Abstract
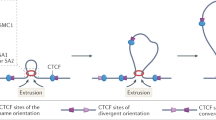
Nuclear processes, such as V(D)J recombination, are orchestrated by the three-dimensional organization of chromosomes at multiple levels, including compartments1 and topologically associated domains (TADs)2,3 consisting of chromatin loops4. TADs are formed by chromatin-loop extrusion5,6,7, which depends on the loop-extrusion function of the ring-shaped cohesin complex8,9,10,11,12. Conversely, the cohesin-release factor Wapl13,14 restricts loop extension10,15. The generation of a diverse antibody repertoire, providing humoral immunity to pathogens, requires the participation of all V genes in V(D)J recombination16, which depends on contraction of the 2.8-Mb-long immunoglobulin heavy chain (Igh) locus by Pax517,18. However, how Pax5 controls Igh contraction in pro-B cells remains unknown. Here we demonstrate that locus contraction is caused by loop extrusion across the entire Igh locus. Notably, the expression of Wapl is repressed by Pax5 specifically in pro-B and pre-B cells, facilitating extended loop extrusion by increasing the residence time of cohesin on chromatin. Pax5 mediates the transcriptional repression of Wapl through a single Pax5-binding site by recruiting the polycomb repressive complex 2 to induce bivalent chromatin at the Wapl promoter. Reduced Wapl expression causes global alterations in the chromosome architecture, indicating that the potential to recombine all V genes entails structural changes of the entire genome in pro-B cells.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
The RNA-seq, ChIP–seq, ATAC-seq, VDJ-seq, 3C-seq and Hi-C data reported in this study (Supplementary Table 5) are available at the Gene Expression Omnibus repository under the accession number GSE140975. Source data are provided with this paper.
References
Lieberman-Aiden, E. et al. Comprehensive mapping of long-range interactions reveals folding principles of the human genome. Science 326, 289–293 (2009).
Dixon, J. R. et al. Topological domains in mammalian genomes identified by analysis of chromatin interactions. Nature 485, 376–380 (2012).
Nora, E. P. et al. Spatial partitioning of the regulatory landscape of the X-inactivation centre. Nature 485, 381–385 (2012).
Rao, S. S. et al. A 3D map of the human genome at kilobase resolution reveals principles of chromatin looping. Cell 159, 1665–1680 (2014).
Nasmyth, K. Disseminating the genome: joining, resolving, and separating sister chromatids during mitosis and meiosis. Annu. Rev. Genet. 35, 673–745 (2001).
Sanborn, A. L. et al. Chromatin extrusion explains key features of loop and domain formation in wild-type and engineered genomes. Proc. Natl Acad. Sci. USA 112, E6456–E6465 (2015).
Fudenberg, G. et al. Formation of chromosomal domains by loop extrusion. Cell Rep. 15, 2038–2049 (2016).
Rao, S. S. P. et al. Cohesin loss eliminates all loop domains. Cell 171, 305–320.e24 (2017).
Schwarzer, W. et al. Two independent modes of chromatin organization revealed by cohesin removal. Nature 551, 51–56 (2017).
Wutz, G. et al. Topologically associating domains and chromatin loops depend on cohesin and are regulated by CTCF, WAPL, and PDS5 proteins. EMBO J. 36, 3573–3599 (2017).
Davidson, I. F. et al. DNA loop extrusion by human cohesin. Science 366, 1338–1345 (2019).
Kim, Y., Shi, Z., Zhang, H., Finkelstein, I. J. & Yu, H. Human cohesin compacts DNA by loop extrusion. Science 366, 1345–1349 (2019).
Kueng, S. et al. Wapl controls the dynamic association of cohesin with chromatin. Cell 127, 955–967 (2006).
Tedeschi, A. et al. Wapl is an essential regulator of chromatin structure and chromosome segregation. Nature 501, 564–568 (2013).
Haarhuis, J. H. I. et al. The cohesin release factor WAPL restricts chromatin loop extension. Cell 169, 693–707 (2017).
Alt, F. W., Zhang, Y., Meng, F.-L., Guo, C. & Schwer, B. Mechanisms of programmed DNA lesions and genomic instability in the immune system. Cell 152, 417–429 (2013).
Fuxa, M. et al. Pax5 induces V-to-DJ rearrangements and locus contraction of the immunoglobulin heavy-chain gene. Genes Dev. 18, 411–422 (2004).
Medvedovic, J. et al. Flexible long-range loops in the VH gene region of the Igh locus facilitate the generation of a diverse antibody repertoire. Immunity 39, 229–244 (2013).
Johnston, C. M., Wood, A. L., Bolland, D. J. & Corcoran, A. E. Complete sequence assembly and characterization of the C57BL/6 mouse Ig heavy chain V region. J. Immunol. 176, 4221–4234 (2006).
Proudhon, C., Hao, B., Raviram, R., Chaumeil, J. & Skok, J. A. Long-range regulation of V(D)J recombination. Adv. Immunol. 128, 123–182 (2015).
Jhunjhunwala, S. et al. The 3D structure of the immunoglobulin heavy-chain locus: implications for long-range genomic interactions. Cell 133, 265–279 (2008).
Zhang, Y. et al. The fundamental role of chromatin loop extrusion in physiological V(D)J recombination. Nature 573, 600–604 (2019).
Jain, S., Ba, Z., Zhang, Y., Dai, H. Q. & Alt, F. W. CTCF-binding elements mediate accessibility of RAG substrates during chromatin scanning. Cell 174, 102–116 (2018).
