Abstract
Radial glial progenitor cells (RGPs) are the major neural progenitor cells that generate neurons and glia in the developing mammalian cerebral cortex1,2,3,4. In RGPs, the centrosome is positioned away from the nucleus at the apical surface of the ventricular zone of the cerebral cortex5,6,7,8. However, the molecular basis and precise function of this distinctive subcellular organization of the centrosome are largely unknown. Here we show in mice that anchoring of the centrosome to the apical membrane controls the mechanical properties of cortical RGPs, and consequently their mitotic behaviour and the size and formation of the cortex. The mother centriole in RGPs develops distal appendages that anchor it to the apical membrane. Selective removal of centrosomal protein 83 (CEP83) eliminates these distal appendages and disrupts the anchorage of the centrosome to the apical membrane, resulting in the disorganization of microtubules and stretching and stiffening of the apical membrane. The elimination of CEP83 also activates the mechanically sensitive yes-associated protein (YAP) and promotes the excessive proliferation of RGPs, together with a subsequent overproduction of intermediate progenitor cells, which leads to the formation of an enlarged cortex with abnormal folding. Simultaneous elimination of YAP suppresses the cortical enlargement and folding that is induced by the removal of CEP83. Together, these results indicate a previously unknown role of the centrosome in regulating the mechanical features of neural progenitor cells and the size and configuration of the mammalian cerebral cortex.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
The datasets generated during and/or analysed during the current study are available from the corresponding authors on reasonable request.
References
Florio, M. & Huttner, W. B. Neural progenitors, neurogenesis and the evolution of the neocortex. Development 141, 2182–2194 (2014).
Homem, C. C., Repic, M. & Knoblich, J. A. Proliferation control in neural stem and progenitor cells. Nat. Rev. Neurosci. 16, 647–659 (2015).
Kriegstein, A. & Alvarez-Buylla, A. The glial nature of embryonic and adult neural stem cells. Annu. Rev. Neurosci. 32, 149–184 (2009).
Rakic, P. Developmental and evolutionary adaptations of cortical radial glia. Cereb. Cortex 13, 541–549 (2003).
Wang, X. et al. Asymmetric centrosome inheritance maintains neural progenitors in the neocortex. Nature 461, 947–955 (2009).
Tsai, J. W., Lian, W. N., Kemal, S., Kriegstein, A. R. & Vallee, R. B. Kinesin 3 and cytoplasmic dynein mediate interkinetic nuclear migration in neural stem cells. Nat. Neurosci. 13, 1463–1471 (2010).
Bultje, R. S. et al. Mammalian Par3 regulates progenitor cell asymmetric division via notch signaling in the developing neocortex. Neuron 63, 189–202 (2009).
Chenn, A., Zhang, Y. A., Chang, B. T. & McConnell, S. K. Intrinsic polarity of mammalian neuroepithelial cells. Mol. Cell. Neurosci. 11, 183–193 (1998).
Conduit, P. T., Wainman, A. & Raff, J. W. Centrosome function and assembly in animal cells. Nat. Rev. Mol. Cell Biol. 16, 611–624 (2015).
Lüders, J. & Stearns, T. Microtubule-organizing centres: a re-evaluation. Nat. Rev. Mol. Cell Biol. 8, 161–167 (2007).
Bettencourt-Dias, M. & Glover, D. M. Centrosome biogenesis and function: centrosomics brings new understanding. Nat. Rev. Mol. Cell Biol. 8, 451–463 (2007).
Kobayashi, T. & Dynlacht, B. D. Regulating the transition from centriole to basal body. J. Cell Biol. 193, 435–444 (2011).
Hu, D. J.-K. et al. Dynein recruitment to nuclear pores activates apical nuclear migration and mitotic entry in brain progenitor cells. Cell 154, 1300–1313 (2013).
Higginbotham, H. et al. Arl13b-regulated cilia activities are essential for polarized radial glial scaffold formation. Nat. Neurosci. 16, 1000–1007 (2013).
Paridaen, J. T. M. L., Wilsch-Bräuninger, M. & Huttner, W. B. Asymmetric inheritance of centrosome-associated primary cilium membrane directs ciliogenesis after cell division. Cell 155, 333–344 (2013).
Higginbotham, H. R. & Gleeson, J. G. The centrosome in neuronal development. Trends Neurosci. 30, 276–283 (2007).
