Abstract
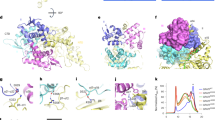
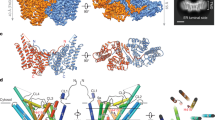
Diacylglycerol O-acyltransferase 1 (DGAT1) synthesizes triacylglycerides and is required for dietary fat absorption and fat storage in humans1. DGAT1 belongs to the membrane-bound O-acyltransferase (MBOAT) superfamily, members of which are found in all kingdoms of life and are involved in the acylation of lipids and proteins2,3. How human DGAT1 and other mammalian members of the MBOAT family recognize their substrates and catalyse their reactions is unknown. The absence of three-dimensional structures also hampers rational targeting of DGAT1 for therapeutic purposes. Here we present the cryo-electron microscopy structure of human DGAT1 in complex with an oleoyl-CoA substrate. Each DGAT1 protomer has nine transmembrane helices, eight of which form a conserved structural fold that we name the MBOAT fold. The MBOAT fold in DGAT1 forms a hollow chamber in the membrane that encloses highly conserved catalytic residues. The chamber has separate entrances for each of the two substrates, fatty acyl-CoA and diacylglycerol. DGAT1 can exist as either a homodimer or a homotetramer and the two forms have similar enzymatic activity. The N terminus of DGAT1 interacts with the neighbouring protomer and these interactions are required for enzymatic activity.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Data availability
The atomic coordinates of human DGAT1 have been deposited in the Protein Data Bank (http://www.rcsb.org) under the accession code 6VP0. The corresponding electron microscopy maps have been deposited in the Electron Microscopy Data Bank (https://www.ebi.ac.uk/pdbe/emdb/) under the accession code EMD-21302
References
Cases, S. et al. Identification of a gene encoding an acyl CoA:diacylglycerol acyltransferase, a key enzyme in triacylglycerol synthesis. Proc. Natl Acad. Sci. USA 95, 13018–13023 (1998).
Das, A., Davis, M. A. & Rudel, L. L. Identification of putative active site residues of ACAT enzymes. J. Lipid Res. 49, 1770–1781 (2008).
Nusse, R. Wnt signaling in disease and in development. Cell Res. 15, 28–32 (2005).
Yen, C.-L. E., Nelson, D. W. & Yen, M.-I. Intestinal triacylglycerol synthesis in fat absorption and systemic energy metabolism. J. Lipid Res. 56, 489–501 (2015).
Yen, C.-L. E., Stone, S. J., Koliwad, S., Harris, C. & Farese, R. V. Jr. Thematic review series: glycerolipids. DGAT enzymes and triacylglycerol biosynthesis. J. Lipid Res. 49, 2283–2301 (2008).
Bhatt-Wessel, B., Jordan, T. W., Miller, J. H. & Peng, L. Role of DGAT enzymes in triacylglycerol metabolism. Arch. Biochem. Biophys. 655, 1–11 (2018).
Kühn, C. et al. Evidence for multiple alleles at the DGAT1 locus better explains a quantitative trait locus with major effect on milk fat content in cattle. Genetics 167, 1873–1881 (2004).
Smith, S. J. et al. Obesity resistance and multiple mechanisms of triglyceride synthesis in mice lacking Dgat. Nat. Genet. 25, 87–90 (2000).
Chen, H. C. et al. Increased insulin and leptin sensitivity in mice lacking acyl CoA:diacylglycerol acyltransferase 1. J. Clin. Invest. 109, 1049–1055 (2002).
Zhao, G. et al. Validation of diacyl glycerolacyltransferase I as a novel target for the treatment of obesity and dyslipidemia using a potent and selective small molecule inhibitor. J. Med. Chem. 51, 380–383 (2008).
McFie, P. J., Stone, S. L., Banman, S. L. & Stone, S. J. Topological orientation of acyl-CoA:diacylglycerol acyltransferase-1 (DGAT1) and identification of a putative active site histidine and the role of the N terminus in dimer/tetramer formation. J. Biol. Chem. 285, 37377–37387 (2010).
Yang, J., Brown, M. S., Liang, G., Grishin, N. V. & Goldstein, J. L. Identification of the acyltransferase that octanoylates ghrelin, an appetite-stimulating peptide hormone. Cell 132, 387–396 (2008).
Ma, D. et al. Crystal structure of a membrane-bound O-acyltransferase. Nature 562, 286–290 (2018).
Cheng, D. et al. Human acyl-CoA:diacylglycerol acyltransferase is a tetrameric protein. Biochem. J. 359, 707–714 (2001).
Zhang, J. et al. Monoacylglycerol acyltransferase-2 is a tetrameric enzyme that selectively heterodimerizes with diacylglycerol acyltransferase-1. J. Biol. Chem. 289, 10909–10918 (2014).