Hu, J. et al. Chromosomal loop domains direct the recombination of antigen receptor genes. Cell 163, 947–959 (2015).
Parelho, V. et al. Cohesins functionally associate with CTCF on mammalian chromosome arms. Cell 132, 422–433 (2008).
Wendt, K. S. et al. Cohesin mediates transcriptional insulation by CCCTC-binding factor. Nature 451, 796–801 (2008).
Chovanec, P. et al. Unbiased quantification of immunoglobulin diversity at the DNA level with VDJ-seq. Nat. Protoc. 13, 1232–1252 (2018).
Margueron, R. & Reinberg, D. The Polycomb complex PRC2 and its mark in life. Nature 469, 343–349 (2011).
Ji, Y. et al. The in vivo pattern of binding of RAG1 and RAG2 to antigen receptor loci. Cell 141, 419–431 (2010).
Durand, N. C. et al. Juicer provides a one-click system for analyzing loop-resolution Hi-C experiments. Cell Syst. 3, 95–98 (2016).
Urbánek, P., Wang, Z.-Q., Fetka, I., Wagner, E. F. & Busslinger, M. Complete block of early B cell differentiation and altered patterning of the posterior midbrain in mice lacking Pax5/BSAP. Cell 79, 901–912 (1994).
Horcher, M., Souabni, A. & Busslinger, M. Pax5/BSAP maintains the identity of B cells in late B lymphopoiesis. Immunity 14, 779–790 (2001).
McManus, S. et al. The transcription factor Pax5 regulates its target genes by recruiting chromatin-modifying proteins in committed B cells. EMBO J. 30, 2388–2404 (2011).
Fuxa, M. & Busslinger, M. Reporter gene insertions reveal a strictly B lymphoid-specific expression pattern of Pax5 in support of its B cell identity function. J. Immunol. 178, 3031–3037 (2007).
Shinkai, Y. et al. RAG-2-deficient mice lack mature lymphocytes owing to inability to initiate V(D)J rearrangement. Cell 68, 855–867 (1992).
Krimpenfort, P. et al. p15Ink4b is a critical tumour suppressor in the absence of p16Ink4a. Nature 448, 943–946 (2007).
Tallquist, M. D. & Soriano, P. Epiblast-restricted Cre expression in MORE mice: a tool to distinguish embryonic vs. extra-embryonic gene function. Genesis 26, 113–115 (2000).
McCormack, M. P., Forster, A., Drynan, L., Pannell, R. & Rabbitts, T. H. The LMO2 T-cell oncogene is activated via chromosomal translocations or retroviral insertion during gene therapy but has no mandatory role in normal T-cell development. Mol. Cell. Biol. 23, 9003–9013 (2003).
Hobeika, E. et al. Testing gene function early in the B cell lineage in mb1-cre mice. Proc. Natl Acad. Sci. USA 103, 13789–13794 (2006).
Seibler, J. et al. Rapid generation of inducible mouse mutants. Nucleic Acids Res. 31, e12 (2003).
de Boer, J. et al. Transgenic mice with hematopoietic and lymphoid specific expression of Cre. Eur. J. Immunol. 33, 314–325 (2003).
Rodríguez, C. I. et al. High-efficiency deleter mice show that FLPe is an alternative to Cre-loxP. Nat. Genet. 25, 139–140 (2000).
Anastassiadis, K. et al. Dre recombinase, like Cre, is a highly efficient site-specific recombinase in E. coli, mammalian cells and mice. Dis. Model. Mech. 2, 508–515 (2009).
Singla, V. et al. Floxin, a resource for genetically engineering mouse ESCs. Nat. Methods 7, 50–52 (2010).
Yang, H. et al. One-step generation of mice carrying reporter and conditional alleles by CRISPR/Cas-mediated genome engineering. Cell 154, 1370–1379 (2013).
Poser, I. et al. BAC TransgeneOmics: a high-throughput method for exploration of protein function in mammals. Nat. Methods 5, 409–415 (2008).
Adams, B. et al. Pax-5 encodes the transcription factor BSAP and is expressed in B lymphocytes, the developing CNS, and adult testis. Genes Dev. 6, 1589–1607 (1992).
Nutt, S. L., Urbánek, P., Rolink, A. & Busslinger, M. Essential functions of Pax5 (BSAP) in pro-B cell development: difference between fetal and adult B lymphopoiesis and reduced V-to-DJ recombination at the IgH locus. Genes Dev. 11, 476–491 (1997).
Minnich, M. et al. Multifunctional role of the transcription factor Blimp-1 in coordinating plasma cell differentiation. Nat. Immunol. 17, 331–343 (2016).
Holzmann, J. et al. Absolute quantification of cohesin, CTCF and their regulators in human cells. eLife 8, e46269 (2019).
Buenrostro, J. D., Giresi, P. G., Zaba, L. C., Chang, H. Y. & Greenleaf, W. J. Transposition of native chromatin for fast and sensitive epigenomic profiling of open chromatin, DNA-binding proteins and nucleosome position. Nat. Methods 10, 1213–1218 (2013).
Revilla-i-Domingo, R. et al. The B-cell identity factor Pax5 regulates distinct transcriptional programmes in early and late B lymphopoiesis. EMBO J. 31, 3130–3146 (2012).
Langmead, B. Aligning short sequencing reads with Bowtie. Curr. Protoc. Bioinformatics Chapter 11, Unit 11.7 (2010).
Zhang, Y. et al. Model-based analysis of ChIP–seq (MACS). Genome Biol. 9, R137 (2008).
