Abstract
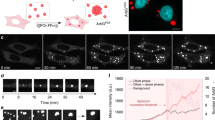
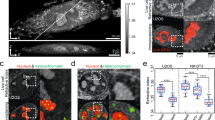
Intracellular bodies such as nucleoli, Cajal bodies and various signalling assemblies represent membraneless organelles, or condensates, that form via liquid–liquid phase separation (LLPS)1,2. Biomolecular interactions—particularly homotypic interactions mediated by self-associating intrinsically disordered protein regions—are thought to underlie the thermodynamic driving forces for LLPS, forming condensates that can facilitate the assembly and processing of biochemically active complexes, such as ribosomal subunits within the nucleolus. Simplified model systems3,4,5,6 have led to the concept that a single fixed saturation concentration is a defining feature of endogenous LLPS7,8,9, and has been suggested as a mechanism for intracellular concentration buffering2,7,8,10. However, the assumption of a fixed saturation concentration remains largely untested within living cells, in which the richly multicomponent nature of condensates could complicate this simple picture. Here we show that heterotypic multicomponent interactions dominate endogenous LLPS, and give rise to nucleoli and other condensates that do not exhibit a fixed saturation concentration. As the concentration of individual components is varied, their partition coefficients change in a manner that can be used to determine the thermodynamic free energies that underlie LLPS. We find that heterotypic interactions among protein and RNA components stabilize various archetypal intracellular condensates—including the nucleolus, Cajal bodies, stress granules and P-bodies—implying that the composition of condensates is finely tuned by the thermodynamics of the underlying biomolecular interaction network. In the context of RNA-processing condensates such as the nucleolus, this manifests in the selective exclusion of fully assembled ribonucleoprotein complexes, providing a thermodynamic basis for vectorial ribosomal RNA flux out of the nucleolus. This methodology is conceptually straightforward and readily implemented, and can be broadly used to extract thermodynamic parameters from microscopy images. These approaches pave the way for a deeper understanding of the thermodynamics of multicomponent intracellular phase behaviour and its interplay with the nonequilibrium activity that is characteristic of endogenous condensates.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Shin, Y. & Brangwynne, C. P. Liquid phase condensation in cell physiology and disease. Science 357, eaaf4382 (2017).
Banani, S. F., Lee, H. O., Hyman, A. A. & Rosen, M. K. Biomolecular condensates: organizers of cellular biochemistry. Nat. Rev. Mol. Cell Biol. 18, 285–298 (2017).
Nott, T. J. et al. Phase transition of a disordered nuage protein generates environmentally responsive membraneless organelles. Mol. Cell 57, 936–947 (2015).
Shin, Y. et al. Spatiotemporal control of intracellular phase transitions using light-activated optoDroplets. Cell 168, 159–171.e14 (2017).
Wang, J. et al. A molecular grammar governing the driving forces for phase separation of prion-like RNA binding proteins. Cell 174, 688–699.e16 (2018).
Bracha, D. et al. Mapping local and global liquid phase behavior in living cells using photo-oligomerizable seeds. Cell 175, 1467–1480.e13 (2018).
Holehouse, A. S. & Pappu, R. V. Functional implications of intracellular phase transitions. Biochemistry 57, 2415–2423 (2018).
Alberti, S., Gladfelter, A. & Mittag, T. Considerations and challenges in studying liquid-liquid phase separation and biomolecular condensates. Cell 176, 419–434 (2019).
McSwiggen, D. T., Mir, M., Darzacq, X. & Tjian, R. Evaluating phase separation in live cells: diagnosis, caveats, and functional consequences. Genes Dev. 33, 1619–1634 (2019).
Oltsch, F., Klosin, A., Julicher, F., Hyman, A. A. & Zechner, C. Phase separation provides a mechanism to reduce noise in cells. Science 367, 464–468 (2019).
Feric, M. et al. Coexisting liquid phases underlie nucleolar subcompartments. Cell 165, 1686–1697 (2016).
Mitrea, D. M. et al. Nucleophosmin integrates within the nucleolus via multi-modal interactions with proteins displaying R-rich linear motifs and rRNA. eLife 5, e13571 (2016).
