Abstract
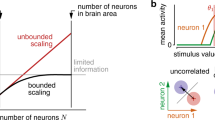
How the brain processes information accurately despite stochastic neural activity is a longstanding question1. For instance, perception is fundamentally limited by the information that the brain can extract from the noisy dynamics of sensory neurons. Seminal experiments2,3 suggest that correlated noise in sensory cortical neural ensembles is what limits their coding accuracy4,5,6, although how correlated noise affects neural codes remains debated7,8,9,10,11. Recent theoretical work proposes that how a neural ensemble’s sensory tuning properties relate statistically to its correlated noise patterns is a greater determinant of coding accuracy than is absolute noise strength12,13,14. However, without simultaneous recordings from thousands of cortical neurons with shared sensory inputs, it is unknown whether correlated noise limits coding fidelity. Here we present a 16-beam, two-photon microscope to monitor activity across the mouse primary visual cortex, along with analyses to quantify the information conveyed by large neural ensembles. We found that, in the visual cortex, correlated noise constrained signalling for ensembles with 800–1,300 neurons. Several noise components of the ensemble dynamics grew proportionally to the ensemble size and the encoded visual signals, revealing the predicted information-limiting correlations12,13,14. Notably, visual signals were perpendicular to the largest noise mode, which therefore did not limit coding fidelity. The information-limiting noise modes were approximately ten times smaller and concordant with mouse visual acuity15. Therefore, cortical design principles appear to enhance coding accuracy by restricting around 90% of noise fluctuations to modes that do not limit signalling fidelity, whereas much weaker correlated noise modes inherently bound sensory discrimination.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
The data that support the findings of this study are available from the corresponding authors upon reasonable request.
Code availability
We used open source software routines for image registration44 (http://bigwww.epfl.ch/thevenaz/turboreg/) and partial least squares analysis (https://www.mathworks.com/matlabcentral/fileexchange/18760-partial-least-squares-and-discriminant-analysis). Software code for extracting individual neurons and their Ca2+ activity traces from Ca2+ videos using principal component and then independent component analyses35,48 is freely available (https://www.mathworks.com/matlabcentral/fileexchange/25405-emukamel-cellsort), although for convenience we used a commercial version of these routines (Mosaic software, version 0.99.17; Inscopix). We wrote all other analysis routines in MATLAB (Mathworks; version 2017b). The primary software code used to support the findings of the study is available at Zenodo.org (https://zenodo.org/record/3593520#.XgWPu-hKg2w).
References
von Neumann, J. The Computer and the Brain 2nd edn (Yale Univ. Press, 1958).
Britten, K. H., Shadlen, M. N., Newsome, W. T. & Movshon, J. A. The analysis of visual motion: a comparison of neuronal and psychophysical performance. J. Neurosci. 12, 4745–4765 (1992).
Newsome, W. T., Britten, K. H. & Movshon, J. A. Neuronal correlates of a perceptual decision. Nature 341, 52–54 (1989).
Zohary, E., Shadlen, M. N. & Newsome, W. T. Correlated neuronal discharge rate and its implications for psychophysical performance. Nature 370, 140–143 (1994).
Averbeck, B. B., Latham, P. E. & Pouget, A. Neural correlations, population coding and computation. Nat. Rev. Neurosci. 7, 358–366 (2006).
Cohen, M. R. & Kohn, A. Measuring and interpreting neuronal correlations. Nat. Neurosci. 14, 811–819 (2011).
Sompolinsky, H., Yoon, H., Kang, K. & Shamir, M. Population coding in neuronal systems with correlated noise. Phys. Rev. E 64, 051904 (2001).
Abbott, L. F. & Dayan, P. The effect of correlated variability on the accuracy of a population code. Neural Comput. 11, 91–101 (1999).
Shamir, M. & Sompolinsky, H. Implications of neuronal diversity on population coding. Neural Comput. 18, 1951–1986 (2006).
Ecker, A. S., Berens, P., Tolias, A. S. & Bethge, M. The effect of noise correlations in populations of diversely tuned neurons. J. Neurosci. 31, 14272–14283 (2011).
Oram, M. W., Földiák, P., Perrett, D. I. & Sengpiel, F. The ‘Ideal Homunculus’: decoding neural population signals. Trends Neurosci. 21, 259–265 (1998).
Kanitscheider, I., Coen-Cagli, R. & Pouget, A. Origin of information-limiting noise correlations. Proc. Natl Acad. Sci. USA 112, E6973–E6982 (2015).
Moreno-Bote, R. et al. Information-limiting correlations. Nat. Neurosci. 17, 1410–1417 (2014).
Pitkow, X., Liu, S., Angelaki, D. E., DeAngelis, G. C. & Pouget, A. How can single sensory neurons predict behavior? Neuron 87, 411–423 (2015).
Prusky, G. T., West, P. W. & Douglas, R. M. Behavioral assessment of visual acuity in mice and rats. Vision Res. 40, 2201–2209 (2000).
Baylor, D. A., Lamb, T. D. & Yau, K. W. Responses of retinal rods to single photons. J. Physiol. (Lond.) 288, 613–634 (1979).
Barlow, H. B. Retinal noise and absolute threshold. J. Opt. Soc. Am. 46, 634–639 (1956).
Siebert, W. M. Some implications of the stochastic behavior of primary auditory neurons. Kybernetik 2, 206–215 (1965).
Yatsenko, D. et al. Improved estimation and interpretation of correlations in neural circuits. PLoS Comput. Biol. 11, e1004083 (2015).
Kanitscheider, I., Coen-Cagli, R., Kohn, A. & Pouget, A. Measuring Fisher information accurately in correlated neural populations. PLoS Comput. Biol. 11, e1004218 (2015).
Ecker, A. S. et al. Decorrelated neuronal firing in cortical microcircuits. Science 327, 584–587 (2010).
Reich, D. S., Mechler, F. & Victor, J. D. Independent and redundant information in nearby cortical neurons. Science 294, 2566–2568 (2001).
Renart, A. et al. The asynchronous state in cortical circuits. Science 327, 587–590 (2010).
Stirman, J. N., Smith, I. T., Kudenov, M. W. & Smith, S. L. Wide field-of-view, multi-region, two-photon imaging of neuronal activity in the mammalian brain. Nat. Biotechnol. 34, 857–862 (2016).
Chen, J. L., Voigt, F. F., Javadzadeh, M., Krueppel, R. & Helmchen, F. Long-range population dynamics of anatomically defined neocortical networks. eLife 5, e14679 (2016).
Sofroniew, N. J., Flickinger, D., King, J. & Svoboda, K. A large field of view two-photon mesoscope with subcellular resolution for in vivo imaging. eLife 5, e14472 (2016).
Tsai, P. S. et al. Ultra-large field-of-view two-photon microscopy. Opt. Express 23, 13833–13847 (2015).
Niell, C. M. & Stryker, M. P. Modulation of visual responses by behavioral state in mouse visual cortex. Neuron 65, 472–479 (2010).
Bonin, V., Histed, M. H., Yurgenson, S. & Reid, R. C. Local diversity and fine-scale organization of receptive fields in mouse visual cortex. J. Neurosci. 31, 18506–18521 (2011).
Averbeck, B. B. & Lee, D. Effects of noise correlations on information encoding and decoding. J. Neurophysiol. 95, 3633–3644 (2006).
Cover, T. M. & Thomas, J. A. Elements of Information Theory 2nd edn, (John Wiley & Sons, 2006).