Salmon-Divon, M., Dvinge, H., Tammoja, K. & Bertone, P. PeakAnalyzer: genome-wide annotation of chromatin binding and modification loci. BMC Bioinformatics 11, 415 (2010).
Slater, G. S. & Birney, E. Automated generation of heuristics for biological sequence comparison. BMC Bioinformatics 6, 31 (2005).
Machanick, P. & Bailey, T. L. MEME-ChIP: motif analysis of large DNA datasets. Bioinformatics 27, 1696–1697 (2011).
Liao, Y., Smyth, G. K. & Shi, W. featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30, 923–930 (2014).
Wagner, G. P., Kin, K. & Lynch, V. J. Measurement of mRNA abundance using RNA-seq data: RPKM measure is inconsistent among samples. Theory Biosci. 131, 281–285 (2012).
Davies, J. O. et al. Multiplexed analysis of chromosome conformation at vastly improved sensitivity. Nat. Methods 13, 74–80 (2016).
Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet J. 17, 10–12 (2011).
Thongjuea, S., Stadhouders, R., Grosveld, F. G., Soler, E. & Lenhard, B. r3Cseq: an R/Bioconductor package for the discovery of long-range genomic interactions from chromosome conformation capture and next-generation sequencing data. Nucleic Acids Res. 41, e132 (2013).
Wingett, S. et al. HiCUP: pipeline for mapping and processing Hi-C data. F1000Res. 4, 1310 (2015).
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Heinz, S. et al. Simple combinations of lineage-determining transcription factors prime cis-regulatory elements required for macrophage and B cell identities. Mol. Cell 38, 576–589 (2010).
Acknowledgements
We thank W. C. Skarnes for advice and help with the Floxin method, M. Leeb for assistance with ES cell derivation, E. Lieberman Aiden for discussion, R. Stocsits for advice with Hi-C analysis, K. Aumayr’s team for flow cytometric sorting, C. Theussl’s team for generating gene-modified mice and A. Sommer’s team at the Vienna BioCenter Core Facilities for Illumina sequencing. This research was supported by Boehringer Ingelheim, the European Research Council (ERC) under the European Union’s Horizon 2020 research and innovation programme (grant agreement 740349-PlasmaCellControl and 693949-CohesinMolMech), the Austrian Industrial Research Promotion Agency (Headquarter Grant FFG-852936), the Human Frontier Science Program grant RGP0057/2018 (to J.-M.P.) and a long-term fellowship from the Human Frontier Science Program LT001527/2017 (to K.N.).
Author information
Authors and Affiliations
Contributions
L.H. performed most of the experiments; A.E. generated and analysed the Eed mutant mouse; G.W. generated the Hi-C data; K.N. performed the iFRAP analyses; H.T. performed ATAC-seq analyses and contributed to 3C-seq experiments; D.K.-P. performed RT–qPCR analyses; K.S. generated the IghCBE/+ and IghCBE-inv/+ mice and inserted the VH8-8 gene into Igh∆890/+ ES cells; Q.S. generated the transgenic Smc3-gfp mouse and contributed to the generation of the Ighfl-890-fl/+ mouse; P.B. performed the PRC2-Pax5 recruitment experiments; M.J. performed most bioinformatic analyses (RNA-seq, VDJ-seq, 3C-seq and Hi-C data); M.F. analysed the Eed RNA-seq data; J.-M.P. provided advice on cohesin biology and the design of iFRAP and Hi-C experiments; L.H. and M.B. planned the project, designed the experiments and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature thanks Feilong Meng and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Generation and characterization of the Igh890-inv allele.
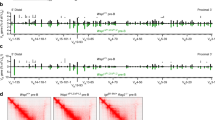
a, Orientation of the CTCF-binding sites in the Igh locus. The CTCF-binding pattern was determined by ChIP-seq of Rag2−/− pro-B cells. The locations of forward (red) and reverse (blue) CTCF motifs detected at the summit of the CTCF peaks are shown together with all predicted CTCF motifs, identified based on the forward and reverse consensus CTCF-binding motifs shown. The annotation of the C57BL/6 Igh locus indicates the distinct VH gene families (different colours) in the distal, middle and proximal VH gene regions19 and the 3′-proximal Igh domain containing the DH, JH and CH elements as well as the Eμ and 3′RR enhancers (red). b, Schematic diagram of the Ighifl-890-fl and Igh890-inv alleles. The indicated selection cassettes were used for introducing the upstream inverted lox71 site (ifl) and downstream loxP site (fl) by sequential ES cell targeting. c, Flow cytometric analysis of bone marrow cells from Igh890-inv/890-inv and Igh+/+ mice. The relative frequencies of the indicated cell types (defined in Methods) are shown as mean values with s.e.m. d, Flow cytometric determination of the ratio of immature IgMb (B6) to IgMa (129) B cells from Igh890-inv(B6)/+(129) and Igh+(B6)/+(129) mice, which were generated by crossing Igh890-inv/+ mice on the C57BL/6 (B6) background with Igh+/+ mice of the 129/Sv (129) strain. The rearranged Igh alleles of the C57BL/6 and 129/Sv strains give rise to expression of the IgMb and IgMa isotypes, respectively. e, VH gene expression from the Igh890-inv(B6) or Igh+(B6) allele in immature IgMb (B6) B cells sorted from Igh890-inv(B6)/+(129) or Igh+(B6)/+(129) mice, respectively. The RNA-seq profiles are shown with the Igh annotation (see a) and inverted region (bar). One of two experiments is shown. RPM, reads per million mapped sequence reads. f, g, VDJ-seq analysis of pro-B cells from the bone marrow of Igh890-inv/890-inv and Igh+/+ mice. f, The percentages of uniquely identified DJH and VDJH sequences are indicated. g, The relative usage of the distal VH genes at the Igh 5′ end is shown as mean percentage of all VDJH recombination events with s.e.m. A horizontal bar indicates the 5′ end of the inverted region. Statistical data are shown as mean value with s.e.m. and were analysed either by multiple t-tests (unpaired and two-tailed with Holm–Sidak correction; c, f) or by the Student’s t-test (unpaired and two-tailed; d). n, number of mice (c, d) or experiments (f, g). Each dot corresponds to one mouse.