Louvi, A. & Grove, E. A. Cilia in the CNS: the quiet organelle claims center stage. Neuron 69, 1046–1060 (2011).
Insolera, R., Bazzi, H., Shao, W., Anderson, K. V. & Shi, S. H. Cortical neurogenesis in the absence of centrioles. Nat. Neurosci. 17, 1528–1535 (2014).
Englund, C. et al. Pax6, Tbr2, and Tbr1 are expressed sequentially by radial glia, intermediate progenitor cells, and postmitotic neurons in developing neocortex. J. Neurosci. 25, 247–251 (2005).
Götz, M., Stoykova, A. & Gruss, P. Pax6 controls radial glia differentiation in the cerebral cortex. Neuron 21, 1031–1044 (1998).
Tanos, B. E. et al. Centriole distal appendages promote membrane docking, leading to cilia initiation. Genes Dev. 27, 163–168 (2013).
Joo, K. et al. CCDC41 is required for ciliary vesicle docking to the mother centriole. Proc. Natl Acad. Sci. USA 110, 5987–5992 (2013).
Ran, F. A. et al. Double nicking by RNA-guided CRISPR Cas9 for enhanced genome editing specificity. Cell 154, 1380–1389 (2013).
Gorski, J. A. et al. Cortical excitatory neurons and glia, but not GABAergic neurons, are produced in the Emx1-expressing lineage. J. Neurosci. 22, 6309–6314 (2002).
Graser, S. et al. Cep164, a novel centriole appendage protein required for primary cilium formation. J. Cell Biol. 179, 321–330 (2007).
Greig, L. C., Woodworth, M. B., Galazo, M. J., Padmanabhan, H. & Macklis, J. D. Molecular logic of neocortical projection neuron specification, development and diversity. Nat. Rev. Neurosci. 14, 755–769 (2013).
Besse, L. et al. Primary cilia control telencephalic patterning and morphogenesis via Gli3 proteolytic processing. Development 138, 2079–2088 (2011).
Willaredt, M. A. et al. A crucial role for primary cilia in cortical morphogenesis. J. Neurosci. 28, 12887–12900 (2008).
Han, Y. G. et al. Hedgehog signaling and primary cilia are required for the formation of adult neural stem cells. Nat. Neurosci. 11, 277–284 (2008).
Snedeker, J. et al. Unique spatiotemporal requirements for intraflagellar transport genes during forebrain development. PLoS One 12, e0173258 (2017).
Haycraft, C. J. et al. Intraflagellar transport is essential for endochondral bone formation. Development 134, 307–316 (2007).
Wang, L., Hou, S. & Han, Y.-G. Hedgehog signaling promotes basal progenitor expansion and the growth and folding of the neocortex. Nat. Neurosci. 19, 888–896 (2016).
Jeong, J., Mao, J., Tenzen, T., Kottmann, A. H. & McMahon, A. P. Hedgehog signaling in the neural crest cells regulates the patterning and growth of facial primordia. Genes Dev. 18, 937–951 (2004).
Haubensak, W., Attardo, A., Denk, W. & Huttner, W. B. Neurons arise in the basal neuroepithelium of the early mammalian telencephalon: a major site of neurogenesis. Proc. Natl Acad. Sci. USA 101, 3196–3201 (2004).
Noctor, S. C., Martínez-Cerdeño, V., Ivic, L. & Kriegstein, A. R. Cortical neurons arise in symmetric and asymmetric division zones and migrate through specific phases. Nat. Neurosci. 7, 136–144 (2004).
Wang, X., Tsai, J.-W., LaMonica, B. & Kriegstein, A. R. A new subtype of progenitor cell in the mouse embryonic neocortex. Nat. Neurosci. 14, 555–561 (2011).
Hansen, D. V., Lui, J. H., Parker, P. R. L. & Kriegstein, A. R. Neurogenic radial glia in the outer subventricular zone of human neocortex. Nature 464, 554–561 (2010).
Shitamukai, A., Konno, D. & Matsuzaki, F. Oblique radial glial divisions in the developing mouse neocortex induce self-renewing progenitors outside the germinal zone that resemble primate outer subventricular zone progenitors. J. Neurosci. 31, 3683–3695 (2011).
Reillo, I., de Juan Romero, C., García-Cabezas, M. A. & Borrell, V. A role for intermediate radial glia in the tangential expansion of the mammalian cerebral cortex. Cereb. Cortex 21, 1674–1694 (2011).