Stringer, C., Michaelos, M. & Pachitariu, M. High precision coding mouse visual cortex. Preprint at https://www.biorxiv.org/content/10.1101/679324v1 (2019).
Prusky, G. T. & Douglas, R. M. Characterization of mouse cortical spatial vision. Vision Res. 44, 3411–3418 (2004).
Glickfeld, L. L., Histed, M. H. & Maunsell, J. H. Mouse primary visual cortex is used to detect both orientation and contrast changes. J. Neurosci. 33, 19416–19422 (2013).
Lecoq, J. et al. Visualizing mammalian brain area interactions by dual-axis two-photon calcium imaging. Nat. Neurosci. 17, 1825–1829 (2014).
Pologruto, T. A., Sabatini, B. L. & Svoboda, K. ScanImage: flexible software for operating laser scanning microscopes. Biomed. Eng. Online 2, 13 (2003).
Madisen, L. et al. Transgenic mice for intersectional targeting of neural sensors and effectors with high specificity and performance. Neuron 85, 942–958 (2015).
Wekselblatt, J. B., Flister, E. D., Piscopo, D. M. & Niell, C. M. Large-scale imaging of cortical dynamics during sensory perception and behavior. J. Neurophysiol. 115, 2852–2866 (2016).
Chettih, S. N. & Harvey, C. D. Single-neuron perturbations reveal feature-specific competition in V1. Nature 567, 334–340 (2019).
Harvey, C. D., Coen, P. & Tank, D. W. Choice-specific sequences in parietal cortex during a virtual-navigation decision task. Nature 484, 62–68 (2012).
Chen, T. W. et al. Ultrasensitive fluorescent proteins for imaging neuronal activity. Nature 499, 295–300 (2013).
Huber, D. et al. Multiple dynamic representations in the motor cortex during sensorimotor learning. Nature 484, 473–478 (2012).
Kim, K. H. et al. Multifocal multiphoton microscopy based on multianode photomultiplier tubes. Opt. Express 15, 11658–11678 (2007).
Thévenaz, P., Ruttimann, U. E. & Unser, M. A pyramid approach to subpixel registration based on intensity. IEEE Trans. Image Process. 7, 27–41 (1998).
Preibisch, S., Saalfeld, S. & Tomancak, P. Globally optimal stitching of tiled 3D microscopic image acquisitions. Bioinformatics 25, 1463–1465 (2009).
Brown, M. & Lowe, D. G. Automatic panoramic image stitching using invariant features. Int. J. Comput. Vis. 74, 59–73 (2007).
Pnevmatikakis, E. A. & Giovannucci, A. NoRMCorre: An online algorithm for piecewise rigid motion correction of calcium imaging data. J. Neurosci. Methods 291, 83–94 (2017).
Mukamel, E. A., Nimmerjahn, A. & Schnitzer, M. J. Automated analysis of cellular signals from large-scale calcium imaging data. Neuron 63, 747–760 (2009).
Vogelstein, J. T. et al. Fast nonnegative deconvolution for spike train inference from population calcium imaging. J. Neurophysiol. 104, 3691–3704 (2010).
Bishop, C. M. Pattern Recognition and Machine Learning Vol. 1 (Springer, 2007).
Geladi, P. & Kowalski, B. R. Partial least-squares regression: a tutorial. Anal. Chim. Acta 185, 1–17 (1986).
Hastie, T., Tibshirani, R. & Friedman, J. The Elements of Statistcal Learning (Springer, 2009).
Podgorski, K. & Ranganathan, G. Brain heating induced by near-infrared lasers during multiphoton microscopy. J. Neurophysiol. 116, 1012–1023 (2016).
Graner, M. W., Cumming, R. I. & Bigner, D. D. The heat shock response and chaperones/heat shock proteins in brain tumors: surface expression, release, and possible immune consequences. J. Neurosci. 27, 11214–11227 (2007).
Kalmbach, A. S. & Waters, J. Brain surface temperature under a craniotomy. J. Neurophysiol. 108, 3138–3146 (2012).
Wang, H. et al. Brain temperature and its fundamental properties: a review for clinical neuroscientists. Front. Neurosci. 8, 307 (2014).
Talan, M. Body temperature of C57BL/6J mice with age. Exp. Gerontol. 19, 25–29 (1984).
Greenberg, D. S., Houweling, A. R. & Kerr, J. N. D. Population imaging of ongoing neuronal activity in the visual cortex of awake rats. Nat. Neurosci. 11, 749–751 (2008).
Karimipanah, Y., Ma, Z., Miller, J. K., Yuste, R. & Wessel, R. Neocortical activity is stimulus- and scale-invariant. PLoS ONE 12, e0177396 (2017).
Acknowledgements
We acknowledge a Stanford Graduate Fellowship (O.I.R.), research support from the Howard Hughes Medical Institute (M.J.S.), the Stanford CNC Program (M.J.S.), DARPA (M.J.S.), an NSF CAREER Award (S.G.), and the Burroughs-Wellcome (S.G.), McKnight (S.G.), James S. McDonnell (S.G.) and Simons (S.G.) foundations. NIH grants MH085500 and DA028298 to H.Z. funded development of the GCaMP6f-tTA-dCre and Rasgrf2-2A-dCre mice. NIH grant R24NS098519 (M.J.S.) supports our effort to make the 16-beam two-photon microscope an open resource available to other laboratories. We thank T. Moore, P. Jercog, J. C. Jung, D. Vucinic, B. F. Grewe, E. T. W. Ho, H. Kim, X. Pitkow and T. Zhang for discussions, D. Flickinger and K. Svoboda for providing design files for the water-immersion objective lens, C. Niell for providing tetO-GCaMP6 s/CaMK2a-tTA mice, and C. Irimia for animal husbandry.
Author information
Authors and Affiliations
Contributions
O.I.R., S.G. and M.J.S. designed experiments and analyses. O.I.R., J.A.L. and J.S. designed and built the microscope. O.I.R., J.A.L., O.H., Y.Z., R.C. and J.L. acquired and analysed data. S.G. developed theory and analysed data. H.Z. provided transgenic mice. O.I.R., S.G. and M.J.S. wrote the paper. All authors edited the paper. S.G. and M.J.S. supervised the research.
Corresponding authors
Ethics declarations
Competing interests
M.J.S. is a scientific co-founder of Inscopix, which produces the Mosaic software used to identify individual neurons in the Ca2+ videos. J.A.L. is also an Inscopix stockholder.
Additional information
Peer review information Nature thanks Stefano Panzeri and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 The discriminability of two sensory stimuli based on the activity patterns of two or more cells depends on the statistical relationship between the mean responses of the cells and their noise correlations, which in turn depends on visual neural circuitry.
