Abstract
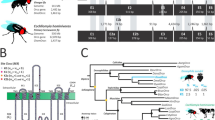
The evolution of animal behaviour is poorly understood1,2. Despite numerous correlations between interspecific divergence in behaviour and nervous system structure and function, demonstrations of the genetic basis of these behavioural differences remain rare3,4,5. Here we develop a neurogenetic model, Drosophila sechellia, a species that displays marked differences in behaviour compared to its close cousin Drosophila melanogaster6,7, which are linked to its extreme specialization on noni fruit (Morinda citrifolia)8,9,10,11,12,13,14,15,16. Using calcium imaging, we identify olfactory pathways in D. sechellia that detect volatiles emitted by the noni host. Our mutational analysis indicates roles for different olfactory receptors in long- and short-range attraction to noni, and our cross-species allele-transfer experiments demonstrate that the tuning of one of these receptors is important for species-specific host-seeking. We identify the molecular determinants of this functional change, and characterize their evolutionary origin and behavioural importance. We perform circuit tracing in the D. sechellia brain, and find that receptor adaptations are accompanied by increased sensory pooling onto interneurons as well as species-specific central projection patterns. This work reveals an accumulation of molecular, physiological and anatomical traits that are linked to behavioural divergence between species, and defines a model for investigating speciation and the evolution of the nervous system.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data, availability
All relevant data supporting the findings of this study are available from the corresponding authors upon request or, for behavioural experiments, are included with the paper as Source Data for Figs. 1b, c, 2c–e, 3d, f, Extended Data Figs. 1c, e, f, g, h, 7e–g. Supplementary Table 7 lists the exact n and mean values for all electrophysiological data.
Code availability
Code used for analyses and all unique biological materials generated in this study (for example, mutant and transgenic Drosophila strains) are available from the corresponding authors upon request.
References
Arguello, J. R. & Benton, R. Open questions: tackling Darwin’s “instincts”: the genetic basis of behavioral evolution. BMC Biol. 15, 26 (2017).
Bendesky, A. & Bargmann, C. I. Genetic contributions to behavioural diversity at the gene–environment interface. Nat. Rev. Genet. 12, 809–820 (2011).
Bendesky, A. et al. The genetic basis of parental care evolution in monogamous mice. Nature 544, 434–439 (2017).
Ding, Y., Berrocal, A., Morita, T., Longden, K. D. & Stern, D. L. Natural courtship song variation caused by an intronic retroelement in an ion channel gene. Nature 536, 329–332 (2016).
Weber, J. N., Peterson, B. K. & Hoekstra, H. E. Discrete genetic modules are responsible for complex burrow evolution in Peromyscus mice. Nature 493, 402–405 (2013).
Garrigan, D. et al. Genome sequencing reveals complex speciation in the Drosophila simulans clade. Genome Res. 22, 1499–1511 (2012).
Schrider, D. R., Ayroles, J., Matute, D. R. & Kern, A. D. Supervised machine learning reveals introgressed loci in the genomes of Drosophila simulans and D. sechellia. PLoS Genet. 14, e1007341 (2018).
Amlou, M., Moreteau, B. & David, J. R. Genetic analysis of Drosophila sechellia specialization: oviposition behavior toward the major aliphatic acids of its host plant. Behav. Genet. 28, 455–464 (1998).
Cobb, M., Burnet, B., Blizard, R. & Jallon, J. M. Courtship in Drosophila sechellia – its structure, functional aspects, and relationship to those of other members of the Drosophila melanogaster species subgroup. J. Insect Behav. 2, 63–89 (1989).
Coyne, J. A. Genetics of sexual isolation in females of the Drosophila simulans species complex. Genet. Res. 60, 25–31 (1992).
Dekker, T., Ibba, I., Siju, K. P., Stensmyr, M. C. & Hansson, B. S. Olfactory shifts parallel superspecialism for toxic fruit in Drosophila melanogaster sibling, D. sechellia. Curr. Biol. 16, 101–109 (2006).
Higa, I. & Fuyama, Y. Genetics of food preference in Drosophila sechellia. I. Responses to food attractants. Genetica 88, 129–136 (1993).
Ibba, I., Angioy, A. M., Hansson, B. S. & Dekker, T. Macroglomeruli for fruit odors change blend preference in Drosophila. Naturwissenschaften 97, 1059–1066 (2010).
Matsuo, T., Sugaya, S., Yasukawa, J., Aigaki, T. & Fuyama, Y. Odorant-binding proteins OBP57d and OBP57e affect taste perception and host-plant preference in Drosophila sechellia. PLoS Biol. 5, e118 (2007).
Prieto-Godino, L. L. et al. Evolution of acid-sensing olfactory circuits in Drosophilids. Neuron 93, 661–676 (2017).
R’Kha, S., Capy, P. & David, J. R. Host-plant specialization in the Drosophila melanogaster species complex: a physiological, behavioral, and genetical analysis. Proc. Natl Acad. Sci. USA 88, 1835–1839 (1991).
Yalcin, B. et al. Genetic dissection of a behavioral quantitative trait locus shows that Rgs2 modulates anxiety in mice. Nat. Genet. 36, 1197–1202 (2004).
Bendesky, A., Tsunozaki, M., Rockman, M. V., Kruglyak, L. & Bargmann, C. I. Catecholamine receptor polymorphisms affect decision-making in C. elegans. Nature 472, 313–318 (2011).
Bumbarger, D. J., Riebesell, M., Rödelsperger, C. & Sommer, R. J. System-wide rewiring underlies behavioral differences in predatory and bacterial-feeding nematodes. Cell 152, 109–119 (2013).
Markow, T. A. & O’Grady, P. Reproductive ecology of Drosophila. Funct. Ecol. 22, 747–759 (2008).
Linz, J. et al. Host plant-driven sensory specialization in Drosophila erecta. Proc. R. Soc. Lond. B 280, 20130626 (2013).
Seeholzer, L. F., Seppo, M., Stern, D. L. & Ruta, V. Evolution of a central neural circuit underlies Drosophila mate preferences. Nature 559, 564–569 (2018).
Dweck, H. K. et al. Olfactory channels associated with the Drosophila maxillary palp mediate short- and long-range attraction. eLife 5, e14925 (2016).
Matute, D. R. & Ayroles, J. F. Hybridization occurs between Drosophila simulans and D. sechellia in the Seychelles archipelago. J. Evol. Biol. 27, 1057–1068 (2014).
Vosshall, L. B. & Stocker, R. F. Molecular architecture of smell and taste in Drosophila. Annu. Rev. Neurosci. 30, 505–533 (2007).
