Abstract
Toll-like receptors (TLRs) have a crucial role in the recognition of pathogens and initiation of immune responses1,2,3. Here we show that a previously uncharacterized protein encoded by CXorf21—a gene that is associated with systemic lupus erythematosus4,5—interacts with the endolysosomal transporter SLC15A4, an essential but poorly understood component of the endolysosomal TLR machinery also linked to autoimmune disease4,6,7,8,9. Loss of this type-I-interferon-inducible protein, which we refer to as ‘TLR adaptor interacting with SLC15A4 on the lysosome’ (TASL), abrogated responses to endolysosomal TLR agonists in both primary and transformed human immune cells. Deletion of SLC15A4 or TASL specifically impaired the activation of the IRF pathway without affecting NF-κB and MAPK signalling, which indicates that ligand recognition and TLR engagement in the endolysosome occurred normally. Extensive mutagenesis of TASL demonstrated that its localization and function relies on the interaction with SLC15A4. TASL contains a conserved pLxIS motif (in which p denotes a hydrophilic residue and x denotes any residue) that mediates the recruitment and activation of IRF5. This finding shows that TASL is an innate immune adaptor for TLR7, TLR8 and TLR9 signalling, revealing a clear mechanistic analogy with the IRF3 adaptors STING, MAVS and TRIF10,11. The identification of TASL as the component that links endolysosomal TLRs to the IRF5 transcription factor via SLC15A4 provides a mechanistic explanation for the involvement of these proteins in systemic lupus erythematosus12,13,14.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
TAP-MS proteomics data have been deposited to the ProteomeXchange Consortium77 via the PRIDE78 partner repository with the dataset identifier PXD014254 and 10.6019/PXD014254. RNA-seq data have been deposited to the Gene Expression Omnibus repository (GSE133317). Source data for immunoblots are provided in Supplementary Fig. 1.
References
Blasius, A. L. & Beutler, B. Intracellular Toll-like receptors. Immunity 32, 305–315 (2010).
Pelka, K., Shibata, T., Miyake, K. & Latz, E. Nucleic acid-sensing TLRs and autoimmunity: novel insights from structural and cell biology. Immunol. Rev. 269, 60–75 (2016).
Kawai, T. & Akira, S. The role of pattern-recognition receptors in innate immunity: update on Toll-like receptors. Nat. Immunol. 11, 373–384 (2010).
Bentham, J. et al. Genetic association analyses implicate aberrant regulation of innate and adaptive immunity genes in the pathogenesis of systemic lupus erythematosus. Nat. Genet. 47, 1457–1464 (2015).
Odhams, C. A. et al. Interferon inducible X-linked gene CXorf21 may contribute to sexual dimorphism in systemic lupus erythematosus. Nat. Commun. 10, 2164 (2019).
Han, J. W. et al. Genome-wide association study in a Chinese Han population identifies nine new susceptibility loci for systemic lupus erythematosus. Nat. Genet. 41, 1234–1237 (2009).
Blasius, A. L. et al. Slc15a4, AP-3, and Hermansky–Pudlak syndrome proteins are required for Toll-like receptor signaling in plasmacytoid dendritic cells. Proc. Natl Acad. Sci. USA 107, 19973–19978 (2010).
Sasawatari, S. et al. The solute carrier family 15A4 regulates TLR9 and NOD1 functions in the innate immune system and promotes colitis in mice. Gastroenterology 140, 1513–1525 (2011).
Kobayashi, T. et al. The histidine transporter SLC15A4 coordinates mTOR-dependent inflammatory responses and pathogenic antibody production. Immunity 41, 375–388 (2014).
Liu, S. et al. Phosphorylation of innate immune adaptor proteins MAVS, STING, and TRIF induces IRF3 activation. Science 347, aaa2630 (2015).
Zhao, B. et al. Structural basis for concerted recruitment and activation of IRF-3 by innate immune adaptor proteins. Proc. Natl Acad. Sci. USA 113, E3403–E3412 (2016).
Tsokos, G. C. Systemic lupus erythematosus. N. Engl. J. Med. 365, 2110–2121 (2011).
Moulton, V. R. et al. Pathogenesis of human systemic lupus erythematosus: a cellular perspective. Trends Mol. Med. 23, 615–635 (2017).
Tsokos, G. C., Lo, M. S., Costa Reis, P. & Sullivan, K. E. New insights into the immunopathogenesis of systemic lupus erythematosus. Nat. Rev. Rheumatol. 12, 716–730 (2016).
Kieser, K. J. & Kagan, J. C. Multi-receptor detection of individual bacterial products by the innate immune system. Nat. Rev. Immunol. 17, 376–390 (2017).