a–f, Schematics of the distributions of responses by two cells to two distinct stimuli in six different cases. Cyan dots indicate joint responses of the cell pair to stimulus 1; orange dots indicate responses to stimulus 2. Ellipses convey the shapes of the statistical distributions of the responses to each stimulus. Three types of noise correlation are depicted. In a and d, the two cells have statistically independent noise correlations. In b and e, the cells share positively correlated noise fluctuations. In c and f, the cells share negatively correlated noise fluctuations. In all six cases, dashed lines indicate optimal linear boundaries for stimulus discrimination. The information in a–f is based on similar plots published previously5,11,30. a–c, When both neurons have similar stimulus-response properties (for example, as schematized, when both cells have a smaller mean response to stimulus 1 than stimulus 2), positively correlated noise fluctuations (b) increase the overlap between the two response distributions and thereby impair stimulus discrimination, whereas negatively correlated noise fluctuations (c) improve stimulus discrimination as compared to the case with independent noise fluctuations (a). d–f, When both neurons have opposite stimulus tuning (for example, as schematized, when neuron 1 responds more vigorously to stimulus 1 and neuron 2 responds more vigorously to stimulus 2), positively correlated noise fluctuations (e) decrease the overlap between the two response distributions as compared to the case with independent noise fluctuations (d) and thereby improve stimulus discrimination, whereas negatively correlated noise fluctuations (f) impair stimulus discrimination by increasing the overlap of the two response distributions. g, Cells in visual cortical areas, denoted by red circles, integrate signals from earlier stages of the visual pathway, as schematized by the input connections to two example cortical neurons. Thus, as visual information propagates through neural circuitry, noise fluctuations become correlated between cells with similar receptive fields, leading to an upper bound on the amount of information that a neural ensemble can encode. h, Example receptive fields for cells in g. Cells in early stages of the visual processing pathway have relatively simple receptive fields. Integration of their activity patterns leads to more complex visual receptive fields in downstream visual areas. Dashed boxes enclose receptive fields (right) for the two example cells marked in g, as well as the receptive fields of cells providing visual inputs (left). i, A network’s pattern of synaptic connectivity constrains the dimensionality of the activity in downstream visual circuits12. Left, in the early layers of the visual pathway, the dimensionality of ensemble activity is about the same order of magnitude as the number of photoreceptors. In downstream visual areas, due to the extraction of visual features, neural activity is constrained to a manifold of lower dimensionality (indicated by the red-shaded manifold in the space of all possible photoreceptor inputs). This manifold is determined by the set of receptive fields and hence the visual features that the downstream visual area detects. Grey ellipses (left) depict the distributions of photoreceptor responses to two distinct visual stimuli; after propagating through the visual circuitry these distributions are confined to the lower-dimensional manifold (red ellipses). Right, for a family of visual stimuli parameterized by a single variable, the mean neural ensemble responses lie along a corresponding tuning curve. Noise in the input circuitry propagates to downstream areas and leads to noise fluctuations in downstream neurons that are statistically correlated for cells with similar receptive fields. This, in turn, implies that the magnitude of noise fluctuations along the neural tuning curve becomes proportional to the number of cells in a neural ensemble and indistinguishable from the encoded visual signals, which also increase in proportion to the number of cells. This proportional growth of noise and signal ultimately limits the ability to discriminate two visual stimuli. Thus, for neural ensembles with more than a certain number of cells, the encoded information reaches an upper bound. j, We simulated a two-layer, linear feedforward neural network, to illustrate that information-limiting correlations are intrinsic to feed-forward neural networks with overlapping receptive fields12. Top, for three example output cells, the plot shows the synaptic weights of the inputs from cells in the first layer of the network. Bottom, diagram of connections between the two layers of the network. Symbols are defined as follows: x is the mean activity of cells in the first layer in response to a given stimulus; n is the noise in the activity of the input cells; r is the activity of the output cells. k, Digitized plots of spike counts for simulated activity in the network of j, for the two example input cells (yellow and black) and three example output cells (red, green, blue). The noise traces for the input cells came from independent Poisson random processes. External inputs to the network selectively drove either the yellow or the black cell, but owing to the presence of noise the two cells are occasionally active concurrently. l, Frequency plots of pairwise activity levels (rounded to the nearest integer) for pairs of output cells in the network of j. Yellow and black circles denote which of the two corresponding input cells received external input. The diameter of each circle denotes the number of time bins with a given pair of activity levels in the two cells. Σ values are noise correlation coefficients and are larger for pairs of output cells with greater overlap in their receptive fields. m, Plot of the distribution of activity responses in the output cell layer, for the three example cells coloured green, red and blue in j. Data points are coloured either yellow or black, to indicate whether the output activity is a response to stimulation of the yellow- or black-coloured cell in the input layer. The red plane denotes the optimal linear classification boundary between the two stimulation conditions.
Extended Data Fig. 2 Spatiotemporal multiplexing of the illumination beams permits imaging of large fields of view at fast frame rates without thermal damage to brain tissue.
a, Computer-assisted design of the mechanical layout of the two-photon microscope. Scale bar, 0.5 m. b, In the pixel multiplexing mode of imaging, each of the 16 beams are assigned to one of four different temporal phases within each cycle of the pixel clock (Extended Data Fig. 3b). Alternatively, in the line-multiplexing mode of imaging, only 8 of the 16 beam paths are used (Methods). In neither imaging mode are neighbouring beams ever active concurrently (Extended Data Fig. 3c), minimizing fluorescence scattering between active image tiles and allowing scattering into inactive image tiles to be corrected computationally (Extended Data Figs. 3d, e, 4a–g). c, To switch between the different sets of active beams, square-wave electronic signals control a set of three electro-optic modulators (EOMs). d, A Ti:sapphire laser provides ultrashort-pulsed infrared illumination. A half-wave (λ/2) plate and a polarizing beam-splitter enable power control. Three pairs of EOMs and polarizing beam-splitters direct the light into one of four main optical paths, with only one path illuminated during each of the four multiplexing phases. In each of these four main paths, three 50:50 beam-splitters create four beams of equal intensity, yielding up to 16 total beams but with only four on at any instant. A chopper blocks all light during the turnaround portion of the galvanometer scanning cycle. e, Seventy-five example fluorescence traces of Ca2+ activity in layer 2/3 pyramidal cells of an awake mouse. f, Maintaining brain temperature within physiological ranges during in vivo two-photon imaging requires a proper balance between heat loss through the cranial window and heating induced by the laser illumination53,55. To directly verify that our cranial window preparation and imaging conditions properly balanced these two opposing effects, we measured brain temperature during two-photon imaging with the 16-beam microscope. For these studies we used an implanted thermocouple53 and either the highest (blue trace) or lowest (green trace) time-averaged laser illumination intensity used for Ca2+ imaging elsewhere in this study (Methods). Consistent with previous work, before laser illumination commenced the brain temperature was about 9 °C below normal mouse body temperature55, a state that is considered to be neuroprotective56. By about 100 s after the start of imaging, brain temperatures attained steady-state values within the physiological range of C57BL/6 mice57 (grey shaded region; 36.3 °C–38.7 °C). Each trace is an average of three bouts of imaging for each of three separate mice. Coloured shading denotes the s.d. across the 9 individual measurements acquired at each illumination intensity. g–i, Fluorescence immunohistochemical analyses of tissue damage markers. To check whether in vivo imaging of brain tissue with the 16-beam instrument (4 mm2 field of view) induced any tissue damage, we immunostained post-mortem brain tissue sections using antibodies to two different damage markers, glial fibrillary activation protein (GFAP) and heat shock protein 70 (HSP70), previously identified as indicators of laser-induced tissue damage53. We also stained the sections with DAPI, which labels cell nuclei. We compared positive control tissue sections (g) that we had deliberately damaged in vivo with high-power (2,680 mW mm−2) laser illumination, negative control sections (h) that received no laser illumination, and experimental tissue sections (i) that had undergone in vivo two-photon imaging at the highest level of laser illumination (80 mW mm−2) used in this study for tracking Ca2+ dynamics in neocortical layer 2/3 pyramidal neurons. Together, these analyses verified the functionality of the antibodies and revealed no signs of tissue damage from two-photon imaging. To image neurons in cortical layers deeper than layer 2/3, users have several options for doing so without delivering excess heat to the brain (Supplementary Video 3, Supplementary Note). Scale bars, 500 μm. Results shown are representative of those from 8 cerebral hemispheres of 4 different mice. j, k, Comparisons between recent large-scale two-photon microscopes24,26. The performance of a laser-scanning microscope closely relates to four main parameters: the scanner speed, image-frame acquisition rate, field of view, and pixel size (Supplementary Note). For microscopes that use a single laser beam to sweep in two dimensions across the field of view, these parameters obey the relationship FOV = d × v × f −1, where FOV is the field-of-view area, d is the spacing between adjacent image lines (or equivalently the pixel width along the slow-axis of laser-scanning), v is the speed at which the beam is swept across the specimen by the fast-axis scanner, and f is the image-frame acquisition rate. By comparison, our approach using four active beams leads to an expression for the maximal field of view, FOV = 4 × d × v × f −1. These relationships enable performance comparisons with other recently published large-scale two-photon microscopes24,26. To illustrate, j shows a plot of the image-frame acquisition rate against the field-of-view area, given a line spacing of d = 1.15 μm. k shows how the image-frame acquisition rate depends on d for a 4 mm2 field of view. Solid red circles denote the performance of our microscope in its line-multiplexing imaging mode using an 8-kHz resonant galvanometer (Methods). Black data points denote performance options of another large two-photon microscope, which uses pair of laser beams with temporally interleaved pulses24, as calculated on the basis of its published capabilities. Blue data points and associated blue dashed lines show performance options for a third large-scale microscope26, as calculated on the basis of its published capabilities.