Extended Data Fig. 2 Dependence of VH-DJH recombination on chromatin looping and VH gene orientation.
a, ChIP-seq analysis of CTCF binding in the distal 890-kb region in short-term-cultured pro-B cells from Igh890-inv/890-inv and Igh+/+ mice. Vertical bars indicated CTCF peaks defined by ‘peak calling’. The ChIP-seq data of the Igh890-inv allele were aligned on the wild-type Igh sequences. One of two experiments per genotype is shown. b, Igh positions of the 3C-seq viewpoints, which are referred to by the nearest VH gene, the hypersensitive site 5 (HS5) in the 3′CBE region or the CBE array insertion (Supplementary Table 1c). c, Interactions from the 5′ (VH1-86; left) and 3′ (HS5; right) viewpoints in short-term-cultured pro-B cells from Igh890-inv/890-invRag2−/− and Igh+/+Rag2−/− mice. The 3C-seq reads along the Igh locus (top two panels) are shown as RPM values with the respective Q values, which were calculated by the r3Cseq program based on two 3C-seq experiments per genotype. The bottom panel shows the fold-change of the RPM values for interaction differences of >2-fold (dashed line) between experimental (Exp) Igh890-inv/890-invRag2−/− and control (Ctrl) Igh+/+Rag2−/− pro-B cells. d, Schematic summarizing the loop formation at the Igh890-inv allele in pro-B cells. The CBEs with their orientation are indicated by red and blue arrowheads, VH genes by grey arrows and loops by arches. The new loop domain is indicated in red. e, Schematic of the Igh∆890, IghV8-8 and IghV8-8-inv alleles. The IghV8-8 and IghV8-8-inv alleles were generated by insertion of the VH8-8 gene with 500 bp of its 5′ and 3′ flanking sequences (lacking any CBE) in the forward or reverse orientation at the deletion point (117,126,667 / 116,237,220; mm9, Chr. 12) of the Igh∆890 allele (Methods). The distances from the 3′ end of the VH8-8 gene to the next CBEs and VH genes are indicated. The sequence of the inserted VH8-8 with its flanking DNA sequences is shown in Supplementary Table 1b. f, RNA-seq profile of the Igh locus in immature B cells from the bone marrow of IghV8-8/∆890, IghV8-8-inv/∆890 and Igh+/∆890 mice. The VH8-8 gene position and Igh annotation are indicated. The data of one of two RNA-seq experiments per genotype is shown. g, VH8-8 mRNA expression in immature B cells of the indicated genotypes is shown as mean TPM value. Each dot corresponds to one experiment.
Extended Data Fig. 3 Characterization of Igh alleles with inserted CBE arrays.
a, Insertion of an array of 20 functional CBEs in forward (red) or inverted (inv, blue) orientation at position 116,242,836 (mm9, Chr. 12) in the IghCBE or IghCBE-inv allele, respectively. The positions of the Igh CBEs used for the generation of the CBE arrays (Supplementary Table 1c) are indicated below a map of all CBEs. b, CTCF binding at the Igh locus, as determined by ChIP-seq analysis of short-term-cultured pro-B cells from the bone marrow of IghCBE-inv/CBE-inv and IghCBE/CBE mice. One of two experiments is shown. c, Flow cytometric determination of the ratio of immature IgMb (B6) to IgMa (129) B cells in the bone marrow of IghCBE-inv(B6)/+(129), IghCBE(B6)/+(129) and Igh+(B6)/+(129) mice (Extended Data Fig. 1d). The IgMb/IgMa ratios are shown relative to that of immature B cells of Igh+(B6)/+(129) mice (set to 1). Statistical data are shown as mean value with s.e.m. and were analysed by one-way ANOVA with Tukey’s post hoc test. d, VH gene expression from the IghCBE-inv(B6), IghCBE(B6) and Igh+(B6) alleles in immature IgMb (B6) B cells sorted from IghCBE-inv(B6)/+(129), IghCBE(B6)/+(129) and Igh+(B6)/+(129) mice, respectively (Extended Data Fig. 1d). The expression (TPM) value of each VH gene is shown as shared (grey) and unique expression of the IghCBE-inv(B6) (blue), IghCBE(B6) (green) and Igh+(B6) (black) alleles. Mean TPM values with s.e.m. were determined by the DESeq2 program and are based on two (IghCBE-inv(B6), IghCBE(B6)) and three (Igh+(B6)) RNA-seq experiments. The VH genes are aligned according to their Igh position. e, f, VDJ-seq analysis of ex vivo-sorted pro-B cells from the bone marrow of Igh+/+, IghCBE/CBE and IghCBE-inv/CBE-inv mice. e, The percentages of uniquely identified DJH and VDJH sequences are shown as mean percentage with s.e.m. f, Pairwise comparisons of VDJ-seq results obtained with Igh+/+ (black), IghCBE/CBE (green) and IghCBE-inv/CBE-inv (blue) pro-B cells. The relative usage of each VH gene was determined as percentage of all VDJH recombination events and is shown as mean percentage with s.e.m. n, number of mice. Each dot corresponds to one mouse (c, e).