Cao, J. et al. Targeting acyl-CoA:diacylglycerol acyltransferase 1 (DGAT1) with small molecule inhibitors for the treatment of metabolic diseases. J. Biol. Chem. 286, 41838–41851 (2011).
von Heijne, G. & Gavel, Y. Topogenic signals in integral membrane proteins. Eur. J. Biochem. 174, 671–678 (1988).
Caldo, K. M. P. et al. Diacylglycerol acyltransferase 1 is regulated by its N-terminal domain in response to allosteric effectors. Plant Physiol. 175, 667–680 (2017).
Weselake, R. J. et al. Acyl-CoA-binding and self-associating properties of a recombinant 13.3 kDa N-terminal fragment of diacylglycerol acyltransferase-1 from oilseed rape. BMC Biochem. 7, 24 (2006).
Panigrahi, R. et al. Intrinsic disorder in the regulatory N-terminal domain of diacylglycerol acyltransferase 1 from Brassica napus. Sci. Rep. 8, 16665 (2018).
Guo, Z., Cromley, D., Billheimer, J. T. & Sturley, S. L. Identification of potential substrate-binding sites in yeast and human acyl-CoA sterol acyltransferases by mutagenesis of conserved sequences. J. Lipid Res. 42, 1282–1291 (2001).
Bai, Y. et al. X-ray structure of a mammalian stearoyl-CoA desaturase. Nature 524, 252–256 (2015).
Mastronarde, D. N. Automated electron microscope tomography using robust prediction of specimen movements. J. Struct. Biol. 152, 36–51 (2005).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Grant, T. & Grigorieff, N. Measuring the optimal exposure for single particle cryo-EM using a 2.6 Å reconstruction of rotavirus VP6. eLife 4, e06980 (2015).
Zhang, K. Gctf: real-time CTF determination and correction. J. Struct. Biol. 193, 1–12 (2016).
Scheres, S. H. Semi-automated selection of cryo-EM particles in RELION-1.3. J. Struct. Biol. 189, 114–122 (2015).
Scheres, S. H. RELION: implementation of a Bayesian approach to cryo-EM structure determination. J. Struct. Biol. 180, 519–530 (2012).
Kimanius, D., Forsberg, B. O., Scheres, S. H. & Lindahl, E. Accelerated cryo-EM structure determination with parallelisation using GPUs in RELION-2. eLife 5, e18722 (2016).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Rosenthal, P. B. & Henderson, R. Optimal determination of particle orientation, absolute hand, and contrast loss in single-particle electron cryomicroscopy. J. Mol. Biol. 333, 721–745 (2003).
Chen, S. et al. High-resolution noise substitution to measure overfitting and validate resolution in 3D structure determination by single particle electron cryomicroscopy. Ultramicroscopy 135, 24–35 (2013).
Kucukelbir, A., Sigworth, F. J. & Tagare, H. D. Quantifying the local resolution of cryo-EM density maps. Nat. Methods 11, 63–65 (2014).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D 66, 486–501 (2010).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D 66, 213–221 (2010).
Barad, B. A. et al. EMRinger: side chain-directed model and map validation for 3D cryo-electron microscopy. Nat. Methods 12, 943–946 (2015).
Rana, M. S. et al. Fatty acyl recognition and transfer by an integral membrane S-acyltransferase. Science 359, eaao6326 (2018).
Stanley, C. J. & Perham, R. N. Purification of 2-oxo acid dehydrogenase multienzyme complexes from ox heart by a new method. Biochem. J. 191, 147–154 (1980).
Ren, Z. et al. Structure of an EIIC sugar transporter trapped in an inward-facing conformation. Proc. Natl Acad. Sci. USA 115, 5962–5967 (2018).
Baker, N. A., Sept, D., Joseph, S., Holst, M. J. & McCammon, J. A. Electrostatics of nanosystems: application to microtubules and the ribosome. Proc. Natl Acad. Sci. USA 98, 10037–10041 (2001).
Wallace, A. C., Laskowski, R. A. & Thornton, J. M. LIGPLOT: a program to generate schematic diagrams of protein-ligand interactions. Protein Eng. Des. Sel. 8, 127–134 (1995).
Laskowski, R. A. & Swindells, M. B. LigPlot+: multiple ligand-protein interaction diagrams for drug discovery. J. Chem. Inf. Model. 51, 2778–2786 (2011).
Sievers, F. et al. Fast, scalable generation of high-quality protein multiple sequence alignments using Clustal Omega. Mol. Syst. Biol. 7, 539 (2011).
Robert, X. & Gouet, P. Deciphering key features in protein structures with the new ENDscript server. Nucleic Acids Res. 42, W320–W324 (2014).
Landau, M. et al. ConSurf 2005: the projection of evolutionary conservation scores of residues on protein structures. Nucleic Acids Res. 33, W299–W302 (2005).
