proteins form amyloid-like aggregates and contribute to tissue aging in Caenorhabditis elegans. eLife 8, e43059 (2019).
Bischof, L. J., Huffman, D. L. & Aroian, R. V. Assays for toxicity studies in C. elegans with Bt crystal proteins. Methods Mol. Biol. 351, 139–154 (2006).
Kim, D. et al. TopHat2: accurate alignment of transcriptomes in the presence of insertions, deletions and gene fusions. Genome Biol. 14, R36 (2013).
Trapnell, C. et al. Transcript assembly and quantification by RNA-Seq reveals unannotated transcripts and isoform switching during cell differentiation. Nat. Biotechnol. 28, 511–515 (2010).
Johnson, L. S., Eddy, S. R. & Portugaly, E. Hidden Markov model speed heuristic and iterative HMM search procedure. BMC Bioinformatics 11, 431 (2010).
Hawrylycz, M. J. et al. An anatomically comprehensive atlas of the adult human brain transcriptome. Nature 489, 391–399 (2012).
Piñero, J. et al. DisGeNET: a discovery platform for the dynamical exploration of human diseases and their genes. Database (Oxford) 2015, bav028 (2015).
Kibbe, W. A. et al. Disease Ontology 2015 update: an expanded and updated database of human diseases for linking biomedical knowledge through disease data. Nucleic Acids Res. 43, D1071–D1078 (2015).
Kelley, L. A., Mezulis, S., Yates, C. M., Wass, M. N. & Sternberg, M. J. The Phyre2 web portal for protein modeling, prediction and analysis. Nat. Protoc. 10, 845–858 (2015).
Venables, W. N. & Ripley, B. D. Modern Applied Statistics with S (Springer-Verlag New York, 2002).
Han, S. K. et al. OASIS 2: online application for survival analysis 2 with features for the analysis of maximal lifespan and healthspan in aging research. Oncotarget 7, 56147–56152 (2016).
Yang, W., Dierking, K. & Schulenburg, H. WormExp: a web-based application for a Caenorhabditis elegans-specific gene expression enrichment analysis. Bioinformatics 32, 943–945 (2016).
Acknowledgements
We thank M. Schölling for help with statistics based on the ordinal logistic regression model. We thank C. Kenyon for providing some C. elegans strains and J. Fares for providing NP717. JM103 E. coli strains were provided by R. V. Aroian. Some C. elegans strains and Microbacterium nematophilum were provided by the C. elegans Genetics Center, which is funded by NIH Office of Research Infrastructure Programs (P40 OD010440), the International C. elegans Gene Knockout Consortium and the National BioResource Project (NBRP). This work was supported by funding from the DZNE (D.C.D.), Max Planck Society (R.J.S.), the Deutsche Forschungsgemeinschaft (DFG, German Research Foundation) (SFB 1035 to M. H., DA 1906/4-1 to D.C.D.) and a Marie Curie International Reintegration Grant (322120 to D.C.D.).
Author information
Authors and Affiliations
Contributions
I.G., A.S, M.H., C.H. and D.C.D. designed the experiments and interpreted data. I.G., A.S., R.J. and D.C.D. performed cloning and transgenic animal generation. R.J., A.S. and D.C.D. performed insolubility analysis of LBP-2 and R.J. carried out analysis of total LBP-2 levels with RNAi treatment. A.S. performed RNAi screen, LBP-2 aggregate counting and endocytosis evaluation. I.G. conducted LBP-2 aggregate counting, lifespan and survival analysis. A.S and I.G. independently confirmed the effects of ECR knockdown by RNAi on LBP-2 aggregation. J.L.B. and I.G. carried out aggregate counting of LYS-7. I.G. and D.C.D. performed image acquisition and analysis. M.H. conducted co-purification and western blots to detect interaction of LBP-2 and C36C5.5. I.G. collected animals for RT–qPCR and S.A., R.J. and C.H. performed RT–qPCR. W.R. carried out RNA sequencing. C.R. performed RNA-sequencing analysis. R.J.S. provided advice on RNA sequencing. M.P. performed homology analysis and structure prediction. D.C.D. and I.G. wrote the manuscript with contributions from R.J.S. and input from the other authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature thanks Carmen Nussbaum and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Absence of coelomocytes causes LBP-2 to accumulate in pseudocoelom together with secreted GFP.