Extended Data Fig. 4 New loop formation upon insertion of an inverted CBE array in the Igh locus.
a–d, 3C-seq analysis of interactions across the Igh locus from the 3′ (HS5; a, b) or 5′ (CBE; c, d) viewpoint in short-term-cultured IghCBE-inv/CBE-inv, IghCBE/CBE and Igh+/+ pro-B cells. a, c, Mapping of the 3C-seq reads as normalized counts across the Igh locus. One of two experiments per genotype is shown. b, d, Quantification of the interactions from the 3′ (HS5, b) or 5′(CBE, d) viewpoint. The 3C-seq reads in regions A and B were quantified as mean RPM value per region, based on two experiments per genotype. e, f, Interactions from the 3′ (HS5; e) or 5′ (CBE; f) viewpoint in Igh+/+, IghCBE/CBE and IghCBE-inv/CBE-inv pro-B cells. The 3C-seq reads along the Igh locus (top two panels) are shown as RPM values with the respective Q values, calculated by the r3Cseq program based on two experiments per genotype. Arrows highlight a strong interaction from the 3′ viewpoint (HS5) to a region immediately downstream of the inserted CBE array (e), and a bar denotes the 600-kb region exhibiting reduced interactions in IghCBE-inv/CBE-inv pro-B cells (e). The bottom panel shows the fold-change of the RPM values for interaction differences of >2-fold (dashed line; e) or >1.5-fold (f) between the experimental (Exp) and control (Ctrl) pro-B cells of the indicated genotypes. g, Schematic summarizing the loop formation at the IghCBE-inv allele in pro-B cells. The CBEs with their orientation are indicated by red and blue arrowheads, VH genes by grey arrows and loops by arches. The new loop domain is indicated in red. A thicker blue arch indicates the enhanced interaction from the 3′CBE to a region immediately downstream of the inserted CBE array.
Extended Data Fig. 5 Cohesin stabilization on chromatin by Pax5-dependent Wapl repression in pro-B cells.
a, Wapl mRNA expression in developing and mature B and T cells of wild-type mice. The indicated B and T cell types were sorted from the bone marrow (pro-B, pre-B cells), thymus (DN, DP, CD4+, CD8+ T cells) or spleen (mature B, CD4+ and CD8+ T cells). The Wapl mRNA expression of the different cell types was determined by RT–qPCR analysis relative to Tbp expression and is shown relative to that of pro-B cells (set to 1). The expression data are shown as mean values with s.e.m., based on four RT–qPCR experiments per cell type. b, Wapl protein expression in ex vivo-sorted Rag2−/− pro-B cells from the bone marrow and wild-type mature B cells from the spleen and lymph nodes, as determined by immunoblotting of threefold serially diluted whole-cell extracts with antibodies detecting Wapl or TBP. One of two experiments is shown with marker proteins (kilodaltons). c, B cell development in the bone marrow of Smc3-gfp transgenic (green) and wild-type (black) mice, as determined by flow cytometry. The relative frequencies of the indicated cell types are shown as mean values with s.e.m. (left). GFP expression in selected cell types is shown (right). d, Interaction of the Smc3–GFP protein with other cohesin subunits in short-term-cultured Smc3-gfp pro-B cells. Endogenous Smc1, Scc1 (Rad21), Wapl, Pds5a and Pds5b proteins were co-precipitated with Smc3–GFP from whole-cell extracts of Smc3-gfp or wild-type pro-B cells with an anti-GFP antibody. The input (1/10) and protein precipitate were analysed by immunoblotting with antibodies detecting the indicated cohesin proteins. Smc3ac, acetylated Smc3. One experiment was performed. e, Images of one of 16 or 24 iFRAP experiments performed with Smc3-gfp pro-B or mature B cells, respectively. The fluorescence intensities of SiR-Hoechst (pink) and GFP (green) are false-coloured. Scale bar, 10 μm. f, Identification of open chromatin regions at the Wapl locus by ATAC-seq of the indicated progenitor and B cell types. MPPs, ALPs, pro-B and mature B cells were sorted from wild-type bone marrow or lymph nodes (mature B). Activated B cells and plasmablasts were generated by in vitro LPS stimulation of mature B cells. hCD2− (Pax5−) and hCD2+ (Pax5+) BLPs were sorted from bone marrow of Pax5ihCd2/ihCd2 mice, and Pax5∆/∆ progenitors from Vav-Cre Pax5fl/flRag2−/− mice. n, number of mice. Each dot corresponds to one mouse (a, c). The different cell types are defined in Methods.
