Abstract
The initiation of an intestinal tumour is a probabilistic process that depends on the competition between mutant and normal epithelial stem cells in crypts1. Intestinal stem cells are closely associated with a diverse but poorly characterized network of mesenchymal cell types2,3. However, whether the physiological mesenchymal microenvironment of mutant stem cells affects tumour initiation remains unknown. Here we provide in vivo evidence that the mesenchymal niche controls tumour initiation in trans. By characterizing the heterogeneity of the intestinal mesenchyme using single-cell RNA-sequencing analysis, we identified a population of rare pericryptal Ptgs2-expressing fibroblasts that constitutively process arachidonic acid into highly labile prostaglandin E2 (PGE2). Specific ablation of Ptgs2 in fibroblasts was sufficient to prevent tumour initiation in two different models of sporadic, autochthonous tumorigenesis. Mechanistically, single-cell RNA-sequencing analyses of a mesenchymal niche model showed that fibroblast-derived PGE2 drives the expansion οf a population of Sca-1+ reserve-like stem cells. These express a strong regenerative/tumorigenic program, driven by the Hippo pathway effector Yap. In vivo, Yap is indispensable for Sca-1+ cell expansion and early tumour initiation and displays a nuclear localization in both mouse and human adenomas. Using organoid experiments, we identified a molecular mechanism whereby PGE2 promotes Yap dephosphorylation, nuclear translocation and transcriptional activity by signalling through the receptor Ptger4. Epithelial-specific ablation of Ptger4 misdirected the regenerative reprogramming of stem cells and prevented Sca-1+ cell expansion and sporadic tumour initiation in mutant mice, thereby demonstrating the robust paracrine control of tumour-initiating stem cells by PGE2–Ptger4. Analyses of patient-derived organoids established that PGE2–PTGER4 also regulates stem-cell function in humans. Our study demonstrates that initiation of colorectal cancer is orchestrated by the mesenchymal niche and reveals a mechanism by which rare pericryptal Ptgs2-expressing fibroblasts exert paracrine control over tumour-initiating stem cells via the druggable PGE2–Ptger4–Yap signalling axis.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
All data that support the findings of this study are available within the paper and its Supplementary Information files. All Drop-seq data that support the findings of this study have been deposited in the Gene Expression Omnibus (GEO) repository with the accession code GSE142431.
Code availability
The code used for single-cell RNA-seq data analysis is available in GitHub (https://github.com/KlugerLab/Scripts_Roulis_et_al_2020).
References
Vermeulen, L. & Snippert, H. J. Stem cell dynamics in homeostasis and cancer of the intestine. Nat. Rev. Cancer 14, 468–480 (2014).
Powell, D. W., Pinchuk, I. V., Saada, J. I., Chen, X. & Mifflin, R. C. Mesenchymal cells of the intestinal lamina propria. Annu. Rev. Physiol. 73, 213–237 (2011).
Kinchen, J. et al. Structural remodeling of the human colonic mesenchyme in inflammatory bowel disease. Cell 175, 372–386 (2018).
Hamilton, T. G., Klinghoffer, R. A., Corrin, P. D. & Soriano, P. Evolutionary divergence of platelet-derived growth factor alpha receptor signaling mechanisms. Mol. Cell. Biol. 23, 4013–4025 (2003).
Molotkov, A., Mazot, P., Brewer, J. R., Cinalli, R. M. & Soriano, P. Distinct requirements for FGFR1 and FGFR2 in primitive endoderm development and exit from pluripotency. Dev. Cell. 41, 511–526 (2017).
Smyth, E. M., Grosser, T., Wang, M., Yu, Y. & FitzGerald, G. A. Prostanoids in health and disease. J. Lipid Res. 50 (Suppl), S423–S428 (2009).
Wang, D. & DuBois, R. N. The role of anti-inflammatory drugs in colorectal cancer. Annu. Rev. Med. 64, 131–144 (2013).
Barker, N. et al. Crypt stem cells as the cells-of-origin of intestinal cancer. Nature 457, 608–611 (2009).
Chulada, P. C. et al. Genetic disruption of Ptgs-1, as well as Ptgs-2, reduces intestinal tumorigenesis in Min mice. Cancer Res. 60, 4705–4708 (2000).
Cherukuri, D. P. et al. Targeted Cox2 gene deletion in intestinal epithelial cells decreases tumorigenesis in female, but not male, Apc Min/+ mice. Mol. Oncol. 8, 169–177 (2014).
Xia, D., Wang, D., Kim, S. H., Katoh, H. & DuBois, R. N. Prostaglandin E2 promotes intestinal tumor growth via DNA methylation. Nat. Med. 18, 224–226 (2012).
Wang, D., Fu, L., Sun, H., Guo, L. & DuBois, R. N. Prostaglandin E2 promotes colorectal cancer stem cell expansion and metastasis in mice. Gastroenterology 149, 1884–1895 (2015).
Bygdeman, M. Pharmacokinetics of prostaglandins. Best Pract. Res. Clin. Obstet. Gynaecol. 17, 707–716 (2003).