Extended Data Fig. 3 Data acquisition and post-processing for two-photon imaging with 16 time-multiplexed excitation beams.
a, Block diagram of the electronics for data acquisition and instrument control. PMT, photomultiplier tube; Pre-amp, pre-amplifier; ADC, analogue-to-digital converter; FPGA, field-programmable gate array; EOM, electro-optic modulator. b, Computer simulation of signal sampling in different stages of the pipeline in a. The ADC samples the analogue, pre-amplified and low-pass filtered signals (blue) from one of the PMTs at a rate of 5 × 107 samples per second. In each of the four temporal phases, the FPGA sums the digitized signals (red) from the ADC to yield the fluorescence intensity values of each image pixel (grey). c, Raw fluorescence images for each of the four excitation phases, acquired in an awake mouse expressing GCaMP6f in layer 2/3 cortical pyramidal cells and averaged over 100 frames (7.23 Hz acquisition rate). In each of the four phases, a distinct set of four PMTs detects most of the fluorescence emissions, creating four active image tiles within the 4 × 4 array. (Each of the four PMTs corresponds to one of the four laser beams that is active in that phase.) To illustrate, the four active tiles within the phase I image are shaded with a different colour (shaded large square regions). However, close to the boundaries of each active tile, some fluorescence photons are detected by the other 12 PMTs. During signal unmixing these photons are reassigned to corresponding pixels in the correct adjacent active image tile. For instance, within the phase I image photons detected in the areas outlined in colour (rectangles and small squares) are reassigned to the colour-corresponding active tiles. d, An image compiling the four sets of four active image tiles from the panels in c. e, During signal un-mixing, we re-assign scattered fluorescence photons to their correct pixels of origin, using the method shown in c, by reassigning the boundary regions of 128 pixels width. The resulting image is displayed with the mean contrast equalized across tiles. Scale bars: c, e, 500 μm.
Extended Data Fig. 4 Crosstalk un-mixing procedure for reconstructing the full field-of-view enables accurate estimation of neural activity traces.
a, To quantify the extent of fluorescence scattering across image tiles, we acquired images in two distinct configurations that enabled us to distinguish fluorescence signals from any crosstalk due to fluorescence scattering across image tiles. Using an awake mouse expressing GCaMP6f in layer 2/3 cortical pyramidal cells, we first imaged with only one active laser beam and its corresponding PMT; the other 15 beams were blocked (configuration 1). In this configuration, there is no fluorescence scattering into the active image tile from the other 15 tiles, only the signals from the active tile. In configuration 2, we blocked the beam that had previously been active, unblocked the other 15 beams, operated the microscope with the normal multiplexing approach, and again sampled signals from all 16 PMTs. To estimate the extent of scattering into the tile with the blocked beam, we applied the computational un-mixing procedure to the raw image data. To estimate how much scattered fluorescence affects cell sorting, we first extracted individual cells and their Ca2+ activity traces from the first dataset, attained in configuration 1 without crosstalk. We then summed the images, frame by frame, from the two datasets, to create a mock dataset comprising unscattered plus scattered fluorescence signals, from which we again computationally extracted cells and their activity traces. This enabled a direct comparison between two datasets containing the exact same patterns of neural activity, with and without fluorescence scattering from other image tiles. b, Activity traces for four example cells, enabling comparisons of the Ca2+ activity traces (top), ΔF(t)/F0, and the resulting traces of the estimated spike counts (bottom), between the datasets with (red traces) and without (black traces) inter-tile scattering. The traces with and without inter-tile scattered fluorescence signals are nearly indistinguishable by eye. c, Histogram of the ratio of estimated spikes for the two datasets constructed in a, for all time bins (0.14 s per time bin) with an estimated spike count greater than 0.5. The mean ratio is 1.0 ± 0.06 (mean ± s.d.; N = 31 cells). Total number of time bins, 5,865. d–g, Studies of fluorescence scattering between the active image tiles in one temporal phase (Extended Data Fig. 2b) of the multiplexing scheme used for two-photon imaging. Throughout the paper, we corrected computationally for fluorescence scattering from active to inactive image tiles within each temporal phase of imaging (Extended Data Fig. 3c, Methods). This approach neglects the small amount of fluorescence scattering from active tiles to other active tiles, which in principle could also be computationally corrected using a more sophisticated method than the one we adopted. Hence, we examined experimentally the validity of our computational approach and the extent to which scattering between active tiles can be justifiably neglected. The amplitude of scattering between active tiles (d) varies with the location of each laser beam and its proximity to a tile boundary. We used fixed cortical tissue slices from adult GCaMP6f-tTA-dCre mice to measure the amplitude of such scattering effects when imaging at different depths within brain tissue. An image (e) of the spatial distribution of two-photon fluorescence excited 500 μm deep within a tissue slice shows that a majority of scattered fluorescence photons exits the brain tissue relatively near to the laser focus. By averaging over 100 different laser foci positions in each of 3 different brain slices, we determined the mean cross-sectional spatial profiles (f) of scattered fluorescence excited at different depths in tissue, as a function of the lateral displacement, x, from the laser focus. Profiles are shown normalized to unity at x = 0. The inset of f shows a magnified view of these cross-sectional profiles for x ∈ [–1,000 μm, –500 μm], that is, up to 1 mm away from the laser focus. We used these empirically determined scattering profiles to compute the probability (mean ± s.d.; N = 300 laser focus positions) (g) that a fluorescence photon originating in one active image tile would scatter into an adjacent active tile. Even when the laser focus is on the boundary of an image tile, this probability remains less than 0.02 for all tissue depths ≤ 600 μm. For our studies of layer 2/3 cortical pyramidal cells in live mice, the probability of a fluorescence photon scattering between active tiles is less than 0.01. In conclusion, computational corrections for fluorescence scattering that account solely for scattering from active to inactive tiles—and neglect scattering between different active tiles—are empirically well justified.