Extended Data Fig. 6 Generation and analysis of Wapl mutant mice.
a, Mutations introduced at the Pax5-binding sites P1 and P2 in the Wapl∆P1,2, Wapl∆P1 and Wapl∆P2 alleles by CRISPR–Cas9-mediated mutagenesis. The consensus Pax5 motif and the extent of the deletions (red) are indicated. b, Loss of Pax5 binding at the mutated P1 site in Wapl∆P1/∆P1 pro-B cells. One of two ChIP-seq experiments is shown. c. Wapl mRNA expression in short-term-cultured Wapl∆P1,2/∆P1,2 and Wapl+/+ pro-B cells is shown as mean TPM value with s.e.m. n, number of RNA-seq experiments. d, Wapl protein expression in short-term-cultured Wapl∆P1,2/∆P1,2Rag2−/− and Wapl+/+Rag2−/− pro-B cells was determined by immunoblotting of twofold serially diluted whole-cell extracts with antibodies detecting Wapl or TBP. One of two experiments is shown with marker proteins (kilodaltons). e, Wapl mRNA expression in ex vivo-sorted Wapl∆P1,2/∆P1,2 (blue), Wapl∆P1,2/+ (light blue) and Wapl+/+ (black) pro-B and pre-B cells, Pax5∆/∆ progenitors (green; Vav-Cre Pax5fl/fl) as well as Wapl∆P1,2/∆P1,2 and Wapl+/+ T cell subsets from the thymus and spleen, as determined by RT–qPCR analysis relative to Tbp expression. The Wapl expression of each cell type is indicated relative to that of the Wapl+/+ cells (set to 1) and is shown as mean value with s.e.m., based on 2–4 independent RT–qPCR experiments for each cell type and genotype. Each dot (c, e) corresponds to one mouse. The different cell types are defined in Methods.
Extended Data Fig. 7 B cell development in Wapl mutant mice.
a, b, Wapl expression in ex vivo-sorted Wapl∆P1/∆P1 (blue), Wapl∆P1/+ (light blue) and Wapl+/+ (black) pro-B and pre-B cells (a) as well as in Wapl∆P2/∆P2 (blue) and Wapl+/+ (black) pro-B cells (b), as determined by RT–qPCR analysis relative to Tbp expression. The Wapl expression of each cell type is shown as mean value with s.e.m. relative to that of Wapl+/+ cells (set to 1). Each dot corresponds to one RT–qPCR experiment performed with sorted cells from one mouse. c, d, Flow cytometric analysis of bone marrow from Wapl∆P2/∆P2 (blue), Wapl∆P2/+ (light blue) and Wapl+/+ (black) mice (c). The frequencies of the indicated cell types are shown as mean values with s.e.m. (d). e, Flow cytometric analysis of bone marrow from Cd79acre/+Waplfl/fl and Cd79acre/+Wapl+/+ mice. f, g, Frequencies of total B, pro-B and pre-B cells are shown for Cd79acre/+Waplfl/fl, Cd79acre/+Waplfl/+ and control mice (f) as well as for Rag1cre/+Waplfl/fl, Rag1cre/+Waplfl/+ and control mice (g). The control mice in f were 3 × Cd79acre/+Wapl+/+ and 5 × Waplfl/+ mice, and the control mice in g were 2 × Rag1cre/+Wapl+/+, 2 × Waplfl/fl, 1 × Waplfl/+ and 1 × Wapl+/+. Statistical data (a, b, d, f, g) are shown as mean values with s.e.m. and were analysed by multiple t-tests (unpaired and two-tailed with Holm–Sidak correction). One of 5 (c) or 4 (e) experiments is shown. n, number of mice. Each dot corresponds to one mouse.
Extended Data Fig. 8 VH–DJH recombination and looping in Wapl mutant mice.
a, VDJ-seq analysis of Igh rearrangements in Wapl∆P1,2/∆P1,2, Wapl∆P1,2/+, Wapl∆P1/∆P1, Wapl∆P2/∆P2 and Wapl+/+ pro-B cells as well as in Pax5∆/∆ progenitors (Vav-Cre Pax5fl/fl), which were performed in 4 different experimental series. The percentages of uniquely identified DJH and VDJH sequences are shown as mean values with s.e.m. and were analysed by multiple t-tests (unpaired and two-tailed with Holm–Sidak correction). b, VH gene recombination in Wapl+/+ and mutant pro-B cells, as determined by pairwise VDJ-seq experiments. The VDJ-seq data of Wapl+/+ and mutant pro-B cells are shown in the top and bottom part, respectively. The VH gene usage is shown as mean percentage of all DJH and VDJH rearrangement events with s.e.m. c, Wapl mRNA expression in in vitro cultured v-Abl transformed pro-B cells of the wild-type (light grey) or Rag2−/− (grey) genotype as well as in ex vivo-sorted Wapl∆P1,2/∆P1,2 (blue) and wild-type Wapl+/+ (black) pro-B cells. The Wapl expression was determined by RT–qPCR analysis relative to the Tbp expression and is shown a mean value relative to that of the Wapl+/+ pro-B cells (set to 1). d, 3C-seq analysis of interactions across the Igh locus from the 5′ viewpoints (VH1-83/81) in short-term-cultured Wapl∆P1,2/∆P1,2Rag2−/− and Wapl+/+Rag2−/− pro-B cells. The 3C-seq reads are mapped as normalized counts. One of two experiments is shown. n, number of experiments. Each dot corresponds to one mouse.