Larsson, M. C. et al. Or83b encodes a broadly expressed odorant receptor essential for Drosophila olfaction. Neuron 43, 703–714 (2004).
Abuin, L. et al. Functional architecture of olfactory ionotropic glutamate receptors. Neuron 69, 44–60 (2011).
Benton, R., Sachse, S., Michnick, S. W. & Vosshall, L. B. Atypical membrane topology and heteromeric function of Drosophila odorant receptors in vivo. PLoS Biol. 4, e20 (2006).
Stensmyr, M. C., Dekker, T. & Hansson, B. S. Evolution of the olfactory code in the Drosophila melanogaster subgroup. Proc. R. Soc. Lond. B 270, 2333–2340 (2003).
Dobritsa, A. A., van der Goes van Naters, W., Warr, C. G., Steinbrecht, R. A. & Carlson, J. R. Integrating the molecular and cellular basis of odor coding in the Drosophila antenna. Neuron 37, 827–841 (2003).
Couto, A., Alenius, M. & Dickson, B. J. Molecular, anatomical, and functional organization of the Drosophila olfactory system. Curr. Biol. 15, 1535–1547 (2005).
Shiao, M. S. et al. Expression divergence of chemosensory genes between Drosophila sechellia and its sibling species and its implications for host shift. Genome Biol. Evol. 7, 2843–2858 (2015).
Yao, C. A., Ignell, R. & Carlson, J. R. Chemosensory coding by neurons in the coeloconic sensilla of the Drosophila antenna. J. Neurosci. 25, 8359–8367 (2005).
Ai, M. et al. Acid sensing by the Drosophila olfactory system. Nature 468, 691–695 (2010).
Ruta, V. et al. A dimorphic pheromone circuit in Drosophila from sensory input to descending output. Nature 468, 686–690 (2010).
Grabe, V. & Sachse, S. Fundamental principles of the olfactory code. Biosystems 164, 94–101 (2018).
Butterwick, J. A. et al. Cryo-EM structure of the insect olfactory receptor Orco. Nature 560, 447–452 (2018).
Aguadé, M. Nucleotide and copy-number polymorphism at the odorant receptor genes Or22a and Or22b in Drosophila melanogaster. Mol. Biol. Evol. 26, 61–70 (2009).
Goldman-Huertas, B. et al. Evolution of herbivory in Drosophilidae linked to loss of behaviors, antennal responses, odorant receptors, and ancestral diet. Proc. Natl Acad. Sci. USA 112, 3026–3031 (2015).
Nozawa, M. & Nei, M. Evolutionary dynamics of olfactory receptor genes in Drosophila species. Proc. Natl Acad. Sci. USA 104, 7122–7127 (2007).
Yassin, A. et al. Recurrent specialization on a toxic fruit in an island Drosophila population. Proc. Natl Acad. Sci. USA 113, 4771–4776 (2016).
Ambrose, D., Ellender, J. H., Lee, E. B., Sprake, C. H. S. & Townsend, R. Thermodynamic properties of organic oxygen compounds XXXVIII. Vapour pressures of some aliphatic ketones. J. Chem. Thermodyn. 7, 453–472 (1975).
Daubert, T. E. & Danner, R. P. Physical and Thermodynamic Properties of Pure Chemicals: Data Compilation (Taylor & Francis, 1997).
Port, F., Chen, H. M., Lee, T. & Bullock, S. L. Optimized CRISPR/Cas tools for efficient germline and somatic genome engineering in Drosophila. Proc. Natl Acad. Sci. USA 111, E2967–E2976 (2014).
Port, F. & Bullock, S. L. Augmenting CRISPR applications in Drosophila with tRNA-flanked sgRNAs. Nat. Methods 13, 852–854 (2016).
Barolo, S., Carver, L. A. & Posakony, J. W. GFP and β-galactosidase transformation vectors for promoter/enhancer analysis in Drosophila. Biotechniques 29, 726–732 (2000).
Gratz, S. J. et al. Highly specific and efficient CRISPR/Cas9-catalyzed homology-directed repair in Drosophila. Genetics 196, 961–971 (2014).
Stern, D. L. Tagmentation-based mapping (TagMap) of mobile DNA genomic insertion sites. Preprint at bioRxiv https://doi.org/10.1101/037762 (2017).
Croset, V. et al. Ancient protostome origin of chemosensory ionotropic glutamate receptors and the evolution of insect taste and olfaction. PLoS Genet. 6, e1001064 (2010).
Bateman, J. R. & Wu, C. T. A simple polymerase chain reaction-based method for the construction of recombinase-mediated cassette exchange donor vectors. Genetics 180, 1763–1766 (2008).
Groth, A. C., Olivares, E. C., Thyagarajan, B. & Calos, M. P. A phage integrase directs efficient site-specific integration in human cells. Proc. Natl Acad. Sci. USA 97, 5995–6000 (2000).
Bischof, J., Maeda, R. K., Hediger, M., Karch, F. & Basler, K. An optimized transgenesis system for Drosophila using germ-line-specific φC31 integrases. Proc. Natl Acad. Sci. USA 104, 3312–3317 (2007).
Han, C., Jan, L. Y. & Jan, Y. N. Enhancer-driven membrane markers for analysis of nonautonomous mechanisms reveal neuron–glia interactions in Drosophila. Proc. Natl Acad. Sci. USA 108, 9673–9678 (2011).
Arnoult, L. et al. Emergence and diversification of fly pigmentation through evolution of a gene regulatory module. Science 339, 1423–1426 (2013).
Gratz, S. J. et al. Genome engineering of Drosophila with the CRISPR RNA-guided Cas9 nuclease. Genetics 194, 1029–1035 (2013).
Zhang, X., Koolhaas, W. H. & Schnorrer, F. A versatile two-step CRISPR- and RMCE-based strategy for efficient genome engineering in Drosophila. G3 (Bethesda) 4, 2409–2418 (2014).
Gohl, D. M. et al. A versatile in vivo system for directed dissection of gene expression patterns. Nat. Methods 8, 231–237 (2011).
Knecht, Z. A. et al. Distinct combinations of variant ionotropic glutamate receptors mediate thermosensation and hygrosensation in Drosophila. eLife 5, e17879 (2016).
Silbering, A. F. et al. Complementary function and integrated wiring of the evolutionarily distinct Drosophila olfactory subsystems. J. Neurosci. 31, 13357–13375 (2011).
Schindelin, J. et al. Fiji: an open-source platform for biological-image analysis. Nat. Methods 9, 676–682 (2012).