Extended Data Fig. 5 Pipeline of offline data processing and procedures for reducing the dimensionality of the neural ensemble activity data and calculating the decoding accuracy.
a, Pipeline of the offline procedures we applied to the acquired fluorescence signals to attain traces of neural activity. Steps coloured purple involve algorithms that use raw or processed image data. Steps coloured yellow involve algorithms that use cells’ spatial filters as their input arguments. Steps coloured green involve algorithms that use cells’ activity traces as their inputs. Purple steps, starting from the raw photocurrents from each of the 16 PMTs (sampled at 50 MHz and assigned to individual image pixels corresponding to a 400-ns laser dwell time), we normalized the photocurrent signals by the gain of each individual PMT, to equalize the image intensity scale across the entire image. We then un-mixed scattered fluorescence, as shown in Extended Data Fig. 3, and applied an image registration routine (TurboReg44) to the videos from the individual image tiles. To highlight Ca2+ transients against baseline fluctuations, we used the fact that the two-photon fluorescence increases of GCaMP6 during Ca2+ transients are many times the s.d. of background noise. Thus, we converted the fluorescence trace of each pixel, F(t), into a trace of z-scores, ΔF(t)/σ. Here ΔF(t) = F(t) – F0 denotes the deviation of the pixel from its mean value, F0, and σ denotes the background noise of the pixel, which we estimated by taking the minimum of all standard deviation values calculated within a sliding 10-s window35. After transforming the movie data into this ΔF(t)/σ form, we identified neural cell bodies and processes using an established cell-sorting algorithm that sequentially applies principal and independent component analyses (PCA and ICA) to extract the spatial filters and time traces of individual cells48. Yellow steps, for all spatial filters corresponding to individual cell bodies, we thresholded the filters at 5% of each filter’s maximum intensity and set to zero any filter components with non-zero weights outside the soma. To attain neural activity traces, we then reapplied the set of resulting filters to the ΔF(t)/F0 movies. Green steps, to estimate the most likely number of spikes fired by each cell in each time bin, we applied a fast non-negative deconvolution algorithm to the ΔF/F0 trace of the cell49. For each neuron, we down-sampled (2×) the activity traces to time bins of 0.275 s by averaging the values within adjacent time bins. To make comparisons across similar behavioural states, we removed all trials during which the mouse was moving. b, Neural responses for each visual stimulus (A and B) are represented as matrices of size Nneurons × Ntrials × Ntime bins. To calculate the accuracy of stimulus discrimination, we first randomly chose a subset of neurons from the dataset. For decoding using the ‘instantaneous’ strategy (Fig. 3, Extended Data Figs. 7–10), we then chose a specific time bin, whereas for the ‘cumulative’ decoding strategy we treated all the different time bins up to a specific time, t, as independent dimensions of the population activity vector. We then split the trials in half, into a training set and a test set, each with equal numbers of trials with the A and B stimuli. We took the neural activity traces in the training set and normalized them by the s.d. of the cell’s activity about its mean, to create to a set of z-score traces. We then performed PLS analysis to identify a low-dimensional basis that well captured the separation between the neural responses to the two sensory stimuli. Using the activity data in the test set, we applied the same normalization and dimensional reduction procedures and values as for the training set. We used the resulting distributions of responses to calculate d′ values and the eigenvectors of the noise covariance matrix. For each mouse we repeated this entire procedure for 100 different randomly chosen subsets of neurons.
Extended Data Fig. 6 Distributions of pairwise noise correlation coefficients do not differ significantly between pyramidal neurons in area V1 and higher-order visual areas.
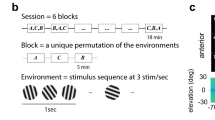
a, Anatomical maps of visual cortical neurons that responded to each of the two stimuli. For these maps (but for no other analyses in the paper), we denoted a cell as responsive to one of the stimuli if, in at least one time bin during the 2-s-stimulation period (0.275 s per bin), the difference between the cell’s mean response and its mean activity trace during the inter-trial intervals was more than twice the sum of the s.e.m. values for these two traces. Cells that responded to stimulus A only are shown red, those that responded only to stimulus B only are shown blue, those that responded to both stimuli are shown purple. b, Mean Ca2+ responses (ΔF/F) of 25 example neurons to the two different moving grating stimuli, oriented at ± 30°. Ca2+ activity traces are shown coloured during the stimulation period (marked with light grey shading) and black otherwise. Coloured shading about each trace denotes the s.e.m. over 217 trials of each type. The inset shows a schematic of the two stimuli, which appeared for 2 s per trial and were presented in random order. c, d, Histograms of the estimated mean spiking rates of individuals neurons during visual stimulation (c) and the absolute values of the differential responses of the individual neurons to the two visual stimuli, |RA – RB| / (RA + RB) (d), where RA and RB denote the mean responses of a cell to stimuli A and B, respectively. The distributions of cells’ activity rates and preferences for one stimulus over the other were consistent with previous studies of rodent visual cortical neurons28,29,38,58,59. Data shown are for N = 8,029 individual cells from N = 5 mice. Error bars are s.d. as estimated on the basis of counting errors. e, Histogram of noise correlation coefficients, r, between pairs of layer 2/3 pyramidal neurons, computed as in Fig. 2d, for V1 cell pairs (dashed lines) and cells pairs in higher-order visual areas (solid lines). The histograms show mean values across the two different visual stimuli for both the real neural activity traces, and for trial-shuffled data in which each cell’s responses to each stimulus presentation were randomly permuted across the set of all presentations of the same stimulus. r values were computed on the basis of cells’ responses integrated over t = [0.5 s, 2 s] from the start of each trial. Histogram bin, 0.01. (N = 1,331,109 V1 cell pairs from 5 mice; N = 2,428,437 cell pairs from higher-order visual areas in 5 mice). f, Box-and-whisker plots of the mean and FWHM values of the distributions in e (real data only). Both statistical metrics are similar for the two classes of visual cortical neurons. Open circles denote individual data points for N = 5 mice. g, h, Histograms (g) and cumulative probability distributions (h) of noise correlation coefficients for all cell pairs (based on all recorded V1 and higher-order visual cortical neurons) with similar or differently tuned mean evoked responses to the two visual stimuli. Unlike Fig. 2e, which shows these distributions for only the most active cells (the highest decile), here the distributions include all cell pairs with either positively (red curves) or negatively (blue curves) correlated mean responses to the two stimuli. Within these two groups of cell pairs, we computed the noise correlation coefficient, r, for each cell pair. Owing to the extremely large number of cell pairs, the two distributions of r values differed significantly (***P < 10−13 for all 5 individual mice; two-tailed Kolmogorov–Smirnov test; 3,482,186 positively correlated cell pairs in total; 3,464,094 negatively correlated pairs), even though the effect size was tiny and the two distributions were nearly identical. This result shows the difficulty of detecting information-limiting correlations by measuring pairwise noise correlations, because the variance in the individual r values is much greater than the difference between the mean values of the two distributions. i, Box-and-whisker plots of the mean values of the correlation coefficients in g, h. Open circles mark individual data points for N = 5 mice. b–i are based on 217–332 trials per stimulus condition in each of 5 mice. In f, i, boxes cover the middle 50% of values, horizontal lines denote medians, and whiskers span the full range of the data.