Extended Data Fig. 9 Generation and analysis of conditional Eed mutant mice.
a, Generation of a floxed (fl) Eed allele. The Eedfl-Neo allele was generated by replacing exon 6 of the Eed gene with the following sequences in the 5′ to 3′ direction: (i) a loxP-flanked Eed cDNA fragment (from exon 6 to the 5′ part of the 3′UTR) linked to six copies of the SV40 polyadenylation (pA) region, (ii) a DNA fragment containing the 3′ splice site and 5′ sequences of exon 6 fused in-frame to the coding sequence of gfp followed by an SV40 pA sequence and (iii) a frt-flanked DNA fragment containing the mouse phosphoglycerate kinase 1 (Pgk1) promoter linked to the neomycin (Neo) resistance gene. Brackets indicate the two homology regions used for ES cell recombination. The frt (blue) and loxP (red) sites (arrowheads) of the Eedfl-Neo allele were used to generate the Eedfl and Eed∆ alleles by sequential Flpe- and Cre-mediated deletion in vivo. The six SV40 pA sequences downstream of the last Eed exon prevented RNA splicing to the gfp exon before Cre-mediated deletion, as demonstrated by flow cytometry (e, right). A genotyping gel image of Eed+/+, Eedfl/+, and Eedfl/fl mice is shown to the right. b, Selective loss of H3K27me3 at the Wapl promoter in Wapl∆P1/∆P1 pro-B cells (light blue) compared to Wapl+/+ pro-B cells (black). The H3K27me3 signal is precipitously lost 220 bp upstream of the P1 site. One of two ChIP-seq experiments is shown. c, Loss of Ezh2 and Pax5 binding at the Wapl promoter in mature B cells after two days of activation. d, Co-precipitation of Myc-tagged Ezh2 or IRF4 by streptavidin (SA) pulldown of biotinylated Pax5-Bio from nuclear extracts prepared from HEK-293T cells that were transiently transfected with expression vectors encoding Pax5-Bio-IRES-BirA, Myc-Ezh2, Eed and Suz12 or Pax5-Bio-IRES-BirA and Myc-IRF4. The input (1/100) and protein precipitates were analysed by immunoblotting with antibodies detecting Myc and Pax5. The band indicated by an asterisk may correspond to endogenous Myc. One of four experiments is shown with marker proteins (kilodaltons). e, Flow cytometric analysis of bone marrow from Rag1cre/+Eedfl/fl (blue) and control (black) mice. Rag1cre/+Eedfl/+, Rag1cre/+, Eedfl/+ and Eed+/+ mice were used as control. The relative frequencies of the indicated cell types (left), which were determined in six experiments, are shown as mean values with s.e.m. and were analysed by multiple t-tests (unpaired and two-tailed with Holm–Sidak correction). GFP expression (right) is shown for Rag1cre/+Eedfl/fl pro-B cells in contrast to control Eedfl/+ pro-B cells. f, Scatter plot of gene expression differences between Eed-deficient and control pro-B cells. Eed-activated (blue) and Eed-repressed (red) genes were defined by an expression difference of >3-fold, an adjusted P value of <0.05 and a TPM value of >5 in Eed-deficient or control pro-B cells, respectively (Supplementary Table 2). The expression data are based on 5 (Rag1cre/+Eedfl/fl) and 4 (control; 3 × Rag1cre/+Eedfl/+, 1 × Eedfl/fl) RNA-seq experiments. g, Functional classification and quantification of the proteins encoded by Eed-activated and Eed-repressed genes (Supplementary Table 2). The bar size indicates the percentage of activated or repressed genes in each functional class relative to the total activated or repressed genes, respectively. Numbers in the bars refer to the genes in each functional class. h, Expression of selected genes in Eed-deficient (grey) and Eed-expressing (black) pro-B cells. The expression of the indicated genes, which are not differentially expressed according to the definition in f, is shown as mean TPM value with s.e.m. i, j, VDJ-seq analysis of Rag1cre/+Eedfl/fl and control pro-B cells. i, The mean percentages of uniquely identified DJH and VDJH sequences were determined based on 4 (Rag1cre/+Eedfl/fl) and 2 (control; 1 × Eedfl/+, 1 × Eed+/+) experiments. j, The VH gene usage is shown as mean percentage of all VDJH rearrangement events with s.e.m. n, number of experiments. Each dot corresponds to one mouse (e, h, i) or one gene (f).
Extended Data Fig. 10 Changes of chromosomal architecture and gene expression in Wapl+/+ and Wapl∆P1,2/∆P1,2 pro-B cells.
a, iFRAP analysis of splenic mature B cells from Wapl∆P1,2/+ (blue) and Wapl+/+ (pink) mice carrying the Smc3-gfp transgene. The difference in fluorescence intensity between bleached and unbleached regions is plotted against time. The mean values are indicated by lines and the s.d. by shading. n, number of cells analysed in three experiments per genotype. b, Frequency distribution of intrachromosomal contacts as a function of the genomic distance using logarithmically increased genomic distance bins, as determined by Homer analysis of Hi-C data generated with short-term-cultured pro-B cells (grey) and ex vivo-sorted splenic mature B cells (pink) from wild-type mice. c, Hi-C contact matrices of a zoomed-in region on chromosome 12 (mm9: 72,500,000–78,500,000; upper row) and 16 (21,500,000–28,500,000; lower row), displayed at a 10-kb bin resolution for Pax5∆/∆ progenitors, Wapl+/+ and Wapl∆P1,2/∆P1,2 pro-B cells. Black dots indicate loop anchors identified with Juicebox, and the intensity of each pixel represents the normalized number of contacts between a pair of loci30. The maximum intensity is indicated in the lower left of each panel. d, Density distribution of the loop length in Pax5∆/∆ progenitors, Wapl+/+ and Wapl∆P1,2/∆P1,2 pro-B cells, as determined with HiCCUPS of Juicer. The median loop length (in kb) is shown for each genotype. e, Scatter plot of gene expression differences, based on two RNA-seq experiments per genotype. Genes upregulated (red) or downregulated (blue) in Wapl∆P1,2/∆P1,2 pro-B cell compared to Wapl+/+ pro-B cells were defined by an expression difference of >2-fold, an adjusted P value of <0.05 and a TPM value of >5 in at least one of the two pro-B cell types (Supplementary Table 3). f, Functional classification and quantification of the proteins encoded by upregulated (red) and downregulated (blue) genes in Wapl∆P1,2/∆P1,2 pro-B cells relative to Wapl+/+ pro-B cells (Supplementary Table 3). See Extended Data Fig. 9g for detailed explanation. g, Expression of selected genes in Wapl∆P1,2/∆P1,2 (grey) and Wapl+/+ (black) pro-B cells. The expression of the indicated genes, which are not differentially expressed according to the definition in e, is shown as mean TPM value. h, Schematic depicting a ‘stable’ loop formed by loop extrusion across the entire Igh locus in pro-B cells. The forward CBEs (in the VH gene cluster and IGCR region) and the reverse CBEs (in the IGCR and 3′CBE regions) are indicated by red and blue arrows, respectively. The RAG-bound recombination centre, which is located at the DJH-rearranged gene segment29, is indicated in grey. The convergent orientation of the 12-RSS (recognition signal sequence with a 12-bp spacer, black arrowhead) of the DH segment and the 23-RSS (with a 23-bp spacer, green arrowhead) of the VH genes is essential for mediating RAG-cleavage and deletional joining16. The loop-extruding cohesin ring (orange) is arrested at convergent CBEs. In a ‘stable’ loop, the correct alignment of the RSS elements of a VH gene and the DJH-rearranged gene segment probably occurs by local diffusion. Arrowheads symbolize the RSS element consisting of the heptamer, nonamer and intervening spacer. i, Correct alignment of the RSS elements of a VH gene and the DJH-rearranged gene segment may be mediated by the ongoing process of loop extrusion. Orange and black arrows indicate the direction of movement of cohesin and DNA, respectively. j, Misalignment of the RSS elements of an inverted VH gene and the DJH-rearranged gene segment during loop extrusion, which prevents RAG-mediated cleavage.
Supplementary information
41586_2020_2454_MOESM1_ESM.pdf
Supplementary Figure 1 Original source images for Western blots shown in Figure 2d, Extended Data Figure 5b, Extended Data Figure 5d, Extended Data Figure 6d and Extended Data Figure 9d. Images are in uncropped form and contain molecular weight markers, loading controls and an indication of how the gels were cropped for the final figure. Controls of the blots shown in Figure 2d, Extended Data Figure 5b and Extended Data Figure 6d were run on the same gel as loading controls. Gating strategies used for cell sorting of immature IgMb B cells, pro-B cells, pre-B cells and Pax5-deficient progenitors from the bone marrow, and T cell populations from the spleen and thymus. Representative pre-sort and post-sort flow cytometry plots are shown.
Supplementary Table 1
Supplementary Table S1a: List of VH genes in chromosomal order; Supplementary Table S1b: VH8-8 gene insertion and genomic context; Supplementary Table S1c: CBE array sequence and genomic context.
Supplementary Table 2
Activated and repressed genes determined by RNA-seq of ex vivo sorted control [Eed-expressing] and Eed KO [Rag1(Cre/+) Eed(fl/fl)] pro-B cells. The expression data, which were analysed with the DESeq2 program, are based on 4 (control; 3× Rag1Cre/+ Eedfl/+ and 1× Eedfl/fl) and 5 (Rag1Cre/+ Eedfl/fl) independent RNA-seq experiments. Eed-activated genes and Eed-repressed genes were defined by an expression difference of > 3-fold, an adjusted P value of < 0.05 and a TPM value of > 5 in Eed-deficient or control B cells, respectively.
Supplementary Table 3
Upregulated and downregulated genes determined by RNA-seq of ex vivo sorted Wapl(+/+) and Wapl(∆P1,2/∆P1,2) pro-B cells. The expression data, which were analysed with the DESeq2 program, are based on 2 (control: Wapl+/+) and 2 (Wapl∆P1,2/∆P1,2) independent RNA-seq experiments. Genes upregulated or downregulated in Wapl∆P1,2/∆P1,2 pro-B cell compared to Wapl+/+ pro-B cells were defined by an expression difference of > 2-fold, an adjusted P value of < 0.05 and a TPM value of > 5 in at least one of the two pro-B cell types.
Supplementary Table 4
Oligonucleotide sequence information. Lists the oligonucleotides that were used for genotyping of alleles generated in this study, for gene expression analysis, for the CRISPR/Cas9-mediated generation of new alleles and for VDJ-seq and 3C-seq analysis.
Supplementary Table 5
Description of all Illumina sequencing experiments generated for this study (GSE140975) and of previously published Illumina sequencing experiments.
Source data
Rights and permissions
About this article
Cite this article
Hill, L., Ebert, A., Jaritz, M. et al. Wapl repression by Pax5 promotes V gene recombination by Igh loop extrusion. Nature 584, 142–147 (2020). https://doi.org/10.1038/s41586-020-2454-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-020-2454-y
This article is cited by
-
RNA processing mechanisms contribute to genome organization and stability in B cells
Oncogene (2024)
-
BRWD1 orchestrates small pre-B cell chromatin topology by converting static to dynamic cohesin
Nature Immunology (2024)
-
Enhancer-instructed epigenetic landscape and chromatin compartmentalization dictate a primary antibody repertoire protective against specific bacterial pathogens
Nature Immunology (2023)
-
Three-dimensional genome organization in immune cell fate and function
Nature Reviews Immunology (2023)
-
An Igh distal enhancer modulates antigen receptor diversity by determining locus conformation
Nature Communications (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.