Abstract
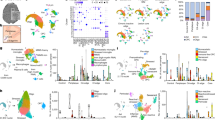
Multiple sclerosis is a chronic inflammatory disease of the CNS1. Astrocytes contribute to the pathogenesis of multiple sclerosis2, but little is known about the heterogeneity of astrocytes and its regulation. Here we report the analysis of astrocytes in multiple sclerosis and its preclinical model experimental autoimmune encephalomyelitis (EAE) by single-cell RNA sequencing in combination with cell-specific Ribotag RNA profiling, assay for transposase-accessible chromatin with sequencing (ATAC–seq), chromatin immunoprecipitation with sequencing (ChIP–seq), genome-wide analysis of DNA methylation and in vivo CRISPR–Cas9-based genetic perturbations. We identified astrocytes in EAE and multiple sclerosis that were characterized by decreased expression of NRF2 and increased expression of MAFG, which cooperates with MAT2α to promote DNA methylation and represses antioxidant and anti-inflammatory transcriptional programs. Granulocyte–macrophage colony-stimulating factor (GM-CSF) signalling in astrocytes drives the expression of MAFG and MAT2α and pro-inflammatory transcriptional modules, contributing to CNS pathology in EAE and, potentially, multiple sclerosis. Our results identify candidate therapeutic targets in multiple sclerosis.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
Sequencing data have been deposited into the Gene Expression Omnibus (GEO) under the SuperSeries accession number GSE130119. Clinical data for patient samples can be found in Supplementary Table 7. All other data and code that support the findings of this study are available from the corresponding author on reasonable request. Data from Schirmer et al.20 were accessed at PRJNA544731 and https://cells.ucsc.edu/?ds=ms. Data from Jäkel et al.17 were accessed at GSE118257. Data from Lake et al.40 were accessed at GSE97942.
References
Reich, D. S., Lucchinetti, C. F. & Calabresi, P. A. Multiple sclerosis. N. Engl. J. Med. 378, 169–180 (2018).
Baecher-Allan, C., Kaskow, B. J. & Weiner, H. L. Multiple sclerosis: mechanisms and immunotherapy. Neuron 97, 742–768 (2018).
Allen, N. J. & Lyons, D. A. Glia as architects of central nervous system formation and function. Science 362, 181–185 (2018).
Sofroniew, M. V. Astrocyte barriers to neurotoxic inflammation. Nat. Rev. Neurosci. 16, 249–263 (2015).
Colonna, M. & Butovsky, O. Microglia function in the central nervous system during health and neurodegeneration. Annu. Rev. Immunol. 35, 441–468 (2017).
Fontana, A., Fierz, W. & Wekerle, H. Astrocytes present myelin basic protein to encephalitogenic T-cell lines. Nature 307, 273–276 (1984).
Ousman, S. S. et al. Protective and therapeutic role for αB-crystallin in autoimmune demyelination. Nature 448, 474–479 (2007).
Sun, D. & Wekerle, H. Ia-restricted encephalitogenic T lymphocytes mediating EAE lyse autoantigen-presenting astrocytes. Nature 320, 70–72 (1986).
Wheeler, M. A. & Quintana, F. J. Regulation of astrocyte functions in multiple sclerosis. Cold Spring Harb. Perspect. Med. 9, a029009 (2019).
Liddelow, S. A. & Barres, B. A. Reactive astrocytes: production, function, and therapeutic potential. Immunity 46, 957–967 (2017).
Rothhammer, V. et al. Microglial control of astrocytes in response to microbial metabolites. Nature 557, 724–728 (2018).
Rothhammer, V. et al. Type I interferons and microbial metabolites of tryptophan modulate astrocyte activity and central nervous system inflammation via the aryl hydrocarbon receptor. Nat. Med. 22, 586–597 (2016).
Wheeler, M. A. et al. Environmental control of astrocyte pathogenic activities in CNS inflammation. Cell 176, 581–596.e18 (2019).
Giladi, A. & Amit, I. Single-cell genomics: a stepping stone for future immunology discoveries. Cell 172, 14–21 (2018).
Stubbington, M. J. T., Rozenblatt-Rosen, O., Regev, A. & Teichmann, S. A. Single-cell transcriptomics to explore the immune system in health and disease. Science 358, 58–63 (2017).
Tanay, A. & Regev, A. Scaling single-cell genomics from phenomenology to mechanism. Nature 541, 331–338 (2017).
Jäkel, S. et al. Altered human oligodendrocyte heterogeneity in multiple sclerosis. Nature 566, 543–547 (2019).
Jordão, M. J. C. et al. Single-cell profiling identifies myeloid cell subsets with distinct fates during neuroinflammation. Science 363, eaat7554 (2019).