Acknowledgements
This work was supported by grants from the NIH (DK122784, HL086392 and GM098878 to M.Z.), the Cancer Prevention and Research Institute of Texas (R1223 to M.Z.), the Robert Welch Foundation (Q1279 to B.V.V.P.), the Ara Parseghian Medical Research Foundation (to N.Y. and Y.H.) and the New Jersey Council for Cancer Research (to H.Q.). N.Y. is supported by the Shirley M. Tilghman endowed professorship from Princeton University. We thank P. Shao for technical support during the acquisition of electron microscope images, and A. Banerjee for advice on the enzymatic assay and purification of αKDH. We acknowledge the use of the Imaging and Analysis Center at Princeton University, which is partially supported by the Princeton Center for Complex Materials, and the National Science Foundation (NSF)-MRSEC program (DMR-1420541).
Author information
Authors and Affiliations
Contributions
M.Z., L.W., Y.N. and Z.R. conceived the project and designed the experiments. L.W., Y.N. and H.Z. expressed and purified human DGAT1. L.W. and Y.N. measured functions of DGAT1. H.Q., Y.H., L.W., Z.R. and H.Z. prepared cryo-EM grids and collected and analysed cryo-EM data. L.W. and Z.R. built and refined the human DGAT1 structure. L.H. and B.V.V.P. advised on model building and refinement. A.L. examined the lipid content of DGAT1. L.W., Z.R., N.Y. and M.Z. analysed data. L.W., Z.R. and M.Z. wrote the initial draft and all authors participated in revising the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature thanks David Drew, Savvas N. Savvides and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Side-by-side comparison of human DGAT1 and DltB.
Both human DGAT1 and DltB have an acyl donor and an acyl acceptor. In the acyl-donor row, the red dashed lines indicate the bonds that are broken during acyl-transfer reactions. In the acyl-acceptor row, the hydroxyl groups are highlighted in red. In the substrates distribution row, DGAT1 and the DltB–DltC complex are shown as cartoon and the membrane as dashed lines. The position of the catalytic histidine in each protein is marked with a yellow star. In human DGAT1, the acyl-CoA comes from the intracellular side whereas the DAG comes from the hydrophobic core of the membrane. In DltB, the 4′-phosphopantetheine-DltC is intracellular whereas the lipoteichoic acid (LTA) is extracellular. In the MBOAT fold row, the MBOAT folds of DGAT1 and DltB are shown in cartoon representation and viewed from the same orientation. Equivalent helices have the same colour. The tunnel row shows the cut-away surface illustrations of DGAT1 and DltB, showing their cytosolic tunnels. The position of the conserved histidine residue in each protein is marked with a yellow star. In DltB, the intracellular loops are placed more towards the centre of the membrane and as a result, the MBOAT fold in DltB does not carve out a reaction chamber in the membrane. Overall, DltB is shaped similarly to an hourglass that allows the two substrates to approach the reaction centre from either side of the membrane, and the transfer of an acyl group across the membrane. These observations highlight the versatility of the MBOAT fold.
Extended Data Fig. 2 Purification and functional characterization of human DGAT1.
a, Size-exclusion chromatograph profile of human DGAT1 extracted with LMNG. Elution volumes of membrane proteins of known molecular weight—bcMalT (about 100 kDa, green)39 and mouse (m)SCD1 (about 40 kDa, blue)22—are marked with arrows. Inset, SDS–PAGE of the purified DGAT1. DGAT1 has a main peak with an elution volume of around 11.7 ml that corresponds to a dimer and a minor peak at around 10.4 ml that corresponds to a tetramer. b, Size-exclusion chromatography profiles of DGAT1 extracted with LMNG (left) or GDN (right). The main peak from the first run (red trace) was collected and reinjected onto the same column after 1 h. In both a and b, the detergent in the mobile phase is GDN. c, A white layer of fat appeared after membrane solubilization and centrifugation, indicating that the heterologously expressed DGAT1 is active in cells. d, The DGAT1 reaction is coupled to that of αKDH to monitor production of coenzyme A (CoA-SH) in real time. e, Fluorescence of NADH plotted versus time. Rapid production of coenzyme A occurs in the presence of oleoyl-CoA, 1,2-dioleoyl-sn-glycerol (1,2-DAG) and the purified dimeric DGAT1. By contrast, production of coenzyme A was not observed when either 1,2-DAG or DGAT1 was omitted from the reaction mixture, indicating that hydrolysis of oleoyl-CoA is tightly coupled to the enzymatic reaction. In addition, coenzyme A production was almost completely suppressed in the presence of 5 μM T863, a known DGAT1 inhibitor16. f, Production of triacylglyceride over time, detected by thin-layer chromatography. The first lane from the left is a triacylglyceride standard. g–i, Initial rate of reaction versus oleoyl-CoA concentration measured using the purified dimeric DGAT1 (g), tetrameric DGAT1 (h) or DGAT1 in cell membrane (i). j, k, Initial rate of reaction versus DAG concentration measured using the dimeric (j) or tetrameric (k) DGAT1. Data are mean ± s.e.m. derived from three independent repeats. Experiments were repeated independently 10 times with similar results (a, c) or 3 times with similar results (b, e, f).