Extended Data Fig. 7 Temporal integration of neural activity improves decoding performance, but quadratic and linear decoding yield identical biological conclusions.
a–c, To identify how many PLS dimensions were needed to determine d′ accurately, we divided data from each of 5 mice into three equally sized portions. We performed PLS analysis using trials in the first third. Onto the PLS dimensions thereby identified, we projected the neural ensemble activity in the second third of the data (training data). We retained only the first NR dimensions of this projection and computed d′ in the reduced space (magenta data points) by identifying a hyperplane for optimal stimulus discrimination. Finally, we applied this discrimination strategy to the remaining third of the data (test data) and again calculated d′ (grey points). Plots show mean values of d′ as a function of NR for the interval [0.83 s, 1.11 s] from stimulus onset (N = 5 mice; error bars denote s.d. across 100 different subsets of 1,000 neurons per mouse). We normalized d′ values to that found for NR = 5 on the test dataset. For NR > 5, discrimination performance declines owing to overfitting for all discrimination strategies: instantaneous (a), cumulative (b) and integrated (c). Hence, throughout the rest of the study we used NR = 5 for all calculations of d′. d, Pearson correlation coefficients between the optimal linear decoding weights attained using instantaneous decoding at different time bins after the onset of grating stimuli (±30° orientations). These weights were highly correlated for different time bins, especially across the interval [0.5 s, 2 s], during which d′ reaches a plateau. Further, optimal decoders for each time bin yielded nearly equivalent decoding performance when applied to data from other time bins. For instance, the optimal decoder for the fourth time bin (t = 0.97 s), when applied to any other of the last five time bins, yielded a performance within less than 2% of that of the optimal instantaneous decoder in all mice. When applied to the first and second time bins, the decoder from the fourth time bin yielded decoding performances that were, respectively, 83 ± 11% and 90 ± 3% (mean ± s.d.; N = 5 mice; 217–232 trials per stimulus) of that of the optimal decoders. e, Plots of d′ versus time after stimulus onset, for instantaneous and cumulative decoding strategies (Fig. 3). For each mouse that viewed gratings oriented at ±30°, we chose 100 random subsets of 1,000 cells and normalized d′ values by those obtained using a time-integrated decoding strategy, which involved optimal linear discrimination over one interval, [0.28 s, 1.94 s], covering most of the visual stimulation period. Green traces, mean d′ values for individual mice using a time bin of 275 ms. Error bars, s.d. across 5 mice. f, In the five-dimensional space used after truncating ensemble neural responses to the five leading PLS dimensions, the distributions of noise in the responses to the two stimuli were highly similar. Specifically, non-diagonal elements, Σij, of the noise covariance matrices for the two stimulus conditions were highly correlated (r: 0.81 ± 0.16; mean ± s.d.; N = 5 mice), as computed for the interval [0.83 s, 1.11 s] after stimulus onset. This similarity argues that a linear discrimination strategy to classify the two sets of ensemble neural responses is near optimal, as confirmed in h. Values of Σij are plotted as mean ± s.d., computed across 100 different randomly chosen subsets of 1,000 neurons per mouse. g, Using optimal linear decoding, d′ values saturated as the number of trials analysed increased. Colours denote individual mice. Data points were calculated for the interval [0.83 s, 1.11 s] after stimulus onset. Error bars, s.d. across 100 different randomly chosen subsets of 1,000 cells per mouse and stimulation trials. h, To check whether our results depended on our use of linear decoding, we tested whether quadratic decoding might yield different conclusions. We examined the KL divergence31, a generalization of (d′)2 that makes no assumption about the statistical distributions under consideration. We computed the KL divergence, which equals (d′)2 for linear decoders, by using Gaussian approximations to the distributions of ensemble neural responses to the two different stimuli, and we plotted the results as a function of the number of cells, n, in the ensemble. First, to recapitulate our determinations of (d′)2 (magenta data points), we computed the KL divergence under the assumption the two different response distributions had distinct means but identical noise covariance matrices, which we estimated as the mean noise covariance matrix averaged over the two different stimulus conditions. This is equivalent to computing (d′)2. Next, we relaxed the assumption that the two noise covariance matrices were equal and computed the KL divergence between the distributions of neural responses to stimulus B relative to those to stimulus A (blue points), and vice versa (red points) (Methods). For all mice, KL divergence values saturated with increasing n and, except in one mouse, were not much larger than (d′)2 values. Thus, quadratic decoders (which are optimal for discriminating two Gaussian distributions with different means and covariances) will yield the same basic conclusions as linear decoders (which are optimal for discriminating two Gaussian distributions with the same covariance matrix). Data points and error bars denote mean ± s.d. values computed in each mouse across 50 different randomly chosen subsets of cells and assignments of visual stimulation trials to decoder training and testing (Extended Data Fig. 5b). i, Mean neural responses, averaged across all cells, to stimuli A (top) and B (bottom) for the first and second halves of the experimental trials in each mouse. Error bars, s.d. across the set of trials. j, d′ values computed for each mouse using instantaneous decoders trained on the first half of the trials and tested on the second half (x axis), plotted with d′ values for an instantaneous decoder trained on the second half of the trials and tested on the first half (y axis). a–j are based on 217–332 trials per stimulus condition in each of 5 mice.
Extended Data Fig. 8 PLS-based decoding methods are robust to multiplicative gain modulation and common mode fluctuations in the neural ensemble dynamics and yield identical conclusions to regularized regression.
a, b, To test whether PLS analysis and dimensionality reduction might lead to underestimates of d′, we compared d′ values determined using an L2-regularized regression (L2RR) performed in the full space of neural responses (a) to those found by PLS analysis (b). The two methods yielded similar estimates of d′, which both saturated with increasing numbers of neurons. Plots show d′ values (mean ± s.d.) for neural responses within [0.83 s, 1.11 s] after stimulus onset, computed across 100 different randomly chosen subsets of neurons and visual stimulation trials (Extended Data Fig. 5b). For PLS analyses, we used half of the trials in each subset for decoder training and the other half for testing. For L2RR we used 90% of the trials in each subset to determine the regression vector and the other 10% to determine d′. We varied the regularization parameter, k, within [1, 105] and used the maximum d′ value so obtained, as determined independently for each mouse, subset of neurons, and subset of trials (217–332 trials per stimulus condition in each of 5 mice). c–h, The conclusions of our study depend on comparisons of decoding performance between real and trial-shuffled datasets. Thus, we checked whether our PLS-based decoding methods would robustly detect information-limiting correlations in models in which such correlations were present but weak; avoid reporting information-limiting correlations in models lacking such correlations; and be robust to the potential presence of other strong sources of neural trial-to-trial variability—such as common mode fluctuations and multiplicative gain modulation—even when they make an order-of-magnitude greater contribution to neural variability than the information-limiting noise fluctuations. We studied these issues using two different computational models (Methods). For both models we plotted empirically determined (d′)2 values as a function of the number of neurons in the ensemble. We compared determinations of (d′)2 using PLS-based decoding and those made using L2RR to the actual ground truth values of (d′)2 in each model. In each panel, the top and bottom plots show results