Mustata, R. C. et al. Identification of Lgr5-independent spheroid-generating progenitors of the mouse fetal intestinal epithelium. Cell Rep. 5, 421–432 (2013).
Haber, A. L. et al. A single-cell survey of the small intestinal epithelium. Nature 551, 333–339 (2017).
Ayyaz, A. et al. Single-cell transcriptomes of the regenerating intestine reveal a revival stem cell. Nature 569, 121–125 (2019).
Miyoshi, H. et al. Prostaglandin E2 promotes intestinal repair through an adaptive cellular response of the epithelium. EMBO J. 36, 5–24 (2017).
Gregorieff, A., Liu, Y., Inanlou, M. R., Khomchuk, Y. & Wrana, J. L. Yap-dependent reprogramming of Lgr5+ stem cells drives intestinal regeneration and cancer. Nature 526, 715–718 (2015).
Hong, A. W., Meng, Z. & Guan, K. L. The Hippo pathway in intestinal regeneration and disease. Nat. Rev. Gastroenterol. Hepatol. 13, 324–337 (2016).
Cai, J., Maitra, A., Anders, R. A., Taketo, M. M. & Pan, D. β-Catenin destruction complex-independent regulation of Hippo–YAP signaling by APC in intestinal tumorigenesis. Genes Dev. 29, 1493–1506 (2015).
Kim, H. B. et al. Prostaglandin E2 activates YAP and a positive-signaling loop to promote colon regeneration after colitis but also carcinogenesis in mice. Gastroenterology 152, 616–630 (2017).
Yu, F. X. et al. Regulation of the Hippo–YAP pathway by G-protein-coupled receptor signaling. Cell 150, 780–791 (2012).
Liu-Chittenden, Y. et al. Genetic and pharmacological disruption of the TEAD–YAP complex suppresses the oncogenic activity of YAP. Genes Dev. 26, 1300–1305 (2012).
Mills, J. C. & Sansom, O. J. Reserve stem cells: differentiated cells reprogram to fuel repair, metaplasia, and neoplasia in the adult gastrointestinal tract. Sci. Signal. 8, re8 (2015).
Nusse, Y. M. et al. Parasitic helminths induce fetal-like reversion in the intestinal stem cell niche. Nature 559, 109–113 (2018).
Huyghe, J. R. et al. Discovery of common and rare genetic risk variants for colorectal cancer. Nat. Genet. 51, 76–87 (2019).
Ishikawa, T. O. & Herschman, H. R. Conditional knockout mouse for tissue-specific disruption of the cyclooxygenase-2 (Cox-2) gene. Genesis 44, 143–149 (2006).
Armaka, M. et al. Mesenchymal cell targeting by TNF as a common pathogenic principle in chronic inflammatory joint and intestinal diseases. J. Exp. Med. 205, 331–337 (2008).
Muzumdar, M. D., Tasic, B., Miyamichi, K., Li, L. & Luo, L. A global double-fluorescent Cre reporter mouse. Genesis 45, 593–605 (2007).
Moser, A. R., Pitot, H. C. & Dove, W. F. A dominant mutation that predisposes to multiple intestinal neoplasia in the mouse. Science 247, 322–324 (1990).
Schneider, A. et al. Generation of a conditional allele of the mouse prostaglandin EP4 receptor. Genesis 40, 7–14 (2004).
Zhang, N. et al. The Merlin/NF2 tumor suppressor functions through the YAP oncoprotein to regulate tissue homeostasis in mammals. Dev. Cell 19, 27–38 (2010).
Madison, B. B. et al. Cis elements of the villin gene control expression in restricted domains of the vertical (crypt) and horizontal (duodenum, cecum) axes of the intestine. J. Biol. Chem. 277, 33275–33283 (2002).
Barker, N. et al. Identification of stem cells in small intestine and colon by marker gene Lgr5. Nature 449, 1003–1007 (2007).
Madisen, L. et al. A robust and high-throughput Cre reporting and characterization system for the whole mouse brain. Nat. Neurosci. 13, 133–140 (2010).
Roulis, M. et al. Intestinal myofibroblast-specific Tpl2–Cox-2–PGE2 pathway links innate sensing to epithelial homeostasis. Proc. Natl Acad. Sci. USA 111, E4658–E4667 (2014).
Masoodi, M. & Nicolaou, A. Lipidomic analysis of twenty-seven prostanoids and isoprostanes by liquid chromatography/electrospray tandem mass spectrometry. Rapid Commun. Mass Spectrom. 20, 3023–3029 (2006).
Macosko, E. Z. et al. Highly parallel genome-wide expression profiling of individual cells using nanoliter droplets. Cell 161, 1202–1214 (2015).
Shekhar, K. et al. Comprehensive classification of retinal bipolar neurons by single-cell transcriptomics. Cell 166, 1308–1323 (2016).
Linderman, G. C., Zhao, J. & Kluger, Y. Zero-preserving imputation of scRNA-seq data using low-rank approximation. Preprint at https://www.bioRxiv.org/content/10.1101/397588v1 (2018).
van der Maaten, L. & Hinton, G. Viualizing data using t-SNE. J. Mach. Learn. Res. 9, 2579–2605 (2008).
