Abstract
The mammalian immune system implements a remarkably effective set of mechanisms for fighting pathogens1. Its main components are haematopoietic immune cells, including myeloid cells that control innate immunity, and lymphoid cells that constitute adaptive immunity2. However, immune functions are not unique to haematopoietic cells, and many other cell types display basic mechanisms of pathogen defence3,4,5. To advance our understanding of immunology outside the haematopoietic system, here we systematically investigate the regulation of immune genes in the three major types of structural cells: epithelium, endothelium and fibroblasts. We characterize these cell types across twelve organs in mice, using cellular phenotyping, transcriptome sequencing, chromatin accessibility profiling and epigenome mapping. This comprehensive dataset revealed complex immune gene activity and regulation in structural cells. The observed patterns were highly organ-specific and seem to modulate the extensive interactions between structural cells and haematopoietic immune cells. Moreover, we identified an epigenetically encoded immune potential in structural cells under tissue homeostasis, which was triggered in response to systemic viral infection. This study highlights the prevalence and organ-specific complexity of immune gene activity in non-haematopoietic structural cells, and it provides a high-resolution, multi-omics atlas of the epigenetic and transcriptional networks that regulate structural cells in the mouse.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
Raw and processed sequencing data (RNA-seq, ATAC-seq and H3K4me2 ChIPmentation) are available from the NCBI Gene Expression Omnibus (GEO) repository under accession number GSE134663. In addition, the dataset is provided as an online resource on a supplementary website (http://structural-immunity.computational-epigenetics.org), which includes links to raw and processed sequencing data, further analysis results and genome browser tracks for interactive visualization of the RNA-seq, ATAC-seq and ChIPmentation profiles. Source data are provided with this paper.
Code availability
The analysis source code underlying the final version of the paper is openly available from http://structural-immunity.computational-epigenetics.org.
References
Kotas, M. E. & Medzhitov, R. Homeostasis, inflammation, and disease susceptibility. Cell 160, 816–827 (2015).
Abbas, A. K., Lichtman, A. H. & Pillai, S. Cellular and Molecular Immunology 9th edn (Elsevier, 2017).
Buckley, C. D., Barone, F., Nayar, S., Bénézech, C. & Caamaño, J. Stromal cells in chronic inflammation and tertiary lymphoid organ formation. Annu. Rev. Immunol. 33, 715–745 (2015).
Pober, J. S. & Sessa, W. C. Evolving functions of endothelial cells in inflammation. Nat. Rev. Immunol. 7, 803–815 (2007).
Schleimer, R. P., Kato, A., Kern, R., Kuperman, D. & Avila, P. C. Epithelium: at the interface of innate and adaptive immune responses. J. Allergy Clin. Immunol. 120, 1279–1284 (2007).
Pawlina, W. & Ross, M. H. Histology: a Text and Atlas, International Edition: with Correlated Cell and Molecular Biology (Wolters Kluwer Law & Business, 2019).
Humbert, M., Hugues, S. & Dubrot, J. Shaping of peripheral T cell responses by lymphatic endothelial cells. Front. Immunol. 7, 684 (2017).
Malhotra, D., Fletcher, A. L. & Turley, S. J. Stromal and hematopoietic cells in secondary lymphoid organs: partners in immunity. Immunol. Rev. 251, 160–176 (2013).
Nowarski, R., Jackson, R. & Flavell, R. A. The stromal intervention: Regulation of immunity and inflammation at the epithelial-mesenchymal barrier. Cell 168, 362–375 (2017).
Perez-Shibayama, C., Gil-Cruz, C. & Ludewig, B. Fibroblastic reticular cells at the nexus of innate and adaptive immune responses. Immunol. Rev. 289, 31–41 (2019).
Turley, S. J., Cremasco, V. & Astarita, J. L. Immunological hallmarks of stromal cells in the tumour microenvironment. Nat. Rev. Immunol. 15, 669–682 (2015).
Davis, M. M., Tato, C. M. & Furman, D. Systems immunology: just getting started. Nat. Immunol. 18, 725–732 (2017).
Villani, A. C., Sarkizova, S. & Hacohen, N. Systems immunology: learning the rules of the immune system. Annu. Rev. Immunol. 36, 813–842 (2018).
Picelli, S. et al. Full-length RNA-seq from single cells using Smart-seq2. Nat. Protocols 9, 171–181 (2014).