for unshuffled and trial-shuffled datasets, respectively. Data points and error bars denote mean ± s.d. values across 30 different simulations. To examine the combined effects of information-limiting noise correlations and common mode fluctuations (c–f) we studied a model of neural ensemble responses in which the noise covariance matrix exhibited information-limiting noise correlations via a single eigenvector f, the eigenvalue of which grew linearly with the number of cells in the ensemble. In addition to this rank 1 component, we included a noise term that was uncorrelated between different cells, as well as a common mode fluctuation, yielding a noise covariance matrix with the form Σ* = σ2I + εcommonJ + ε fT f, where σ2 = 1 is the amplitude of uncorrelated noise, I is the identity matrix, J is a rank 1 matrix of all ones, reflecting a common mode fluctuation, and f is the information-limiting direction, a vector that we chose randomly in each individual simulation from a multi-dimensional Gaussian distribution with unity variance in each dimension. The amplitude of information-limiting correlations was ε = 0.002, approximately matching the level observed in the experimental data. We chose the difference in the means of the two stimulus response distributions, Δμ, to be aligned with f (Fig. 3a) and to have a magnitude of 0.2 so that the asymptotic value of d′ for large numbers of cells approximately matched that of the data. We compared decoding results attained with and without the presence of the common mode fluctuations in the neural responses. In the version of the model without common mode fluctuations, we set εcommon to zero. In this case (c) both PLS- and L2RR-based decoders correctly detected the saturation of information in the real data but not in trial-shuffled datasets. (See Extended Data Fig. 10h, k for theoretical results showing how the accuracy of d′ estimates from PLS analysis depends on the numbers of neurons and experimental trials in this particular model.) To verify that our methods would not incorrectly report an information saturation when it was in fact absent, we next set ε = 0 and confirmed that in the absence of information-limiting noise correlations (d), neither decoder detected a saturation of information in the real or shuffled data. In the version of the model with common mode fluctuations, we set εcommon = 0.02, ten times the value of ε = 0.002. In this case (e), both PLS- and L2RR-based decoders correctly detected the information saturation in the real but not in the shuffled data. To verify that common mode fluctuations alone cannot induce an illusory saturation of information (f), we set ε = 0 while maintaining εcommon = 0.02 and confirmed that neither PLS- nor L2RR-based decoders reported an illusory information saturation. Overall, these results indicate that our methods accurately detect the presence of weak information-limiting correlations buried within common mode noise that can be an order of magnitude larger, without falsely detecting information-limiting correlations when they are absent. To study the possible effects of multiplicative gain modulation (g, h), we compared two versions of a model in which the responses of the V1 neural population either were or were not subject to a multiplicative stochastic gain modulation but were otherwise statistically equivalent. We modelled the V1 cell population as a set of Gabor filters (see Appendix section 5). In the model version with gain modulation, on each visual stimulation trial we multiplied the output of each Gabor filter by a randomly chosen factor, uniformly distributed between 50%–150%, the value of which was the same for all cells but varied from trial to trial. In the model version without gain modulation (g) both PLS- and L2RR-based decoders detected the information saturation in the real but not in the trial-shuffled datasets. When we added global gain modulation to the model (h) both decoders correctly found the information saturation in the real but not in the shuffled datasets.
Extended Data Fig. 9 Moving grating visual stimuli oriented at ±6° are harder to distinguish on the basis of their evoked neural ensemble responses than gratings oriented at ±30°, but also reveal the saturation of information signalling in large neural populations.
a, (d′)2 values determined using an ‘instantaneous’ decoder for the interval [0.70 s, 0.94 s] from visual stimulation onset, plotted as a function of the number of cells, n, in the ensemble in mice presented moving gratings oriented at ±6°. Data points represent mean values determined across 100 different subsets of cells, and the shading represents s.e.m. As in Fig. 3f, g, we fit the (d′)2 values as a function of n using a one-parameter fit, (d′)2 = (d′)2shuffled/(1 + ε × n), where (d′)2shuffled (n) is the empirically determined value of (d′)2 for the same number of cells in the shuffled data, and ε is the fit parameter. For each mouse, for both real and trial-shuffled data we normalized (d′)2 values by the value of (d′shuffled)2 for n = 1,000 neurons. Goodness of fit: R2 = 0.41 ± 0.17 (s.d). N = 5 mice. ε = 0.0021 ± 0.0008 (s.d.), 122–167 trials per stimulus condition for each mouse. b, Same as a, but using the ‘cumulative’ decoding strategy over the [0 s, 0.94 s] time interval. c, Box-and-whisker plots of the asymptotic values of d′ in the limit of many neurons (right) and the number of cells at which (d′)2 attains half its asymptotic value (left) as determined from parametric fits to the data of a and b for the instantaneous (open boxes) and cumulative (filled boxes) decoding strategies. Optimal linear decoders (green data) slightly but significantly outperformed diagonal decoders (black data) (**P < 0.0001; one-tailed Wilcoxon rank sum test; N = 100 different randomly chosen assignments of trials to decoder training and test sets in each mouse; 122–167 trials per stimulus condition for each mouse; open circles denote mean values from N = 5 individual mice). d, e, Histograms for the real (unshuffled) and shuffled datasets of the ensemble neural responses to each of the two visual stimuli, projected onto the direction of the optimal decoding vector determined by PLS analysis, as computed in each mouse viewing moving gratings oriented either at ±30° (d) or ±6° (e), using all imaged neurons and the instantaneous decoding approach. Error bars denote counting errors. Values on the x axes are plotted for each mouse in units of the s.d. of its neural ensemble responses along the decoding vector for the shuffled data. For each mouse, the histograms have approximately equal shapes for the two visual stimuli, are unimodal and approximately symmetric about their mean values, bolstering the use of linear decoding and d′. This analysis involved 217–232 trials per stimulus condition per mouse in d and 122–167 trials per stimulus condition per mouse in e.
Extended Data Fig. 10 Hundreds of experimental trials sufficed to estimate the statistical structure of signals and noise in visual cortical coding.
a, b, PLS analysis represents ensemble neural responses in a low-dimensional subspace that helps for understanding visual discrimination (Fig. 4). On the basis of Extended Data Fig. 7a–c, computations here used the five most informative PLS dimensions. Each column shows results from an individual mouse that viewed gratings oriented at ±30° (217–332 trials per stimulus). Each colour denotes a different eigenvector, eα, of the noise covariance matrix in the five-dimensional subspace. α denotes the dimension index, {1,2,3,4,5}. As illustrated in Fig. 4e, each mouse had multiple eigenvalues, λα, of the noise covariance matrix that increased with the number of cells, n, used for analysis. As shown in Fig. 4f, visual signals—defined as the mean separation, Δμ, between the two response distributions—also increased with n. a, b show eigenvalues λα (a) and signal components |Δμ · eα| (b) plotted against the number of trials analysed. Both signal and noise estimates plateau, indicating that there were sufficient trials to accurately estimate signal and noise structure in the reduced five-dimensional space. Throughout a–d, lines and shading denote mean ± s.d. across 100 different randomly chosen subsets of cells and assignments of trials to decoder training and testing, except in a, b we used all cells from each mouse and 30 different assignments of trials. c, d, The statistical relationships between visual signals and noise show the largest noise mode is not information-limiting. Each mouse had multiple eigenvalues, λα, of the