Ester, M., Kriegel, H.-P., Sander, J. & Xu, X. A density-based algorithm for discovering clusters in large spatial databases with noise. KDD 96, 226–231 (1996).
Hänzelmann, S., Castelo, R. & Guinney, J. GSVA: gene set variation analysis for microarray and RNA-seq data. BMC Bioinformatics 14, 7 (2013).
Acknowledgements
We thank C. Lieber, J. Alderman and E. Hughes-Picard for administrative assistance; the Yale Pathology Tissue Service–Tissue Procurement and Distribution Facility for providing human tissue samples; D. Gonzalez for assistance in two-photon imaging; M. Graham for assistance in electron microscopy; M. Samiotaki and T. Wu for assistance in mass spectrometry; and Flavell laboratory members R. Jackson and W. Bailis for discussions. M.R. is supported by a Crohn’s and Colitis Foundation Career Development Award (510777); M.S., L.-S.F., M.S.K. and M.B. were supported by an Austrian Marshall Plan Foundation Master’s Fellowship. This work was supported in part by ERC project MCs-inTEST (340217) (G.K.), the National Natural Science Foundation of China (31930035, 91942311) (B.S.), the Blavatnik Family Foundation and the Howard Hughes Medical Institute (R.A.F.).
Author information
Authors and Affiliations
Contributions
M.R. conceived and designed the study and wrote the manuscript. M.R. designed, performed and analysed experiments in Figs. 1–4 and Extended Data Figs. 1–10, assisted by M.S., L.-S.F., M.S.K., M.B., H.N.B. and J.R.B.; A.K. performed experiments in Fig. 2 and Extended Data Figs. 3, 9, assisted by V.K., N.C. and A.H. P.B. implemented Drop-seq. J.Z., R.Q. and Y.K. analysed Drop-seq data. E.K. and V.A. implemented HPLC–MS/MS analyses. X.Z. and B.S. performed in situ hybridization, H.R.H. and J.J. contributed Ptgs2f/f and Ptgs2LSL mice, R.M.B. contributed Ptger4f/f mice and D.J. contributed human FFPE tissues. G.K. and R.A.F. supervised all research, participated in the interpretation of results and edited the manuscript.
Corresponding authors
Ethics declarations
Competing interests
R.A.F. is a scientific advisor to GlaxoSmithKline and a shareholder and consultant for Zai Lab. All other authors declare no competing interests.
Additional information
Peer review information Nature thanks Garret A. FitzGerald, Dominic Grün, Kun-Liang Guan, Don W. Powell, Omer Yilmaz and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Mesenchymal Ptgs2 expression in the microenvironment of crypt stem cells.
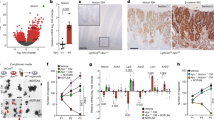
a, Transmission electron microscopy photograph of the base of a mouse ileum crypt. P, Paneth cell; S, columnar basal stem cell; F, fibroblast. Scale bar, 5 μm. Indicative of independent observations in two experiments. b, Immunostaining for Lgr5–eGFP and Vimentin in the ileum of an Lgr5-eGFP-IRES-creERT2 mouse. Scale bar, 20 μm. Indicative of independent observations in one experiment. c, PTGS2 relative gene expression (RE) in intestinal epithelial cells (IECs) and stromal cells isolated from healthy human colonic tissues (n = 6 individuals). Statistical comparison performed with two-tailed Wilcoxon matched-pairs signed-rank test. d, Ptgs2 gene expression in IECs and stromal cells isolated from the ileum and the colon of wild-type mice (n = 4). Statistical significance was determined by two-tailed paired t-test. e, Biological replicates of the Drop-seq experiment shown in Fig. 1a visualized on the respective t-SNE plot depicting n = 3,179 mesenchymal cells. Mesenchymal cells were independently isolated from two groups of wild-type mice (biological replicates 1 and 2). From each of these isolations up to three independent Drop-seq samples were collected (A to C) for a total of five samples. f, All Ptgs2-expressing single cells (n = 1,136) detected in the experiment shown in Fig. 1a, c were analysed separately and re-clustered. Cluster annotations are visualized on a t-SNE plot. Violin plots display the entire distribution of gene expression levels per single cell in each cluster for key mesenchymal marker genes. F, fibroblasts. g, Schematic representation of the arachidonic acid metabolism pathway. For each mesenchymal cluster shown in Fig. 1a, violin plots display the entire distribution of gene expression levels per single cell for six genes involved in the metabolism of arachidonic acid to prostanoids. Data from n = 3,179 single mesenchymal cells are shown. h, Analysis of single-cell RNA-seq data (GSE11434) from the healthy human colonic mesenchyme3. Clustering results for n = 4,348 cells and cluster annotations are visualized on a t-SNE plot. The annotations of stromal populations are matched with the ones reported by Kinchen et al.3 on the basis of the respective markers. Expression levels of PTGS2 per single cell are visualized on a t-SNE plot. The entire range of gene expression levels per single cell for PTGS1, PTGS2 and key mesenchymal marker genes is displayed in violin plots. Data are mean ± s.e.m.; ns, non-significant; *P < 0.05; **P < 0.01.
