Abstract
The D2 dopamine receptor (DRD2) is a therapeutic target for Parkinson’s disease1 and antipsychotic drugs2. DRD2 is activated by the endogenous neurotransmitter dopamine and synthetic agonist drugs such as bromocriptine3, leading to stimulation of Gi and inhibition of adenylyl cyclase. Here we used cryo-electron microscopy to elucidate the structure of an agonist-bound activated DRD2–Gi complex reconstituted into a phospholipid membrane. The extracellular ligand-binding site of DRD2 is remodelled in response to agonist binding, with conformational changes in extracellular loop 2, transmembrane domain 5 (TM5), TM6 and TM7, propagating to opening of the intracellular Gi-binding site. The DRD2–Gi structure represents, to our knowledge, the first experimental model of a G-protein-coupled receptor–G-protein complex embedded in a phospholipid bilayer, which serves as a benchmark to validate the interactions seen in previous detergent-bound structures. The structure also reveals interactions that are unique to the membrane-embedded complex, including helix 8 burial in the inner leaflet, ordered lysine and arginine side chains in the membrane interfacial regions, and lipid anchoring of the G protein in the membrane. Our model of the activated DRD2 will help to inform the design of subtype-selective DRD2 ligands for multiple human central nervous system disorders.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
Structural data have been deposited in the PDB with coordinate accession number 6vms, and maps have been deposited in the Electron Microscopy Data Bank (EMDB) with accession numbers EMD-21243, EMD-21244 and EMD-21245. All other data generated or analysed during this study are included in this published article or are available from the corresponding authors on reasonable request.
References
Sibley, D. R. & Monsma, F. J. Jr. Molecular biology of dopamine receptors. Trends Pharmacol. Sci. 13, 61–69 (1992).
Seeman, P. Targeting the dopamine D2 receptor in schizophrenia. Expert Opin. Ther. Targets 10, 515–531 (2006).
Michael Besser, G., Pfeiffer, R. F. & Thorner, M. O. Anniversary review: 50 years since the discovery of bromocriptine. Eur. J. Endocrinol. 179, R69–R75 (2018).
Dickson, D. W. et al. Neuropathological assessment of Parkinson’s disease: refining the diagnostic criteria. Lancet Neurol. 8, 1150–1157 (2009).
Seo, E.-J., Sugimoto, Y., Greten, H. J. & Efferth, T. Repurposing of bromocriptine for cancer therapy. Front. Pharmacol. 9, 1030 (2018).
Madras, B. K. History of the discovery of the antipsychotic dopamine D2 receptor: a basis for the dopamine hypothesis of schizophrenia. J. Hist. Neurosci. 22, 62–78 (2013).
Civelli, O., Bunzow, J. R. & Grandy, D. K. Molecular diversity of the dopamine receptors. Annu. Rev. Pharmacol. Toxicol. 33, 281–307 (1993).
Moritz, A. E., Free, R. B. & Sibley, D. R. Advances and challenges in the search for D2 and D3 dopamine receptor-selective compounds. Cell. Signal. 41, 75–81 (2018).
Wang, S. et al. Structure of the D2 dopamine receptor bound to the atypical antipsychotic drug risperidone. Nature 555, 269–273 (2018).
Chien, E. Y. T. et al. Structure of the human dopamine D3 receptor in complex with a D2/D3 selective antagonist. Science 330, 1091–1095 (2010).
Wang, S. et al. D4 dopamine receptor high-resolution structures enable the discovery of selective agonists. Science 358, 381–386 (2017).
Koehl, A. et al. Structure of the µ-opioid receptor–Gi protein complex. Nature 558, 547–552 (2018).
Krishna Kumar, K. et al. Structure of a signaling cannabinoid receptor 1–G protein complex. Cell 176, 448–458.e12 (2019).
Senogles, S. E., Amlaiky, N., Falardeau, P. & Caron, M. G. Purification and characterization of the D2-dopamine receptor from bovine anterior pituitary. J. Biol. Chem. 263, 18996–19002 (1988).
Whorton, M. R. et al. A monomeric G protein-coupled receptor isolated in a high-density lipoprotein particle efficiently activates its G protein. Proc. Natl Acad. Sci. USA 104, 7682–7687 (2007).