Janeway, C. A. Jr & Medzhitov, R. Innate immune recognition. Annu. Rev. Immunol. 20, 197–216 (2002).
O’Neill, L. A., Golenbock, D. & Bowie, A. G. The history of Toll-like receptors – redefining innate immunity. Nat. Rev. Immunol. 13, 453–460 (2013).
Blasius, A. L., Krebs, P., Sullivan, B. M., Oldstone, M. B. & Popkin, D. L. Slc15a4, a gene required for pDC sensing of TLR ligands, is required to control persistent viral infection. PLoS Pathog. 8, e1002915 (2012).
Baccala, R. et al. Essential requirement for IRF8 and SLC15A4 implicates plasmacytoid dendritic cells in the pathogenesis of lupus. Proc. Natl Acad. Sci. USA 110, 2940–2945 (2013).
Dosenovic, P. et al. Slc15a4 function is required for intact class switch recombination to IgG2c in response to TLR9 stimulation. Immunol. Cell Biol. 93, 136–146 (2015).
Griffith, A. D. et al. A requirement for Slc15a4 in imiquimod-induced systemic inflammation and psoriasiform inflammation in mice. Sci. Rep. 8, 14451 (2018).
Langefeld, C. D. et al. Transancestral mapping and genetic load in systemic lupus erythematosus. Nat. Commun. 8, 16021 (2017).
Pollard, K. M. et al. Induction of systemic autoimmunity by a xenobiotic requires endosomal TLR trafficking and signaling from the late endosome and endolysosome but not type I IFN. J. Immunol. 199, 3739–3747 (2017).
Rebsamen, M. et al. SLC38A9 is a component of the lysosomal amino acid sensing machinery that controls mTORC1. Nature 519, 477–481 (2015).
Pichlmair, A. et al. Viral immune modulators perturb the human molecular network by common and unique strategies. Nature 487, 486–490 (2012).
Nakamura, N. et al. Endosomes are specialized platforms for bacterial sensing and NOD2 signalling. Nature 509, 240–244 (2014).
Harris, V. M., Harley, I. T. W., Kurien, B. T., Koelsch, K. A. & Scofield, R. H. Lysosomal pH is regulated in a sex dependent manner in immune cells expressing CXorf21. Front. Immunol. 10, 578 (2019).
Libert, C., Dejager, L. & Pinheiro, I. The X chromosome in immune functions: when a chromosome makes the difference. Nat. Rev. Immunol. 10, 594–604 (2010).
Klijn, C. et al. A comprehensive transcriptional portrait of human cancer cell lines. Nat. Biotechnol. 33, 306–312 (2015).
FANTOM Consortium and the RIKEN PMI and CLST. A promoter-level mammalian expression atlas. Nature 507, 462–470 (2014).
Arazi, A. et al. The immune cell landscape in kidneys of patients with lupus nephritis. Nat. Immunol. 20, 902–914 (2019).
Muskardin, T. L. W. & Niewold, T. B. Type I interferon in rheumatic diseases. Nat. Rev. Rheumatol. 14, 214–228 (2018).
Hu, Y., Song, F., Jiang, H., Nuñez, G. & Smith, D. E. SLC15A2 and SLC15A4 mediate the transport of bacterially derived di/tripeptides to enhance the nucleotide-binding oligomerization domain-dependent immune response in mouse bone marrow-derived macrophages. J. Immunol. 201, 652–662 (2018).
Lee, J. et al. pH-dependent internalization of muramyl peptides from early endosomes enables Nod1 and Nod2 signaling. J. Biol. Chem. 284, 23818–23829 (2009).
Chow, K. T. et al. Differential and overlapping immune programs regulated by IRF3 and IRF5 in plasmacytoid dendritic cells. J. Immunol. 201, 3036–3050 (2018).
Graham, R. R. et al. A common haplotype of interferon regulatory factor 5 (IRF5) regulates splicing and expression and is associated with increased risk of systemic lupus erythematosus. Nat. Genet. 38, 550–555 (2006).
Schoenemeyer, A. et al. The interferon regulatory factor, IRF5, is a central mediator of Toll-like receptor 7 signaling. J. Biol. Chem. 280, 17005–17012 (2005).
Takaoka, A. et al. Integral role of IRF-5 in the gene induction programme activated by Toll-like receptors. Nature 434, 243–249 (2005).
Eames, H. L., Corbin, A. L. & Udalova, I. A. Interferon regulatory factor 5 in human autoimmunity and murine models of autoimmune disease. Transl. Res. 167, 167–182 (2016).