Extended Data Fig. 2 Location of major fibroblast populations in the mouse intestine.
a, Detection of Pdgfra-expressing mesenchymal cells in the intestine of adult Pdgfra-H2B-eGPF-knockin mice4. Two distinct populations of Pdgfrahigh and Pdgfralow mesenchymal cells were detected in fixed tissue sections by direct eGFP fluorescence (green) and confocal microscopy. Nuclei are stained with DAPI (blue). Pdgfrahigh cells are located under the epithelium along the crypt–villus axis and in the muscularis propria. They form clusters at the tips of villi and the apical part of the colonic mucosa. Pdgfralow cells are located in the inner part of the villi, the pericryptal area and the submucosa. Filled arrows indicate pericryptal Pdgfralow cells. Open arrows indicate subepithelial Pdgfrahigh cells. M, mucosa; V, villus; SM, submucosa; MP, muscularis propria. Scale bars, 20 μm. Data are representative of six independent experiments. b, Detection of Pdgfrahigh and Pdgfralow fibroblasts in the fresh, intact intestine of adult Pdgfra-H2B-eGPF-knockin mice4 by two-photon microscopy. The cells were detected by direct eGFP fluorescence (green). Pdgfrahigh cells are predominant in the muscularis propria, whereas Pdgfralow cells are predominant in the submucosa. Both populations are present in the mucosa. Data are representative of independent observations from one experiment. Scale bars, 100 μm. c, Detection of Pdgfrahigh and Pdgfralow fibroblasts in the intestine of Pdgfra-H2B-eGPF-knockin embryos on embryonic day 15 (E15.0) and in early postnatal development. E15.0: clusters of Pdgfrahigh cells in early villi are indicated by white arrows. Pdgfralow mesenchymal cells occupy the rest of the mesenchyme (asterisks). P0: Pdgfrahigh cells are observed in the villi (V) and Pdgfralow cells are observed both in the villi and in the rest of the mesenchyme (asterisks). P15: Pdgfralow cells surround an early crypt (C) and Pdgfrahigh cells are located at the edges of the crypt (open white arrows). Pdgfralow cells occupy the inner mesenchyme (asterisks). Data are representative of independent observations from one experiment per developmental stage. Scale bars, 20 μm. d, The location of Fgfr2-expressing mesenchymal cells was determined in the intestine of an Fgfr2-T2A-H2B-mCherry-knockin mouse5, by detecting direct mCherry fluorescence (red) in the nucleus (blue, DAPI). The arrows indicate pericryptal Fgfr2+ fibroblasts. Data are representative of independent observations from one experiment. Scale bars, 20 μm. e, Immunostaining for laminin A1 (encoded by Lama1), the epithelial marker E-cadherin and the mesenchymal marker vimentin in the normal mouse intestines shows that laminin A1 is detected specifically at the mesenchymal–epithelial interface. Data are representative of two independent experiments. Scale bars, 5 μm. f, In-situ hybridization analysis showing the location of Rspo1-expressing cells in the normal mouse colon. The position of Rspo1-expressing cells along the crypt axis was quantified in 40 × 80 μm2 sub-epithelial areas at the base, middle and top sections of n = 9 crypts. Unpaired two-tailed Student’s t-test. Mean ± s.e.m. **P < 0.01.
Extended Data Fig. 3 Mice with fibroblast-specific ablation or fibroblast-restricted expression of Cox-2.