Buenrostro, J. D., Giresi, P. G., Zaba, L. C., Chang, H. Y. & Greenleaf, W. J. Transposition of native chromatin for fast and sensitive epigenomic profiling of open chromatin, DNA-binding proteins and nucleosome position. Nat. Methods 10, 1213–1218 (2013).
Schmidl, C., Rendeiro, A. F., Sheffield, N. C. & Bock, C. ChIPmentation: fast, robust, low-input ChIP-seq for histones and transcription factors. Nat. Methods 12, 963–965 (2015).
Wang, Y., Li, X. & Hu, H. H3K4me2 reliably defines transcription factor binding regions in different cells. Genomics 103, 222–228 (2014).
Su, A. I. et al. A gene atlas of the mouse and human protein-encoding transcriptomes. Proc. Natl Acad. Sci. USA 101, 6062–6067 (2004).
Han, X. et al. Mapping the mouse cell atlas by microwell-seq. Cell 172, 1091–1107.e17 (2018).
The Tabula Muris Consortium. Single-cell transcriptomics of 20 mouse organs creates a Tabula Muris. Nature 562, 367–372 (2018).
Bock, C. et al. Reference maps of human ES and iPS cell variation enable high-throughput characterization of pluripotent cell lines. Cell 144, 439–452 (2011).
Baazim, H. et al. CD8+ T cells induce cachexia during chronic viral infection. Nat. Immunol. 20, 701–710 (2019).
Guerrero-Juarez, C. F. et al. Single-cell analysis reveals fibroblast heterogeneity and myeloid-derived adipocyte progenitors in murine skin wounds. Nat. Commun. 10, 650 (2019).
Kinchen, J. et al. Structural remodeling of the human colonic mesenchyme in inflammatory bowel disease. Cell 175, 372–386 (2018).
Mizoguchi, F. et al. Functionally distinct disease-associated fibroblast subsets in rheumatoid arthritis. Nat. Commun. 9, 789 (2018).
Montoro, D. T. et al. A revised airway epithelial hierarchy includes CFTR-expressing ionocytes. Nature 560, 319–324 (2018).
Parikh, K. et al. Colonic epithelial cell diversity in health and inflammatory bowel disease. Nature 567, 49–55 (2019).
Plasschaert, L. W. et al. A single-cell atlas of the airway epithelium reveals the CFTR-rich pulmonary ionocyte. Nature 560, 377–381 (2018).
Haber, A. L. et al. A single-cell survey of the small intestinal epithelium. Nature 551, 333–339 (2017).
Rodda, L. B. et al. Single-cell RNA sequencing of lymph node stromal cells reveals niche-associated heterogeneity. Immunity 48, 1014–1028 (2018).
Yan, K. S. et al. Non-equivalence of Wnt and R-spondin ligands during Lgr5+ intestinal stem-cell self-renewal. Nature 545, 238–242 (2017).
Ahmed, R., Salmi, A., Butler, L. D., Chiller, J. M. & Oldstone, M. B. Selection of genetic variants of lymphocytic choriomeningitis virus in spleens of persistently infected mice. Role in suppression of cytotoxic T lymphocyte response and viral persistence. J. Exp. Med. 160, 521–540 (1984).
Bergthaler, A. et al. Viral replicative capacity is the primary determinant of lymphocytic choriomeningitis virus persistence and immunosuppression. Proc. Natl Acad. Sci. USA 107, 21641–21646 (2010).
ImmGen Consortium. Final ImmGen sorting SOP; http://www.immgen.org/Protocols/ImmGen%20Cell%20prep%20and%20sorting%20SOP.pdf (accessed 24 March 2020).
Krausgruber, T. et al. T-bet is a key modulator of IL-23-driven pathogenic CD4+ T cell responses in the intestine. Nat. Commun. 7, 11627 (2016).
Saluzzo, S. et al. First-breath-induced type 2 pathways shape the lung immune environment. Cell Rep. 18, 1893–1905 (2017).
Lercher, A. et al. Type I interferon signaling disrupts the hepatic urea cycle and alters systemic metabolism to suppress T cell function. Immunity 51, 1074–1087.e9 (2019).
Corces, M. R. et al. Lineage-specific and single-cell chromatin accessibility charts human hematopoiesis and leukemia evolution. Nat. Genet. 48, 1193–1203 (2016).