Benton, R. & Dahanukar, A. Electrophysiological recording from Drosophila olfactory sensilla. Cold Spring Harb. Protoc. 2011, 824–838 (2011).
Saina, M. & Benton, R. Visualizing olfactory receptor expression and localization in Drosophila. Methods Mol. Biol. 1003, 211–228 (2013).
Prieto-Godino, L. L. et al. Olfactory receptor pseudo-pseudogenes. Nature 539, 93–97 (2016).
Benton, R., Vannice, K. S., Gomez-Diaz, C. & Vosshall, L. B. Variant ionotropic glutamate receptors as chemosensory receptors in Drosophila. Cell 136, 149–162 (2009).
Sánchez-Alcañiz, J. A., Zappia, G., Marion-Poll, F. & Benton, R. A mechanosensory receptor required for food texture detection in Drosophila. Nat. Commun. 8, 14192 (2017).
Ostrovsky, A., Cachero, S. & Jefferis, G. Clonal analysis of olfaction in Drosophila: immunochemistry and imaging of fly brains. Cold Spring Harb. Protoc. 2013, 342–346 (2013).
Jefferis, G. S. et al. Comprehensive maps of Drosophila higher olfactory centers: spatially segregated fruit and pheromone representation. Cell 128, 1187–1203 (2007).
Manton, J. D. et al. Combining genome-scale Drosophila 3D neuroanatomical data by bridging template brains. Preprint at bioRxiv https://doi.org/10.1101/006353 (2014).
Cachero, S., Ostrovsky, A. D., Yu, J. Y., Dickson, B. J. & Jefferis, G. S. Sexual dimorphism in the fly brain. Curr. Biol. 20, 1589–1601 (2010).
Feng, L., Zhao, T. & Kim, J. neuTube 1.0: a new design for efficient neuron reconstruction software based on the SWC format. eNeuro 2, ENEURO.0049-14.2014 (2015).
Caron, S. J., Ruta, V., Abbott, L. F. & Axel, R. Random convergence of olfactory inputs in the Drosophila mushroom body. Nature 497, 113–117 (2013).
Camacho, C. et al. BLAST+: architecture and applications. BMC Bioinformatics 10, 421 (2009).
Birney, E., Clamp, M. & Durbin, R. GeneWise and Genomewise. Genome Res. 14, 988–995 (2004).
Abascal, F., Zardoya, R. & Telford, M. J. TranslatorX: multiple alignment of nucleotide sequences guided by amino acid translations. Nucleic Acids Res. 38, W7–W13 (2010).
Ronquist, F. et al. MrBayes 3.2: efficient Bayesian phylogenetic inference and model choice across a large model space. Syst. Biol. 61, 539–542 (2012).
Yang, Z. PAML 4: phylogenetic analysis by maximum likelihood. Mol. Biol. Evol. 24, 1586–1591 (2007).
Signor, S. A., New, F. N. & Nuzhdin, S. A large panel of Drosophila simulans reveals an abundance of common variants. Genome Biol. Evol. 10, 189–206 (2018).
Grenier, J. K. et al. Global diversity lines – a five-continent reference panel of sequenced Drosophila melanogaster strains. G3 (Bethesda) 5, 593–603 (2015).
Huang, W. et al. Natural variation in genome architecture among 205 Drosophila melanogaster genetic reference panel lines. Genome Res. 24, 1193–1208 (2014).
Danecek, P. et al. The variant call format and VCFtools. Bioinformatics 27, 2156–2158 (2011).
Grabe, V. et al. Elucidating the neuronal architecture of olfactory glomeruli in the Drosophila antennal lobe. Cell Rep. 16, 3401–3413 (2016).
Pellegrino, M., Steinbach, N., Stensmyr, M. C., Hansson, B. S. & Vosshall, L. B. A natural polymorphism alters odour and DEET sensitivity in an insect odorant receptor. Nature 478, 511–514 (2011).
Adams, M. D. et al. The genome sequence of Drosophila melanogaster. Science 287, 2185–2195 (2000).
Acknowledgements
We thank Y. Bellaïche, B. Deplancke, K. S. Douglas, S. Lavista-Llanos, K. O’Connor-Giles, J. Posakony, V. Ruta, D. Stern, G. Suh, A. Yassin, the Bloomington Drosophila Stock Center (National Institute of Health (NIH) P40OD018537) and the Developmental Studies Hybridoma Bank (NICHD of the NIH, University of Iowa) for reagents, B. Prud’homme for instruction on Drosophila microinjections, B. Sutcliffe and S. Cachero for advice on the generation of reference brains, I. Alali and K. Weniger for technical assistance, J. Simpson for details of the nSyb promoter construct; P. C. Chai for assistance with glomerular identification; and I. Rentero Rebollo, J. Sánchez-Alcañiz, L. Prieto-Godino and members of the Benton laboratory for discussions and comments on the manuscript. T.O.A. is supported by a Human Frontier Science Program Long-Term Fellowship (LT000461/2015-L). M.A.K., B.S.H. and M.K. are supported by the Max Planck Society. K.E. and S.J.C.C. are supported by a NIH Award (1 R01 NS 167970) and a Eunice Kennedy Shriver National Institute of Child Health & Human Development Award of the NIH (5 T32 HD 007491). J.R.A. is supported by a Swiss National Science Professorship Grant (PP00P3 176956). G.S.X.E.J. is supported by an ERC Consolidator Grant (649111) and the MRC (MC_U105188491). Research in R.B.’s laboratory is supported by ERC Consolidator and Advanced Grants (615094 and 833548, respectively), the Swiss National Science Foundation and the Fondation Herbette.
Author information
Authors and Affiliations
Contributions
T.O.A. and R.B. conceived the project. All authors contributed to experimental design, analysis and interpretation of results. T.O.A. generated all molecular reagents, and new drosophilid mutants and transgenic lines. Other experimental contributions were as follows: T.O.A. contributed to experiments shown in Figs. 1c, 2a, b, d, e, 3a–c, e, 4a–d, g, Extended Data Figs. 1a–f, 2c, 3, 4c, e, 5, 6, 7a–f, 8, 9, 10a–d, 11f–m, 12; M.A.K., with input from B.S.H. and M.K., contributed to experiments shown in Figs. 1b, d, 2c, 3d, f, Extended Data Figs. 1f–h, 2a, b, d, e, 7g, Supplementary Table 1; G.Z. contributed to experiments shown in Figs. 1c, 2d, 4b, Extended Data Figs. 1a, b, 3b, 5c, e, g, 6, 7a, b, f, 10b, 11k; A.F.S. contributed to experiments shown in Fig. 1e, f, Extended Data Fig. 4d, f, g; K.E., with input from S.J.C.C., contributed to experiments shown in Fig. 4e–g, Extended Data Fig. 10e; R.Á.-O. contributed to experiments shown in Extended Data Fig. 4a, b; J.R.A. contributed to experiments shown in Extended Data Fig. 11d, e; G.S.X.E.J. contributed to experiments shown in Fig. 4c; and R.B. contributed to experiments shown in Fig. 4c. T.O.A. and R.B. wrote the paper with input from all other authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature thanks Lindy McBride and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Species-specific short- and long-range behavioural responses to diverse fruit stimuli.