Maeda, T. et al. A novel plasmacytoid dendritic cell line, CAL-1, established from a patient with blastic natural killer cell lymphoma. Int. J. Hematol. 81, 148–154 (2005).
Steinhagen, F. et al. IRF-5 and NF-κB p50 co-regulate IFN-β and IL-6 expression in TLR9-stimulated human plasmacytoid dendritic cells. Eur. J. Immunol. 43, 1896–1906 (2013).
Newstead, S. Recent advances in understanding proton coupled peptide transport via the POT family. Curr. Opin. Struct. Biol. 45, 17–24 (2017).
Kelley, L. A., Mezulis, S., Yates, C. M., Wass, M. N. & Sternberg, M. J. The Phyre2 web portal for protein modeling, prediction and analysis. Nat. Protocols 10, 845–858 (2015).
Chen, W. et al. Insights into interferon regulatory factor activation from the crystal structure of dimeric IRF5. Nat. Struct. Mol. Biol. 15, 1213–1220 (2008).
Lopez-Pelaez, M. et al. Protein kinase IKKβ-catalyzed phosphorylation of IRF5 at Ser462 induces its dimerization and nuclear translocation in myeloid cells. Proc. Natl Acad. Sci. USA 111, 17432–17437 (2014).
Ren, J., Chen, X. & Chen, Z. J. IKKβ is an IRF5 kinase that instigates inflammation. Proc. Natl Acad. Sci. USA 111, 17438–17443 (2014).
Doench, J. G. et al. Optimized sgRNA design to maximize activity and minimize off-target effects of CRISPR–Cas9. Nat. Biotechnol. 34, 184–191 (2016).
Bigenzahn, J. W. et al. LZTR1 is a regulator of RAS ubiquitination and signaling. Science 362, 1171–1177 (2018).
Brinkman, E. K., Chen, T., Amendola, M. & van Steensel, B. Easy quantitative assessment of genome editing by sequence trace decomposition. Nucleic Acids Res. 42, e168 (2014).
Troegeler, A. et al. An efficient siRNA-mediated gene silencing in primary human monocytes, dendritic cells and macrophages. Immunol. Cell Biol. 92, 699–708 (2014).
Rudashevskaya, E. L. et al. A method to resolve the composition of heterogeneous affinity-purified protein complexes assembled around a common protein by chemical cross-linking, gel electrophoresis and mass spectrometry. Nat. Protocols 8, 75–97 (2013).
Varjosalo, M. et al. Interlaboratory reproducibility of large-scale human protein-complex analysis by standardized AP-MS. Nat. Methods 10, 307–314 (2013).
Olsen, J. V. et al. Parts per million mass accuracy on an Orbitrap mass spectrometer via lock mass injection into a C-trap. Mol. Cell. Proteomics 4, 2010–2021 (2005).
Chambers, M. C. et al. A cross-platform toolkit for mass spectrometry and proteomics. Nat. Biotechnol. 30, 918–920 (2012).
Apweiler, R. et al. UniProt: the Universal Protein knowledgebase. Nucleic Acids Res. 32, D115–D119 (2004).
Cox, J. & Mann, M. MaxQuant enables high peptide identification rates, individualized p.p.b.-range mass accuracies and proteome-wide protein quantification. Nat. Biotechnol. 26, 1367–1372 (2008).
Wright, J. C. & Choudhary, J. S. DecoyPyrat: fast non-redundant hybrid decoy sequence generation for large scale proteomics. J. Proteomics Bioinform. 9, 176–180 (2016).
Kim, S. & Pevzner, P. A. M. S.-G. F. MS-GF+ makes progress towards a universal database search tool for proteomics. Nat. Commun. 5, 5277 (2014).
Elias, J. E. & Gygi, S. P. Target-decoy search strategy for increased confidence in large-scale protein identifications by mass spectrometry. Nat. Methods 4, 207–214 (2007).
The, M., MacCoss, M. J., Noble, W. S. & Käll, L. Fast and accurate protein false discovery rates on large-scale proteomics data sets with percolator 3.0. J. Am. Soc. Mass Spectrom. 27, 1719–1727 (2016).
Choi, H. et al. Analyzing protein-protein interactions from affinity purification-mass spectrometry data with SAINT. Curr. Protoc. Bioinformatics 39, 8.15.1–8.15.23 (2012).
Shannon, P. et al. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res. 13, 2498–2504 (2003).
Mellacheruvu, D. et al. The CRAPome: a contaminant repository for affinity purification-mass spectrometry data. Nat. Methods 10, 730–736 (2013).
UniProt Consortium. UniProt: a worldwide hub of protein knowledge. Nucleic Acids Res. 47 (D1), D506–D515 (2019).