Fietz, S. A. et al. OSVZ progenitors of human and ferret neocortex are epithelial-like and expand by integrin signaling. Nat. Neurosci. 13, 690–699 (2010).
Betizeau, M. et al. Precursor diversity and complexity of lineage relationships in the outer subventricular zone of the primate. Neuron 80, 442–457 (2013).
Aragona, M. et al. A mechanical checkpoint controls multicellular growth through YAP/TAZ regulation by actin-processing factors. Cell 154, 1047–1059 (2013).
Halder, G., Dupont, S. & Piccolo, S. Transduction of mechanical and cytoskeletal cues by YAP and TAZ. Nat. Rev. Mol. Cell Biol. 13, 591–600 (2012).
Kasioulis, I., Das, R. M. & Storey, K. G. Inter-dependent apical microtubule and actin dynamics orchestrate centrosome retention and neuronal delamination. eLife 6, e26215 (2017).
Mota, B. & Herculano-Houzel, S. Cortical folding scales universally with surface area and thickness, not number of neurons. Science 349, 74–77 (2015).
Nagasaka, A. et al. Differences in the mechanical properties of the developing cerebral cortical proliferative zone between mice and ferrets at both the tissue and single-cell levels. Front. Cell Dev. Biol. 4, 139 (2016).
Failler, M. et al. Mutations of CEP83 cause infantile nephronophthisis and intellectual disability. Am. J. Hum. Genet. 94, 905–914 (2014).
Reginensi, A. et al. Yap- and Cdc42-dependent nephrogenesis and morphogenesis during mouse kidney development. PLoS Genet. 9, e1003380 (2013).
Caspary, T., Larkins, C. E. & Anderson, K. V. The graded response to Sonic Hedgehog depends on cilia architecture. Dev. Cell 12, 767–778 (2007).
Liboska, R., Ligasová, A., Strunin, D., Rosenberg, I. & Koberna, K. Most anti-BrdU antibodies react with 2′-deoxy-5-ethynyluridine—the method for the effective suppression of this cross-reactivity. PLoS One 7, e51679 (2012).
Aigouy, B., Umetsu, D. & Eaton, S. Segmentation and quantitative analysis of epithelial tissues. Methods Mol. Biol. 1478, 227–239 (2016).
Legland, D., Arganda-Carreras, I. & Andrey, P. MorphoLibJ: integrated library and plugins for mathematical morphology with ImageJ. Bioinformatics 32, 3532–3534 (2016).
Mori, S. & Zhang, J. Principles of diffusion tensor imaging and its applications to basic neuroscience research. Neuron 51, 527–539 (2006).
Wu, D. et al. In vivo high-resolution diffusion tensor imaging of the mouse brain. Neuroimage 83, 18–26 (2013).
Chuang, N. et al. An MRI-based atlas and database of the developing mouse brain. Neuroimage 54, 80–89 (2011).
Ceritoglu, C. et al. Multi-contrast large deformation diffeomorphic metric mapping for diffusion tensor imaging. Neuroimage 47, 618–627 (2009).
Zhang, J. et al. Longitudinal characterization of brain atrophy of a Huntington’s disease mouse model by automated morphological analyses of magnetic resonance images. Neuroimage 49, 2340–2351 (2010).
Chen, E. J., Novakofski, J., Jenkins, W. K. & Brien, W. D. O. Young’s modulus measurements of soft tissues with application to elasticity imaging. IEEE Trans. Ultrason. Ferroelectr. Freq. Control 43, 191–194 (1996).
Acknowledgements
We thank B. Yoder, J. Wrana and A. McMahon for providing Ift88fl/fl, Yapfl/fl and R26-LSL-SmoM2 mice, respectively; M. Brendel and K. Manova-Todorova at the Molecular Cytology Core and Y. Wang and W. Mark at the Mouse Genetics Core Facility of Memorial Sloan Kettering Cancer Centre, K. Uryu at the Electron Microscopy Resource Centre and J. Gao and C. Yang at the Gene Targeting Resource Centre of Rockefeller University for their technical support; and members of the H.S. and S.-H.S. laboratories for their input and comments on the manuscript. This work was supported by grants from the NIH (R01DA024681 and R01NS085004 to S.-H.S., and P30CA008748 to Memorial Sloan Kettering Cancer Centre), the Howard Hughes Medical Institute (to S.-H.S.) and the Beijing Outstanding Young Scientist Program (BJJWZYJH01201910003012 to S.-H.S.).
