Abstract
Clonally expanded blood cells that contain somatic mutations (clonal haematopoiesis) are commonly acquired with age and increase the risk of blood cancer1,2,3,4,5,6,7,8,9. The blood clones identified so far contain diverse large-scale mosaic chromosomal alterations (deletions, duplications and copy-neutral loss of heterozygosity (CN-LOH)) on all chromosomes1,2,5,6,9, but the sources of selective advantage that drive the expansion of most clones remain unknown. Here, to identify genes, mutations and biological processes that give selective advantage to mutant clones, we analysed genotyping data from the blood-derived DNA of 482,789 participants from the UK Biobank10. We identified 19,632 autosomal mosaic chromosomal alterations and analysed these for relationships to inherited genetic variation. We found 52 inherited, rare, large-effect coding or splice variants in 7 genes that were associated with greatly increased vulnerability to clonal haematopoiesis with specific acquired CN-LOH mutations. Acquired mutations systematically replaced the inherited risk alleles (at MPL) or duplicated them to the homologous chromosome (at FH, NBN, MRE11, ATM, SH2B3 and TM2D3). Three of the genes (MRE11, NBN and ATM) encode components of the MRN–ATM pathway, which limits cell division after DNA damage and telomere attrition11,12,13; another two (MPL and SH2B3) encode proteins that regulate the self-renewal of stem cells14,15,16. In addition, we found that CN-LOH mutations across the genome tended to cause chromosomal segments with alleles that promote the expansion of haematopoietic cells to replace their homologous (allelic) counterparts, increasing polygenic drive for blood-cell proliferation traits. Readily acquired mutations that replace chromosomal segments with their homologous counterparts seem to interact with pervasive inherited variation to create a challenge for lifelong cytopoiesis.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Data availability
Mosaic event calls are available in Supplementary Data in anonymized form. The mCA call set has also been returned to UK Biobank (as Return 2062) to enable individual-level linkage to approved UK Biobank applications. Access to the UK Biobank Resource is available by application (http://www.ukbiobank.ac.uk/).
Code availability
A standalone software implementation (MoChA) of the algorithm used to call mCAs is available at https://github.com/freeseek/mocha. Code used to perform the specific analyses in this study is available from the authors upon request (but unlike MoChA, this code is not immediately portable to other computing environments).
References
Jacobs, K. B. et al. Detectable clonal mosaicism and its relationship to aging and cancer. Nat. Genet. 44, 651–658 (2012).
Laurie, C. C. et al. Detectable clonal mosaicism from birth to old age and its relationship to cancer. Nat. Genet. 44, 642–650 (2012).
Genovese, G. et al. Clonal hematopoiesis and blood-cancer risk inferred from blood DNA sequence. N. Engl. J. Med. 371, 2477–2487 (2014).
Jaiswal, S. et al. Age-related clonal hematopoiesis associated with adverse outcomes. N. Engl. J. Med. 371, 2488–2498 (2014).
Machiela, M. J. et al. Characterization of large structural genetic mosaicism in human autosomes. Am. J. Hum. Genet. 96, 487–497 (2015).
Vattathil, S. & Scheet, P. Extensive hidden genomic mosaicism revealed in normal tissue. Am. J. Hum. Genet. 98, 571–578 (2016).
Zink, F. et al. Clonal hematopoiesis, with and without candidate driver mutations, is common in the elderly. Blood 130, 742–752 (2017).
Abelson, S. et al. Prediction of acute myeloid leukaemia risk in healthy individuals. Nature 559, 400–404 (2018).
Loh, P.-R. et al. Insights into clonal haematopoiesis from 8,342 mosaic chromosomal alterations. Nature 559, 350–355 (2018).
Bycroft, C. et al. The UK Biobank resource with deep phenotyping and genomic data. Nature 562, 203–209 (2018).
Uziel, T. et al. Requirement of the MRN complex for ATM activation by DNA damage. EMBO J. 22, 5612–5621 (2003).
Lee, J.-H. & Paull, T. T. ATM activation by DNA double-strand breaks through the Mre11-Rad50-Nbs1 complex. Science 308, 551–554 (2005).
Deng, Y., Guo, X., Ferguson, D. O. & Chang, S. Multiple roles for MRE11 at uncapped telomeres. Nature 460, 914–918 (2009).
Kimura, S., Roberts, A. W., Metcalf, D. & Alexander, W. S. Hematopoietic stem cell deficiencies in mice lacking c-Mpl, the receptor for thrombopoietin. Proc. Natl Acad. Sci. USA 95, 1195–1200 (1998).