Larkin, M. A. et al. Clustal W and Clustal X version 2.0. Bioinformatics 23, 2947–2948 (2007).
Drozdetskiy, A., Cole, C., Procter, J. & Barton, G. J. JPred4: a protein secondary structure prediction server. Nucleic Acids Res. 43, W389–W394 (2015).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Derr, A. et al. End Sequence Analysis Toolkit (ESAT) expands the extractable information from single-cell RNA-seq data. Genome Res. 26, 1397–1410 (2016).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Marbach, D. et al. Tissue-specific regulatory circuits reveal variable modular perturbations across complex diseases. Nat. Methods 13, 366–370 (2016).
Hochberg, Y. & Benjamini, Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J. R. Stat. Soc. 57, 289–300 (1995).
Huang, W., Sherman, B. T. & Lempicki, R. A. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat. Protocols 4, 44–57 (2009).
Blaszczyk, M., Ciemny, M. P., Kolinski, A., Kurcinski, M. & Kmiecik, S. Protein-peptide docking using CABS-dock and contact information. Brief. Bioinform. 20, 2299–2305 (2019).
London, N., Raveh, B., Cohen, E., Fathi, G. & Schueler-Furman, O. Rosetta FlexPepDock web server—high resolution modeling of peptide–protein interactions. Nucleic Acids Res. 39, W249–W253 (2011).
Colas, C. et al. An improved flow cytometry assay to monitor phagosome acidification. J. Immunol. Methods 412, 1–13 (2014).
Carpenter, A. E. et al. CellProfiler: image analysis software for identifying and quantifying cell phenotypes. Genome Biol. 7, R100 (2006).
Deutsch, E. W. et al. The ProteomeXchange consortium in 2017: supporting the cultural change in proteomics public data deposition. Nucleic Acids Res. 45, D1100–D1106 (2017).
Perez-Riverol, Y. et al. The PRIDE database and related tools and resources in 2019: improving support for quantification data. Nucleic Acids Res. 47, D442–D450 (2019).
Acknowledgements
We thank all members of the G.S.-F. laboratory, T. Pemovska, G. Boehmelt, T. Krausgruber and E. Salzer for discussions and feedback; the Proteomics and Metabolomics Facility (CeMM) for the proteomics analyses; the Biomedical Sequencing Facility (CeMM/Medical University of Vienna) for the NGS sequencing; the Core Facility Imaging of the Medical University of Vienna; and T. Maeda for kindly providing the CAL-1 cells. This work was supported by Boehringer Ingelheim (Research Collaboration Agreement BI-CeMM 238114 to L.X.H., UK, K.P., A.S. and M.R.), the Austrian Academy of Sciences (to G.S.-F., F.K., V.S. and A.C.-R.), the European Research Council (ERC AdG 695214 GameofGates to G.S.-F., M.R., S.S., P.E., U.G., M.D.P., A.B. and E.G.) and Austrian Science Fund (FWF SFB F4711, to J.W.B.). Plasmids obtained through Addgene were a gift from F. Zhang, D. Trono and D. Golenbock.
Author information
Authors and Affiliations
Contributions
L.X.H., J. Lee, E.G., M.L.M., C.E.W., M.R. and G.S.-F. designed research; L.X.H., J. Lee, U.K., K.P., F.K., P.E., A.S., S.S., A.C.M., F.J.K., M.D.P., E.G. and M.R. performed research; V.S., A.C.-R. and U.G. performed bioinformatic analysis; J.W.B., A.B. and J. Li generated reagents and provided scientific insight, L.X.H., M.R. and G.S.-F. analysed and interpreted the data; L.X.H., M.R. and G.S.-F. wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
J. Lee, J. Li, J.K., M.L.M. and C.E.W. are employed in the Immunology and Respiratory department, Drug Concept Discovery group of Boehringer-Ingelheim. The work in the G.S.-F. laboratory is supported by Boehringer-Ingelheim (grant agreement 238114).
Additional information
Peer review information Nature thanks Zhijian (James) Chen and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Subcellular localization of tagged SLC15A4 and related constructs in THP1 cells.
a, Gene expression levels of SLC15A1, SLC15A2, SLC15A3 and SLC15A4 in THP1 cells from a previous publication29. b, Domain organization of SLC15A4 protein. TM, transmembrane domain. c, d, Confocal microscopy of indicated THP1 cells. Red, anti-HA; green, anti-LAMP1; blue, DAPI. Scale bars, 10 μm. e, Lysates from indicated THP1 cells untreated or treated with PNGase F were analysed by immunoblotting. c–e, Data representative of two independent experiments. For gel source data, see Supplementary Fig. 1.