Author information
Authors and Affiliations
Contributions
W.S. and S.-H.S. conceived and H.S. and S.-H.S. supervised the project; W.S. and J.Y. performed most of the experiments with help from Z.Y., M.H. and X.-Y.Y.; C.H.L. and J.Z. performed the MRI experiment; A.L.J., K.V.A. and M.-F.B.T. advised on the project; W.S., J.Y., H.S. and S.-H.S. wrote the manuscript with input from all authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature thanks Gordon Fishell, Anthony Wynshaw-Boris and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Deletion of Cep83 in cortical RGPs.
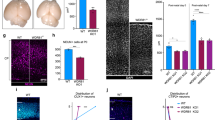
a, Schematic diagram of the generation of the Cep83cKO mouse using a CRISPR–Cas9-mediated double-nicking strategy. The DNA sequence at the top depicts the sites targeted by a pair of guide RNAs (outlined in blue) downstream of the critical exon 3 in the Cep83 gene. Green boxes represent exons, red triangles represent loxP sites and yellow triangles represent FRT sites. NeoR, neomycin-resistance gene cassette. b, Representative Southern blot showing the correct gene targeting against the 5′-homology arm of the Cep83 floxed allele with the presence of the deletion-specific 3.5-kb band (n = 3). c, Representative images of wild-type and Cep83cKO cortices at E12.5, E13.5 and E15.5, stained for PCNT (green) and ARL13B (a primary cilium marker; red), and with DAPI (blue) (n = 3). High-magnification images of individual centrosomes are shown in the insets. Note the loss of primary cilia in the Cep83cKO cortex by E13.5. Scale bars, 10 μm (main image); 1 μm (inset). d, Representative images of P1 wild-type (n = 11) and Cep83cKO (n = 12) cortices stained for CTIP2 (green) and CUX1 (red), and with DAPI (blue). The asterisk indicates the folding in the medial region of the Cep83cKO cortex. High-magnification images of the folding are shown to the right. Scale bars, 500 μm (left); 200 μm (right). e, Representative images of the medial region of P1 wild-type and Cep83cKO cortices stained for brain lipid-binding protein (BLBP; green) and with DAPI (blue). High-magnification images are shown to the right. Scale bars, 500 μm (left); 100 μm (right).
Extended Data Fig. 2 Deletion of Cep83 in RGPs leads to increased neurogenesis and gliogenesis.
a, Representative images of the medial regions of P21 wild-type and Cep83cKO cortices stained for FOXP2 (green) and SATB2 (red), and with DAPI (blue). Scale bar, 100 μm. b, Quantification of the number of SATB2+ (left) and FOXP2+ (right) neurons per 250-μm column (wild type, n = 8 brains; Cep83cKO, n = 8 brains). c, Representative images of the dorsal regions of P21 wild-type and Cep83cKO cortices stained for FOXP2 (green) and SATB2 (red), and with DAPI (blue). Scale bar, 100 μm. d, Quantification of the number of SATB2+ (left) and FOXP2+ (right) neurons per 250-μm column (wild type, n = 6 brains; Cep83cKO, n = 5 brains). e, Representative images of P21 wild-type and Cep83cKO cortices stained for OLIG2 (an oligodendrocyte marker; green) and S100 (an astrocyte marker; red), and with DAPI (blue). Scale bar, 100 μm. f, Quantification of the number of OLIG2+ oligodendrocytes (n = 10 regions from 5 brains for each genotype) and S100+ astrocytes (n = 6 regions from 3 brains for each genotype) per 650-μm column. A two-sided Mann–Whitney U test was used for statistical analysis. Bar charts show mean + s.e.m. Box plots as in Fig. 1.
Extended Data Fig. 3 Deletion of Ift88 in RGPs does not lead to any obvious defect in centrosome appendages, centrosome membrane anchorage or cortical development.