Whorton, M. R. et al. Efficient coupling of transducin to monomeric rhodopsin in a phospholipid bilayer. J. Biol. Chem. 283, 4387–4394 (2008).
Bayburt, T. H. & Sligar, S. G. Membrane protein assembly into nanodiscs. FEBS Lett. 584, 1721–1727 (2010).
Gao, Y., Cao, E., Julius, D. & Cheng, Y. TRPV1 structures in nanodiscs reveal mechanisms of ligand and lipid action. Nature 534, 347–351 (2016).
Koehl, A. et al. Structural insights into the activation of metabotropic glutamate receptors. Nature 566, 79–84 (2019).
Staus, D. P. et al. Structure of the M2 muscarinic receptor–β-arrestin complex in a lipid nanodisc. Nature 579, 297–302 (2020).
Westfield, G. H. et al. Structural flexibility of the Gαs α-helical domain in the β2-adrenoceptor Gs complex. Proc. Natl Acad. Sci. USA 108, 16086–16091 (2011).
Seeman, P. & Van Tol, H. H. Dopamine receptor pharmacology. Trends Pharmacol. Sci. 15, 264–270 (1994).
Denisov, I. G., Grinkova, Y. V., Lazarides, A. A. & Sligar, S. G. Directed self-assembly of monodisperse phospholipid bilayer nanodiscs with controlled size. J. Am. Chem. Soc. 126, 3477–3487 (2004).
Rosenbaum, D. M. et al. GPCR engineering yields high-resolution structural insights into β2-adrenergic receptor function. Science 318, 1266–1273 (2007).
Mansour, A. et al. Site-directed mutagenesis of the human dopamine D2 receptor. Eur. J. Pharmacol. 227, 205–214 (1992).
Wiens, B. L., Nelson, C. S. & Neve, K. A. Contribution of serine residues to constitutive and agonist-induced signaling via the D2S dopamine receptor: evidence for multiple, agonist-specific active conformations. Mol. Pharmacol. 54, 435–444 (1998).
Cho, W., Taylor, L. P., Mansour, A. & Akil, H. Hydrophobic residues of the D2 dopamine receptor are important for binding and signal transduction. J. Neurochem. 65, 2105–2115 (1995).
Shi, L. & Javitch, J. A. The second extracellular loop of the dopamine D2 receptor lines the binding-site crevice. Proc. Natl Acad. Sci. USA 101, 440–445 (2004).
Goldstein, M. et al. Interaction of pergolide with central dopaminergic receptors. Proc. Natl Acad. Sci. USA 77, 3725–3728 (1980).
Sibley, D. R. & Creese, I. Interactions of ergot alkaloids with anterior pituitary D-2 dopamine receptors. Mol. Pharmacol. 23, 585–593 (1983).
DeVree, B. T. et al. Allosteric coupling from G protein to the agonist-binding pocket in GPCRs. Nature 535, 182–186 (2016).
McCorvy, J. D. et al. Structure-inspired design of β-arrestin-biased ligands for aminergic GPCRs. Nat. Chem. Biol. 14, 126–134 (2018).
Sanchez-Soto, M. et al. A structural basis for how ligand binding site changes can allosterically regulate GPCR signaling and engender functional selectivity. Sci. Signal. 13, eaaw5885 (2020).
Latorraca, N. R., Venkatakrishnan, A. J. & Dror, R. O. GPCR dynamics: structures in motion. Chem. Rev. 117, 139–155 (2017).
Rasmussen, S. G. F. et al. Crystal structure of the β2 adrenergic receptor–Gs protein complex. Nature 477, 549–555 (2011).
Wall, M. A. et al. The structure of the G protein heterotrimer Giα1β1γ2. Cell 83, 1047–1058 (1995).
Kang, Y. et al. Cryo-EM structure of human rhodopsin bound to an inhibitory G protein. Nature 558, 553–558 (2018).
Mumby, S. M., Heukeroth, R. O., Gordon, J. I. & Gilman, A. G. G-protein alpha-subunit expression, myristoylation, and membrane association in COS cells. Proc. Natl Acad. Sci. USA 87, 728–732 (1990).
Mumby, S. M., Kleuss, C. & Gilman, A. G. Receptor regulation of G-protein palmitoylation. Proc. Natl Acad. Sci. USA 91, 2800–2804 (1994).