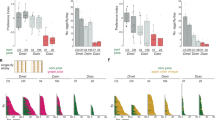
a, Data reproduced from Fig. 1c. Behavioural responses in a trap assay testing preferences between noni and grape, or between noni juice and grape juice. n = 15–27 experiments, 22–25 female flies per experiment. Comparisons to responses of Dsec.07 flies to noni juice are shown. In a–d, h (right), pairwise Wilcoxon rank-sum test and P values adjusted for multiple comparisons using the Benjamini and Hochberg method. b, Proportion of flies (mean ± s.e.m.) in each stimulus trap for the assays shown in a. Comparisons to responses of Dsec.07 flies are shown. c, Behavioural responses in a trap assay testing preferences between noni juice and diverse fruit juices or fruits for D. sechellia, D. simulans and D. melanogaster. n = 9–11 experiments, 22–25 female flies per experiment. Comparisons to responses of Dsec.07 flies are shown. d, Proportion of flies (mean ± s.e.m.) in each stimulus trap for the assays shown in c. Comparisons to responses of Dsec.07 flies are shown. e, Radar plot showing the mean percentage of flies per trap in a multiple-choice trap assay with eight different stimuli for D. sechellia, D. simulans and D. melanogaster. n = 11 experiments, 22–25 female flies per experiment. ACV, apple cider vinegar. f, Left, behavioural responses to noni juice in a wind tunnel assay of D. sechellia reared on standard food with (+) and without (−) noni supplement. Kruskal–Wallis test. Right, behavioural responses in a trap assay testing preferences between noni juice and grape juice for D. sechellia reared on standard food with (+) and without (−) noni supplement. Pairwise Wilcoxon rank-sum test. g, Behavioural responses to apple cider vinegar, mango juice, pineapple or fig in a wind tunnel assay of D. sechellia, D. simulans and D. melanogaster. n = 10–12 experiments, 10 female flies per experiment. Comparisons to responses of Dsec.07 flies are shown. In g, h (left), Kruskal–Wallis test with Dunn’s post hoc correction. h, Behavioural responses in a wind tunnel assay testing preference between noni fruit and apple cider vinegar of D. sechellia, D. simulans and D. melanogaster. n = 15 experiments, 10 female flies per experiment. Left, total number of flies landing on an odour source. Comparisons to responses of Dsec.07 flies are shown. Right, attraction index calculated as: (flies landing on apple cider vinegar − flies landing on noni)/flies landing on either source. Comparisons to responses of Dsec.07 flies are shown. NS, not significant (P > 0.05); *P < 0.05; ***P < 0.001.
Extended Data Fig. 2 Chemicals emitted by natural odour sources and odour bouquet changes in behavioural assays.
a, Principal constituents of the odour bouquet of noni fruit at different stages of ripening and commercial noni juice, as determined by gas chromatography–mass spectrometry. AU, arbitrary units. b, Chemical composition of the odour bouquet of noni fruit at different stages of ripening, noni juice, grape juice and apple cider vinegar. Representative gas chromatograms are shown on the right. Numbers correspond to compounds as listed in Supplementary Table 1 (not all identified peaks are shown). c, Dose-dependent electrophysiological responses of Or22a, Or85c/b and Ir75b neurons in Dsec.07 to their best odour agonists. Mean ± s.e.m. and individual data points, n = 7–20, female flies. The contribution of Or35a neurons (the spiking of which is difficult to separate from Ir75b neurons in ac3I) to hexanoic-acid responses is likely to be minimal (Fig. 2b). D. sechellia Or22a and Or85c/b neuron dose–response data are replotted from Fig. 3a. d, Chemical profile of odours collected by SPME at the release and landing platforms in the wind tunnel assay within the first 10 min of noni-juice application. e, Chemical profile of odours collected by SPME in the trap assay arena within 5 min of the placement of a trap (that is, t = 0 h), and after 5 h and 10 h, using noni juice as stimulus; 0% indicates that only trace proportions of the compound were detected.
Extended Data Fig. 3 OSN Gal4 driver lines in D. sechellia.
a, Schematic of the Gal4 reporter-allele generation strategy, through CRISPR–Cas9-mediated integration of an attP site (marked by 3×P3:DsRed) into the desired Or or Ir gene (Extended Data Figs. 5, 6 provide details of specific alleles), followed by introduction of a Gal4 ORF via φC31-mediated transgenesis. b, Coexpression of the indicated OrGal4-driven, IrGal4-driven or control background GCaMP6f signal (detected by anti-GFP) with the corresponding receptor protein or RNA in whole-mount antennae. Arrowheads point to examples of colabelled cells. Scale bars, 25 μm (main panels), 5 μm (insets). Whereas DsecOr22aGal4 and DsecIr64aGal4 flies completely recapitulate endogenous receptor expression, DsecOrcoGal4 and DsecOr85c/bGal4 flies lack expression in some receptor-expressing neurons. DsecOr35aGal4 and DsecIr75bGal4 might be expressed in ectopic cells (as shown in c, d) or the protein or RNA signal for these receptor genes could be below the detection threshold. c, Expression of the indicated OrGal4-driven, IrGal4-driven or control background GCaMP6f signal (detected by anti-GFP) in glomeruli of whole-mount antennal lobes. Three focal planes of the neuropil (visualized with nc82 (magenta)) are shown. Images were registered to a D. sechellia reference brain (Methods) for better comparison of antennal lobe structure. Scale bar, 25 μm. d, Summary of the glomerular labelling by OrGal4 or IrGal4 drivers as characterized in c (dark blue indicates GCaMP6f signal was detected in at least 3/3 independent brains). Glomeruli are organized by the compartmentalization of the corresponding OSN populations into different classes of sensilla (based on data in D. melanogaster81). ab, antennal basiconic; at, antennal trichoid; ai, antennal intermediate; ac, antennal coeloconic; pb, palp basiconic; sac, sacculus; ?, OSN population unknown. DsecOrcoGal4 is expressed in most—but not all (for example, Or67d DA1)—of the expected OSN populations; DsecOr35aGal4 and DsecOr85c/bGal4 display some ectopic expression, as inferred from their labelling of more than one glomerulus.