a, Representative images of E13.5 wild-type and Ift88cKO ventricular zone surface stained for PCNT (green) and ARL13B (red), and with DAPI (blue) (n = 3). High-magnification images of individual centrosomes are shown in the insets. Note the complete loss of primary cilia in the Ift88cKO cortex by E13.5. Scale bars, 10 μm (main image); 1 μm (inset). b, Representative ssTEM images of E15.5 wild-type (top) and Ift88cKO (bottom) ventricular zone surface showing individual centrosomes of RGPs in the apical endfeet. High-magnification images (white dashed boxes) are shown on the right. Note that the Ift88cKO mother centriole possesses the DAPs that are anchored at the apical membrane (red arrows) and the sDAPs (yellow arrows), but does not support any microtubule-based ciliary axoneme (wild type, n = 9 centrosomes; Ift88cKO, n = 20 centrosomes). All wild-type mother centrioles were anchored to the apical membrane with microtubule-based cilia. All Ift88cKO mother centrioles were anchored to the apical membrane, but none possessed microtubule-based cilia. Scale bars, 800 nm (left); 200 nm (right). c, Representative images of E15.5 wild-type and Ift88cKO ventricular zone surface stained for PCNT (green) and CEP164 (red), and with DAPI (blue) (n = 3). High-magnification images of individual centrosomes are shown in the insets. Note the normal presence of CEP164 at the centrosome in the Ift88cKO cortex. Scale bars, 10 μm (main images); 0.5 μm (insets). d, Representative images of E15.5 wild-type (n = 6) and Ift88cKO (n = 6) ventricular zone surface stained for PCNT (green) and ODF2 (an sDAP marker; red), and with DAPI (blue). High-magnification images of individual centrosomes are shown in the insets. Note the normal presence of ODF2 at the centrosome in the Ift88cKO cortex. Scale bars, 5 μm (main image); 1 μm (inset). e, Representative whole-mount images of P21 wild-type and Ift88cKO brains. Scale bar, 0.5 cm. f, Quantification of the projected cortical area (n = 6 brains for each genotype). g, Images of P21 wild-type and Ift88cKO brain sections stained for CTIP2 (green) and CUX1 (red), and with DAPI (blue). Scale bar, 0.5 mm. h, Quantification of the cortical area (wild type, n = 4 brains; Ift88cKO, n = 4 brains). i, Images of the dorsal regions of P21 wild-type and Ift88cKO cortices stained for CTIP2 (green) and CUX1 (red), and with DAPI (blue). Scale bar, 100 μm. j, Quantification of the number of CUX1+ (left) and CTIP2+ (right) neurons per 250-μm column (n = 8 brains for each genotype). k, Representative images of P21 wild-type and Ift88cKO brain sections along the rostrocaudal axis, stained with DAPI (grey) (n = 5). Note that there is no obvious hydrocephalus in the Ift88cKO brain. Scale bar, 1 mm. A two-sided Mann–Whitney U test was used for statistical analysis. Bar charts show mean + s.e.m. Box plots as in Fig. 1.
Extended Data Fig. 4 Expression of SmoM2 in RGPs leads to cortical dysplasia.
a, Representative whole-mount images of P1 wild-type and SmoM2 brains (n = 5). Arrowheads indicate the agenesis of the olfactory bulb in the SmoM2 brain. Scale bar, 0.5 cm. b, Representative images of P1 wild-type and SmoM2 brain sections stained for CTIP2 (green) and CUX1 (red), and with DAPI (blue) (n = 5). Arrowheads indicate the absence of corpus callosum in the SmoM2 brain. The asterisk indicates the agenesis of the hippocampus in the SmoM2 brain. Scale bar, 0.5 mm. c, Representative images of P1 wild-type and SmoM2 cortices stained for CTIP2 (green) and CUX1 (red), and with DAPI (blue) (n = 4). Note the drastic disorganization of the SmoM2 cortex. Scale bar, 100 μm. d, Representative images of E15.5 wild-type and SmoM2 ventricular zone surface stained for PCNT (green) and ARL13B (red), and with DAPI (blue) (n = 3). High-magnification images of individual centrosomes are shown in the insets. Note the presence of the primary cilium at the SmoM2 centrosome. Scale bars, 5 μm (main image); 1 μm (inset). e, Representative images of E15.5 wild-type and SmoM2 ventricular zone surface stained for PCNT (green) and CEP89 (a DAP marker; red), and with DAPI (blue) (n = 3). High-magnification images of individual centrosomes are shown in the insets. Note the normal presence of CEP89 at the SmoM2 centrosome. Scale bars, 5 μm (main image); 1 μm (inset). f, Representative images of E15.5 wild-type and SmoM2 ventricular zone surface (n = 5 brains) stained for PCNT (green) and ODF2 (red), and with DAPI (blue) (n = 3 brains). High-magnification images of individual centrosomes are shown in the insets. Note the normal presence of ODF2 at the SmoM2 centrosome. Scale bars, 5 μm (main image); 1 μm (inset).