Extended Data Fig. 2 TASL is an immune-cell-restricted protein that is conserved in vertebrates.
a, Multiple sequence alignment of TASL protein from representative vertebrate species. UniProt entry names: CX021_HUMAN, H2QYF9_PANTR, CX021_BOVIN, CX021_MOUSE, H0Z9M3_TAEGU, A0A1L1RS25_CHICK, A0A1U7RX84_ALLSI, G1KG99_ANOCA, F7BWY0_XENTR, W5NMP6_LEPOC, I3KXE0_ORENI. Boxes above the alignment indicate consensus prediction from JPred4. Red, helix; yellow, β sheet. b, c, Expression levels of SLC15A4 and CXorf21 in primary human cells as measured by CAGE by FANTOM530 (b) and in human cancer cell lines measured by RNA-seq from a previous publication29 (c). Circles represent individual samples; in box plots, bars indicate the median, boxes indicate the first to third quartiles. The top whisker extends from hinge to largest value no further than 1.5× interquartile range (IQR) from the hinge, and the bottom whisker extends from the hinge to smallest value at most 1.5× IQR of the hinge.
Extended Data Fig. 3 Type-I-interferon-inducible TASL forms a complex with SLC15A4.
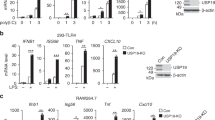
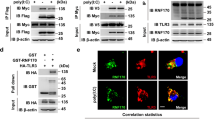
a, Normalized mRNA expression of SLC15A4 and CXorf21 relative to GAPDH in THP1 cells treated as indicated. Data show mean (n = 3 technical replicates). b, Immunoblots of lysates of THP1 cells stimulated with interferon-β or interferon-γ (20 ng ml−1, for 16 h) treated with PNGase F as indicated. c, e, Lysates from KBM7 cells transduced (c) or HEK293T cells transiently transfected (e), as indicated, were subjected to HA immunoprecipitation. Immunoprecipitates and whole-cell extracts were analysed by immunoblotting. d, Overview of deletion mutants used in Fig. 1g. f, Lysates from indicated THP1 cells were subjected to HA immunoprecipitation and treated or not with λ phosphatase. Immunoprecipitates and whole-cell extract were analysed by immunoblotting. g, Immunoblots of indicated THP1 cells treated with PNGase F. h, Multiple sequence alignment of human and mouse SLC15A3 and SLC15A4. UniProt entry names: S15A3_HUMAN, S15A3_MOUSE, S15A4_HUMAN, S15A4_MOUSE. In a–c, e–g, data are representative of two independent experiments. For gel source data, see Supplementary Fig. 1.
Extended Data Fig. 4 SLC15A4 controls TASL protein abundance and subcellular localization.
a, b, Flow cytometry (a) and immunoblot (b) of indicated THP1 cells. c, Confocal live microscopy of indicated THP1 cells. Green, TASL–GFP; red, Lysotracker; blue, Hoechst33342. Scale bars, 10 μm. d, Top, confocal microscopy of indicated formaldehyde-fixed THP1 cells. Green, TASL–GFP; red, HA; blue, DAPI. Scale bars, 10 μm. Bottom, profiles of signal intensity of TASL–GFP (green) and HA (red) along the red lines shown in microscopy images (top). In a–d, data are representative of two independent experiments. For gel source data, see Supplementary Fig. 1.
Extended Data Fig. 5 TASL mirrors SLC15A4 requirement for TLR7 and TLR8 activation.