a, Immunofluorescence of ileum and colon sections from Col6-cre-Rosa26tdTomato/+ mice (scale bar, 20 μm) and of a small intestinal tumour section from an ApcMin/+-Col6-cre-Rosa26tdTomato/+ mouse (scale bar, 150 μm). Data are representative of two experiments. b, Efficiency of Col6-cre-mediated recombination of a lox-stop-lox tdTomato reporter in Pdgfrahigh and Pdgfralow Cd45− cells determined by flow cytometry in intestinal mesenchymal and lamina propria cells isolated from the small intestine and the colon of Col6-cre-Rosa26tdTomato/+PdgfraeGFP/+ mice in one experiment. c, Ptgs2 relative gene expression (RE) in whole tissue, isolated IECs, FACS-sorted Col6Cre+ fibroblasts (CD45−tdTomato+) and Col6Cre− mesenchymal cells (CD45−tdTomato−) from the small intestine of Col6-cre-Rosa26tdTomato/+ mice (n = 3, pooled). Representative of two experiments. d, Efficiency of Col6-cre-mediated Ptgs2 gene ablation in Col6-Cre+ mesenchymal cells determined by RT–qPCR analysis of Ptgs2 expression in FACS-sorted Col6-cre+ fibroblasts (eGFP+) from the small intestine of Col6-cre-Rosa26mT/mGPtgs2f/+ (n = 3) and Col6-cre-Rosa26mT/mGPtgs2f/f (n = 3) mice. Unpaired two-tailed Welch's t-test. e, Expression of the Ptgs2 gene in whole tissue ileum of littermate Ptgs2f/f and Ptgs2ΔFibr mice (n = 7 each). Two-tailed t-test. f, Spleen weight of 5.5-month-old ApcMin/+Ptgs2f/f (n = 8) and ApcMin/+Ptgs2ΔFibr (n = 6) mice. Average spleen weight of (n = 6) normal littermates (Ptgs2f/f) is displayed for comparison. Two-tailed t-test. g, Survival analysis of ApcMin/+Ptgs2f/f (n = 12) and ApcMin/+Ptgs2ΔFibr (n = 12) mice. A two-tailed P = 0.00009687 was calculated by log-rank test. h, Size of 274 adenomas from 5.5-month-old ApcMin/+Ptgs2f/f (n = 16) and ApcMin/+Ptgs2ΔFibr (n = 18) mice. The whiskers extend from minimum to maximum and the box extends from the 25th to 75th percentiles with the median indicated. Two-tailed Mann–Whitney test. i, Generation of knockin mice bearing a lox-stop-lox cassette insertion in intron-3 of the Ptgs2 gene which prevents its expression (Ptgs2OFF). Col6-cre-mediated excision of the lox-stop-lox cassette reactivates Ptgs2 expression specifically in fibroblasts (Ptgs2FibrON). The orange box depicts an frt site remaining from the flp-mediated removal of an frt-flanked PGK-neomycin selection cassette (see Methods). j, Ptgs2f/f (n = 30) and Ptgs2ΔFibr (n = 24) mice were subjected to 10 weekly intraperitoneal injections with 10 mg kg−1 azoxymethane as displayed. Quantification of the number of dysplastic foci and microadenomas per mouse and quantification of tumour size is shown. Statistical significance was tested by two-tailed Mann–Whitney test. k, Quantification of intestinal epithelial populations in the ileum of littermate Ptgs2f/f and Ptgs2ΔFibr mice (n = 3–5 per genotype). Immunostaining was performed for markers of Paneth cells (lysozyme), tuft cells (Dclk1), enteroendocrine cells (chromogranin A) and stem cells (Olfm4). Goblet cells were identified by periodic acid Schiff (PAS) staining and enterocytes were identified by detecting alkaline phosphatase enzymatic activity. Incorporation and immunohistochemical detection of BrdU was used to determine the numbers of cycling cells. Data for each mouse represent mean number of positive cells per crypt or crypt–villus unit as indicated. N = 400–822 crypts and/or villi were evaluated per staining. Statistical comparisons were performed with two-tailed unpaired t-test except for Olfm4+ cells for which unpaired t-test with Welch’s correction was applied. Scale bars, 50 μm. All data represent mean ± s.e.m. unless otherwise indicated. ns, non-significant; *P < 0.05, **P < 0.01, ***P < 0.001.
Extended Data Fig. 4 PGE2-driven spheroids contain more functional stem cells.
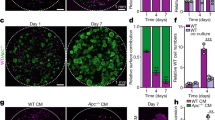
a, Crypts isolated from the small intestine of wild-type mice were grown into organoids by 3D culture with OGM or OGM that was supplemented daily with 0.1 μM 16,16-dimethyl PGE2 (dmPGE2). Indicative images and quantification of the absolute numbers of organoids and spheroids grown per 3D structure are shown. n = 6 3D cultures were evaluated per condition. Scale bar, 100 μm. b, Assessment of stem-cell activity in organoids or PGE2-driven spheroids grown as in a by dissociation into single cells and 3D culture in OGM. Growth of crypts and organoids from the same initial number of cells was quantified on day 14. The results are indicative of five independent experiments starting from independent crypt isolations. c, Normal crypts were grown into organoids with OGM in a 3D co-culture with primary mouse intestinal fibroblasts with or without 10 μM ONO-AE3-208 (Ptger4/EP4 inhibitor). Indicative images and quantification of the absolute numbers of organoids and spheroids grown per 3D structure are shown. n = 6 3D co-cultures were evaluated per condition. Scale bar, 200 μm. d, Separation of n = 2,192 fibroblasts and epithelial cells in single-cell RNA-seq data from fibroblast–crypt organotypic cultures on the basis of the expression of key marker genes. Expression of intestinal epithelial marker genes (Epcam, Atp1b1 and Krt8) and fibroblast marker genes (Sparc, Col1a1 and Col3a1) in single cells from Ptger4-ON and Ptger4–OFF fibroblast–crypt co-cultures is shown projected onto t-SNE plots. e–j, Single-cell data from Ptger4-ON and Ptger4–OFF fibroblast–crypt co-cultures as shown in Fig. 3d, visualized on the respective t-SNE plot depicting n = 1,585 epithelial cells. e, Expression of epithelial population-specific signatures (metagenes) per single epithelial cell. Population signatures were calculated on the basis of single-cell profiling of the mouse intestinal epithelium15. f, Cell cycle analysis of single epithelial cells projected onto the t-SNE plot. g–j, Expression levels of metagenes for the signatures or transcriptional programs of RSC16 (g), β-catenin (h), Yap (i) and early (non-tumour) ApcMin/+ tumorigenesis (j) per single epithelial cell projected onto t-SNE plots. k, Data from n = 1,585 single epithelial cells, visualized in violin plots for each co-culture condition (Ptger4-ON or Ptger4-OFF). The entire range of metagene expression levels per single epithelial cell for the signatures or transcriptional programs of RSCs, β-catenin, Yap and early ApcMin/+ tumorigenesis is displayed. In a, c, two-way ANOVA. Data are mean ± s.e.m. ****P < 0.0001.
