Abstract
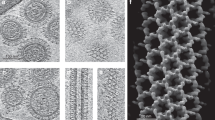
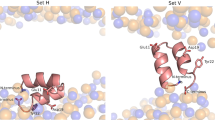
Infection by enveloped viruses involves fusion of their lipid envelopes with cellular membranes to release the viral genome into cells. For HIV, Ebola, influenza and numerous other viruses, envelope glycoproteins bind the infecting virion to cell-surface receptors and mediate membrane fusion. In the case of influenza, the receptor-binding glycoprotein is the haemagglutinin (HA), and following receptor-mediated uptake of the bound virus by endocytosis1, it is the HA that mediates fusion of the virus envelope with the membrane of the endosome2. Each subunit of the trimeric HA consists of two disulfide-linked polypeptides, HA1 and HA2. The larger, virus-membrane-distal, HA1 mediates receptor binding; the smaller, membrane-proximal, HA2 anchors HA in the envelope and contains the fusion peptide, a region that is directly involved in membrane interaction3. The low pH of endosomes activates fusion by facilitating irreversible conformational changes in the glycoprotein. The structures of the initial HA at neutral pH and the final HA at fusion pH have been investigated by electron microscopy4,5 and X-ray crystallography6,7,8. Here, to further study the process of fusion, we incubate HA for different times at pH 5.0 and directly image structural changes using single-particle cryo-electron microscopy. We describe three distinct, previously undescribed forms of HA, most notably a 150 Å-long triple-helical coil of HA2, which may bridge between the viral and endosomal membranes. Comparison of these structures reveals concerted conformational rearrangements through which the HA mediates membrane fusion.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Data availability
Maps and models have been deposited in the Electron Microscopy Data Bank (http://www.ebi.ac.uk/pdbe/emdb/, under accession numbers EMD-10696, EMD-10697, EMD-10698, EMD-10699, EMD-10700, EMD-10701, and EMD-10702). Models have been deposited in the Protein Data Bank (https://www.ebi.ac.uk/pdbe/, under identification codes 6Y5G, 6Y5H, 6Y5I, 6Y5J, 6Y5K and 6Y5L). The raw image for Extended Data Fig. 5c is provided in Supplementary Fig. 1.
References
Cossart, P. & Helenius, A. Endocytosis of viruses and bacteria. Cold Spring Harb. Perspect. Biol. 6, a016972 (2014).
Skehel, J. J. & Wiley, D. C. Receptor binding and membrane fusion in virus entry: the influenza hemagglutinin. Annu. Rev. Biochem. 69, 531–569 (2000).
Brunner, J. Testing topological models for the membrane penetration of the fusion peptide of influenza virus hemagglutinin. FEBS Lett. 257, 369–372 (1989).
Ruigrok, R. W. et al. Electron microscopy of the low pH structure of influenza virus haemagglutinin. EMBO J. 5, 41–49 (1986).
Ruigrok, R. W. H. et al. Studies on the structure of the influenza virus haemagglutinin at the pH of membrane fusion. J. Gen. Virol. 69, 2785–2795 (1988).
Wilson, I. A., Skehel, J. J. & Wiley, D. C. Structure of the haemagglutinin membrane glycoprotein of influenza virus at 3 Å resolution. Nature 289, 366–373 (1981).
Bullough, P. A., Hughson, F. M., Skehel, J. J. & Wiley, D. C. Structure of influenza haemagglutinin at the pH of membrane fusion. Nature 371, 37–43 (1994).
Chen, J., Skehel, J. J. & Wiley, D. C. N- and C-terminal residues combine in the fusion-pH influenza hemagglutinin HA2 subunit to form an N cap that terminates the triple-stranded coiled coil. Proc. Natl Acad. Sci. USA 96, 8967–8972 (1999).
Godley, L. et al. Introduction of intersubunit disulfide bonds in the membrane-distal region of the influenza hemagglutinin abolishes membrane fusion activity. Cell 68, 635–645 (1992).