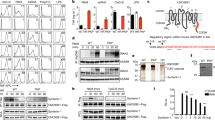
a, Fraction of R848-induced genes affected by SLC15A4 and CXorf21 knockout, related to Fig. 2b. b, Upset plot representing number of R848-induced genes commonly affected by the indicated sgRNAs, related to Fig. 2b. c, CXorf21 gene expression levels in indicated THP1 cells, related to Fig. 2b. Bar graphs shown mean (n = 3 biological replicates), error bars show 95% confidence interval of mean. d, Flow cytometry of PD-L1 surface expression in indicated unstimulated (ns) or R848-stimulated (5 μg ml−1, for 24 h) THP1 cells. e, Immunoblots of indicated THP1 DUAL cells. Lysates treated with PNGase F, as indicated. f–i, k, Indicated THP1 DUAL cells were (co-)treated for 24 h with R848 (5 μg ml−1), CL075 (5 μg ml−1), single-stranded (ss)RNA40 complexed with LyoVec (5 μg ml−1) or inactive control ssRNA41 with LyoVec (5 μg ml−1), C12-iE-DAP (5 μg ml−1), MDP (10 μg ml−1), murabutide (10 μg ml−1), Pam3CSK4 (0.1 μg ml−1), flagellin (0.1 μg ml−1), cGAMP (3 μg ml−1) or interferon-β (20 ng ml−1). h, CRISPR–Cas9 editing efficiency (%) estimated by TIDE. j, Indicated THP1 DUAL cells were primed or not with interferon-γ (0.1 μg ml−1) for 24 h, washed and stimulated or not with MDP (10 μg ml−1, for 24 h). f–k, Supernatants were analysed for ISRE and NF-κB reporter activity. Mean ± s.d. (n = 3 biological replicates). l, Relative mRNA expression of SLC15A4, CXorf21 or MYD88 in siRNA-transfected CD14+ monocytes in comparison to control (siCTRL). Data represent mean ± s.d. from six (MYD88) or seven (SLC15A4 and CXorf21) individual donors. In d–k, data are representative of two independent experiments. For gel source data, see Supplementary Fig. 1.
Extended Data Fig. 6 TASL and SLC15A4 deficiency impairs endosomal TLR-mediated signalling downstream of receptor engagement.
a, Immunoblots of lysates of THP1 cells treated with PNGase F, as indicated. b, c, Cytokine production of indicated THP1 cells unstimulated or stimulated with CpG-A, CpG-B (5 μM) or R848 (5 μg ml−1) for 24 h. Data show mean ± s.d. of biological replicates (TNF and CCL2, n = 3; IFNβ, n = 2). d, Immunoblots of indicated THP1 cells stimulated or not with interferon-γ (0.1 μg ml−1, for 16 h). e, Immunoblots of indicated THP1 cells. f, Indicated THP1::TLR9 cells treated with FITC-labelled CpG-A or CpG-B (1 μM, for 0–120 min) were analysed by flow cytometry. g, Representative flow cytometry scatter plots of phagocytosis assays. Differentiated THP1 cells, treated or not with bafilomycin A1, were incubated with dual-coloured opsonized beads. Using intensities of pH-insensitive (YG) and pH-sensitive (pHrodo-Red, signal increases with decrease in pH) dyes, cells are divided into phagocytosis-negative (PhagoNeg, double-negative), cells that have undergone phagocytosis and phagosome acidification (PhagoLate, double-positive) and early phagocytic cells (PhagoEarly, YG and low pHrodo-Red signal). The marginal intensity distributions are displayed on the sides of the plot. h, Bar graphs show mean ± s.d. (n = 3 biological replicates) of fractions described in g. i, j, Indicated THP1 cells were subjected to phagocytosis assays. i, Bar graphs show mean ± s.d. (n = 3 biological replicates) of fractions described in g. j, Bar graphs represent mean ± s.d. (n = 3 biological replicates) of the mean fluorescence intensities (MFI) of the pHrodo-Red signal acquired in the MFI gate shown in g, focusing on cells having taken up 1–3 beads per cell. k, Flow diagram for quantification of Lysosensor Green intensities in lysosomal compartments by microscopy. Box plots show intensity of Lysosensor signal on Lysotracker-positive lysosomes, as measured by imaging in the indicated THP1 cells. Bars indicate median, boxes indicate the first to third quartiles; the top whisker extends from the hinge to the largest value no further than 1.5× IQR from the hinge; the bottom whisker extends from the hinge to the smallest value at most 1.5× IQR of the hinge. Outliers are shown as circles. sgRen, n = 2,432; sgSLC15A4-1, n = 1,721; sgSLC15A4-2, n = 1,981; sgTASL-1, n = 2,378; sgTASL-2, n = 2,627 quantified speckles. l, Immunoblots of indicated THP1 cells stimulated with R848 (5 μg ml−1, for 0–180 min.). In a–k, data are representative of two(a–f, k, l) or three(g–j) independent experiments. For gel source data, see Supplementary Fig. 1.