Extended Data Fig. 5 Expression of PGE2 receptors in mouse and human tissues.
a, RT–qPCR analysis for Ptger1, Ptger2, Ptger3 and Ptger4 genes across 12 mouse tissues. Expression relative to B2m is displayed as 2−ΔCt. Data represent one experiment. MLN, mesenteric lymph nodes. b, RT–qPCR analysis for Ptger1, Ptger2, Ptger3 and Ptger4 genes in isolated IECs and matched stromal fractions from the small intestine (ileum) and the colon of wild-type mice (n = 4). Statistical comparisons were performed by two-tailed paired t-test. c, Expression levels of Ptger1, Ptger2, Ptger3 and Ptger4 genes determined by RNA-seq in FACS-sorted intestinal epithelial cell populations in 2 or 3 biological replicates and displayed as FPKM (fragments per kilobase of transcript per million mapped reads). Data retrieved from the GSE83394 GEO dataset. d, Expression levels of the human PTGER1, PTGER2, PTGER3 and PTGER4 genes in matched normal colon and tumour tissues from colorectal cancer patients (n = 41), determined by RNA-seq and displayed as FPKM. Data retrieved from The Cancer Genome Atlas for colon adenocarcinoma (TCGA–COAD dataset). Statistical comparisons were performed by two-tailed Wilcoxon matched-pairs signed-rank test. e, Analysis of single-cell RNA-seq data16 (GSE117783) from crypts isolated from the small intestine of normal mice (blue) and mice treated with 12 Gy irradiation (red). n = 6,644 single cells are visualized on t-SNE plots based on the experimental condition (normal, n = 2,882; irradiated, n = 3,762) and the clustering results. Violin plots represent the entire distribution of Ptger4 expression levels per single cell in each cluster and in each condition. The annotations of epithelial populations are matched with the ones reported by Ayyaz et al.16 on the basis of the respective markers. b–d, Mean ± s.e.m.; ns, non-significant; *P < 0.05; **P < 0.01.
Extended Data Fig. 6 PGE2–Ptger4 drive the induction of Yap target genes in intestinal organoids.
a, Volcano plot displaying the results of differential gene-expression analysis performed in single epithelial cells from Ptger4-ON and Ptger4–OFF fibroblast–crypt co-cultures (n = 1,585). Yap1 and Yap target genes18 are indicated. Moderated t-test with false-discovery rate (Benjamini–Hochberg) correction. b, Expression levels of the genes indicated in single epithelial cells from Ptger4-ON and Ptger4–OFF fibroblast–crypt co-cultures (n = 1,585), projected onto t-SNE plots. c, Experimental setup for data shown in d, e. Crypts were grown into organoids or spheroids by 3D culture in OGM or OGM that was supplemented daily with 0.1 μM dmPGE2 for 7 days. Gene expression levels were measured by RT–qPCR on day 7. d, Relative expression of Yap target genes (Ly6a, Clu, Il1rn, Msln and Cxcl16) in day 7 organoids and PGE2-driven spheroids developed from wild-type crypts. N = 3 3D cultures per condition. Two-tailed Welch’s t-test. e, Relative expression of Yap target genes in day 7 organoids and PGE2-driven spheroids developed from crypts isolated from Ptger4f/f and Ptger4ΔIEC mice. n = 3 3D cultures per genotype and condition. One-way ANOVA. f, Correlation between the expression levels of metagenes of a Yap transcriptional program and an early (non-tumour) ApcMin/+ tumorigenesis transcriptional program in single epithelial cells (n = 1,585) from the Ptger4-ON and Ptger4–OFF fibroblast–crypt co-cultures of Fig. 3. g, Small intestinal crypts were grown into organoids or spheroids with OGM or OGM that was supplemented daily with 0.1 μM dmPGE2. Western blot analysis for Yap1 and β-actin was performed in total lysates from untreated organoids, organoids treated with 0.1 μM dmPGE2 for 16 h and untreated spheroids. Data from one organoid and three independent spheroid cultures. h, Relative expression of the Yap1 gene and Yap target genes in wild-type organoid cultures treated with 0.1 μM dmPGE2 for 13 h, as determined by RT–qPCR. n = 3–5 cultures per condition. Statistical comparisons were performed with unpaired two-tailed t-test. For Ly6a, Welch’s correction was applied. i, Western blot analysis for Ser127 pYap and total Yap performed in total lysates from wild-type organoids stimulated with 0.1 μM dmPGE2 for the indicated time-points. Indicative of five independent experiments. j, Relative expression of Yap target genes in wild-type organoids treated with 1 μM verteporfin and 0.1 μM dmPGE2 for 13 h. n = 3–4 cultures per condition. Statistical comparisons were performed with unpaired two-tailed t-test, two-tailed Welch’s t-test or Mann–Whitney test on the basis of the criteria described in Methods. All data are mean ± s.e.m. *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001.