Simonds, W. F., Butrynski, J. E., Gautam, N., Unson, C. G. & Spiegel, A. M. G-protein βγ dimers. Membrane targeting requires subunit coexpression and intact γ C-A-A-X domain. J. Biol. Chem. 266, 5363–5366 (1991).
Gallego, C., Gupta, S. K., Winitz, S., Eisfelder, B. J. & Johnson, G. L. Myristoylation of the Gαi2 polypeptide, a G protein α subunit, is required for its signaling and transformation functions. Proc. Natl Acad. Sci. USA 89, 9695–9699 (1992).
Sensoy, O. & Weinstein, H. A mechanistic role of helix 8 in GPCRs: computational modeling of the dopamine D2 receptor interaction with the GIPC1-PDZ-domain. Biochim. Biophys. Acta 1848, 976–983 (2015).
Loisel, T. P. et al. Activation of the β2-adrenergic receptor-Gαs complex leads to rapid depalmitoylation and inhibition of repalmitoylation of both the receptor and Gαs. J. Biol. Chem. 274, 31014–31019 (1999).
Krishna, A. G., Menon, S. T., Terry, T. J. & Sakmar, T. P. Evidence that helix 8 of rhodopsin acts as a membrane-dependent conformational switch. Biochemistry 41, 8298–8309 (2002).
von Heijne, G. The distribution of positively charged residues in bacterial inner membrane proteins correlates with the trans-membrane topology. EMBO J. 5, 3021–3027 (1986).
Dorairaj, S. & Allen, T. W. On the thermodynamic stability of a charged arginine side chain in a transmembrane helix. Proc. Natl Acad. Sci. USA 104, 4943–4948 (2007).
Feng, X., Ambia, J., Chen, K. M., Young, M. & Barth, P. Computational design of ligand-binding membrane receptors with high selectivity. Nat. Chem. Biol. 13, 715–723 (2017).
Chen, K. M., Keri, D. & Barth, P. Computational design of G protein-coupled receptor allosteric signal transductions. Nat. Chem. Biol. 16, 77–86 (2020).
Cleator, J. H., Mehta, N. D., Kurtz, D. T. & Hildebrandt, J. D. The N54 mutant of Gαs has a conditional dominant negative phenotype which suppresses hormone-stimulated but not basal cAMP levels. FEBS Lett. 443, 205–208 (1999).
Lee, E., Taussig, R. & Gilman, A. G. The G226A mutant of Gs alpha highlights the requirement for dissociation of G protein subunits. J. Biol. Chem. 267, 1212–1218 (1992).
Iiri, T., Bell, S. M., Baranski, T. J., Fujita, T. & Bourne, H. R. A Gsα mutant designed to inhibit receptor signaling through Gs. Proc. Natl Acad. Sci. USA 96, 499–504 (1999).
Draper-Joyce, C. J. et al. Structure of the adenosine-bound human adenosine A1 receptor–Gi complex. Nature 558, 559–563 (2018).
Danev, R. & Baumeister, W. Cryo-EM single particle analysis with the Volta phase plate. eLife 5, e13046 (2016).
Mastronarde, D. N. Automated electron microscope tomography using robust prediction of specimen movements. J. Struct. Biol. 152, 36–51 (2005).
Scheres, S. H. W. RELION: implementation of a Bayesian approach to cryo-EM structure determination. J. Struct. Biol. 180, 519–530 (2012).
Zivanov, J. et al. New tools for automated high-resolution cryo-EM structure determination in RELION-3. eLife 7, e42166 (2018).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Zhang, K. Gctf: real-time CTF determination and correction. J. Struct. Biol. 193, 1–12 (2016).
Scheres, S. H. W. & Chen, S. Prevention of overfitting in cryo-EM structure determination. Nat. Methods 9, 853–854 (2012).
Bai, X.-C., Rajendra, E., Yang, G., Shi, Y. & Scheres, S. H. W. Sampling the conformational space of the catalytic subunit of human γ-secretase. eLife 4, e11182 (2015).
Pettersen, E. F. et al. UCSF Chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Cowtan, K. Fitting molecular fragments into electron density. Acta Crystallogr. D 64, 83–89 (2008).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D 66, 486–501 (2010).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D 66, 213–221 (2010).