Barbey-Martin, C. et al. An antibody that prevents the hemagglutinin low pH fusogenic transition. Virology 294, 70–74 (2002).
Watanabe, A. et al. Antibodies to a conserved influenza head interface epitope protect by an IgG subtype-dependent mechanism. Cell 177, 1124–1135.e16 (2019).
Bangaru, S. et al. A site of vulnerability on the influenza virus hemagglutinin head domain trimer interface. Cell 177, 1136–1152 (2019).
Bizebard, T. et al. Structure of influenza virus haemagglutinin complexed with a neutralizing antibody. Nature 376, 92–94 (1995).
Ward, C. W. & Dopheide, T. A. Influenza virus haemagglutinin. Structural predictions suggest that the fibrillar appearance is due to the presence of a coiled-coil. Aust. J. Biol. Sci. 33, 441–447 (1980).
Carr, C. M. & Kim, P. S. A spring-loaded mechanism for the conformational change of influenza hemagglutinin. Cell 73, 823–832 (1993).
Calder, L. J. & Rosenthal, P. B. Cryomicroscopy provides structural snapshots of influenza virus membrane fusion. Nat. Struct. Mol. Biol. 23, 853–858 (2016).
Daniels, R. S. et al. Fusion mutants of the influenza virus hemagglutinin glycoprotein. Cell 40, 431–439 (1985).
Wharton, S. A., Skehel, J. J. & Wiley, D. C. Studies of influenza haemagglutinin-mediated membrane fusion. Virology 149, 27–35 (1986).
Xu, R. & Wilson, I. A. Structural characterization of an early fusion intermediate of influenza virus hemagglutinin. J. Virol. 85, 5172–5182 (2011).
Collins, P. J. et al. Recent evolution of equine influenza and the origin of canine influenza. Proc. Natl Acad. Sci. USA 111, 11175–11180 (2014).
Lorieau, J. L., Louis, J. M., Schwieters, C. D. & Bax, A. pH-triggered, activated-state conformations of the influenza hemagglutinin fusion peptide revealed by NMR. Proc. Natl Acad. Sci. USA 109, 19994–19999 (2012).
Benton, D. J. et al. Influenza hemagglutinin membrane anchor. Proc. Natl Acad. Sci. USA 115, 10112–10117 (2018).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Rohou, A. & Grigorieff, N. CTFFIND4: fast and accurate defocus estimation from electron micrographs. J. Struct. Biol. 192, 216–221 (2015).
Scheres, S. H. W. RELION: implementation of a Bayesian approach to cryo-EM structure determination. J. Struct. Biol. 180, 519–530 (2012).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Wagner, T. et al. SPHIRE-crYOLO is a fast and accurate fully automated particle picker for cryo-EM. Commun. Biol. 2, 218 (2019).
Zivanov, J. et al. New tools for automated high-resolution cryo-EM structure determination in RELION-3. eLife 7, e42166 (2018).
Zivanov, J., Nakane, T. & Scheres, S. H. W. A Bayesian approach to beam-induced motion correction in cryo-EM single-particle analysis. IUCrJ 6, 5–17 (2019).
Ilca, S. L. et al. Multiple liquid crystalline geometries of highly compacted nucleic acid in a dsRNA virus. Nature 570, 252–256 (2019).
Cardone, G., Heymann, J. B. & Steven, A. C. One number does not fit all: mapping local variations in resolution in cryo-EM reconstructions. J. Struct. Biol. 184, 226–236 (2013).
Rosenthal, P. B. & Henderson, R. Optimal determination of particle orientation, absolute hand, and contrast loss in single-particle electron cryomicroscopy. J. Mol. Biol. 333, 721–745 (2003).
Lin, Y. P. et al. Evolution of the receptor binding properties of the influenza A(H3N2) hemagglutinin. Proc. Natl Acad. Sci. USA 109, 21474–21479 (2012).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D 66, 486–501 (2010).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D 66, 213–221 (2010).