Solar, G. P. et al. Role of c-mpl in early hematopoiesis. Blood 92, 4–10 (1998).
Seita, J. et al. Lnk negatively regulates self-renewal of hematopoietic stem cells by modifying thrombopoietin-mediated signal transduction. Proc. Natl Acad. Sci. USA 104, 2349–2354 (2007).
Loh, P.-R., Palamara, P. F. & Price, A. L. Fast and accurate long-range phasing in a UK Biobank cohort. Nat. Genet. 48, 811–816 (2016).
Loh, P.-R. et al. Reference-based phasing using the Haplotype Reference Consortium panel. Nat. Genet. 48, 1443–1448 (2016).
Auer, P. L. et al. Rare and low-frequency coding variants in CXCR2 and other genes are associated with hematological traits. Nat. Genet. 46, 629–634 (2014).
Schultz, K. A. P. et al. PTEN, DICER1, FH, and their associated tumor susceptibility syndromes: clinical features, genetics, and surveillance recommendations in childhood. Clin. Cancer Res. 23, e76–e82 (2017).
Landrum, M. J. et al. ClinVar: improving access to variant interpretations and supporting evidence. Nucleic Acids Res. 46, D1062–D1067 (2018).
Van Hout, C. V. et al. Whole exome sequencing and characterization of coding variation in 49,960 individuals in the UK Biobank. Preprint at https://www.bioRxiv.org/content/10.1101/572347v1 (2019).
Meuwissen, T. H., Hayes, B. J. & Goddard, M. E. Prediction of total genetic value using genome-wide dense marker maps. Genetics 157, 1819–1829 (2001).
Purcell, S. M. et al. Common polygenic variation contributes to risk of schizophrenia and bipolar disorder. Nature 460, 748–752 (2009).
Khera, A. V. et al. Genome-wide polygenic scores for common diseases identify individuals with risk equivalent to monogenic mutations. Nat. Genet. 50, 1219–1224 (2018).
Loh, P.-R. et al. Efficient Bayesian mixed-model analysis increases association power in large cohorts. Nat. Genet. 47, 284–290 (2015).
Thompson, D. J. et al. Genetic predisposition to mosaic Y chromosome loss in blood. Nature 575, 652–657 (2019).
Jaiswal, S. et al. Clonal hematopoiesis and risk of atherosclerotic cardiovascular disease. N. Engl. J. Med. 377, 111–121 (2017).
Davoli, T. et al. Cumulative haploinsufficiency and triplosensitivity drive aneuploidy patterns and shape the cancer genome. Cell 155, 948–962 (2013).
O’Keefe, C., McDevitt, M. A. & Maciejewski, J. P. Copy neutral loss of heterozygosity: a novel chromosomal lesion in myeloid malignancies. Blood 115, 2731–2739 (2010).
Chase, A. et al. Profound parental bias associated with chromosome 14 acquired uniparental disomy indicates targeting of an imprinted locus. Leukemia 29, 2069–2074 (2015).
Choate, K. A. et al. Mitotic recombination in patients with ichthyosis causes reversion of dominant mutations in KRT10. Science 330, 94–97 (2010).
Tesi, B. et al. Gain-of-function SAMD9L mutations cause a syndrome of cytopenia, immunodeficiency, MDS, and neurological symptoms. Blood 129, 2266–2279 (2017).
Schizophrenia Working Group of the Psychiatric Genomics Consortium. Biological insights from 108 schizophrenia-associated genetic loci. Nature 511, 421–427 (2014).
Sudlow, C. et al. UK biobank: an open access resource for identifying the causes of a wide range of complex diseases of middle and old age. PLoS Med. 12, e1001779 (2015).
Wain, L. V. et al. Novel insights into the genetics of smoking behaviour, lung function, and chronic obstructive pulmonary disease (UK BiLEVE): a genetic association study in UK Biobank. Lancet Respir. Med. 3, 769–781 (2015).
Das, S. et al. Next-generation genotype imputation service and methods. Nat. Genet. 48, 1284–1287 (2016).
Chang, C. C. et al. Second-generation PLINK: rising to the challenge of larger and richer datasets. Gigascience 4, 7 (2015).
Peiffer, D. A. et al. High-resolution genomic profiling of chromosomal aberrations using Infinium whole-genome genotyping. Genome Res. 16, 1136–1148 (2006).