Masuda, T. et al. Spatial and temporal heterogeneity of mouse and human microglia at single-cell resolution. Nature 566, 388–392 (2019).
Schirmer, L. et al. Neuronal vulnerability and multilineage diversity in multiple sclerosis. Nature 573, 75–82 (2019).
Falcão, A. M. Disease-specific oligodendrocyte lineage cells arise in multiple sclerosis. Nat. Med. 24, 1837–1844 (2018).
Ajami, B. et al. Single-cell mass cytometry reveals distinct populations of brain myeloid cells in mouse neuroinflammation and neurodegeneration models. Nat. Neurosci. 21, 541–551 (2018).
Itoh, N. et al. Cell-specific and region-specific transcriptomics in the multiple sclerosis model: focus on astrocytes. Proc. Natl Acad. Sci. USA 115, E302–E309 (2018).
Chao, C.-C. et al. Metabolic control of astrocyte pathogenic activity via cPLA2-MAVS. Cell 179, 1483–1498.e22 (2019).
Macosko, E. Z. et al. Highly parallel genome-wide expression profiling of individual cells using nanoliter droplets. Cell 161, 1202–1214 (2015).
Cuadrado, A. et al. Therapeutic targeting of the NRF2 and KEAP1 partnership in chronic diseases. Nat. Rev. Drug Discov. 18, 295–317 (2019).
Mayo, L. et al. Regulation of astrocyte activation by glycolipids drives chronic CNS inflammation. Nat. Med. 20, 1147–1156 (2014).
Katoh, Y. et al. Methionine adenosyltransferase II serves as a transcriptional corepressor of Maf oncoprotein. Mol. Cell 41, 554–566 (2011).
Lin, C. C. & Edelson, B. T. New insights into the role of IL-1β in experimental autoimmune encephalomyelitis and multiple sclerosis. J. Immunol. 198, 4553–4560 (2017).
Valentin-Torres, A. et al. Sustained TNF production by central nervous system infiltrating macrophages promotes progressive autoimmune encephalomyelitis. J. Neuroinflammation 13, 46 (2016).
Wardyn, J. D., Ponsford, A. H. & Sanderson, C. M. Dissecting molecular cross-talk between Nrf2 and NF-κB response pathways. Biochem. Soc. Trans. 43, 621–626 (2015).
Katsuoka, F. & Yamamoto, M. Small Maf proteins (MafF, MafG, MafK): history, structure and function. Gene 586, 197–205 (2016).
Shavit, J. A. et al. Impaired megakaryopoiesis and behavioral defects in mafG-null mutant mice. Genes Dev. 12, 2164–2174 (1998).
Raghunath, A. et al. Antioxidant response elements: discovery, classes, regulation and potential applications. Redox Biol. 17, 297–314 (2018).
Fang, M., Ou, J., Hutchinson, L. & Green, M. R. The BRAF oncoprotein functions through the transcriptional repressor MAFG to mediate the CpG Island Methylator phenotype. Mol. Cell 55, 904–915 (2014).
Codarri, L. et al. RORγt drives production of the cytokine GM-CSF in helper T cells, which is essential for the effector phase of autoimmune neuroinflammation. Nat. Immunol. 12, 560–567 (2011).
El-Behi, M. et al. The encephalitogenicity of T(H)17 cells is dependent on IL-1- and IL-23-induced production of the cytokine GM-CSF. Nat. Immunol. 12, 568–575 (2011).
Ponomarev, E. D. et al. GM-CSF production by autoreactive T cells is required for the activation of microglial cells and the onset of experimental autoimmune encephalomyelitis. J. Immunol. 178, 39–48 (2007).
Croxford, A. L. et al. The cytokine GM-CSF drives the inflammatory signature of CCR2+ monocytes and licenses autoimmunity. Immunity 43, 502–514 (2015).
Lake, B. B. et al. Integrative single-cell analysis of transcriptional and epigenetic states in the human adult brain. Nat. Biotechnol. 36, 70–80 (2018).
Liddelow, S. A. et al. Neurotoxic reactive astrocytes are induced by activated microglia. Nature 541, 481–487 (2017).
Apetoh, L. et al. The aryl hydrocarbon receptor interacts with c-Maf to promote the differentiation of type 1 regulatory T cells induced by IL-27. Nat. Immunol. 11, 854–861 (2010).
Matcovitch-Natan, O. et al. Microglia development follows a stepwise program to regulate brain homeostasis. Science 353, aad8670 (2016).
Xu, M. et al. c-MAF-dependent regulatory T cells mediate immunological tolerance to a gut pathobiont. Nature 554, 373–377 (2018).