Flory, P. J. Principles of Polymer Chemistry (Cornell Univ. Press, 1953).
Wei, M.-T., Chang, Y.-C., Shimobayashi, S. F., Shin, Y. & Brangwynne, C. P. Nucleated transcriptional condensates amplify gene expression. Preprint at https://www.biorxiv.org/content/10.1101/737387v2 (2019).
Kedersha, N. et al. G3BP–Caprin1–USP10 complexes mediate stress granule condensation and associate with 40S subunits. J. Cell Biol. 212, e201508028 (2016).
Choi, J.-M., Dar, F. & Pappu, R. V. LASSI: A lattice model for simulating phase transitions of multivalent proteins. PLoS Comput. Biol. 15, e1007028 (2019).
Mao, S., Kuldinow, D., Haataja, M. P. & Košmrlj, A. Phase behavior and morphology of multicomponent liquid mixtures. Soft Matter 15, 1297–1311 (2019).
Priftis, D. & Tirrell, M. Phase behaviour and complex coacervation of aqueous polypeptide solutions. Soft Matter 8, 9396–9405 (2012).
Jacobs, W. M. & Frenkel, D. Phase transitions in biological systems with many components. Biophys. J. 112, 683–691 (2017).
Mitrea, D. M. et al. Self-interaction of NPM1 modulates multiple mechanisms of liquid–liquid phase separation. Nat. Commun. 9, 842 (2018).
Lin, Y.-H., Brady, J. P., Forman-Kay, J. D. & Chan, H. S. Charge pattern matching as a ‘fuzzy’ mode of molecular recognition for the functional phase separations of intrinsically disordered proteins. New J. Phys. 19, 115003 (2017).
Banerjee, P. R., Milin, A. N., Moosa, M. M., Onuchic, P. L. & Deniz, A. A. Reentrant phase transition drives dynamic substructure formation in ribonucleoprotein droplets. Angew. Chem. Int. Ed. 56, 11354–11359 (2017).
Ferrolino, M. C., Mitrea, D. M., Michael, J. R. & Kriwacki, R. W. Compositional adaptability in NPM1–SURF6 scaffolding networks enabled by dynamic switching of phase separation mechanisms. Nat. Commun. 9, 5064 (2018).
Banani, S. F. et al. Compositional control of phase-separated cellular bodies. Cell 166, 651–663 (2016).
Geuskens, M. & Bernhard, W. Cytochimie ultrastructurale du nucléole. 3. Action de l’actinomycine D sur le métabolisme du RNA nucléolaire. Exp. Cell Res. 44, 579–598 (1966).
Lazdins, I. B., Delannoy, M. & Sollner-Webb, B. Analysis of nucleolar transcription and processing domains and pre-rRNA movements by in situ hybridization. Chromosoma 105, 481–495 (1997).
Burger, K. et al. Chemotherapeutic drugs inhibit ribosome biogenesis at various levels. J. Biol. Chem. 285, 12416–12425 (2010).
Wei, M.-T. et al. Phase behaviour of disordered proteins underlying low density and high permeability of liquid organelles. Nat. Chem. 9, 1118–1125 (2017).
Zhu, L. et al. Controlling the material properties and rRNA processing function of the nucleolus using light. Proc. Natl Acad. Sci. USA 116, 17330–17335 (2019).
Handwerger, K. E., Cordero, J. A. & Gall, J. G. Cajal bodies, nucleoli, and speckles in the Xenopus oocyte nucleus have a low-density, sponge-like structure. Mol. Biol. Cell 16, 202–211 (2005).
Acknowledgements
We thank members of the Brangwynne laboratory for discussions and comments on this manuscript. This work was supported by the Howard Hughes Medical Institute, the St Jude Collaborative on Membraneless Organelles, and grants from the National Institutes of Health (NIH) 4D Nucleome Program (U01 DA040601) and the Princeton Center for Complex Materials, a Materials Research Science and Engineering Center supported by the National Science Foundation (NSF) (DMR 1420541). L.Z. was supported by an NSF graduate fellowship (DGE-1656466). R.W.K acknowledges support from the NIH (R01 GM115634, R35 GM131891 and P30 CA021765 (to St Jude Children’s Research Hospital)) and ALSAC. M.T. acknowledges support from the NIH (F32 GM131524). Some images were acquired at the St Jude Cell & Tissue Imaging Center, which is supported by St Jude Children’s Research Hospital and the National Cancer Institute (P30 CA021765); we thank V. Frohlich and J. Peters for technical assistance.