Diskin, S. J. et al. Adjustment of genomic waves in signal intensities from whole-genome SNP genotyping platforms. Nucleic Acids Res. 36, e126 (2008).
Manichaikul, A. et al. Robust relationship inference in genome-wide association studies. Bioinformatics 26, 2867–2873 (2010).
Rentzsch, P., Witten, D., Cooper, G. M., Shendure, J. & Kircher, M. CADD: predicting the deleteriousness of variants throughout the human genome. Nucleic Acids Res. 47, D886–D894 (2019).
McLaren, W. et al. The Ensembl Variant Effect Predictor. Genome Biol. 17, 122 (2016).
Thorvaldsdóttir, H., Robinson, J. T. & Mesirov, J. P. Integrative Genomics Viewer (IGV): high-performance genomics data visualization and exploration. Brief. Bioinform. 14, 178–192 (2013).
Pedersen, B. S. & Quinlan, A. R. Mosdepth: quick coverage calculation for genomes and exomes. Bioinformatics 34, 867–868 (2018).
Regier, A. A. et al. Functional equivalence of genome sequencing analysis pipelines enables harmonized variant calling across human genetics projects. Nat. Commun. 9, 4038 (2018).
Loh, P.-R., Kichaev, G., Gazal, S., Schoech, A. P. & Price, A. L. Mixed-model association for biobank-scale datasets. Nat. Genet. 50, 906–908 (2018).
Turner, J. J. et al. InterLymph hierarchical classification of lymphoid neoplasms for epidemiologic research based on the WHO classification (2008): update and future directions. Blood 116, e90–e98 (2010).
Arber, D. A. et al. The 2016 revision to the World Health Organization classification of myeloid neoplasms and acute leukemia. Blood 127, 2391–2405 (2016).
Jones, A. V. et al. JAK2 haplotype is a major risk factor for the development of myeloproliferative neoplasms. Nat. Genet. 41, 446–449 (2009).
Kilpivaara, O. et al. A germline JAK2 SNP is associated with predisposition to the development of JAK2 V617F-positive myeloproliferative neoplasms. Nat. Genet. 41, 455–459 (2009).
Olcaydu, D. et al. A common JAK2 haplotype confers susceptibility to myeloproliferative neoplasms. Nat. Genet. 41, 450–454 (2009).
Koren, A. et al. Genetic variation in human DNA replication timing. Cell 159, 1015–1026 (2014).
Gusev, A. et al. Whole population, genome-wide mapping of hidden relatedness. Genome Res. 19, 318–326 (2009).
Acknowledgements
We thank S. Bakhoum, S. Raychaudhuri, M. Sherman, S. Elledge and C. Terao for discussions. This research was conducted using the UK Biobank Resource under application no. 19808. P.-R.L. was supported by US National Institutes of Health (NIH) grant DP2 ES030554, a Burroughs Wellcome Fund Career Award at the Scientific Interfaces, the Next Generation Fund at the Broad Institute of MIT and Harvard, a Glenn Foundation for Medical Research and AFAR Grants for Junior Faculty award, and a Sloan Research Fellowship. G.G. and S.A.M. were supported by US NIH grant R01 HG006855. G.G. was supported by US Department of Defense Breast Cancer Research Breakthrough Award W81XWH-16-1-0316. Computational analyses were performed on the O2 High Performance Compute Cluster, supported by the Research Computing Group, at Harvard Medical School (http://rc.hms.harvard.edu), and on the Genetic Cluster Computer (http://www.geneticcluster.org) hosted by SURFsara and financially supported by the Netherlands Scientific Organization (NWO 480-05-003 PI: Posthuma) along with a supplement from the Dutch Brain Foundation and the VU University Amsterdam. We thank S. Elledge, B. Ebert, and C. Patil for helpful comments on the manuscript.
Author information
Authors and Affiliations
Contributions
P.-R.L., G.G. and S.A.M. designed the study. P.-R.L. performed computational analyses. P.-R.L., G.G. and S.A.M. wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
Patent application PCT/WO2019/ 079493 has been filed on the mCA detection method used in this work.