Extended Data Fig. 4 Comparative olfactory representations of noni in D. melanogaster and D. sechellia.
a, Representative odour-evoked calcium responses in the axon termini of Orco OSNs in the antennal lobes of D. melanogaster (Orco-Gal4/Orco-Gal4;UAS-GCaMP6f/UAS-GCaMP6f) and D. sechellia (UAS-GCaMP6f/UAS-GCaMP6f;;DsecOrcoGal4/+;), acquired by wide-field imaging. Left, raw fluorescence signals. Right, relative increase in GCaMP6f fluorescence (ΔF/F%) after stimulation with the indicated complex stimuli and single odours. Diagnostic odours: ethyl propionate (10−4 dilution (v/v)) for Or42b neurons (innervating DM1); methyl hexanoate (10−6 dilution) for Or22a or Or22a/b neurons (DM2); 2-heptanone (10−5 dilution) for Or85c/b neurons (VM5d); 2,3-butanedione (10−4 dilution) for Or92a neurons (VA2); and 1-hexanol (10−4 dilution) for Or35a neurons (VC3). Glomerular boundaries and the entire antennal lobe are outlined. Scale bars, 50 μm. b, Quantification of odour-evoked calcium responses for the flies represented in a. Maximum calcium-response amplitudes for each experiment are plotted. n = 5–8 female flies. Significantly different responses of species to the same stimulus are shown. Wilcoxon signed-rank test. c, Combined electrophysiological responses of neurons in the ab3 sensillum in D. melanogaster (top) and D. sechellia (bottom) upon stimulation with increasing concentrations of noni juice or noni fruit extract. Mean ± s.e.m. and individual data points; n = 6, female flies. Significant differences in responses are shown. Pairwise Wilcoxon rank-sum test. Responses of D. sechellia are stronger to noni fruit than noni juice, which may reflect the lower abundance of relevant ligands in the juice. d, Representative odour-evoked calcium responses in the axon termini of Orco OSNs in the antennal lobe of D. sechellia (UAS-GCaMP6f/UAS-GCaMP6f;;DsecOrcoGal4/+) acquired by two-photon imaging. Three focal planes are shown, revealing different glomeruli along the dorsoventral axis. Far left column, raw fluorescence images. Other columns show relative increase in GCaMP6f fluorescence (ΔF/F%) after stimulation with diagnostic odours. Scale bar, 25 μm. e, Electrophysiological responses in the antennal coeloconic (ac) sensilla classes to the indicated stimuli (n = 6–11, female flies) in D. sechellia (DSSC 14021-0248.07) representing the summed, solvent-corrected activities of the two or three neurons that they house. f, Representative odour-evoked calcium responses in the axon termini of Ir64a OSNs in the D. sechellia antennal lobe (UAS-GCaMP6f/UAS-GCaMP6f;;DsecIr64aGal4/+) acquired by wide-field imaging. Left, raw fluorescence signals. Right, relative increase in GCaMP6f fluorescence (ΔF/F%) after stimulation with noni juice (10−2 dilution) or grape juice. Scale bar, 25 μm. g, Quantification of odour-evoked calcium responses for the flies represented in f. Maximum calcium-response amplitudes for each experiment are plotted. n = 7–10 female flies. Comparisons of responses to noni (10−2 dilution) and grape juice are shown. Wilcoxon signed-rank test. *P < 0.05; ** P < 0.01.
Extended Data Fig. 5 Generation and validation of loss-of-function alleles of D. sechellia Or genes.
a, Schematic of the strategy for generating mutant alleles of Or genes, through integration of an eye-expressed fluorescent marker (3×P3:DsRed or 3×P3:GFPnls) into the desired locus via CRISPR–Cas9-cleavage induced homologous recombination. Brown triangles, loxP sites for removal of the fluorescent marker via Cre recombination. b, Schematics depicting Or gene organization, the structure of mutant alleles and the location of the sequences that encode antibody epitopes. For DsecOrco1, the fluorescent marker was integrated into the first coding exon; for DsecOrco2, the marker replaces parts of exons 1 and 3 and the whole of exon 2. DsecOr22aRFP carries the fluorescent marker in the first coding exon close to the start codon. DsecOr35aRFP lacks most of exons 1 and 2. For DsecOr85bGFP, the marker was integrated into exon 1; for DsecOr85c/bRFP, the marker replaces most of the Or85c gene and part of exon 1 of Or85b. c, Immunofluorescence for Orco and Ir25a (as an internal staining control) on whole-mount antennae from wild-type and DsecOrco2 flies. Scale bars, 25 μm (c–g, main panels), 5 μm (c–g, insets). d, RNA FISH for Or22a and Or85b on whole-mount antennae from wild-type, DsecOr22aRFP and DsecOr85bGFP mutant flies. e, Immunofluorescence for Ir75b and RNA FISH for Or35a on whole-mount antennae from wild-type and DsecOr35aRFP mutant flies. Arrowheads indicate Or35a-expressing cells. Or35a neurons also pair with Ir75c neurons in ac3II sensilla15, which is reflected in Or35a-positive cells that are not paired with Ir75b-expressing cells in wild-type antennae. f, Immunofluorescence for Or22a on whole-mount antennae from wild-type and DsecOr22aRFP mutant flies. Arrowheads indicate sensilla that house Or22a neurons. g, Far left, immunofluorescence for Orco and Ir25a (as an internal staining control) on whole-mount antennae from wild-type (same image as shown in c) (top) and DsecOrco1 (bottom) flies. Middle and right, electrophysiological responses in the two neurons of the ab3 sensillum (Fig. 2a) to odours present in noni in wild-type D. sechellia and DsecOrco1 mutants (n = 5–20, female flies). Representative response traces to methyl hexanoate (10−6 dilution) and 2-heptanone (10−6 dilution) are shown. Data points represent the solvent-corrected activities per neuron. Responses of wild-type D. sechellia neurons are replotted from Fig. 2a. Even though Orco expression is undetectable by immunofluorescence, weak electrophysiological responses in ab3 sensilla (and other Orco-dependent sensilla (data not shown)) can be detected. These observations suggest that trace levels of functional Orco are produced from this allele, potentially through use of in-frame start codons downstream of the marker insertion site (as shown in h). h, Schematic depicting the location of the Orco start codon, the fluorescent-marker insertion site of the DsecOrco1 allele and downstream potential alternative in-frame start codons.