Gustafsson, C., De Paepe, A., Schmidl, C. & Månsson, R. High-throughput ChIPmentation: freely scalable, single day ChIPseq data generation from very low cell-numbers. BMC Genomics 20, 59 (2019).
Pinschewer, D. D. et al. Innate and adaptive immune control of genetically engineered live-attenuated arenavirus vaccine prototypes. Int. Immunol. 22, 749–756 (2010).
Kouadjo, K. E., Nishida, Y., Cadrin-Girard, J. F., Yoshioka, M. & St-Amand, J. Housekeeping and tissue-specific genes in mouse tissues. BMC Genomics 8, 127 (2007).
Li, B. et al. A comprehensive mouse transcriptomic BodyMap across 17 tissues by RNA-seq. Sci. Rep. 7, 4200 (2017).
Shay, T. et al. Conservation and divergence in the transcriptional programs of the human and mouse immune systems. Proc. Natl Acad. Sci. USA 110, 2946–2951 (2013).
Lavin, Y. et al. Tissue-resident macrophage enhancer landscapes are shaped by the local microenvironment. Cell 159, 1312–1326 (2014).
Vento-Tormo, R. et al. Single-cell reconstruction of the early maternal–fetal interface in humans. Nature 563, 347–353 (2018).
Ramilowski, J. A. et al. A draft network of ligand–receptor-mediated multicellular signalling in human. Nat. Commun. 6, 7866 (2015).
Acknowledgements
We thank the Core Facility Flow Cytometry of the Medical University of Vienna for cell sorting service; the Biomedical Sequencing Facility at CeMM for assistance with next-generation sequencing; S. Zahalka and S. Knapp for help and advice with the preparation of lung samples; S. Niggemeyer, J. Riede, S. Jungwirth and N. Fleischmann for animal caretaking; and all members of the Bock laboratory for their help and advice. This work was conducted in the context of two Austrian Science Fund (FWF) Special Research Programme grants (FWF SFB F6102; FWF SFB F7001). T.K. is supported by a Lise Meitner fellowship from the Austrian Science Fund (FWF M2403). N.F. is supported by a fellowship from the European Molecular Biology Organization (EMBO ALTF 241-2017). A.L. is supported by a DOC fellowship of the Austrian Academy of Sciences. A.B. is supported by an ERC Starting Grant (European Union’s Horizon 2020 research and innovation programme, grant agreement no. 677006). C.B. is supported by a New Frontiers Group award of the Austrian Academy of Sciences and by an ERC Starting Grant (European Union’s Horizon 2020 research and innovation programme, grant agreement no. 679146).
Author information
Authors and Affiliations
Contributions
T.K. designed the project, performed experiments, analysed data and cowrote the manuscript. N.F. designed and performed the bioinformatic analysis and cowrote the manuscript. V.F.-G., M.S., L.C.S., A.N. and C.S. contributed to sample collection and sequencing library preparation. A.L. contributed to the experimental design and performed in vivo experiments (LCMV infections; cytokine treatments). A.F.R. contributed bioinformatic software. A.B. contributed to the experimental design and supervised the in vivo experiments. C.B. supervised the project and cowrote the manuscript. All authors read, contributed to, and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature thanks Ari M. Melnick and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Standardized identification and purification of structural cells across 12 organs.
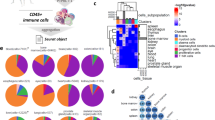
a, Cell-type identification and cell-sorting scheme (top row) with representative flow cytometry plots (selected from n = 4 independent biological replicates) in one representative organ (brain) under homeostatic conditions (bottom row). b, Representative plots (selected from n = 4 independent biological replicates) for gating steps 4–6 of the standardized cell-type identification and cell-sorting scheme (a) across the 12 organs under homeostatic conditions. c, Relative frequencies of structural cell types among non-haematopoietic (CD45−) cells across 12 organs, for cell suspensions obtained by standardized organ dissociation. d, Relative frequencies of structural cell types among non-haematopoietic (CD45−) cells across three organs, for cell suspensions obtained by either standardized organ dissociation or organ-specific dissociation protocols. Shown are mean and s.e.m. values. Sample size: n = 4 (c) and n = 3 (d) independent biological replicates.