a, LBP-2::tagRFP expression pattern in body-wall muscles (day 2, n = 14 worms). b, c, Secreted LBP-2::tagRFP and secreted GFP colocalize in day 2 animals with coelomocytes (n = 16 worms) (b) and accumulate in animals without coelomocytes (n = 6) (c). Asterisk indicates pharyngeal GFP reporter in animal without coelomocytes. b, c, Secreted GFP exposure, 5 times shorter in c versus b, secreted LBP-2::tagRFP, identical exposure. d, LBP-2::tagRFP puncta in tail region (day 2, n = 15 worms; day 8, n = 14). Maximum projection. Scale bar, 20 μm. e, Quantification of LBP-2::tagRFP aggregation with age in the tail (n = 2 independent experiments). P values determined by two-sided Fisher’s exact test (day 8) and chi-square test (day 12). f, LBP-2 aggregates are separate from neurons (n = 27 worms). Scale bar, 20 μm. Single plane. g, Total protein stain of blot in Fig. 1j (n = 2 independent experiments) with fold changes quantified per fraction relative to levels in day 2. For blot source image, see Supplementary Fig. 1.
Extended Data Fig. 2 RNAi targeting ECRs reproducibly enhances LBP-2 aggregation.
a, Pie charts depict results from RNAi screen targeting genes encoding secreted factors and their effect on LBP-2 aggregation. b, Quantification of LBP-2::tagRFP aggregation with RNAi targeting top 13 candidates from egg in non-sterile background at day 4 (n = 1 independent experiment). Ctrl (−) is empty vector; ctrl (+) is rme-1 RNAi. P values determined by ordinal logistic regression and for lys-3 and tag-196 RNAi treatment by two-sided Fisher’s exact test. c, Maximum projection of head region of day 4 transgenics overexpressing LBP-2::tagRFP subjected to RNAi targeting a subset of ECRs (empty vector n = 8 worms, clec-1 n = 2, F56B6.6 n = 5, lys-3 n = 7, C36C5.5 n = 5). Scale bar, 20 μm. Laser intensity 8%. d, e, Downregulation of ECRs by RNAi does not change total levels of LBP-2::tagRFP (n = 2 independent experiments). Western blot detection of LBP-2::tagRFP in total fraction at day 4, 25 °C. Control 1, 2 and 3 are empty vector. Fold changes (in d) are normalized to total protein levels quantified by protein staining (e). For blot source images, see Supplementary Fig. 1.
Extended Data Fig. 3 ECRs regulate LYS-7 aggregation.
a, Quantification of animals without GFP-labelled coelomocytes in transgenic animals expressing secreted GFP (GS1912) treated with RNAi targeting 13 top candidates (n = 2 independent experiments). Ctrl (−) denotes empty vector; ctrl (+) denotes dyn-1 RNAi. b, LYS-7::tagRFP in young whole animal. Arrowheads indicate localization in coelomocytes, and asterisk indicates localization in anterior intestinal cells (top panel, n = 10 worms). LYS-7::tagRFP diffuse localization in head region of young animal (bottom left panel, n = 19) and puncta localized in head region of aged animal (bottom right panel, n = 14). Laser intensity 15%, maximum projection. Scale bar, 20 μm. c, Quantification of LYS-7::tagRFP aggregation with age (n = 2 independent experiments). d, Effect on LYS-7::tagRFP aggregation at day 6, 25 °C, with RNAi targeting top 13 candidates (n = 4 independent experiments). e, LYS-7 aggregation is reduced by CLEC-1 overexpression quantified at day 6 (n = 2 independent experiments). Ctrl indicates CLEC-1 non-overexpressing animals. P values determined by two-sided Fisher’s exact test with Benjamini–Hochberg correction.
Extended Data Fig. 4 ECR overexpression effectively prevents extracellular protein aggregation.
a, Overexpression of ECRs reduces LBP-2 aggregation (ssGFP n = 13 worms, LYS-3 n = 27, F56B6.6 n = 21, CLEC-1 n = 19, C36C5.5 n = 24). Scale bar, 20 μm. Maximum projection, laser intensity 8%. b, Secreted GFP does not accumulate in LBP-2 aggregates (n = 21 worms). Scale bar, 20 μm. Single plane. c, Coomassie staining of co-purification of C36C5.5 with LBP-2 (n = 2 independent experiments). Open arrow indicates LBP-2::tagRFP; closed arrow denotes C36C5.5::mVenus::histag. Three independent co-purification experiments with the same starting material are shown (elution 1–3).
Extended Data Fig. 5 Extracellular proteostasis influences ageing and is differentially regulated during ageing.
a, F56B6.6-overexpressing and LYS-3-overexpressing animals are long-lived compared with non-overexpressing siblings (n = 2 independent experiments). b, Animals lacking coelomocytes are short-lived compared with control animals (n = 3 independent experiments). c, Secreted GFP (n = 2 biologically independent samples) and LBP-2 overexpression (n = 1 independent experiment) do not influence lifespan. P values were determined by log-rank test (a–c). For detailed values, see Extended Data Table 1. d, Changes in expression levels of ECR candidates with age (day 8 versus day 2 at 25 °C). Left, expression level in LBP-2 overexpressing sterile animals. Right, expression level in wild-type sterile animals. Data are means ± s.e.m. of n = 4 biologically independent samples. P values were determined by two-sided unpaired t-test with Welch’s correction.