Extended Data Fig. 6 Generation and validation of loss-of-function alleles of D. sechellia Ir genes.
a, Schematics depicting organization of Ir genes, the structure of mutant alleles and the sequences that encode antibody epitopes. For DsecIr8aRFP or DsecIr8aGFP, the fluorescent marker was integrated into the first coding exon. For DsecIr64aRFP, the marker replaces parts of exon 2. DsecIr75bRFP lacks parts of exons 3 and 4, and for DsecIr75bGFP the marker was integrated into exon 3. For both alleles of Ir75b, the fluorophore was removed via Cre-mediated recombination to produce Ir75b1 and Ir75b2. b, Immunofluorescence for Ir64a and Ir8a (as an internal staining control) on whole-mount antennae from wild-type and DsecIr64aRFP mutant flies. Arrowheads indicate the Ir64a-neuron dendrites that innervate sensilla in the sacculus (sac). Scale bars, 25 μm (main panels), 5 μm (insets). c, Left, immunofluorescence for Ir8a and Ir25a (as an internal staining control) on whole-mount antennae from wild-type and DsecIr8aGFP flies (top and middle). Left, immunofluorescence for Ir75b and RNA FISH for Or35a on whole-mount antennae from DsecIr75b2 mutant flies (bottom). Scale bars, 25 μm (main panels), 5 μm (insets). Right, electrophysiological responses in the ac3I sensillum (neurons housed are indicated in the cartoon) to noni juice, grape juice and odours present in noni (n = 4–11, female flies) in wild-type D. sechellia and olfactory-receptor mutants with the Ir75b neuron affected (DsecIr8aGFP and DsecIr75b2). Data points represent the summed solvent-corrected activities of the sensillum. Responses of wild-type D. sechellia are replotted from Fig. 2b. The Or35a neuron exhibits residual responses to hexanoic acid in the DsecIr8a and DsecIr75b olfactory-receptor mutants (Fig. 2b). d, Left, immunofluorescence for Ir8a and Ir25a (as an internal staining control) on whole-mount antennae from wild-type (same image as shown in c) and DsecIr8aRFP and DsecIr8aGFP (same image as shown in c) flies. Scale bars, 25 μm (main panels), 5 μm (insets). Right, electrophysiological responses in the ac2 sensillum (neurons housed are indicated in the cartoon) to noni juice, grape juice and odours present in noni (n = 3–11, female flies) in wild-type D. sechellia and olfactory-receptor mutants in which the Ir75a neuron is affected (DsecIr8aRFP and DsecIr8aGFP). Data points represent the summed solvent-corrected neuronal activities of the sensillum. Responses of wild-type D. sechellia to noni and grape juice are as shown in Extended Data Fig. 4e. e, Immunofluorescence for Ir75b and RNA FISH for Or35a on whole-mount antennae from wild-type and DsecIr75b1 and DsecIr75b2 (same image as shown in c) flies. Scale bars, 25 μm (main panels), 5 μm (insets).
Extended Data Fig. 7 Genetic and chemical contributions that promote the attraction of D. sechellia to noni.
a, Data reproduced from Fig. 2d. Behavioural responses in a trap assay testing preference of the indicated genotypes for noni juice or grape juice. n = 13–25 experiments, 22–25 female flies per experiment. Comparisons to responses of Dsec.07 flies are shown. In a–d, f, pairwise Wilcoxon rank-sum test and P values adjusted for multiple comparisons using the Benjamini and Hochberg method. Red bars, no significant difference; salmon bars, significantly different responses of D. sechellia genotypes. b, Proportion of flies (mean ± s.e.m.) in each stimulus trap for the assays shown in a. Comparisons to responses of Dsec.07 flies are shown. c, Olfactory responses in a trap assay testing preferences between noni juice and grape juice of wild-type D. sechellia, DsecOrco1 Ir8aGFP double mutants and wild-type D. sechellia with the third antennal segments removed (antennaless). n = 9–15 experiments, 22–25 female or male flies (as indicated) per experiment. These data represent the same experiments shown in Fig. 2e, but attraction indices were calculated here taking only alive flies into account. The percentages of flies alive at the end of the assay are indicated below, revealing the high mortality rate of antennaless flies and DsecOrco1 Ir8aGFP double mutants (DsecOrco2 Ir8aGFP mutants appeared to be nonviable). Normally, trap assay experiments with >25% fly mortality were discarded (Methods). Comparisons to responses of Dsec.07 flies are shown. d, Proportion of flies (mean ± s.e.m.) in each stimulus trap for the assays shown in c. Comparisons to responses of Dsec.07 flies are shown. e, Behavioural responses in a trap assay testing preferences between noni fruit and grape juice of Dsec.07 and DsecOr22aRFP flies. Comparisons to responses of Dsec.07 flies are shown. Pairwise Wilcoxon rank-sum test. f, Behavioural responses in a trap assay testing preferences between grape juice and 10−2 dilutions of the indicated odours in grape juice of D. sechellia, DsecOr22aRFP, D. simulans and D. melanogaster. Comparisons to responses of Dsec.07 flies to methyl hexanoate are shown. g, Behavioural responses in a wind tunnel assay testing attraction of D. sechellia to three single noni odours (10−2 dilution in water), a mix of all three in the approximate proportions of ripe noni fruit (1:0.04:1, methyl hexanoate:2-heptanone:hexanoic acid) and noni juice. n = 10 experiments, 10 female flies per experiment. Comparisons to responses of Dsec.07 flies to noni juice are shown. Kruskal–Wallis test with Dunn’s post hoc correction. NS, not significant (P > 0.05); *P < 0.05; **P < 0.01; ***P < 0.001.