Extended Data Fig. 7 Genetic ablation of Yap prevents spheroid formation and Sca-1+ stem-cell expansion in fibroblast–crypt organotypic co-cultures.
a, Crypts isolated from the small intestines of Yap1f/f and Yap1ΔIEC mice were grown into organoids by 3D culture with OGM supplemented with 0.5 mg ml−1 recombinant mouse epiregulin (Ereg) as previously described18, or in a co-culture with wild-type primary mouse intestinal fibroblasts with OGM without Ereg supplementation. Indicative images and quantification of the percentages of crypts, organoids and spheroids grown per 3D structure are shown. n = 2 cultures per condition. Data are representative of two independent experiments. Scale bars, 100 μm. Two-way ANOVA. Data represent mean ± s.e.m. ****P < 0.0001. b, Intestinal crypts isolated from the small intestines of Yap1f/f and Yap1ΔIEC mice were co-cultured with wild-type primary mouse intestinal fibroblasts. On day 4, these co-cultures and control Yap1f/f organoid cultures were processed into single-cell suspensions and analysed by flow cytometry for Sca-1 expression in Cd24+ epithelial cells. n = 2 cultures per condition. Scale bars, 100 μm. FSC, forward scatter. Unpaired two-tailed t-test. Mean ± s.e.m. ***P < 0.001.
Extended Data Fig. 8 Ptger4 ablation does not affect epithelial lineage differentiation and stem-cell function.
a, Ptger4 gene expression in crypts isolated from the ileum of littermate Ptger4f/f and Ptger4ΔIEC mice (n = 3 mice per genotype) and in organoids grown from these crypts (n = 3 cultures per genotype) determined by RT–qPCR analysis. Two-tailed unpaired t-test. b, Single-cell RNA-seq (Drop-seq) was performed in crypt epithelial cells isolated from littermate Ptger4f/f and Ptger4ΔIEC mice. Data for 2,439 single epithelial cells are shown in a t-SNE plot. c, Biological replicates visualized on a t-SNE plot. Crypt epithelial cells were independently isolated from two groups of mice per genotype (biological replicates 1 and 2). From the first biological replicate, two independent Drop-seq samples were collected (A and B) for a total number of three samples per genotype. d, Clustering and cluster assignments of 2,439 single epithelial cells displayed on a t-SNE plot. e, Proportion of each epithelial cluster among total crypt epithelial cells in Ptger4f/f and Ptger4ΔIEC mice. f, Violin plots showing the entire range of expression levels for a metagene of the β-catenin transcriptional program in n = 2,439 single epithelial cells from Ptger4f/f and Ptger4ΔIEC mice. g, Analysis of all Ptger4-expressing single cells detected (n = 478). Re-clustering results of Ptger4-expressing single cells with cluster annotations are visualized on a t-SNE plot. The expression levels of key marker genes for these clusters are visualized on t-SNE plots. h, Lineage tracing of Ptger4 heterozygous (Ptger4-HET) and Ptger4-knockout (Ptger4-KO) Lgr5+ stem cells. The small intestines of Lgr5-creERT2-Rosa26tdTomato/+Ptger4f/+ (Ptger4-HET) and Lgr5-creERT2-Rosa26tdTomato/+Ptger4f/f (Ptger4-KO) mice were examined for direct tdTomato fluorescence 5 days after a single injection of 2 mg tamoxifen per mouse. The results shown are representative of independent observations from one experiment. Scale bars, 70 μm. i, Quantification of intestinal epithelial populations in the ileum of littermate Ptger4f/f (n = 5) and Ptger4ΔIEC (n = 5) mice. Immunostaining was performed for markers of Paneth cells (lysozyme), tuft cells (Dclk1), enteroendocrine cells (chromogranin A) and stem cells (Olfm4). Scale bars, 20 μm. Goblet cells were identified by PAS staining and enterocytes were detected by alkaline phosphatase enzymatic activity. Scale bars, 50 μm. Incorporation and immunohistochemical detection of BrdU was used to determine the numbers of cycling cells. Scale bars, 50 μm. Data for each mouse represent mean number of positive cells per crypt or crypt–villus unit as indicated. n = 217–565 crypts and/or villi were evaluated per staining. Statistical comparisons were performed with two-tailed unpaired t-test except for PAS+ cells, for which unpaired Welch's t-test was applied. Mean ± s.e.m.; ns, non-significant; **P < 0.01.
Extended Data Fig. 9 Nuclear localization of Yap and activation of Yap target genes in ApcMin/+ and azoxymethane-induced tumorigenesis.