Extended Data Fig. 7 Loss of TASL or SLC15A4 mirrors IRF5 deficiency in perturbing endosomal TLR responses.
a, d, e, THP1 DUAL cells were (co-)treated for 24 h with R848 (5 μg ml−1), LPS (0.1 μg ml−1), Pam3CSK4 (0.1 μg ml−1), cGAMP (3 μg ml−1), flagellin (0.1 μg ml−1) or MDP (10 μg ml−1) as indicated. Supernatants were analysed for ISRE and NF-κB reporter activity. b, c, Immunoblots of indicated THP1 DUAL (b) or THP1::TLR9 (c) cells. f, TNF production of indicated THP1::TLR9 cells stimulated with CpG-B (2 μM, for 24 h). g, Immunoblots of indicated THP1 cells stimulated or not with R848 (5 μg ml−1, for 3 h). h, Upset plot representing number of CpG-B-induced genes (2 μM, for 6 h) (DESeq2 adjusted P value < 0.05, n = 3 biological replicates) in comparison to control (sgRen) commonly affected by indicated sgRNAs. No gene was significantly affected by sgIRF7-1. i, Principal component analysis plot of transcriptional profiles of untreated and CpG-B-treated (2 μM, for 6 h) THP1::TLR9 cells (n = 3 biological replicates) shown in Fig. 3d. j, Heat map representing 20 most-induced genes by CpG-B in control THP1::TLR9 cells and not affected by SLC15A4, CXorf21 or IRF5 knockout, related to Fig. 3d, e. k, Transcription factor enrichment analysis (two-sided Fisher’s exact test, P value adjusted for multiple testing) of genes upregulated upon CpG-B treatment in control THP1 cells specifically affected (DESeq2 adjusted P value < 0.05, n = 3 biological replicates) (left) or not (right) by SLC15A4 and CXorf21 knockout, related to Fig. 3d, e. Background sets are defined as all genes upregulated by CpG-B treatment or all expressed genes (counts per million > 1) respectively. In a, d–f, data are mean ± s.d. (n = 3 biological replicates). In a–g, data are representative of two independent experiments. For gel source data, see Supplementary Fig. 1.
Extended Data Fig. 8 The SLC15A4–TASL complex is required for IRF5-dependent signalling in human CAL-1 plasmacytoid dendritic cells.
a–c, Immunoblots of lysates of CAL-1 cells, with PNGase F treatment as indicated. d, Immunoblots of CAL-1 or THP1::TLR9 cells stimulated with R848 (5 μg ml−1), CpG-B (5 μM) or cGAMP (3 μg ml−1) as indicated. e, f, Cytokine production of indicated CAL-1 cells stimulated with R848 (5 μg ml−1) or imiquimod (IMQ) (5 μg ml−1) for 24 h. f, CRISPR–Cas9 editing efficiency (%) estimated by TIDE. g, Immunoblots of CAL-1 cells stimulated with CpG-B (5 μM, for 0–180 min) as indicated. h, Lysates from indicated CAL-1 cells were subjected to HA immunoprecipitation and analysed by immunoblotting. i, TNF production of cells described in h upon stimulation with R848 (5 μg ml−1, for 24 h). j, Immunoblots show expression levels (whole-cell extracts) and HA immunoprecipitates from indicated THP1 cells. Bar graphs represent TNF production upon R848 stimulation (5 μg ml−1, for 24 h). In e, f, i, j, bar graphs show mean ± s.d. (n = 3 biological replicates). In a–j, data are representative of two independent experiments. For gel source data, see Supplementary Fig. 1.
Extended Data Fig. 9 Mutagenesis of TASL identifies functional elements and reveals a pLxIS motif required for IRF5 activation.
a, Overview of mutants used in Fig. 4b; changes to alanine indicated by red circles. b, f, Immunoblots of indicated THP1 cells. Bar graphs represent TNF levels following R848 stimulation (5 μg ml−1, for 24 h). In f, the dashed line indicates cropping of unrelated lanes from the same blots. c, Immunoblots of indicated reconstituted TASL-deficient THP1 cells. d, Normal expression levels, but reduced detection by anti-TASL antibodies, of TASL mutants targeting amino acids 261–277. Immunoblots of HEK293T cells transiently transfected with indicated cDNAs. e, Lysates from indicated THP1 cells were subjected to immunoprecipitation. Immunoprecipitates and whole-cell extracts were analysed by immunoblotting. g, Abundance of indicated proteins determined by mass spectrometry in V5 immunoprecipitates from THP1::TLR9 cells stimulated with CpG-B (5 μM, for 2 h) as indicated, related to Fig. 4g. Three biological replicates are shown. h, Crystal structures of IRF3 bound to phosphorylated pLxIS-containing peptides from STING (pink, PDB ID: 5JEJ), MAVS (green, PDB ID: 5JEK) and TRIF (blue, PDB ID: 5JEL). Residues in STING peptides are shown as sticks. i, Superposition of peptide-bound IRF3 (PDB ID: 5JEJ) and dimeric IRF5 (one monomer shown, PDB ID: 3DSH), showing highly similar folds. j, Model of phosphomimetic pLxID-containing peptide from TASL (pmTASL, ISTPSLHIDQYSNV, yellow) bound to IRF5. Residues corresponding to the pLxID motif shown as sticks. k, Comparison of binding mode of pLxIS-containing peptides to IRF proteins. Only IRF5 is shown for clarity. l, Immunoblots of indicated THP1 cells unstimulated or stimulated with R848 (5 μg ml−1, for 2 h). m, TNF production of cells described in l, stimulated with R848 (5 μg ml−1, for 24 h). In b, f, m, bar graphs show mean ± s.d. (n = 3 biological replicates). In b–f, l, m, data are representative of two independent experiments. For gel source data, see Supplementary Fig. 1.