Pettersen, E. F. et al. UCSF Chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Rosenthal, P. B. et al. Structure of the haemagglutinin-esterase-fusion glycoprotein of influenza C virus. Nature 396, 92–96 (1998).
Acknowledgements
We thank A. Nans of the Structural Biology Science Technology Platform for assistance with data collection; P. Walker and A. Purkiss of the Structural Biology Science Technology Platform; the Scientific Computing Science Technology Platform for computational support; and L. Calder for discussions. This work was funded by the Francis Crick Institute, which receives its core funding from Cancer Research UK (grant numbers FC001078 and FC001143), the UK Medical Research Council (FC001078 and FC001143), and the Wellcome Trust (FC001078 and FC001143).
Author information
Authors and Affiliations
Contributions
D.J.B carried out research and collected and analysed data. All authors conceived and designed the research and wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature thanks Stephen Harrison, Juha Huiskonen and Yorgo Modis for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Features of intermediate states.
a, Displacement measurements between dilated HA1 domains. Measurements are the position of the centroid of HA1 residues 45–310, with structures aligned on the invariant parts of HA2 (residues 76–98). b, An interaction can be seen between the glycan attached to Asn165 (orange) and Trp222 (green) on the neighbouring HA1 monomer (blue), crosslinking the HA1 domains. This is present in states I, II and III. c, Cryo-EM density (grey) and model, with HA1 in blue and HA2 in red, for the short helix of HA2 (residues 38–55) and the 30-loop (HA1 residues 22–37). Density extends beyond the top of the helix, indicating elongation of this helix by residues from the interhelical loop. d, Example of typical density (grey) for the same section of an HA2 helix from states I and IV (green).
Extended Data Fig. 2 Resolution estimates of obtained cryo-EM maps.
Local-resolution estimates, global-resolution FSC curves and orientation-distribution plots for different conformations of HA. FSC curves have values for FSC of 0.143, shown as the resolution criterion. Reconstruction of state V has a resolution of 9.9 Å and has little variability in local resolution. See Extended Data Fig. 9 for the FSC curve for state V.
Extended Data Fig. 3 Flexibility of the membrane-distal domains.
The membrane-distal regions of HA1 (roughly residues 45–310) can adopt a range of different orientations compared with the C3 symmetrized structure (state IV). The remainder of HA1 and HA2 adopt a structure with less evidence of flexibility. We examined the flexibility in ten maps generated by asymmetric classification. a, The generated models are shown aligned to the symmetrized version of the protein (yellow). b, Examples of the most-populated classes are shown on a molecular surface, with HA1 in blue and HA2 in red to emphasize the different locations of these domains when compared with the symmetrized version. c, The displacements of the locations of the centroids of each of these mobile domains were measured to the nearest symmetrized monomer, giving 30 data points. These adopt a range of locations, with the main flexibility being in a rotation angle around the threefold axis, with little lateral movement towards and away from the threefold axis. The angles of displacement vary from −15° to +25°.
Extended Data Fig. 4 The 30-loop.
a, Potential interactions in the 30-loop are similar in both the neutral-pH (state I) and the extended HA2 (state IV) conformations. There are, however, several changes in the side chains involved in this interaction, owing to the relocation of the short helix of HA2 in the neutral-pH structure. This helix relocation removes an HA2 salt bridge between Arg54 of the short helix and Glu97 of the long helix, as well as the interaction of Lys51 with His106. These rearrangements permit new potential interactions between Thr30 of HA1 with Gln105 of HA2 and His106 of the adjacent HA2 chain. A salt bridge also forms between Lys27 of HA1 and Glu97 of HA2. b, Density of the 30-loop and interacting regions of the long helices of HA2 in the extended subtracted structure. c, The location and architecture of the 30-loop is conserved in all HAs, including influenza C HEF37. In HA, a cluster of conserved hydrophobic-loop residues at positions 26, 34 and 36 packs against the strictly conserved Ile108 in the long α-helix; the strictly conserved Asn104 forms hydrogen bonds with loop residue Lys27 and Lys315 of HA1; and HA2 residue 105, conserved as Gln or Glu, interacts with Thr30. Amino-acid substitutions in the loop (Thr30 to Ser), the short α-helix (Arg54 to Lys) and the long α-helix (Gln105 to Lys and His106 to Ala) that interact with loop residues 28, 29 and 30, respectively, destabilize the mutant HAs, as shown by their elevated fusion pH17,20. These observations indicate the functional importance of the loop and suggest its involvement in membrane fusion. Formation of the 180° turn in the extended helix requires removal of the 30-loop from its interactions with Gln105 and His106. The observed unfolding of the loop and its acquisition of susceptibility to protease digestion in stage V are consistent with this requirement and with the suggested role of the loop in supporting the extended helix in states II, III and IV.