Davis, I. W. et al. MolProbity: all-atom contacts and structure validation for proteins and nucleic acids. Nucleic Acids Res. 35, W375–W383 (2007).
Acknowledgements
Cryo-EM data were collected at the University of Texas Southwestern Medical Center Cryo-EM Facility, which is funded by the CPRIT Core Facility Support Award RP170644. We thank E. Ross for providing plasmids for G-protein expression. This project was supported by the Edward Mallinckrodt, Jr. Foundation (Scholar Award to D.M.R.), the Welch Foundation (grant I-1770 to D.M.R. and grant I-1944-20180324 to X.-c.B.), the EPFL (to P.B.), the Swiss National Science Foundation (grant 31003A_182263 to P.B.), the Ludwig Institute for Cancer research (to P.B.), the Virginia Murchison Linthicum Scholar in Medical Research at University of Texas Southwestern (to X.-c.B.), CPRIT (RR160082 to X.-c.B.) and the US National Institutes of Health (R01-GM097207 to P.B. and R01-GM083118 to R.K.S.).
Author information
Authors and Affiliations
Contributions
J.Y. expressed and purified the DRD2–Gi complex, collected cryo-EM data and carried out the cryo-EM reconstructions. K.-Y.M.C. carried out the computational design calculations on DRD2 mutants and performed in vitro experimental characterization of the designed DRD2 constructs. M.J.C. collected functional data on DRD2 constructs. M.H. carried out the molecular dynamics simulations of DRD2. P.K. assisted with DRD2 purification and testing of mutants. X.-c.B. supervised cryo-EM data collection and analysis. R.K.S. supervised the collection of functional data on DRD2 constructs and helped to write the manuscript. P.B. developed the strategy for computational design of a stabilized DRD2 active-state construct and supervised the computational design calculations. D.M.R. supervised the overall project, assisted with cryo-EM data analysis and wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
P.B. is an inventor on a patent application (EP19189259.5) submitted by Ecole Polytechnique Fédérale de Lausanne on the design methods and designed protein variants herein. All other authors declare no competing interests.
Additional information
Peer review information Nature thanks R. Benjamin Free, Amy Moritz, David Sibley and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Rational design of receptor variants shifted in the active state.
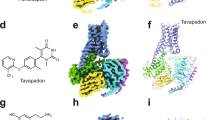
a, Energy landscape of three receptor variants with various energy differences in the resting (ligand-free) state ΔGresting, in the allosteric coupling controlling the inverse agonist efficacy ΔGinverse agonist and the stabilization of the receptor inactive state upon ligand stimulus. b, Active state occupancy as a function of ΔGresting and ΔGinverse agonist (the energy of inverse agonist binding), as determined by the Boltzmann law. c, Sensitivity of the receptors to inverse agonist binding. d–f, Rational design of DRD2 stabilized in the active state. DRD2 (D2) wild-type active state homology model (d). The 120-residue-long ICL3 is represented as a dotted blue line. DRD2 designed with residue microswitches shown in spheres selectively stabilizing the active state structure (e). DRD2 further stabilized in the active state through a de novo designed seven-residue stable ICL3 (f).
Extended Data Fig. 2 Effect of T4L, modified ICL3 and scFv16 on hDRD2WT ligand-binding properties.