Extended Data Fig. 2 Surface marker profiling of structural cells under homeostatic conditions.
a, Gating strategy for the flow cytometry-based validation of the structural cell sorting scheme. Identification of structural cells starts with gating for intact cells (1), single cells (2), live cells (3) and non-haematopoietic cells (4). From the resulting non-haematopoietic (CD45−) cell population, potential epithelial cells (5.1) are gated for epithelial cell markers (5.2). Similarly, potential endothelial cells and fibroblasts (6.1, 6.2) are gated for endothelial cell markers (6.3) and fibroblast markers (6.4). b, Relative frequencies of potential structural cell types based on gates 5.2, 6.3 and 6.4 (from a), comparing the selected markers with alternative markers. c, Expression of the selected surface markers of structural cell types (top row) and potential alternative markers for cells gated as in Extended Data Fig. 1a. Shown are mean and s.e.m. values. Sample size (all panels): n = 3 independent biological replicates.
Extended Data Fig. 3 Comparison of the structural cell transcriptomes to published reference data.
a, Overlap of the identified cell-type-specific and organ-specific marker genes (derived from the RNA-seq experiments in the current study) with tissue-specific gene sets from a microarray-based expression atlas (two-sided Fisher’s exact test with multiple-testing correction). b, Gene expression across cell types and organs (from the current study) aggregated across marker genes of structural cell clusters in a single-cell RNA-seq atlas of the mouse19. c, Gene expression across cell types and organs (from the current study) plotted for a manually curated list of commonly used markers of structural cells. d, Hierarchical clustering of structural cells across cell types and organs based on the transcriptome profiles from the current study. Sample size (all panels): n = 3 independent biological replicates.
Extended Data Fig. 4 Inference of cell–cell interactions across cell types and organs.
a, Enrichment analysis for potential cell–cell interactions between structural cells and haematopoietic immune cells, based on gene expression of known receptor–ligand pairs (two-sided Fisher’s exact test with multiple-testing correction). For each combination of one structural cell type and one haematopoietic immune cell type, the analysis assesses whether all pairs of marker genes between the two cell types are enriched for annotated receptor–ligand pairs. b, Differently expressed genes across cell types and organs, based on a manually curated list of receptors and ligands (Supplementary Table 4). Sample size (all panels): n = 3 independent biological replicates.
Extended Data Fig. 5 Analysis of immune gene expression among structural cells in an independent dataset.
a, Relative frequencies of single-cell transcriptomes classified as endothelium, epithelium and fibroblasts in selected organs according to the Tabula Muris dataset20. b, Expression of immune gene signatures in structural cells according to the Tabula Muris dataset, jointly normalized across all plots (for comparability). c, Expression of immune gene signatures in haematopoietic immune cells according to the Tabula Muris dataset, normalized as in b. d, Expression of selected immune genes in structural cells and in haematopoietic immune cells according to the Tabula Muris dataset. Sample size: n = 7 (all panels) independent biological replicates, comprising 4 male and 3 female mice.
Extended Data Fig. 6 Analysis of transcription regulation in structural cells.
a, Multidimensional scaling analysis of the similarity of chromatin profiles across cell types, organs and replicates based on ATAC-seq (top) and H3K4me2 ChIPmentation (bottom). b, Correlation of chromatin profiles across cell types and organs for ATAC-seq (left) and H3K4me2 ChIPmentation (right). c, Transcriptional regulators of the inferred gene-regulatory network for structural cells, arranged by similarity using multidimensional scaling. d, Motif enrichment for transcriptional regulators among differential chromatin peaks, shown separately for each regulator (one-sided hypergeometric test with multiple-testing correction). e, Gene expression of the transcriptional regulators across cell types and organs (genes discussed in the text are in bold). Sample size (all panels): n = 2 independent biological replicates.
Extended Data Fig. 7 Detection and analysis of genes with unrealized epigenetic potential.
a, Scatterplot showing the correlation between chromatin accessibility in promoter regions and the corresponding gene expression levels in structural cells across cell types and organs. Genes with significant unrealized epigenetic potential (calculated as the difference between normalized ATAC-seq and RNA-seq signals) are highlighted in blue. b, Enrichment of immune-related gene sets among the genes with unrealized epigenetic potential (two-sided Fisher’s exact test with multiple-testing correction). Sample size (all panels): n = 2 independent biological replicates.