Extended Data Fig. 6 Impairing extracellular proteostasis accelerates intoxication-related mortality.
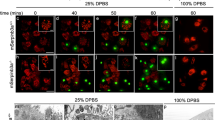
a, Expression level of LBP-2::tagRFP is not reduced after 3 h exposure to 100% Cry5B. Data are mean ± s.e.m. of n = 4 biologically independent samples. b, Quantification of LBP-2::tagRFP aggregation upon exposure to Microbacterium nematophilum and Bacillus atrophaeus at day 6 (n = 2 independent experiments). c, Expression level of four selected ECR candidates in unchallenged conditions with control pQE9 empty vector in kgb-1(km21) mutant versus wild-type background. d, Expression levels of eight selected ECRs with vhp-1 versus control RNAi. Data are mean ± s.e.m. of n = 3 (c) and n = 4 (d) biologically independent samples. e, Survival analysis of LBP-2::tagRFP transgenics on jnk-1(gk7) versus wild-type background subjected to 50% Cry5B (n = 2 independent experiments). f, Survival analysis of LBP-2::tagRFP transgenics subjected to 25% Cry5B with RNAi targeting selected ECRs (n = 2 independent experiments). g, Survival analysis of secreted GFP transgenics with versus without coelomocytes, subjected to 50% Cry5B (n = 4 independent experiments). h, Survival analysis of LBP-2::tagRFP transgenics versus N2 wild-type subjected to 50% Cry5B (n = 1 independent experiment). i, Survival analysis of LBP-2::tagRFP transgenics with and without secreted GFP overexpression subjected to 50% Cry5B (n = 2 biologically independent samples). j, Increased LBP-2::tagRFP aggregation at day 4 of adulthood during exposure to 25% Cry5B and treatment with RNAi targeting selected ECRs compared to empty vector (n = 2 independent experiments). P values determined by two-sided unpaired t-test with Welch’s correction (a, c, d), chi-square test (b) and two-sided Fisher’s exact test with Benjamini–Hochberg correction (j), and log-rank test with Bonferroni correction (e–i). For detailed values see Extended Data Tables 2, 3.
Extended Data Fig. 7 ECR overexpression does not induce a global immune response.
Heat map of z-score-normalized expression data (rows correspond to genes, columns indicate samples). The dendrogram on the top shows that RNA-seq samples cluster by treatment (Cry5B versus control).
Supplementary information
Supplementary Information
This file contains Supplementary Table 4 (List of 57 extracellular regulators of protein aggregation and predicted human orthologs); Supplementary Table 5 (the transgenic strains and alleles used); Supplementary Table 7 (Primer sequences for qRT-PCR); and Supplementary Figure 1 (Uncropped scans with size marker indications).
Supplementary Table 1
| Bioinformatic functional analysis of ECR candidates and their aggregation score.
Supplementary Table 2
| Enrichment analysis of ECRs among genes regulated by microbes. Excel list of ECRs significantly enriched in genes differentially regulated in response to microbes. Analysis was performed with WormExp (one-sided Fisher’s exact test).
Supplementary Table 3
| RNA sequencing of ECR overexpressing transgenics. Excel file with genes differentially regulated in at least two ECR overexpressing transgenics compared to control in Cry5B challenged conditions (sheet 1) or un-challenged conditions (sheet 2) (see Methods for statistics), enrichment analysis of genes upregulated during Cry5B related to microbes (sheet 3, WormExp one-sided Fisher’s exact test).
Supplementary Table 6
| List of predicted secreted coding genes tested by RNAi. Excel file list of genes with predicted signal peptide and without transmembrane domains knocked-down by RNAi to investigate their effect on LBP-2 aggregation.
Supplementary Video 1
| LBP-2 aggregates localize outside body-wall muscles. Confocal z-stack of C. elegans head with LBP-2 aggregates (magenta) and F-actin stained with phalloidin (green) at day 6 of adulthood. Representative video of n = 16 worms.
Rights and permissions
About this article
Cite this article
Gallotta, I., Sandhu, A., Peters, M. et al. Extracellular proteostasis prevents aggregation during pathogenic attack. Nature 584, 410–414 (2020). https://doi.org/10.1038/s41586-020-2461-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-020-2461-z
This article is cited by
-
Small-sample learning reveals propionylation in determining global protein homeostasis
Nature Communications (2023)
-
Extracellular protein homeostasis in neurodegenerative diseases
Nature Reviews Neurology (2023)
-
Protein homeostasis from the outside in
Nature Cell Biology (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.