Extended Data Fig. 5 Deletion of Cep83 does not affect the densities of RGPs and intermediate progenitor cells at E13.5, but leads to increased densities of these cells in the dorsomedial cortex at E15.5.
a, Representative images of E13.5 wild-type and Cep83cKO cortices stained for PAX6 (green) and TBR2 (red), and with DAPI (blue). Scale bar, 50 μm. b, c, Quantification of the number of PAX6+ (b) and TBR2+ (c) cells per 250-μm column in a. Wild type, n = 8 brains; Cep83cKO, n = 8 brains. d, Images of E12.5 wild-type and Cep83cKO cortices (dorsomedial region) that were subjected to EdU (red) and BrdU (green) sequential pulse-chase labelling. Cortices were stained for PAX6 (grey), and with DAPI (blue). Example regions (white dashed boxes) are shown at the bottom. Scale bars, 25 μm. e, f, Quantification of the percentage of EdU+BrdU+ cells among the total EdU+ cells in the ventricular zone (e), and the number of BrdU+ cells in the ventricular zone per 250-μm column (f) (wild type, n = 6 brains; Cep83cKO, n = 6 brains). g, Images of E15.5 wild-type and Cep83cKO cortices (dorsomedial region) stained for PAX6 (green) and TRB2 (red), and with DAPI (blue). Scale bar, 50 μm. h, i, Quantification of the number of PAX6+ (h) or TBR2+ (i) cells per 250-μm column (wild type, n = 8 brains; Cep83cKO, n = 6 brains). j, Quantification of the distribution width of TBR2+ cells in wild-type or Cep83cKO cortices (E13.5, n = 4 brains for each genotype; E15.5, n = 5 brains for each genotype). A two-sided Mann–Whitney U test was used for statistical analysis. Box plots as in Fig. 1.
Extended Data Fig. 6 Increased mitotic cells in the subventricular zone of the Cep83cKO cortex are predominantly intermediate progenitor cells.
a, Images of E15.5 wild-type and Cep83cKO cortices stained for P-HH3 (a mitotic cell marker; red), and with DAPI (blue). Scale bar, 50 μm. b, d, Images of E15.5 wild-type and Cep83cKO cortices stained for P-HH3 (red) and TBR2 (green) (b) or SOX2 (an RGP marker; green) (d), and with DAPI (blue). High-magnification images of individual P-HH3+ cells are shown at the bottom. Note that P-HH3+ cells in the subventricular zone of the Cep83cKO cortex are predominantly TBR2+ but SOX2−. Scale bars, 25 μm (top); 10 μm (bottom). c, Images of E15.5 wild-type and Cep83cKO cortices stained for P-HH3 (red) and phospho-vimentin (P-VIM; green), and with DAPI (blue). High-magnification images of individual P-HH3+ cells are shown at the bottom. Scale bars, 25 μm (top); 10 μm (bottom). e, f, Quantification of the number of apical (e) and basal (f) P-HH3+ cells per 250-μm column (wild type, n = 16 brains; Cep83cKO, n = 14 brains). g, h, Quantification of the percentage of P-HH3+ cells in the subventricular zone that are TBR2+ (g; lateral, n = 4 brains for each genotype; medial, n = 3 brains for each genotype) or SOX2− (h; n = 4 brains for each genotype). i, Quantification of the percentage of P-HH3+ cells without a P-VIM labelled basal radial glial fibre (lateral, n = 4 brains for each genotype; medial, n = 3 brains for each genotype). j–l, Representative images of E15.5 wild-type and Cep83cKO cortices stained for PAX6 (green) and three previously suggested markers of outer subventricular zone RGPs (HOPX (j), PTPRZ1 (k) or TNC (l)) (red), and with DAPI (blue) (n = 4). Note that there is no obvious increase in the expression of HOPX, PTPRZ1 or TNC in the Cep83cKO cortex and low expression in both wild-type and Cep83cKO cortices. Scale bars, 50 μm. A two-sided Mann–Whitney U test was used for statistical analysis. Box plots as in Fig. 1.