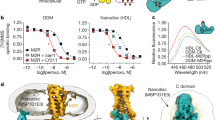
a, Competition binding of dopamine for [3H]spiperone on membranes containing hDRD2WT (blue) or hDRD2 modified with nT4L and de novo-designed ICL3 (nT4L-hDRD2(ΔICL3); magenta). The log-transformed inhibition constant (Ki) ± s.e.m. = −4.6 ± 0.04 for hDRD2WT. log(Ki ± s.e.m.) = −4.6 ± 0.04 for nT4L-hDRD2(ΔICL3). n = 4 biologically independent experiments on Sf9 cell membranes. b, Agonist-stimulated [35S]GTPγS binding of nT4L-hDRD2(ΔICL3) (magenta) compared to hDRD2WT (blue). Dose–responses using dopamine (closed magenta circles) or bromocriptine (BCT; open magenta circles) show log(EC50 ± s.e.m.) = −7.5 ± 0.02 and −7.8 ± 0.06, respectively, for nT4L-hDRD2(ΔICL3) compared to log(EC50 ± s.e.m.) = −7.0 ± 0.02 for dopamine on hDRD2WT (open blue circles). n = 3 biologically independent experiments on HEK293T cell membranes. c, Saturation binding of [3H]spiperone to membranes containing hDRD2WT with or without scFv16. Dissociation constant (Kd) ± s.e.m. = 0.15 ± 0.03 nM for control. Kd ± s.e.m. = 0.14 ± 0.02 nM with scFv16. n = 4 biologically independent experiments on Sf9 cell membranes. d, Competition binding of dopamine for [3H]spiperone on membranes containing DRD2WT with or without scFv16. log(Khigh ± s.e.m.) = −6.4 ± 0.3 and log(Klow ± s.e.m.) = −4.3 ± 0.08 for control. log(Khigh ± s.e.m.) = −6.5 ± 0.2 and log(Klow ± s.e.m.) = −4.3 ± 0.07 with scFv16. n = 4 biologically independent experiments on Sf9 cell membranes. In all panels, each data point is displayed as the mean with the error bars showing ± s.e.m., with individual data points for each repeat represented in small symbols.
Extended Data Fig. 3 Function and thermostability of hDRD2EM.
a, Gi stimulation by dopamine in Sf9 cells transformed with Gi and hDRD2 baculoviruses. log(EC50 ± s.e.m.) = −6.3 ± 0.1 for hDRD2WT (blue). log(EC50 ± s.e.m.) = −6.6 ± 0.3 for hDRD2EM (orange). n = 4 biologically independent experiments on Sf9 cell membranes. b, Inhibition of dopamine-stimulated (10 μM) Gi activation by the inverse agonist spiperone. log( IC50 ± s.e.) = −8.6 ± 0.1 for hDRD2WT (blue). log(IC50 ± s.e.m.) = −6.3 ± 0.2 for hDRD2EM (orange). n = 4 biologically independent experiments on Sf9 cell membranes. c, Modulation of basal Gi activation by dopamine (open circles), BCT (closed circles) or spiperone (squares) with the hDRD2EM (orange symbols) relative to hDRD2WT (blue symbols). log(EC50 ± s.e.m.) for dopamine (−6.6 ± 0.2), BCT (−7.6 ± 0.1) and spiperone (−7.6 ± 0.2) for hDRD2EM. log(EC50 ± s.e.m.) for dopamine (−7.0 ± 0.1) and BCT (−7.8 ± 0.2) for hDRD2WT. n = 3 biologically independent experiments on Sf9 cell membranes. d, Apparent functional stability of agonist-bound hDRD2 constructs assessed by measuring the fraction of partially purified receptor-activating Gi as a function of incubation time at 37 °C. Half-life ± s.e.m. = 27.6 ± 1.0 min for hDRD2WT (blue). Half-life ± s.e.m. = 49.3 ± 1.1 min for hDRD2 with five designed mutations and designed truncated ICL3 (hDRD2(ΔICL3-5mut); orange). n = 3 biologically independent experiments. e, Apparent thermostability of agonist-bound hDRD2 constructs assessed by measuring the fraction of partially purified receptor-binding agonist as a function of temperature. Melting temperature (Tm) ± s.e.m. = 30.9 ± 0.9 °C for hDRD2WT (blue). Tm ± s.e.m. = 39.6 ± 0.3 °C for hDRD2(ΔICL3-5mut) (orange). n = 3 biologically independent experiments. In all panels, each data point is displayed as the mean with the error bars showing ± s.e.m., with individual data points for each repeat represented in small symbols.
Extended Data Fig. 4 Purification and reconstitution of the hDRD2EM–Gi complex.
a, Superose 6 gel-filtration profile of hDRD2EM–Gi-scFv16-HDL purified by M1 Flag affinity chromatography. b, Coomassie-stained PAGE of the isolated peak fraction from gel filtration. c, Overlaid Superose 200 gel-filtration profiles (in LMNG detergent) of different hDRD2 constructs purified by M1 Flag chromatography with saturating bromocriptine present. nT4L-hDRD2-5mut has the five thermostabilizing mutations of the EM construct, but has full-length ICL3. nT4L-hDRD2-ΔICL3 has the wild-type transmembrane region (no mutations) but the truncated ICL3. Receptors were purified as in Methods, but in the absence of co-transduced G-protein baculovirus.