Additional information
Peer review information Nature thanks Paul Scheet, George Vassiliou and John Witte for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
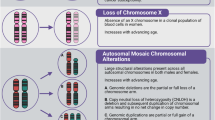
Extended Data Fig. 1 Mosaic chromosomal alterations detected among 482,789 UK Biobank participants.
a, Each horizontal line corresponds to an mCA; a total of 19,632 autosomal events in 17,111 unique individuals are displayed. Detected events are colour-coded by copy number of the affected chromosome or segment (orange, LOH; blue, loss/deletion; red, gain/duplication). Focal deletions are labelled in blue with the names of putative target genes. Loci containing inherited variants influencing somatic events in cis are labelled in the same colour as the corresponding mCA (orange for CN-LOH-associated loci, blue for losses). b, Sex and age distributions of individuals with detected mosaic events. Marker size and colour intensity increase with event frequency. Error bars denote 95% confidence intervals. Sample sizes are provided in Supplementary Table 1 and numeric data are provided in Supplementary Table 4. Three events with unusual sex biases (gains on chromosome 15, 16p11.2 deletions and 10q terminal deletions) were previously reported9, all of which replicated here. We have not identified a mechanism that could explain the sex biases. The overall tendency of male enrichment for most mCAs raises the possibility that environmental exposures could result in genomic insults that lead to mCAs; however, the heterogeneity of the level of male enrichment across different mCAs suggests that the mechanisms producing sex biases may be event-specific. c, Enrichment of mosaic chromosomal alterations in individuals with anomalously high blood indices. Different mCAs are significantly enriched (FDR of 0.05; one-sided Fisher’s exact test) among n = 455,009 individuals with anomalous blood counts in different blood lineages (adjusted for age, sex and smoking status). Events were grouped by chromosome and copy number, with loss and CN-LOH events subdivided by p-arm versus q-arm. (We did not subdivide gain events by arm because most gain events are whole-chromosome trisomies.) Numeric data are provided in Supplementary Table 5.
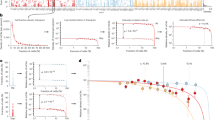
Extended Data Fig. 2 Copy number determination and quality control of mosaic chromosomal alteration calls.
a–d, Total versus relative allelic intensities of mCAs detected on each chromosome. Mean log2(R ratio) (LRR) of each detected mCA is plotted against estimated change in B allele frequency at heterozygous sites (|ΔBAF|). The data exhibit the characteristic ‘arrowhead’ pattern in which LRR/|ΔBAF| approximately equals a positive constant for gain events, zero for CN-LOH events, and a negative constant for loss events. Possible constitutional duplications were filtered according to thresholds on LRR and |ΔBAF| defined in Supplementary Note 1. Constitutional duplications have expected |ΔBAF| = 1/6 and have LRR values of approximately 0.36 in this dataset. We chose exclusion thresholds to conservatively discard all calls that might belong to this cluster, applying more stringent filtering to shorter events because (i) most constitutional duplications are short; and (ii) shorter events have noisier LRR and |ΔBAF| estimates. e, Estimation of FDR using age distributions of individuals with mCA calls. We generated age distributions for (i) ‘high confidence’ events passing a permutation-based FDR threshold of 0.01 (bright green); (ii) ‘medium confidence’ events below the FDR threshold of 0.01 but passing an FDR threshold of 0.05 (darker green); and (iii) ‘low confidence’ events below the FDR threshold of 0.05 but passing an FDR threshold of 0.10 (darkest green; excluded from our call set but plotted for context). We compared these distributions to the overall age distribution of UK Biobank participants (grey). On the basis of the numbers of events in each category, approximately 32% of medium-confidence detected events are expected to be false positives. To estimate our true FDR, we regressed the medium-confidence age distribution on the high-confidence and overall age distributions, reasoning that the medium-confidence age distribution should be a mixture of correctly called events (with age distribution similar to that of the high-confidence events) and spurious calls (with age distribution similar to the overall cohort). We observed a regression weight of 0.44 for the component corresponding to spurious calls, in good agreement with expectation, and indicating a true FDR of 6.6% (4.5–8.6%, 95% confidence interval based on regression fit on n = 6 age bins). f, Fractions of individuals with at least one detected autosomal mCA stratified by age and sex. Error bars denote 95% confidence intervals. Numeric data are provided in Supplementary Table 3.
Extended Data Fig. 3 Principal component plot of UK Biobank participants.
Individuals are plotted by their first two genetic principal component coordinates as computed by UK Biobank10 and coloured according to self-reported ethnic background. Red circles indicate individuals identified in our exome analyses (of self-reported white individuals with mosaic CN-LOH events) as carriers of rare coding or splice variants in frequently-targeted genes. Marginal density histograms stratified by self-reported ethnic background are provided next to the PC1 and PC2 axes.
Extended Data Fig. 4 Quantile–quantile plots of P values produced by association analyses.