Extended Data Fig. 8 Odour-tuning properties of drosophilid Or85c/b and Or22a/b neurons, and genomic modifications of the Or22a/b loci.
a, Left, dose-dependent electrophysiological responses of Or85c/b neurons (ab3B) in Dsec.07 to 2-heptanone and 1-hexanol. Mean ± s.e.m. and individual data points; n = 11–20, female flies. Right, dose-dependent electrophysiological responses of Or22a neurons (ab3A) in Dsec.07 to methyl butanoate, methyl hexanoate and methyl octanoate. Mean ± s.e.m. and individual data points; n = 11–20, female flies. The dose–response curves for 2-heptanone and methyl hexanoate are replotted from Fig. 3a. b, Schematics depicting the arrangement of wild-type, mutant and rescue allele versions of DsecOr22a (top) and DmelOr22a/b (bottom). Asterisk, stop codon that prevents read-through from the endogenous Or22a ORF. c, Schematics depicting the arrangement of wild-type and mutant alleles of DsimOr22a and DsimOr22b. d, RNA FISH for Or22a on whole-mount antennae from wild-type D. simulans (DSSC 14021-0251.195 (Dsim.195)), DsimOr22aRFP and DsimOr22a/bRFP mutant flies. As Or22a shares 85% sequence similarity with Or22b, the Or22a probe hybridizes with transcripts from both genes. Arrowheads indicate Or22b-expressing cells in DsimOr22aRFP. Scale bar, 25 μm (main panels), 5 μm (insets). e, Electrophysiological responses of Or22a/b neurons to different esters in wild-type D. simulans and olfactory-receptor mutants (DsimOr22aRFP and DsimOr22a/bRFP). n = 6–10, female flies. Representative response traces to methyl hexanoate (10−6 dilution) and ethyl butanoate (10−2 dilution) are shown to the left. f, Heat maps of the data shown in e, together with the data of the DsimOr22aWT response profile when expressed in DsecOr22aRFP (replotted from Fig. 3c). The receptors expressed in the analysed neurons are listed to the right. Significant differences to responses of wild-type D. simulans are shown. Pairwise Wilcoxon rank-sum test and P values adjusted for multiple comparisons using the Benjamini and Hochberg method. The equivalent responses to ethyl butanoate of wild-type D. simulans and Or22a-mutant neurons (but complete loss in Or22a/b-mutant neurons) suggests that this odour is detected principally by Or22b. g, Box plots with individual data points of the electrophysiological data presented in Fig. 3b. h, Box plots with individual data points of the electrophysiological data presented in Fig. 3c. i, Box plots with individual data points of the electrophysiological data presented in Fig. 3e. NS, not significant (P > 0.05); *P < 0.05; **P < 0.01; ***P < 0.001.
Extended Data Fig. 9 Mapping of odour-specificity determinants of Or22a.
a, Electrophysiological responses of D. melanogaster Or22a/b-mutant neurons expressing DsecOr22aWT or DmelOr22aWT upon stimulation with increasing concentrations of noni fruit extract. Mean ± s.em. and individual data points; n = 9, female flies. Significantly different values are indicated. Pairwise Wilcoxon rank-sum test. b, Protein sequence alignment of Or22a orthologues of six species within the D. melanogaster species subgroup of drosophilids. Red shading, amino acid differences between D. melanogaster, D. sechellia, D. simulans and D. mauritiana that were analysed by mutagenesis in this study; blue shading, all other sequence differences. Arrowheads, chimaera breakpoints (for chimaeras analysed in c). Predicted transmembrane (TM) domains are indicated with grey lines (location as in a previous publication11). c, Electrophysiological responses to a panel of noni odours conferred by chimeric Or22a proteins encoded by transgenes integrated at the Or22a/b locus of D. melanogaster. n = 5–6, female flies. Schematics on the left indicate the relative proportions of D. sechellia (red) and D. melanogaster (dark grey) sequences in each chimaera (precise chimaera breakpoints are shown in b). Significant differences to responses of DmelOr22aWT-expressing neurons are shown. In c, d, g, pairwise Wilcoxon rank-sum test and P values adjusted for multiple comparisons using the Benjamini and Hochberg method. d, Electrophysiological responses of D. melanogaster Or22a/b-mutant neurons expressing different Or22a variants. n = 5–7, female flies. The location of each mutated residue is indicated in b. Data for responses of Or22a/b-mutant and DmelOr22aWT-expressing neurons are replotted from c. Significant differences to responses of DmelOr22aWT-expressing neurons are shown. e, Box plots with individual data points showing the same data as in c. f, Box plots with individual data points showing the same data as in d. g, Dose-dependent electrophysiological responses of D. sechellia Or22a neurons that express the indicated transgenes to ethyl or methyl hexanoate. Mean ± s.e.m. and individual data points; n = 10–11, female flies. Significant comparisons to the responses of neurons expressing the DmelOr22aWT (left) or the DsecOr22aWT (right) transgene are shown. NS, not significant (P > 0.05); *P < 0.05; **P < 0.01; ***P < 0.001.
Extended Data Fig. 10 Changes in the peripheral and central olfactory circuit in D. sechellia.
a, Quantification of the number of OSNs expressing Or22a/b in antennae of D. sechellia and D. melanogaster (data as shown in Fig. 4b), Or22a/b mutants in both species and rescue lines expressing DsecOr22aWT. n = 9–11, female flies. Comparisons of rescue and wild-type genotypes for each species are shown. Pairwise Wilcoxon rank-sum test. No significant differences in Or22a cell number were observed for different rescue transgenes (data not shown). b, Quantification of the number of OSNs expressing Or13a (ab6), Or98a (ab7) or Or35a (ac3I/II) in D. sechellia, D. simulans and D. melanogaster (n = 10–15, female flies). Comparisons to cell number counts in Dsec.07 flies are shown. Pairwise Wilcoxon rank-sum test and P values adjusted for multiple comparisons using the Benjamini and Hochberg method. c, Immunofluorescence with nc82 (neuropil), anti-Elav (neurons) and anti-GFP in a DsecnSyb-Gal4/UAS-C3PA-GFP transgenic line, which expresses photoactivatable GFP pan-neuronally. The schematic on the left indicates the region of image acquisition. An anterior section through the antennal lobe (AL) is shown to reveal the position of the labelled projection neuron (PN) somas (circled in the right panel). Scale bar, 25 μm. d, Electrophysiological responses of the Or22a neuron to odours present in noni in homozygous DsecOr22aGal4 (mutant) transgenic flies. n = 6, female flies. Data points represent the solvent-corrected activities. Representative response traces to methyl hexanoate (10−6 dilution) in wild-type and transgenic flies are shown on top. e, Tracing of axonal branches in the lateral horn of dye-filled DM2 projection neurons in wild-type D. sechellia and homozygous DsecOr22aGal4 mutant flies. Three representative samples are shown. The circles depict the position of the axonal branch specific to D. sechellia. Scale bar, 10 μm. Samples could not be discriminated by genotype when presented to six independent researchers blindly. NS, not significant (P > 0.05); **P < 0.01; ***P < 0.001.
