Abstract
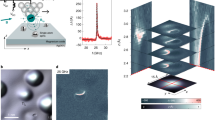
Imaging of biomolecules guides our understanding of their diverse structures and functions1,2. Real-space imaging at sub-nanometre resolution using cryo-electron microscopy has provided key insights into proteins and their assemblies3,4. Direct molecular imaging of glycans—the predominant biopolymers on Earth, with a plethora of structural and biological functions5—has not been possible so far6. The inherent glycan complexity and backbone flexibility require single-molecule approaches for real-space imaging. At present, glycan characterization often relies on a combination of mass spectrometry and nuclear magnetic resonance imaging to provide insights into size, sequence, branching and connectivity, and therefore requires structure reconstruction from indirect information7,8,9. Here we show direct imaging of single glycan molecules that are isolated by mass-selective, soft-landing electrospray ion beam deposition and imaged by low-temperature scanning tunnelling microscopy10. The sub-nanometre resolution of the technique enables the visualization of glycan connectivity and discrimination between regioisomers. Direct glycan imaging is an important step towards a better understanding of the structure of carbohydrates.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
The data that support the findings of this study are available from the authors on reasonable request; see Author contributions for specific datasets.
Change history
15 July 2020
A Correction to this paper has been published: https://doi.org/10.1038/s41586-020-2518-z
References
Unwin, P. N. T. & Henderson, R. Molecular structure determination by electron microscopy of unstained crystalline specimens. J. Mol. Biol. 94, 425–440 (1975).
Blundell T. L. et al. (eds) Celebrating structural biology. Nat. Struct. Mol. Biol. 18, 1304–1316 (2011).
Bai, X.-c., McMullan, G. & Scheres, S. H. W. How cryo-EM is revolutionizing structural biology. Trends Biochem. Sci. 40, 49–57 (2015).
Callaway, E. The revolution will not be crystallized: a new method sweeps through structural biology. Nature 525, 172–174 (2015).
Varki, A. et al. Essentials of Glycobiology 3rd edn (Cold Spring Harbor Laboratory Press, 2017).
Gray, C. J. et al. Advancing solutions to the carbohydrate sequencing challenge. J. Am. Chem. Soc. 141, 14463–14479 (2019).
Mariño, K., Bones, J., Kattla, J. J. & Rudd, P. M. A systematic approach to protein glycosylation analysis: a path through the maze. Nat. Chem. Biol. 6, 713–723 (2010).
Dell, A. & Morris, H. R. Glycoprotein structure determination by mass spectrometry. Science 291, 2351 (2001).
Lundborg, M., Fontana, C. & Widmalm, G. Automatic structure determination of regular polysaccharides based solely on NMR spectroscopy. Biomacromolecules 12, 3851–3855 (2011).
Rauschenbach, S., Ternes, M., Harnau, L. & Kern, K. Mass spectrometry as a preparative tool for the surface science of large molecules. Annu. Rev. Anal. Chem. 9, 473–498 (2016).
Hofmann, J., Hahm, H., Seeberger, P. & Pagel, K. Identification of carbohydrate anomers using ion mobility-mass spectrometry. Nature 526, 241–244 (2015).
Faleh, A. B., Warnke, S. & Rizzo, T. R. Combining ultrahigh-resolution ion-mobility spectrometry with cryogenic infrared spectroscopy for the analysis of glycan mixtures. Anal. Chem. 91, 4876–4882 (2019).
Kley, C. S. et al. Atomic-scale observation of multiconformational binding and energy level alignment of ruthenium-based photosensitizers on TiO2 anatase. Nano Lett. 14, 563–569 (2014).
Deng, Z. et al. A close look at proteins: submolecular resolution of two- and three-dimensionally folded cytochrome c at surfaces. Nano Lett. 12, 2452–2458 (2012).
Longchamp, J.-N. et al. Imaging proteins at the single-molecule level. Proc. Natl Acad. Sci. USA 114, 1474 (2017).
Abb, S., Harnau, L., Gutzler, R., Rauschenbach, S. & Kern, K. Two-dimensional honeycomb network through sequence-controlled self-assembly of oligopeptides. Nat. Commun. 7, 10335 (2016).
Rauschenbach, S. et al. Two-dimensional folding of polypeptides into molecular nanostructures at surfaces. ACS Nano 11, 2420–2427 (2017).
Liu, L. et al. Chaperon-mediated single molecular approach toward modulating Aβ peptide aggregation. Nano Lett. 9, 4066–4072 (2009).
Abb, S. et al. Carbohydrate self-assembly at surfaces: STM imaging of sucrose conformation and ordering on Cu(100). Angew. Chem. Int. Ed. 131, 8424–8428 (2019).
Abb, S. et al. Polymorphism in carbohydrate self-assembly at surfaces: STM imaging and theoretical modelling of trehalose on Cu(100). RSC Adv. 9, 35813–35819 (2019).
Rauschenbach, S. et al. Electrospray ion beam deposition of clusters and biomolecules. Small 2, 540–547 (2006).
Guberman, M. & Seeberger, P. H. Automated glycan assembly: a perspective. J. Am. Chem. Soc. 141, 5581–5592 (2019).
Panza, M., Pistorio, S. G., Stine, K. J. & Demchenko, A. V. Automated chemical oligosaccharide synthesis: novel approach to traditional challenges. Chem. Rev. 118, 8105–8150 (2018).