These plots verify the calibration of the statistical tests we used to identify the genome-wide significant associations reported in Extended Data Table 1 (see legend for details of statistical tests and sample sizes). In each plot, the blue dots correspond to an analysis of all variants tested, and the black dots correspond to an analysis in which regions surrounding significant associations were excluded. Specifically, the plots respectively exclude chr1:35–55 Mb (MPL), chr1:239–244 Mb (FH), chr8:88–93 Mb (NBN), chr9:2.5–7.5 Mb (JAK2), chr11:92–97 Mb (MRE11), chr11:103–113 Mb (ATM), chr12:109–114 Mb (SH2B3), chr14:92.5–102.5 Mb (TCL1A and DLK1) and chr15:100Mb–qter (TM2D3) (hg19 coordinates). In all cases, exclusion of the hit regions (which account for a small fraction of the variants tested) resulted in a distribution close to the expected null.
Extended Data Fig. 5 Identification and validation of an inherited MPL structural variant.
We suspected that an association between rs144279563 and acquired 1p CN-LOH mutations might tag a causal structural variant in MPL. (Although rs144279563 is approximately 1.5 Mb downstream of MPL, it is sufficiently rare to be in linkage disequilibrium with variants several megabases away.) We therefore examined genotyping intensities at MPL from 49,950 individuals typed on the BiLEVE chip (which contains more probes within MPL than the Biobank chip, on which the remaining individuals were typed). a, Mean genotyping intensities over 42 carriers of the rs144279563 rare allele exhibit a sharp increase at the end of MPL exon 9 (1 genotyping probe) followed by a sharp decrease in exon 10 (3 genotyping probes). b, c, Closer inspection of genotyping intensities at the 4 probes across all BiLEVE individuals enabled identification of 27 individuals likely to carry an inherited structural variant (20 of which carry the rs144279563 rare allele). We called this variant in the BiLEVE cohort using two criteria: (i) correct sign of LRR at the 4 probes (+, –, –, –); and (ii) mean signed LRR shift >0.4 over the four probes. d, Read support for a 454-bp deletion spanning MPL exon 10 in exome-sequenced individuals. We used IGV44 to plot paired-end reads aligning in or near MPL exons 9 and 10 in four exome-sequenced individuals imputed to carry the MPL structural variant (and also mosaic for 1p CN-LOH events). Read pairs highlighted in red have unusually long insert sizes, consistent with a deletion of genomic sequence between the aligned reads. Multicoloured read segments indicate clipped reads in which one end of a read stops aligning to the reference genome. On the left side of the deletion, clipped reads align through hg19 base pair 43,814,728 (…AGGGACTGGG; last five matching bases in bold for comparison to sequences below), with mismatches consistently occurring starting from 43,814,729 rightward (hg19: CGCCG…). On the right side of the deletion, clipped reads align starting from 43,815,178 (CTGGGACTCG…), with mismatches starting from 43,815,177 leftward (hg19: …CACCT). Examination of individual clipped reads revealed sequence matching …AGGGACTGGGACTCG…, indicating deletion of 5 bp (CTGGG) in addition to the 449 bp between aligning read segments. In this legend we have used hg19 coordinates for consistency with the rest of this Article; the IGV plot uses hg38 coordinates because reads had been aligned to hg38 (amounting to an offset of −465,671 bp relative to hg19 at MPL). e, f, Decreased read depth at exon 10 in all 32 imputed carriers of the MPL exon 10 deletion who had been exome-sequenced. We used mosdepth45 to compute mean read depth across all 12 MPL exons in the 32 exome-sequenced imputed deletion carriers along with 32 controls. We normalized read depth in each individual by dividing by mean read depth across exons 1–8 and 11–12. All 32 imputed carriers of the exon 10 deletion had lower exon 10 normalized read depths than all 32 controls. We did not observe any evidence of increased read depth in exon 9 in carriers versus controls.
Extended Data Fig. 6 Identity-by-descent graph at MPL among individuals with likely 1p CN-LOH events spanning MPL.
We called identity-by-descent (IBD) tracts using GERMLINE with haplotype extension54. Coloured nodes indicate carriers of the 28 rare coding or splice variants we observed to be independently (and probably causally) associated with 1p CN-LOH mutations (always replacing the rare allele with the reference allele) (Extended Data Table 1, Supplementary Table 7). (The numbers of carriers listed for each variant here are slightly higher than in the ‘allelic shift’ columns of Extended Data Table 1 and Supplementary Table 7 because allelic shifts could only be confidently ascertained for a subset of carriers.) The presence of additional IBD clusters not carrying any of the 28 highlighted variants suggests that even more causal variants in MPL remain to be discovered.