noise covariance matrix (c) that increased with n, the number of cells. Visual signals (d) also increased with n, as shown by decomposing Δμ into components along the five eigenvectors, eα. In every mouse the eigenvector with the largest eigenvalue, e1, was the least well aligned with the signals, Δμ (compare red curves in c, d). e, Plots of noise values, computed as in c, versus signal values, computed as in d, based on all recorded neurons from each mouse and the same 100 subsets of data used in c, d. The largest noise mode (red points) was generally an order of magnitude greater than noise modes that limited neural ensemble signalling (green and yellow points). f–k, In a–e and throughout much of the paper, we analysed populations of up to 2,191 neurons using 217–332 trials with each stimulus, which sufficed to accurately determine the Fisher information, (d′)2, and principal eigenvectors of the noise covariance matrix (Fig. 4). By comparison, there were insufficient trials to accurately determine noise covariance matrix elements—that is, noise correlations between cell pairs (Fig. 2d). To explain this, we derived the accuracy with which d′ and principal noise covariance eigenvectors and eigenvalues can be estimated through PLS analysis of recordings of n neurons across P trials, using the computational model of Extended Data Fig. 8c (Appendix section 6 has derivations of results in f–k). The central idea, illustrated in f, is that one can estimate accurately the principal noise covariance eigenvector, because it has a large eigenvalue, λ, that grows linearly with n (\(\lambda \cong cn\), where c is a constant). The theory predicts that the correlation coefficient, \({\mathscr{C}}\), between estimated and actual eigenvectors is given by \({{\mathscr{C}}}^{2}=\frac{cP-1/(cn)}{cP+1}\), for \({c}^{2}Pn > 1\). Otherwise, \({\mathscr{C}}\) = 0. f shows predictions for \({\mathscr{C}}\) (black curve) versus the number of trials, P, for n = 2,000 and c = 0.005. We chose this c value to fall within the lower range of growth rates for experimentally determined eigenvalues, c. The predicted \({\mathscr{C}}\) values match those describing the accuracy (red points) with which we could estimate the principal noise covariance eigenvector in the computational model. However, correlation coefficients (blue points) between estimated and actual individual elements of the noise covariance matrix were unsatisfactory, even with 800 trials. i shows predicted values of \({\mathscr{C}}\) as a joint function of n and P. Iso-contours of \({\mathscr{C}}\) are hyperbolic, revealing a tradeoff such that recording more cells enables accurate estimation of noise eigenvectors using fewer trials. We also derived how accurately one can estimate eigenvalues of the noise covariance matrix, as quantified using the ratio, \({\Re }_{\lambda }=\lambda /\hat{\lambda }\)where λ = cn is the actual eigenvalue in the model and \(\hat{\lambda }\) is the estimate based on P trials. The theory predicts \({\Re }_{\lambda }=\frac{cP}{cP+1}\) when \({c}^{2}Pn > 1\); otherwise we set \({\Re }_{\lambda }=0\), because we cannot accurately estimate the corresponding eigenvector when \({c}^{2}Pn < 1.\) g plots predictions of \({\Re }_{\lambda }\,\)(black curve) versus P (for n = 2,000 cells and c = 0.005), which match the accuracy with which we estimated the model eigenvalues from simulated data (red dots). j shows \({\Re }_{\lambda }\) predictions as a joint function of n and P. We also studied how well one can estimate the Fisher information, (d′)2, via PLS analysis of data with fewer trials than recorded neurons. We examined the ratio, \(\Re \), of the d′ estimate to its actual value using the model and simulated data of Extended Data Fig. 8c and found \({\Re }^{2}=\,\frac{1+n\varepsilon }{1/{{\mathscr{C}}}_{{\rm{PLS}}}^{2}+n\varepsilon }\), where \({{\mathscr{C}}}_{{\rm{PLS}}}^{2}=\frac{\Delta {s}^{2}P+4(\varepsilon +1/n)}{\Delta {s}^{2}P+4(\varepsilon +1)}\) is the predicted correlation coefficient between the PLS regression vector and the optimal one. Here Δs2 and ε determine the Fisher information in the model of Extended Data Fig. 8c via \({({d}_{{\rm{o}}{\rm{p}}{\rm{t}}}^{{\prime} })}^{2}\,=\,\frac{n\Delta {s}^{2}}{1+n\,\varepsilon }\). As in Extended Data Fig. 8c, we used ε = 0.002 to match the growth rate of (d′)2 in experimental data with increasing n, and Δs2 = 0.04 to approximate the magnitude, \(\frac{\Delta {s}^{2}}{\varepsilon }\), of (d′)2 in the data for large n. \({{\mathscr{C}}}_{{\rm{PLS}}}^{2}\) increases monotonically with P and n, confirming that PLS regression improves as n and P increase. As \({{\mathscr{C}}}_{PLS}^{2}\) nears 1, so does \({\Re }^{2}\), indicating that PLS analysis can accurately estimate (d′)2. h shows predictions for \({\Re }^{2}\) versus P for n = 2,000 cells (black curve). The theory matches the accuracy with which we estimated (d′)2 via PLS analyses of the simulated model data (red dots). k shows predicted \({\Re }^{2}\) values versus n and P. Iso-contours of \({\Re }^{2}\) are hyperbolic, indicating recordings of more neurons permit accurate estimates of (d′)2 based on fewer trials.
Supplementary information
Supplementary Information
Supplementary Appendix | Mathematical derivations and analyses regarding information-limiting noise correlations.
Supplementary Information
Supplementary Note | Technical discussion of large-scale two-photon imaging. References for the Supplementary material.
Supplementary Table
Supplementary Table 1 | Summary of statistical results associated with the figures and extended data figures.
41586_2020_2130_MOESM5_ESM.mp4
Video 1: The 16-beam two-photon microscope enables simultaneous monitoring of Ca2+ dynamics in >2000 cortical neurons in an awake mouse. A two-photon Ca2+ video of the activity of layer 2/3 visual cortical pyramidal neurons expressing GCaMP6f in an awake mouse. 2191 individual cells were identified in the full video dataset from this mouse. The data were recorded at 7.23 Hz and are played back at 8× real-speed (30 fps playback, with each frame the average of two image acquisitions). The field-of-view is 2 mm × 2 mm.
41586_2020_2130_MOESM6_ESM.mov
Video 2: Large-scale Ca2+ dynamics of layer 2/3 neocortical neurons in an awake mouse, recorded at 17.5 Hz over a 2 mm × 2 mm field-of-view. A two-photon Ca2+ video of the activity of layer 2/3 visual cortical pyramidal neurons expressing GCaMP6f in an awake mouse. The data were recorded at 17.5 Hz and are played back at 6.8× real-speed (30 fps playback), with each displayed video frame equaling the average of four image acquisitions. During pre-processing, we stitched together the 16 images tiles, corrected for brain motion artifacts via image registration, and adjusted the contrast to highlight the details (Methods).
41586_2020_2130_MOESM7_ESM.mov
Video 3: Large-scale Ca2+ dynamics of layer 5 neocortical pyramidal cells in an awake mouse imaged with the 16-beam microscope. A two-photon Ca2+ video acquired 500 μm deep below the cortical surface in a transgenic mouse (tetO-GCaMP6s/CaMK2a-tTA) expressing GCaMP6s in a subset of layer 5 cortical pyramidal cells38. The greater Ca2+ affinity and fluorescence output of GCaMP6s as compared to GCaMP6f enabled us to image layer 5 cells using the same total illumination power as the maximum value (320 mW) used elsewhere in the paper for studying layer 2/3 neurons expressing GCaMP6f. The data were recorded at 17.5 Hz, processed in the same way as Supplementary Video 2, and played back at 6.8× real-speed (30 fps playback). The field-of-view is 2 mm × 2 mm.
Rights and permissions
About this article
Cite this article
Rumyantsev, O.I., Lecoq, J.A., Hernandez, O. et al. Fundamental bounds on the fidelity of sensory cortical coding. Nature 580, 100–105 (2020). https://doi.org/10.1038/s41586-020-2130-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-020-2130-2
This article is cited by
-
Large-scale cranial window for in vivo mouse brain imaging utilizing fluoropolymer nanosheet and light-curable resin
Communications Biology (2024)
-
Retinal spike train decoder using vector quantization for visual scene reconstruction
Complex & Intelligent Systems (2024)
-
Rapid detection of neurons in widefield calcium imaging datasets after training with synthetic data
Nature Methods (2023)
-
Macroscopic resting-state brain dynamics are best described by linear models
Nature Biomedical Engineering (2023)
-
Targeted V1 comodulation supports task-adaptive sensory decisions
Nature Communications (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.