Author information
Authors and Affiliations
Contributions
J.A.R., L.Z., D.M.M., R.W.K. and C.P.B. designed the research; in vivo studies were performed and analysed by J.A.R., L.Z., D.W.S. and M.W.; in vitro studies were performed and analysed by M.C.F., M.T. and D.M.M.; J.A.R., L.Z. and C.P.B. wrote the manuscript, which was reviewed and edited by all authors.
Corresponding authors
Ethics declarations
Competing interests
R.W.K. is a consultant for, and D.M.M. has recently become employed by, Dewpoint Therapeutics. The remaining authors declare no competing interests.
Additional information
Peer review information Nature thanks Rohit Pappu and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 NPM1 lacks a fixed Cdil and Cden, suggesting that nucleoli undergo multicomponent-mediated phase separation.
a, b, Dependence of the measured concentration of NPM1 in the relevant dense phase (here ‘den’ refers to the granular component of nucleoli) (a) and the apparent partition coefficient of NPM1 (that is, the ratio of its concentration in the dense and dilute phases) (b) on the total concentration of NPM1 in the nucleus. c, Dependence of the transfer free energy on the concentration of NPM1 in the dilute phase, for mCherry-tagged NPM1 (filled circles) and mGFP-tagged (open circles). The trends for each are similar. Dashed lines represent mean confidence intervals to fits described in the Supplementary Methods; the red lines in a, b represent expected trends for single-biomolecule phase separation.
Extended Data Fig. 2 G3BP1, coilin and DCP1A lack fixed Cdil and Cden in cells.
a–c, Relationship between the approximated total concentration and the dilute concentration in cells expressing variable amounts of fluorescently tagged G3BP1 (a), coilin (b) and DCP1A (c). Points are red in a to indicate that only cells with phase separation in the G3BP1 double knockout line after stress are included. d–f, Relationship between the dilute and dense concentrations for cells expressing variable amounts of fluorescently tagged G3BP1 (d), coilin (e) and DCP1A (f). Dashed lines represent mean confidence intervals to fits described in the Supplementary Methods. Statistical significance (P < 0.01) for these increasing monotonic relationships between the axes are reported in Supplementary Methods. Red points in d and f represent diffraction-limited foci.
Extended Data Fig. 3 In silico validation of the composition dependence of phase separation using Flory–Huggins theory.
Phase separation of two (non-solvent) components, denoted #1 and #2, with their heterotypic interactions being equal, stronger and weaker, than their homotypic interactions shown as black, blue and orange, respectively, for a–c. a, The initial dependence of [#1]dil on [#1]tot at fixed [#2]tot, such that phase separation will occur at the ‘goldilocks point’—when [#1]tot = [#2]tot. The axes are normalized by the initial saturation (init sat) concentration—that is, the lowest [#1]tot at which phase separation emerges. The dashed line is the 1:1 line that would expected without phase separation. b, c, \(\Delta {G}_{\#1}^{{\rm{tr}}}\) (b) and \(\Delta {G}_{\#2}^{{\rm{tr}}}\) (c) as a function of [#1]dil. Circles indicate the location of the goldilocks point under each condition. d, The change in ΔGtr with respect to [#1]dil as a function of the heterotypic interaction strength χ12 (in which more negative implies stronger heterotypic interactions) at the goldilocks point for the transfer free energy of #1 and #2, as indicated.