Extended Data Fig. 5 Cryo-EM data analysis.
a, Representative 2D class averages. Scale bar, 100 Å. b, Gold-standard Fourier shell correlation (FSC) curves of the 3D reconstructions. The dashed lines intercept the y axis at a FSC value of 0.143. c, Image processing procedure with the final maps coloured based on local resolution. Ctf, contrast transfer function.
Extended Data Fig. 6 3D classification with signal subtraction, model validation and representative map density.
a, Focused 3D classification subtracting T4L, the AH domain, scFv16 and rHDL/nanodisc density. b, Focused 3D classification subtracting all but the G protein. In a and b, the numbers below the images indicate the particle number in each 3D class. c, FSC curve showing the model–map correlation (focused map without T4L, the AH domain, scFv16 and rHDL/nanodisc density). The dashed line intercepts the y axis at a FSC value of 0.5. d, Representative hDRD2 transmembrane and ligand density.
Extended Data Fig. 7 Surfaces of the orthosteric pocket in the active and inactive conformations of hDRD2.
a, Solvent-accessible surface for the active conformation (purple) bound to bromocriptine (yellow sticks). b, Solvent-accessible surface for the inactive conformation (orange) bound to risperidone (green sticks), showing the deeper subpocket (PDB: 6cm4).
Extended Data Fig. 8 Interaction stabilizing the designed hDRD2EM active state conformation.
a, Zoomed view of the TM6–TM7 interface on the cytoplasmic side of the receptor. Atomic interactions involving the designed Y6.40 were accurately predicted in the design model. The conformation of R3.50 is displaced in the experimental structure due to the binding of Gαi, which was absent in the model. The experimental structure is in purple and the predicted DRD2 conformation is in green. b, c, Conformational energy landscape of the wild-type (b) and designed (c) DRD2 constructed from multiple all-atom MD simulation trajectories of the receptor active state homology models (orange circle) in the absence of bound agonist. The conformational space is reported along two canonical active state structure metrics (TM3–TM6 distance and NPxxY rmsd to inactive state). The DRD2 wild-type relaxes primarily due to an inactive state conformation, whereas the designed variant remains in partially active state conformation displaying large TM3–TM6 distance. The antagonist-bound DRD2 crystal structure (PDB: 6CM4) is used as a reference for the inactive state (green square).
Extended Data Fig. 9 AH domain density and comparison with rhodopsin–Gi.
a, AH domain density in the current structure. b, AH domain density in the rhodopsin–Gi complex (PDB: 6cmo). c, Alignment of the Gαi subunit in the hDRD2EM–Gi (purple) and the rhodopsin–Gi (blue) complexes. d, Alignment of the AH domain in the hDRD2EM–Gi (purple) and the rhodopsin–Gi (blue) complexes.
Extended Data Fig. 10 Electrostatic surfaces in the hDRD2EM–Gi complex.
a, Surface potential of the hDRD2 (top) and Gi (bottom) components shown separately. b, Surface potential of the complex, with the scale in eV. Calculation was done using the APBS plugin in PyMOL.
Supplementary information
Rights and permissions
About this article
Cite this article
Yin, J., Chen, KY.M., Clark, M.J. et al. Structure of a D2 dopamine receptor–G-protein complex in a lipid membrane. Nature 584, 125–129 (2020). https://doi.org/10.1038/s41586-020-2379-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-020-2379-5
This article is cited by
-
Cryo-EM structure of cell-free synthesized human histamine 2 receptor/Gs complex in nanodisc environment
Nature Communications (2024)
-
Improved green and red GRAB sensors for monitoring dopaminergic activity in vivo
Nature Methods (2024)
-
Sequestration of Gβγ by deubiquitinated arrestins into the nucleus as a novel desensitization mechanism of G protein–coupled receptors
Cell Communication and Signaling (2023)
-
Mapping the conformational landscape of the stimulatory heterotrimeric G protein
Nature Structural & Molecular Biology (2023)
-
Orthosteric ligand selectivity and allosteric probe dependence at Hydroxycarboxylic acid receptor HCAR2
Signal Transduction and Targeted Therapy (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.