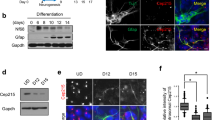
Extended Data Fig. 7 Disruption of other components of the DAP assembly pathway leads to the overproduction of RGPs and intermediate progenitor cells.
a, Diagram of the hierarchical DAP assembly pathway. b–d, Western blot assays to show the efficacy of shRNAs against Cep83 (b), Cep89 (c) or Sclt1 (d) in suppressing protein expression (n = 3). e, Representative images of E16.5 cortices that received in utero electroporation of EGFP (green) together with shRNAs against Cep83 or against Cep89 and Sclt1 at E13.5. Cortices were stained for PAX6 (red, top) and TBR2 (red, bottom), and with DAPI (blue). Note that expression of Cep83 shRNA-a (which effectively suppresses protein expression), but not that of Cep83 shRNA-c (which does not suppress protein expression), leads to a significant increase in both PAX6+ RGPs in the ventricular zone and TBR2+ intermediate progenitor cells in the subventricular zone. Moreover, expression of Cep89 shRNA-c and Sclt1 shRNA-c (which effectively suppress protein expression), but not that of Cep89 shRNA-b and Sclt1 shRNA-b (neither of which suppresses protein expression), results in a similar increase in both PAX6+ RGPs and TBR2+ intermediate progenitor cells. Scale bars, 100 μm. f, g, Quantification of the percentage of EGFP+ cells that are PAX6+ (f) or TBR2+ (g). Note that similar to the Cep83cKO, expression of Cep83 shRNA-a and Cep89 and Sclt1 shRNA-c—but not that of Cep83 shRNA-c or Cep89 and Sclt1 shRNA-b—leads to a significant increase in PAX6+ RGPs and TBR2+ intermediate progenitor cells, indicating that disruption of other DAP components causes excessive production of RGPs and intermediate progenitor cells in a similar manner to the removal of CEP83 (control, n = 4 brains (f) and 5 brains (g); Cep83 shRNA-a, n = 5 brains; Cep89 and Sclt1 shRNA-c, n = 4 brains; Cep83 shRNA-c, n = 6 brains; Cep89 and Sclt1 shRNA-b, n = 4 brains). h, Quantification of the percentage of EGFP+ cells in different cortical regions (control, n = 5 brains; Cep83 shRNA-a, n = 5 brains; Cep89 and Sclt1 shRNA-c, n = 4 brains; Cep83 shRNA-c, n = 7 brains; Cep89 and Sclt1 shRNA-b, n = 5 brains. A two-sided Mann–Whitney U test was used for statistical analysis. Bar charts show mean + s.e.m. Box plots as in Fig. 1.
Extended Data Fig. 8 Detachment of the centrosome from the apical membrane does not disrupt RGP polarity, junction formation or radial glial fibre scaffolding.
a, Representative images of E15.5 wild-type and Cep83cKO cortices in coronal sections stained for α-tubulin (α-TUB; green), and with DAPI (blue) (n = 3). Note that there is no obvious difference in non-apical membrane microtubules between the wild-type and Cep83cKO cortices. Scale bar, 10 μm. b, Representative en face images of mitotic RGPs in E15.5 wild-type and Cep83cKO cortices stained for α-tubulin (green), and with DAPI (blue) (n = 12). Note that there is no obvious difference in microtubule spindles between the wild-type and Cep83cKO RGPs. Scale bar, 2 μm. c, Representative images of E15.5 wild-type and Cep83cKO cortices stained for partitioning defective protein 3 (PARD3; red), an evolutionarily conserved polarity protein, and with DAPI (blue) (n = 3). Scale bar, 10 μm. d, Representative images of E15.5 wild-type and Cep83cKO cortices stained for three junction markers (β-catenin (β-CAT; left), N-cadherin (N-CAD; middle) or ZO-1 (right)) (red), and with DAPI (blue). Scale bar, 50 μm. e–g, Quantification of the staining intensity of β-catenin (e; wild type, n = 8 brains; Cep83cKO, n = 5 brains), N-cadherin (f; wild type, n = 4 brains; Cep83cKO, n = 3 brains) or ZO-1 (g; wild type, n = 8 brains; Cep83cKO, n = 7 brains) at the ventricular zone surface. h, i, Representative images of E15.5 wild-type and Cep83cKO cortices stained for nestin (h) or BLBP (i) (green), and with DAPI (blue) (n = 5). Scale bars, 50 μm. A two-sided Mann–Whitney U test was used for statistical analysis. Box plots as in Fig. 1.