Extended Data Fig. 8 Standardized identification and purification of structural cells after LCMV infection.
a, Cell-type identification and cell-sorting scheme (top row) with representative flow cytometry plots (selected from n = 3 independent biological replicates) in one representative organ (brain) after LCMV infection (bottom row). b, Representative plots (selected from n = 3 independent biological replicates) for gating steps 4–6 of the standardized cell-type identification and cell-sorting scheme (a) across the 12 organs after LCMV infection. c, Change in the relative frequency of structural cells after LCMV infection. Sample size (all panels): n = 3 independent biological replicates.
Extended Data Fig. 9 Analysis of differential gene expression in response to LCMV infection.
a, Number of differentially expressed genes in structural cells after LCMV infection (this includes not only immune genes but also genes associated with the substantial organ-specific tissue damage and other direct and indirect effects of LCMV infection). b, Correlation of the observed changes in gene expression after LCMV infection across cell types and organs. c, Organ-specific viral load at day 8 of LCMV infection, measured by qPCR in whole-tissue samples collected from each organ (without FACS purification of individual cell types). Five reference genes were used for normalization and results were ranked across organs, to make the analysis robust towards tissue-specific differences in the expression of these housekeeping genes. However, the experimental results do not support an absolute quantification of viral load in each organ nor do they account for differences in the relative frequencies of cells that are susceptible to LCMV infection across organs. d, Scatterplot illustrating the low correlation between the activated epigenetic potential and the measured viral load across cell types and organs. e, f, Network analysis (e) and enrichment analysis (f) of potential cell–cell interactions between structural cells and haematopoietic immune cells, inferred from gene expression of known receptor–ligand pairs after LCMV infection (two-sided Fisher’s exact test with multiple-testing correction). For each combination of one structural cell type and one haematopoietic immune cell type, the analysis assesses whether all pairs of marker genes between the two cell types are enriched for annotated receptor–ligand pairs. Sample size (all panels): n = 3 independent biological replicates.
Extended Data Fig. 10 Visualization of differential gene expression in response to in vivo cytokine treatments.
The heat map displays changes in the expression of genes associated with immune functions, plotted across cell types, organs and cytokines (two-sided linear model with multiple-testing correction). Sample size: n = 3 independent biological replicates.
Extended Data Fig. 11 Analysis of differential gene expression in response to in vivo cytokine treatments.
a, Number of differentially expressed genes in response to the individual cytokine treatments. b, Gene expression for the known receptors involved in the response to the individual cytokine treatments, plotted across cell types and organs under homeostatic conditions. Sample size (all panels): n = 3 independent biological replicates.
Supplementary information
Supplementary Table 1
Sequencing statistics for RNA-seq, ATAC-seq and H3K4me2 ChIPmentation.
Supplementary Table 2
Cell-type-specific and organ-specific marker genes based on RNA-seq.
Supplementary Table 3
List of receptor-ligand pairs used to construct the cell-cell interaction network under homeostatic conditions.
Supplementary Table 4
Curated list of immune-related receptors and ligands.
Supplementary Table 5
Frequencies of structural cell types per organ in the Tabula Muris dataset.
Supplementary Table 6
Cell-type-specific and organ-specific marker peaks based on ATAC-seq.
Supplementary Table 7
List of genes with unrealized epigenetic potential in structural cells under homeostatic conditions.
Supplementary Table 8
Curated list of immune-related gene sets used for enrichment analysis.
Supplementary Table 9
List of differentially expressed genes upon LCMV infection.
Supplementary Table 10
List of receptor-ligand pairs used to construct the cell-cell interaction network upon LCMV infection.
Supplementary Table 11
List of differentially expressed genes upon cytokine treatment.
Source data
Rights and permissions
About this article
Cite this article
Krausgruber, T., Fortelny, N., Fife-Gernedl, V. et al. Structural cells are key regulators of organ-specific immune responses. Nature 583, 296–302 (2020). https://doi.org/10.1038/s41586-020-2424-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-020-2424-4
This article is cited by
-
The immunoregulatory roles of non-haematopoietic cells in the kidney
Nature Reviews Nephrology (2024)
-
Mucosal T-cell responses to chronic viral infections: Implications for vaccine design
Cellular & Molecular Immunology (2024)
-
Advance in Multi-omics Research Strategies on Cholesterol Metabolism in Psoriasis
Inflammation (2024)
-
Ecto-5ʹ-nucleotidase (CD73): an emerging role as prognostic factor in allergic sensitization
Inflammation Research (2024)
-
AMPK signaling inhibits the differentiation of myofibroblasts: impact on age-related tissue fibrosis and degeneration
Biogerontology (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.