Extended Data Fig. 3 Cryo-EM data and processing.
a. A representative micrograph of DGAT1 (left), its Fourier transform (right) and representative 2D class averages (middle). Representative particles are highlighted with red circles. b, c, A flow chart for data processing and the final maps of DGAT1 (Methods). d, The gold-standard Fourier shell correlation (FSC) curve for the final map shown in c. e, Fourier shell correlation curves of the atomic model of DGAT1 versus the full map and individual half maps. f, Local-resolution map of DGAT1 shown in two orientations.
Extended Data Fig. 4 Density maps and structural model of DGAT1.
a, The overall map (left) and cartoon representation (right) of DGAT1. b, Individual secondary structures of DGAT1 shown as sticks, contoured in their density (green mesh). The density for oleoyl-CoA (green mesh) is shown at the same contour level as its neighbouring helix, TM7 (red mesh). The tubular density (green mesh) is shown at the same contour level as its neighbouring helix, TM4 (red mesh). c, Detailed view of each detergent or lipid molecule and its density. d, Electrostatic surface representations of the DGAT1 dimer in three orientations. The electrostatic potential was calculated using the APBS plug-in40 from PyMOL.
Extended Data Fig. 5 Binding of the N terminus to the neighbouring protomer and its functional consequences.
a, b, The DGAT1 dimer (cartoon) viewed in two orientations. c–e, Detailed view of the boxed regions shown in a, b. Residues involved in the interactions are shown as sticks. f, Size-exclusion profiles of N-terminal truncations. Experiments in f were repeated independently three times with similar results.
Extended Data Fig. 6 Oleoyl-CoA-binding site.
a–c, Oleoyl-CoA (spheres) bound to the DGAT1 protomer (cartoon) in three orientations. d–g, Detailed views of the boxed regions in a–c. Residues that coordinate oleoyl-CoA are shown as sticks with carbon atoms in magenta. h, LigPlus41,42 plot of the oleoyl-CoA-binding site. i, Normalized enzymatic activity of wild-type DGAT1 and various mutants. Data are mean ± s.e.m. derived from three independent repeats. j, Size-exclusion profiles of DGAT1 mutants. Experiments in i were repeated independently three times with similar results.
Extended Data Fig. 7 DGAT1 sequence alignment.
a, DGAT1 from human (Homo, Uniprot accession number O75907), mouse (Mus, Q9Z2A7), frog (Xenopus, A0A1L8G0L4), fish (Danio, Q6P3J0) and thale cress (Arabidopsis, Q9SLD2) are aligned using the Clustal Omega server43. Secondary structural elements of human DGAT1 are marked above the alignment. Residues are coloured on the basis of their conservation using the ESPript server44. Residues at the acyl-CoA-binding site are marked with green triangles and those at the active site with red stars. b, ConSurf mapped onto the human DGAT1 structure. DGAT1 is shown in different orientations in cartoon (top) and surface (bottom) representations. One DGAT1 monomer is coloured on the basis of the conservation score of each residue, calculated by the ConSurf server45. c, Sequence alignments of the FYXDWWN motif (red rectangle) from DGAT1 of different species and human ACAT1 (Uniprot P35610).
Extended Data Fig. 8 Proposed gateway for DAG entry.
a–c, A large opening to the core of the membrane is framed by TM4 and TM6. A tubular density is observed extending into the reaction chamber. d. Residues that line the opening are shown as magenta sticks.
Supplementary information
Supplementary Figure
This file contains Supplementary Figure 1: source data for the SDS-PAGE.
Rights and permissions
About this article
Cite this article
Wang, L., Qian, H., Nian, Y. et al. Structure and mechanism of human diacylglycerol O-acyltransferase 1. Nature 581, 329–332 (2020). https://doi.org/10.1038/s41586-020-2280-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-020-2280-2
This article is cited by
-
Lipid droplets and cellular lipid flux
Nature Cell Biology (2024)
-
Mechanism of d-alanine transfer to teichoic acids shows how bacteria acylate cell envelope polymers
Nature Microbiology (2023)
-
Lipid droplet biogenesis and functions in health and disease
Nature Reviews Endocrinology (2023)
-
The structure of phosphatidylinositol remodeling MBOAT7 reveals its catalytic mechanism and enables inhibitor identification
Nature Communications (2023)
-
Mechanism of action for small-molecule inhibitors of triacylglycerol synthesis
Nature Communications (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.