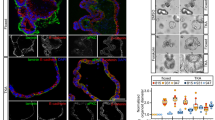
a, Immunostaining for Yap in the small intestine of five-month-old ApcMin/+ and wild-type littermate control mice. Nuclear localization of Yap is displayed on the basis of colocalization with DAPI. Normal (N) and tumour (T) areas of the ApcMin/+ intestine are indicated. Scale bars, 70 μm. Data are indicative of at least ten different tumour areas. b, Immunostaining for Sca-1 and the epithelial marker E-cadherin in normal and tumour areas of the small intestine of five-month-old ApcMin/+ mice. Indicative of two independent experiments. c, Immunostaining for Yap in the colon of wild-type mice subjected to 10 weekly intraperitoneal injections with 10 mg kg−1 azoxymethane as indicated and in untreated controls. Nuclear localization of Yap is displayed on the basis of colocalization with DAPI. Scale bars, 20 μm. Data indicative of three mice analysed. d, Relative expression of the Yap target gene Clu in normal and tumour areas of the colon of wild-type mice (n = 8) subjected to 10 weekly intraperitoneal injections with 10 mg kg−1 azoxymethane as shown in c. Two-tailed Mann–Whitney test. e, Relative expression of Yap1 and Yap target genes in the small intestine of 5-week-old Ptger4f/f (n = 3), ApcMin/+Ptger4f/f (n = 6) and ApcMin/+Ptger4ΔIEC (n = 8) mice. Statistical comparisons were performed with two-tailed t-test for Yap1 and Il1rn and with two-tailed Mann–Whitney test for Ly6a. f, Spleen weight of 5.5-month-old ApcMin/+Ptger4f/f (n = 8) and ApcMin/+Ptger4ΔIEC (n = 7) mice. Average spleen weight of (n = 2) normal littermates (Ptger4f/f) is displayed for comparison. Two-tailed t-test. g, Size of 72 adenomas from 5.5-month-old ApcMin/+Ptger4f/f (n = 6) and ApcMin/+Ptger4ΔIEC (n = 4) mice. The whiskers extend from minimum to maximum and the box extends from the 25th to 75th percentiles with the median indicated. Two-tailed Mann–Whitney test. Mean ± s.e.m.; ns, non-significant; *P < 0.05; **P < 0.01.
Extended Data Fig. 10 PGE2–PTGER4 controls stem-cell function in human colonic crypts and YAP displays a nuclear localization in human colorectal tumours.
a, Human colonic crypts were grown into organoids by 3D culture with OGM or OGM supplemented daily with 0.1 μM dmPGE2, with or without 10 μM ONO-AE3-208 (PTGER4–EP4 inhibitor). Images indicative of three independent experiments with crypts isolated from three patients are shown. Scale bar, 100 μm. b, Immunostaining for YAP in sections of human colorectal adenomas or adenocarcinomas and neighbouring normal tissue areas. Nuclear localization of Yap is displayed on the basis of colocalization with DAPI. Clearly defined normal (N) and tumour (T) areas are indicated wherever applicable. Images shown are representative of specimens obtained and analysed from n = 16 patients with the types of colorectal tumours indicated. Patient characteristics and the type of colorectal tumour per individual are described in the Supplementary Table 3. c, Schematic representation of the mechanism proposed in the present study. TISC, tumour-initiating stem cell.
Supplementary information
Supplementary Figure 1
This file contains the uncropped western blots.
Supplementary Table 1
Pathway analysis in fibroblast clusters. Top 10 KEGG pathways found to be enriched in each fibroblast population of the mouse colon mesenchyme by Gene Set Variation Analysis.
Supplementary Table 2
Quality control metrics of single cell RNA-seq experiments. Quality control metrics for the Drop-seq experiments performed 1) in mouse colonic mesenchymal cells, 2) in intestinal fibroblast/crypt co-cultures and 3) in primary crypt epithelial cells from Ptger4f/f and Ptger4ΔIEC mice.
Supplementary Table 3
Characteristics of human subjects. Characteristics of human subjects/patients from whom fresh normal tissues or Formalin-Fixed Paraffin Embedded tumor tissues were obtained.
Source data
Rights and permissions
About this article
Cite this article
Roulis, M., Kaklamanos, A., Schernthanner, M. et al. Paracrine orchestration of intestinal tumorigenesis by a mesenchymal niche. Nature 580, 524–529 (2020). https://doi.org/10.1038/s41586-020-2166-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-020-2166-3
This article is cited by
-
Beyond genetics: driving cancer with the tumour microenvironment behind the wheel
Nature Reviews Cancer (2024)
-
Towards Full Thickness Small Intestinal Models: Incorporation of Stromal Cells
Tissue Engineering and Regenerative Medicine (2024)
-
The cyclooxygenase-expressing mesenchyme resists intestinal epithelial injury by paracrine signaling
Cell Regeneration (2023)
-
Identification of a prognostic six-immune-gene signature and a nomogram model for uveal melanoma
BMC Ophthalmology (2023)
-
A stromal lineage maintains crypt structure and villus homeostasis in the intestinal stem cell niche
BMC Biology (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.