Extended Data Fig. 4 In vitro destabilization of SURF6N partitioning by increasing the concentration of SURF6N itself or NPM1.
a, b, Changes in the transfer free energy of SURF6N forming into multicomponent droplets as additional SURF6N (a) or additional NPM1 (b) is added on top of NPM1:SURF6N:rRNA ternary droplets as described in the Supplementary Methods. The number of droplets (in order of increasing concentration) are n = 122, 115, 105, 98, 91, 74 and 99. Data are mean ± s.d. c, Phase diagram in vitro in the presence of 25 ng μl−1 wheatgerm rRNA, 5 μM SURF6N, and various concentrations of NPM1. Units shown are absorbance units corrected for background, quantum yield differences between the two phases, and the (nonlinear) fraction labelled of NPM1. d, Changes in the phase diagram as additional NPM1 is added. As in c, NPM1 concentrations in the dense or dilute phases are indicative of total NPM1. Hyperbolic fits shown highlight that the largest changes upon NPM1 addition are from an increase in the dilute phase concentration of NPM1 and a decrease in the dense phase concentration of SURF6N. To assess significance, the y axes in d are shown from zero arbitrary units (AU) to 2.5 times the mean of all points shown.
Extended Data Fig. 5 The change in the transfer free energy for R -proteins and NPM1.
ΔΔGtr of r-proteins RPL23A (left) and RPL5 (right) compared with that for SURF6 as the concentration of NPM1 is increased.
Extended Data Fig. 6 ActD treatment decreases the size of the nucleolus.
The fraction of the image area corresponding to the nucleolus as a function of time after the addition of ActD in individual cells expressing NPM1–mCherry. The colours represent the same cells as in Fig. 3g.
Extended Data Fig. 7 Characterization of Corelet non-ideality and extrapolation from high valence.
a, The transfer free energy for the N-terminal half of NPM1 (NC)-sspB in cells without the core expressed (orange representing ΔGtr) or with the indicated valences after core activation (black indicating ΔΔGtrNC-Core). The ΔΔGtrNC-Core in this case is the energetic difference between the NC and core channels, which is approximately the energetic difference for transferring an additional NC to the core at that valence. b, At valences higher then 24, the transfer free energy is approximated as quadratic and extrapolated back to a valence of 24 to obtain the transfer free energy at this valence. c, Transfer free energy reported from the sspB channel as a function of valence, which is weighted by the number of sspB molecules (owing to the number of mCherry molecules observed being proportional to the valence of each molecule, as opposed to at the core where it is always constant at 24 GFPs). The red point represents the extrapolated value and mean confidence error as determined in b.
Extended Data Fig. 8 Controls for ribosomal mimics.
a, b, SDS–PAGE (a) and denaturing agarose gel (b) detailing the purity of reagents used in the experiments in Fig. 4d, e. c, Microscopy image of 10 μM NPM1-594 droplets formed with 5% PEG without any rRNA. The limited fluorescence indicates that neither NPM1 nor PEG binds SYTO 40 and the droplet environment does not promote the fluorescence of SYTO 40. d, Phase separation assessed by turbidity of the indicated ribosomal substrate (fixed at 50 μg ml−1) as a function of NPM1 concentration. The dashed grey line indicates where phase separation is typically observed in microscopy measurements.
Supplementary information
Supplementary Information
Supplemental Notes: This file contains notes for field specific discussion and clarification.
Supplementary Figure
Figure 1: Uncropped gel images of Extended Data Figure 8A-B. (A) SDS-PAGE of NPM1 and 30S subunit, and (B) denaturing agarose gels of 16S and 30S used in the experiments presented in Figure 4D and E.
Source data
Rights and permissions
About this article
Cite this article
Riback, J.A., Zhu, L., Ferrolino, M.C. et al. Composition-dependent thermodynamics of intracellular phase separation. Nature 581, 209–214 (2020). https://doi.org/10.1038/s41586-020-2256-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-020-2256-2
This article is cited by
-
Physiology and pharmacological targeting of phase separation
Journal of Biomedical Science (2024)
-
Mex-3 RNA binding family member A (MEX3A)/circMPP6 complex promotes colorectal cancer progression by inhibiting autophagy
Signal Transduction and Targeted Therapy (2024)
-
Sequestration within peptide coacervates improves the fluorescence intensity, kinetics, and limits of detection of dye-based DNA biosensors
Communications Chemistry (2024)
-
Biomolecular condensates form spatially inhomogeneous network fluids
Nature Communications (2024)
-
A solid beta-sheet structure is formed at the surface of FUS droplets during aging
Nature Chemical Biology (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.