Brazil, R. Can chemists crack the sugar code? Chemistry World https://www.chemistryworld.com/features/can-chemists-crack-our-cells-sugar-code/4010803.article#/ (2020).
Delbianco, M. et al. Well-defined oligo- and polysaccharides as ideal probes for structural studies. J. Am. Chem. Soc. 140, 5421–5426 (2018).
Acknowledgements
We thank the Max Planck Society and Alexander von Humboldt Foundation for financial support. M.D. thanks the Minerva Fast Track Program and the MPG-FhG Cooperation Project Glyco3Dysplay. The authors acknowledge the Emmy-Noether-Program of the Deutsche Forschungsgemeinschaft.
Author information
Authors and Affiliations
Contributions
S.R., S.A., P.H.S. and K.K. conceived this project. X.W., K.A., S.R., S.A., U.S., K.K., P.H.S. and M.D. designed the experiments. X.W., K.A., T.M., S.S., M.P., U.S. and S.A. carried out the sample preparation and measurements. X.W., S.A. and T.M. analysed the data. M.D., A.P.V. and P.B. carried out the molecule synthesis. All authors contributed to and discussed the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature thanks Steven De Feyter and Sabine Flitsch for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
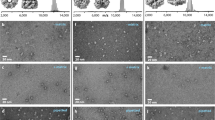
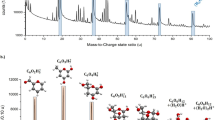
Extended Data Fig. 1 Mass spectra of all deposited glycans.
a–f, The mass spectra show absence of major contaminants in the ion beam after mass filtering. For glycans 1–5, the charge state of the molecule is −1, whereas undecasaccharide 6 has a charge state of −2.
Extended Data Fig. 2 STM overview image of a sample of hexamannoside 4.
Single molecules (marked with yellow circles) are observed as the major species on the terrace. The dark spots are due to impurities on the surface, which is typical for depositions at low temperature. Scale bar, 10 nm.
Extended Data Fig. 3 STM topography image of linear glycans.
a–f, STM images of pentamannoside 1 (a, b), hexamannoside 2 (c, d) and hexaglucoside 3 (e, f) showing different conformations on the surface. Scale bars, 1 nm.
Extended Data Fig. 4 STM topography image of branched hexamannosides.
a, b, STM images of adsorbed hexamannoside 4 showing different conformations. Two branches are clearly visible. c, d, STM images of hexamannoside 5 of different conformations on the surface, with branching clearly visible. The assignment of the linkage is based on the position of the linker (low-height feature) and on the interunit distance, as shown in the inset (1-2 linkage in pink). Scale bars, 1 nm.
Extended Data Fig. 5 Statistical analysis of the feature distance of 1-6 linkages in linear glycans.
a, Pentamannoside 1. b, Hexamannoside 2. c, Hexaglucoside 3. d, Combined histogram for all linear glycans. Values for the mean feature distance of the 1-6 linkages were calculated by fitting with Gaussian peaks, resulting in centre and width values of 0.48 ± 0.09 nm, 0.51 ± 0.08 nm and 0.53 ± 0.04 nm, respectively. The total averaging of the 1-6 linkages (including α1-6 and β1-6) yields a measured 1-6 feature distance of 0.52 ± 0.07 nm. Histogram bin size, 0.05 nm.
Extended Data Fig. 6 Statistical analysis of the feature distance of α1-6 and α1-2 linkages in hexamannoside 4.
The distance between the monosaccharide units connected with α1-6 and α1-2 linkages in hexamannoside 4 was identified on the basis of the linker position and the branching points. The histogram (bin size, 0.02 nm) of the α1-6 (grey) and α1-2 (purple) linkages was fitted with Gaussian peaks at 0.53 ± 0.05 nm and 0.61 ± 0.03 nm, respectively. The distributions show little overlap and thus the feature distance is identified as a reliable parameter for distinguishing between α1-6 and α1-2 linkages. The wide distribution of α1-6 (black) linkages compared to the narrow shape of α1-2 linkages (purple) indicates larger conformational variability, probably due to the greater flexibility of the α1-6 links.
Extended Data Fig. 7 STM topography image of branched undecasaccharide 6.
a, STM images of undecasaccharide 6, clearly showing branching points and individually resolved monosaccharide units. The assignment of the branches is made by the number of units in the branch and by the feature distance between the monosaccharide units. Scale bar, 1 nm. b, Line profiles as indicated in a.
Supplementary information
Supplementary Information
This file contains: (1) General Materials and Methods, (2) Automated Glycan Assembly, and (3) additional References.
Rights and permissions
About this article
Cite this article
Wu, X., Delbianco, M., Anggara, K. et al. Imaging single glycans. Nature 582, 375–378 (2020). https://doi.org/10.1038/s41586-020-2362-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-020-2362-1
This article is cited by
-
Towards glycan foldamers and programmable assemblies
Nature Reviews Materials (2024)
-
Visualizing the structural diversity of glycoconjugates
Nature Chemical Biology (2024)
-
Dinuclear complex-induced DNA melting
Journal of Nanobiotechnology (2023)
-
Soft-landing methods aim to simplify structural biology
Nature (2023)
-
Molecular sensitised probe for amino acid recognition within peptide sequences
Nature Communications (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.