Extended Data Fig. 11 Phylogenetic and functional analysis of odour-specificity determinants in Or22a.
a, Side view of the Orco monomer structure (determined by cryo-electron microscopy37); the approximate location of the plasma membrane is indicated. The location of the residues corresponding to the odour-specificity determinants of Or22a analysed in this study (on the basis of previously generated alignments37) are highlighted as spheres. b, Top view of a cross-section through the putative ligand-binding pocket of the Orco structure shown in a. c, Partial protein sequence alignment of Or22a and Or59b. The equivalent residue to D. melanogaster Or22a M93 in Or59b is V91, which exhibits intraspecific sequence variation that affects odour sensitivity82. d, Results of branch-based models of molecular evolution that tested for changes in the rates of protein evolution among Or22a and Or22b orthologues (Methods, Supplementary Table 8): the rate of protein changes within the Or22a and Or22b phylogenetic tree highlights dN/dS ratios (ω) that differ from the ‘background rate’ (ω = 0.1772). Most branches exhibited low ω, arguing for strong purifying selection to maintain protein function over much of the tree. The two ω values that are >1 indicate an excess of protein changes, consistent with positive selection. The branch leading to D. simulans and D. sechellia Or22a displays nearly equal rates of silent and replacement substitutions, consistent with relaxed constraint during this period. e, Allele frequencies within population datasets for D. melanogaster78,79, D. simulans7,77 and D. sechellia7 at the three sites of Or22a that were functionally characterized in this study. The table displays amino acid (aa) positions 45, 67 and 93 of Or22a and the frequencies at which variants within the corresponding codons are segregating (number of alleles with respective variant/number of alleles analysed). NA (not applicable) indicates that positions within the codon are invariant. Datasets analysed are referenced on the right. Selected Or22a variants from the Drosophila melanogaster Genetic Reference Panel (DGRP)79 were confirmed by sequencing (f) and Or22a-neuron physiology was analysed (g, h). f, Protein sequence alignment of Or22a orthologues of D. melanogaster83, three lines of the DGRP, D. mauritiana (DSSC 14021-0241.151 (Dmau.151)) and D. sechellia. Red shading, amino acid differences (compared to the other analysed sequences) that are shared by DGRP-303, DGRP-304 and D. mauritiana at position 59 and the key odour-specificity determinant at residue 93; blue shading, all other sequence differences. No line within the DGRP with a polymorphism only at position 93 was identified. g, Electrophysiological responses of the Or22a/b neuron to odours present in noni (n = 5–20, female flies) in the strains shown in f. The similarity between the response profiles of DGRP-303, DGRP-304 and D. mauritiana suggests that their only shared polymorphism (at position 59) modifies Or22a-response properties in these strains. Comparisons to responses of Dmel BER flies are shown. Pairwise Wilcoxon rank-sum test and P values adjusted for multiple comparisons using the Benjamini and Hochberg method. D. mauritiana and D. sechellia data are replotted from Fig. 3b. h, Box plots with individual data points showing the same data as in g. D. mauritiana and D. sechellia data are replotted from Extended Data Fig. 8g. i, Protein sequence alignment of Or22a orthologues of the noni-specialized D. yakuba mayottensis (Dyak may.)41 and three other strains of D. yakuba (DSSC 14021-0261.00 (Dyak.00), 14021-0261.40 (Dyak.40) and 14021-0261.49 (Dyak.49)). Blue shading, differences between these sequences. j, Collection sites of D. yakuba strains shown in i. k, Quantification of the number of OSNs that express Or22a/b in D. sechellia, D. simulans, D. melanogaster (data as shown in Fig. 4b) and D. yakuba. n = 10–12 female flies. Comparisons to cell number counts in Dsec.07 flies are shown. In k, l, pairwise Wilcoxon rank-sum test and P values adjusted for multiple comparisons using the Benjamini and Hochberg method. l, Electrophysiological responses to odours present in noni of the Or22a/b neurons in D. sechellia, D. melanogaster and D. yakuba (n = 5–20, female flies). Comparisons to responses of Dsec.07 flies are shown. D. sechellia, D. melanogaster and Dyak.00 data are replotted from Fig. 3b. m, Box plots with individual data points showing the same data as in l. D. sechellia, D. melanogaster and Dyak.00 data are replotted from Extended Data Fig. 8g. NS, not significant (P > 0.05); *P < 0.05; **P < 0.01; ***P < 0.001.
Extended Data Fig. 12 Protein sequence alignments of Or22a and Ir75b.
a, Or22a orthologues of strains used for behavioural assays as well as genome-sequenced strains (‘gen.’) of each species (version: D. sechellia (r1.3), D. simulans (r2.01) and D. melanogaster (r6.28)). Blue or red shading indicates differences between species or strains, respectively. Green boxes, residues tested in this study for their role in defining ester tuning specificity. b, Ir75b orthologues of same strains as shown in a. Blue or red shading indicates differences between species or strains, respectively. Black boxes, residues predicted to be located within the ligand-binding domain that contributes to odour tuning specificity63. The premature stop codon of the Canton-S strain (position 169, marked by an asterisk) does not impair receptor function, as shown in other strains63.
Supplementary information
Supplementary Tables
This file contains Supplementary Tables 2 – 6 and 8.
Supplementary Table
Supplementary Table 1: Gas Chromatography/Mass Spectrometry analysis of headspaces of noni fruit stages, fruit juices and apple cider vinegar. 69 compounds (3 replicates/sample; values represent mean ± standard deviation of peak area covered, ordered according to their retention times) were identified with high probability based on matches to the National Institute of Standards and Technology library. Compounds are named following International Union of Pure and Applied Chemistry standards (for a subset of chemicals, alternative common names used in figures are shown in parentheses) and colour-coded as follows: fruit-specific compounds (orange); juice-specific compounds (blue); apple cider vinegar-specific compounds (green). 18 compounds (asterisks) were newly identified in our study compared to84,85.
Supplementary Table
Supplementary Table 7: Spike counts for electrophysiological experiments.
Rights and permissions
About this article
Cite this article
Auer, T.O., Khallaf, M.A., Silbering, A.F. et al. Olfactory receptor and circuit evolution promote host specialization. Nature 579, 402–408 (2020). https://doi.org/10.1038/s41586-020-2073-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-020-2073-7
This article is cited by
-
Elevated ozone disrupts mating boundaries in drosophilid flies
Nature Communications (2024)
-
Odor-regulated oviposition behavior in an ecological specialist
Nature Communications (2023)
-
Flexible neural control of transition points within the egg-laying behavioral sequence in Drosophila
Nature Neuroscience (2023)
-
The power of Drosophila genetics in studying insect toxicology and chemical ecology
Crop Health (2023)
-
Olfactory navigation in arthropods
Journal of Comparative Physiology A (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.