Extended Data Fig. 9 Centrosome detachment leads to enlargement of the apical membrane, and nuclear expression of YAP is low in TBR2+ intermediate progenitor cells and dissociated RGPs in culture.
a, Representative en face segmented images of wild-type and Cep83cKO ventricular zone surface at E13.5 and E15.5. Each apical domain is colour-coded on the basis of its size: blue colour indicates an apical domain that is relatively larger. Scale bar, 10 μm. b, Quantification of the size of the apical domain of wild-type and Cep83cKO RGPs at E13.5 and E15.5 (E13.5: wild type, n = 5,038 apical domains from 8 embryos; Cep83cKO, n = 2,891 apical domains from 6 embryos; E15.5: wild type, n = 4,780 apical domains from 12 embryos; Cep83cKO, n = 1,959 apical domains from 8 embryos). c, Quantification of the size of the apical domain of interphase and mitotic wild-type and Cep83cKO RGPs at E15.5 (wild type, n = 1,703 interphase apical domains and n = 145 mitotic apical domains from 4 embryos; Cep83cKO, n = 988 interphase apical domains and n = 83 mitotic apical domains from 4 embryos). d, Representative images of the subventricular zone of E15.5 wild-type and Cep83cKO cortices stained for YAP (green) and TBR2 (red), and with DAPI (blue) (n = 5). Individual TBR2+ intermediate progenitor cells are shown in the insets. Note the low expression of YAP in the nuclei of TBR2+ intermediate progenitor cells in the subventricular zone of the wild-type and Cep83cKO cortices. Scale bars, 50 μm (main image); 5 μm (inset). e, Representative images of acutely dissociated cell cultures of E15.5 wild-type and Cep83cKO cortical ventricular zone stained for SOX2 (red) and YAP (green), and with DAPI (blue). Scale bar, 20 μm. f, Quantification of the YAP staining intensity in SOX2+ RGPs (wild type, n = 13 brains; Cep83cKO, n = 8 brains). A two-sided Mann–Whitney U test was used for statistical analysis. Box plots as in Fig. 1.
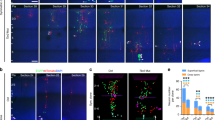
Extended Data Fig. 10 Excessive neurogenesis in the Cep83cKO cortex depends on excessive expression and activation of YAP.
a, Representative high-magnification images of P21 wild-type, YapcKO, Cep83cKO and Cep83YapcDKO cortices stained for CTIP2 (green) and CUX1 (red), and with DAPI (blue). Scale bar, 100 μm. b, Quantification of the number of CUX1+ neurons per 250-μm column (n = 8 brains for each genotype). c, Representative en face images of coronal sections of E15.5 wild-type, YapcKO, Cep83cKO and Cep83YapcDKO cortices stained for α-tubulin (green) and actin (red), and with DAPI (blue). Scale bar, 5 μm. d–f, Quantification of the intensity of microtubules per apical domain (d), individual apical domain size (e) and the intensity of PCNT per apical domain (f) (wild type, n = 1,730 apical domains from 4 embryos; YapcKO, n = 1,074 apical domains from 3 embryos, Cep83cKO, n = 456 apical domains from 3 embryos; Cep83YapcDKO, n = 540 apical domains from 3 embryos). A two-sided Mann–Whitney U test was used for statistical analysis. Bar charts show mean + s.e.m. Box plots as in Fig. 1.
Supplementary information
41586_2020_2139_MOESM1_ESM.pdf
Supplementary Figure Supplementary Figure 1: Uncropped original gel electrophoresis data. The extended Data Figures that include each image are shown above each gel, with a black outline to show the excerpted protion.
Rights and permissions
About this article
Cite this article
Shao, W., Yang, J., He, M. et al. Centrosome anchoring regulates progenitor properties and cortical formation. Nature 580, 106–112 (2020). https://doi.org/10.1038/s41586-020-2139-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-020-2139-6
This article is cited by
-
Upregulation of TUBG1 expression promotes hepatocellular carcinoma development
Medical Oncology (2023)
-
Inhibition of Foxp4 Disrupts Cadherin-based Adhesion of Radial Glial Cells, Leading to Abnormal Differentiation and Migration of Cortical Neurons in Mice
Neuroscience Bulletin (2023)
-
LPA signaling acts as a cell-extrinsic mechanism to initiate cilia disassembly and promote neurogenesis
Nature Communications (2021)
-
Centrosome instability: when good centrosomes go bad
Cellular and Molecular Life Sciences (2021)
-
Molecular causes of primary microcephaly and related diseases: a report from the UNIA Workshop
Chromosoma (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.