Extended Data Fig. 7 Identity-by-descent graph at ATM among individuals with likely 11q CN-LOH events spanning ATM.
We called IBD tracts using GERMLINE with haplotype extension54. Coloured nodes indicate carriers of the eight rare coding or splice variants we observed to be independently (and probably causally) associated with 11q CN-LOH mutations (always making the rare allele homozygous) (Extended Data Table 1, Supplementary Table 7). The presence of additional IBD clusters not carrying any of the highlighted variants suggests that even more causal variants in ATM remain to be discovered. The two carriers of rs786204751 are also carriers of rs587779872, as discussed in Methods.
Extended Data Fig. 8 Variant allele fractions of rare coding or splice variants likely to be targets of CN-LOH mutations in exome-sequenced individuals.
Variant allele fractions (VAF; the number of reads matching the alternative allele divided by the total number of reads matching either the reference or the alternative allele) are plotted for each variant call identified as the potential target of a CN-LOH event (from either association analyses or burden analyses). Error bars denote 95% confidence intervals approximated using binomial standard errors multiplied by 1.96. Allelic read depths for variants identified at DNMT3A, TET2 and JAK2 are broadly indicative of somatic origin (VAF < 0.5), whereas read depths for variants at the seven inherited risk loci are broadly consistent with inherited variation (VAF ≈ 0.5). Read depths were generally insufficient to make a confident assessment of somatic versus inherited origin on a per-variant level, as evidenced by wide VAF error bars; in addition, making this determination is further complicated by mapping bias towards the reference allele, which can produce VAF lower than 0.5 even for inherited variants3.
Extended Data Fig. 9 Tendencies of CN-LOH mutations to modify polygenic scores for 29 blood cell parameters.
For each blood count parameter and each chromosome arm, the heat map reports the z-score for the mean change in polygenic score across all CN-LOH mutations detected on the arm. Among the 29 blood count parameters we considered, some of the parameters corresponding to abundances of blood cell types might be surrogates for enhanced cellular fitness (in many cases of mitotic progenitors rather than the cell types themselves). Other parameters reflect cell size or morphology. Effects of CN-LOH mutations on polygenic scores for these parameters may reflect the production of abnormal cells by biologically altered stem cells, rather than cellular fitness itself (which may be a property of the unobserved haematopoietic stem cells). Columns: platelet count and crit (PLT#, Pct); red blood cell count (RBC#), haemoglobin (Hgb) and haematocrit (Hct) (both strongly correlated with red blood cell count); reticulocyte count and percentage (RET#, RET%); high light scatter reticulocyte count and percent (HLR#, HLR%); immature reticulocyte fraction (IRF); white blood cell count (WBC#); neutrophil count and percentage (NEU#, NEU%); eosinophil count and percentage (EOS#, EOS%); monocyte count and percentage (MON#, MON%); basophil count and percentage (BAS#, BAS%); lymphocyte count and percentage (LYM#, LYM%); platelet distribution width (PDW), mean platelet volume (MPV), RBC distribution width (RDW), mean corpuscular volume (MCV), mean reticulocyte volume (MRV), mean sphered cell volume (MSCV), mean corpuscular haemoglobin (MCH) and mean corpuscular haemoglobin concentration (MCHC).
Supplementary information
Supplementary Information
This file contains Supplementary Notes 1-11 and Supplementary Tables 1-23.
Supplementary Data
This file contains anonymized individual-level mosaic chromosomal alteration calls.
Rights and permissions
About this article
Cite this article
Loh, PR., Genovese, G. & McCarroll, S.A. Monogenic and polygenic inheritance become instruments for clonal selection. Nature 584, 136–141 (2020). https://doi.org/10.1038/s41586-020-2430-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-020-2430-6
This article is cited by
-
Driver mutation zygosity is a critical factor in predicting clonal hematopoiesis transformation risk
Blood Cancer Journal (2024)
-
Protein-altering variants at copy number-variable regions influence diverse human phenotypes
Nature Genetics (2024)
-
A concerted neuron–astrocyte program declines in ageing and schizophrenia
Nature (2024)
-
Inherited polygenic effects on common hematological traits influence clonal selection on JAK2V617F and the development of myeloproliferative neoplasms
Nature Genetics (2024)
-
Multiparameter prediction of myeloid neoplasia risk
Nature Genetics (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.