Extended Data Fig. 10 IKKβ is required for TASL-dependent IRF5 activation.
a, Immunoblots of HEK293T cells transfected as indicated. b, THP1 DUAL cells were pre-treated for 30 min with DMSO or inhibitors, as indicated, and stimulated with R848 (5 μg ml−1, for 24 h). Supernatants were analysed for ISRE and NF-κB reporter activity and normalized to the respective R848-only-treated conditions. Three biological replicates are shown. c, Immunoblots of THP1::TLR9 cells pre-treated for 30 min with DMSO or inhibitors (5 μM) and stimulated with CpG-B (5 μM, for 4 h), as indicated. d, Lysates from THP1::TLR9 cells pre-treated (for 30 min) with DMSO or inhibitors (5 μM) and stimulated with CpG-B (5 μM, for 2 h), as indicated, were subjected to V5 immunoprecipitation. Immunoprecipitates and whole-cell extracts were analysed by immunoblotting. e, Immunoblots of indicated THP1::TLR9 cells. CHUK gene encodes IKKα; MAP3K7 encodes TAK1. f, TNF production of indicated THP1::TLR9 cells stimulated with CpG-B (2 μM, for 24 h). Data are mean ± s.d. (n = 3 biological replicates). g, Immunoblots of indicated THP1::TLR9 cells stimulated with CpG-B (5 μM, for 3 h). h, Schematic representing functional homology of the SLC15A4–TASL module in mediating IRF5 activation in comparison to the IRF3 adaptors STING, MAVS and TRIF. In a–g, data are representative of two independent experiments. For gel source data, see Supplementary Fig. 1.
Supplementary information
Supplementary Figure
Supplementary Figure 1 – Source data (Immunoblots). Uncropped scans of immunoblots presented in the manuscript.
Supplementary Table
Supplementary Table 1 – Data related to interaction proteomics (Fig. 1b, e). TAP-MS/MS analysis of THP1 cells stably expressing StHA-SLC15A4 constructs (n=2 biological replicates).
Supplementary Table
Supplementary Table 2 – Gene expression data (related to Extended Data Fig. 2b). Expression levels of SLC15A4 and CXorf21 in primary human cells as measured by CAGE by FANTOM530.
Supplementary Table
Supplementary Table 3 – Data related to R848-induced transcriptional responses (related to Fig. 2b). Transcriptional profiles of unstimulated and R848 (5 µg/ml, 6h) stimulated THP1 cell lines. Differential expression analysis was performed with DESeq2 on the basis of read counts (n=3 biological replicates).
Supplementary Table
Supplementary Table 4 - Data related to CpG-B-induced transcriptional responses (related to Fig. 3d). Transcriptional profiles of unstimulated and CpG-B (2 µM, 6h)-stimulated THP1::TLR9 cell lines. Differential expression analysis was performed with DESeq2 on the basis of read counts (n=3 biological replicates).
Supplementary Table
Supplementary Table 5 - Data related to interaction proteomics (related to Fig. 4g, Extended Data Fig. 9g). Immunoprecipitates from the indicated THP1::TLR9 cell lines stimulated with CpG-B (5 µM, 2h) were analysed by MS (n=3 biological replicates).
Rights and permissions
About this article
Cite this article
Heinz, L.X., Lee, J., Kapoor, U. et al. TASL is the SLC15A4-associated adaptor for IRF5 activation by TLR7–9. Nature 581, 316–322 (2020). https://doi.org/10.1038/s41586-020-2282-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-020-2282-0
This article is cited by
-
Inhibition of STAT3 alleviates LPS-induced apoptosis and inflammation in renal tubular epithelial cells by transcriptionally down-regulating TASL
European Journal of Medical Research (2024)
-
The conneXion between sex and immune responses
Nature Reviews Immunology (2024)
-
IKKε and TBK1 prevent RIPK1 dependent and independent inflammation
Nature Communications (2024)
-
Future opportunities in solute carrier structural biology
Nature Structural & Molecular Biology (2024)
-
Sinking the carrier
Nature Chemical Biology (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.