Extended Data Fig. 5 2D class averages and representative micrographs for the 10 s, 20 s and 60 s time points.
a, 2D classes fall into two broad groups: neutral-pH-like and extended-intermediate-like. Representative classes are shown for each group for each time point, with the percentages indicating the overall numbers of particles that fall into these two broad groups. There is a general trend towards increasing numbers of extended-intermediate-like classes as the time point increases, with a decrease in numbers of neutral-pH-like classes. There is limited diversity in the orientations seen for the 10 s extended-intermediate-like and the 60 s neutral-pH-like classes. b, Representative micrographs (scale bars, 50 nm). Numbers of micrographs collected: 10 s, n = 4,004; 20 s, n = 8,728; 60 s, n = 2,925. c, SDS–PAGE of the HA sample used for experiments.
Extended Data Fig. 6 Classification scheme for the 10 s time point.
The figure shows the workflow for image processing to determine structures at 10 s.
Extended Data Fig. 7 Classification scheme for the 20 s time point.
The figure shows the workflow for image processing to determine structures at 20 s.
Extended Data Fig. 8 Classification scheme for the 60 s time point.
The figure shows the workflow for image processing to determine structures at 60 s.
Extended Data Fig. 9 Data processing for 30 min time points.
a, 2D classes from the 30 min time point show the presence of the extended HA2 (state IV) and the post-fusion conformations, as well as a minor population of neutral-pH conformations. The post-fusion averages are interpretable only for the end-on view down the threefold axis, where this conformation forms a characteristic dense ring-like shape. This limited range of interpretable views is probably due to the, now flexibly linked, HA1 domains increasing the structural heterogeneity. The interpretability of this species could be improved by the addition of 10 mM of 2-mercaptoethanol to detach the disulfide-linked HA1 domains. b, Classification scheme for the 30-min time point supplemented with 2-mercaptoethanol. c, FSC curve for the final refined structure, with resolution criterion FSC = 0.143 shown.
Supplementary information
Supplementary Figure
Supplementary Figure 1: Raw Image of SDS-PAGE gel in Extended Data Figure 5C.
Rights and permissions
About this article
Cite this article
Benton, D.J., Gamblin, S.J., Rosenthal, P.B. et al. Structural transitions in influenza haemagglutinin at membrane fusion pH. Nature 583, 150–153 (2020). https://doi.org/10.1038/s41586-020-2333-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-020-2333-6
This article is cited by
-
Immune response in influenza virus infection and modulation of immune injury by viral neuraminidase
Virology Journal (2023)
-
Invasion by exogenous RNA: cellular defense strategies and implications for RNA inference
Marine Life Science & Technology (2023)
-
Ligand-switchable nanoparticles resembling viral surface for sequential drug delivery and improved oral insulin therapy
Nature Communications (2022)
-
Visualization of conformational changes and membrane remodeling leading to genome delivery by viral class-II fusion machinery
Nature Communications (2022)
-
Membrane attachment and fusion of HIV-1, influenza A, and SARS-CoV-2: resolving the mechanisms with biophysical methods
Biophysical Reviews (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.