Aziz, A., Soucie, E., Sarrazin, S. & Sieweke, M. H. MafB/c-Maf deficiency enables self-renewal of differentiated functional macrophages. Science 326, 867–871 (2009).
Soucie, E. L. et al. Lineage-specific enhancers activate self-renewal genes in macrophages and embryonic stem cells. Science 351, aad5510 (2016).
Gandhi, R. et al. Activation of the aryl hydrocarbon receptor induces human type 1 regulatory T cell-like and Foxp3+ regulatory T cells. Nat. Immunol. 11, 846–853 (2010).
Huynh, J. L. et al. Epigenome-wide differences in pathology-free regions of multiple sclerosis-affected brains. Nat. Neurosci. 17, 121–130 (2014).
Baranzini, S. E. et al. Genome, epigenome and RNA sequences of monozygotic twins discordant for multiple sclerosis. Nature 464, 1351–1356 (2010).
Wendeln, A. C. et al. Innate immune memory in the brain shapes neurological disease hallmarks. Nature 556, 332–338 (2018).
Ayata, P. et al. Epigenetic regulation of brain region-specific microglia clearance activity. Nat. Neurosci. 21, 1049–1060 (2018).
Garcia, A. D., Doan, N. B., Imura, T., Bush, T. G. & Sofroniew, M. V. GFAP-expressing progenitors are the principal source of constitutive neurogenesis in adult mouse forebrain. Nat. Neurosci. 7, 1233–1241 (2004).
Madisen, L. et al. A robust and high-throughput Cre reporting and characterization system for the whole mouse brain. Nat. Neurosci. 13, 133–140 (2010).
Sanz, E. et al. Cell-type-specific isolation of ribosome-associated mRNA from complex tissues. Proc. Natl Acad. Sci. USA 106, 13939–13944 (2009).
Srinivasan, R. et al. New transgenic mouse lines for selectively targeting astrocytes and studying calcium signals in astrocyte processes in situ and in vivo. Neuron 92, 1181–1195 (2016).
Anderson, M. A. et al. Astrocyte scar formation aids central nervous system axon regeneration. Nature 532, 195–200 (2016).
Jaitin, D. A. et al. Massively parallel single-cell RNA-seq for marker-free decomposition of tissues into cell types. Science 343, 776–779 (2014).
Feng, H., Zhang, X. & Zhang, C. mRIN for direct assessment of genome-wide and gene-specific mRNA integrity from large-scale RNA-sequencing data. Nat. Commun. 6, 7816 (2015).
Zhang, X. et al. Comparative analysis of droplet-based ultra-high-throughput single-cell rna-seq systems. Mol. Cell 73, 130–142.e135 (2019).
Butler, A., Hoffman, P., Smibert, P., Papalexi, E. & Satija, R. Integrating single-cell transcriptomic data across different conditions, technologies, and species. Nat. Biotechnol. 36, 411–420 (2018).
Stuart, T. et al. Comprehensive integration of single-cell data. Cell 177, 1888–1902.e21 (2019).
Finak, G. et al. MAST: a flexible statistical framework for assessing transcriptional changes and characterizing heterogeneity in single-cell RNA sequencing data. Genome Biol. 16, 278 (2015).
Becht, E. et al. Dimensionality reduction for visualizing single-cell data using UMAP. Nat. Biotechnol. 37, 38–44 (2018).
van der Maaten, L. & Hinton, G. Visualizing data using t-SNE. J. Mach. Learn. Res. 9, 2579–2605 (2008).
Qiu, X. et al. Single-cell mRNA quantification and differential analysis with Census. Nat. Methods 14, 309–315 (2017).
Qiu, X. et al. Reversed graph embedding resolves complex single-cell trajectories. Nat. Methods 14, 979–982 (2017).
Trapnell, C. et al. The dynamics and regulators of cell fate decisions are revealed by pseudotemporal ordering of single cells. Nat. Biotechnol. 32, 381–386 (2014).
Takenaka, M. C. et al. Control of tumor-associated macrophages and T cells in glioblastoma via AHR and CD39. Nat. Neurosci. 22, 729–740 (2019).
Sanjana, N. E., Shalem, O. & Zhang, F. Improved vectors and genome-wide libraries for CRISPR screening. Nat. Methods 11, 783–784 (2014).
Chen, S. et al. Genome-wide CRISPR screen in a mouse model of tumor growth and metastasis. Cell 160, 1246–1260 (2015).
Lee, Y., Messing, A., Su, M. & Brenner, M. GFAP promoter elements required for region-specific and astrocyte-specific expression. Glia 56, 481–493 (2008).
Wang, T. et al. Gene essentiality profiling reveals gene networks and synthetic lethal interactions with oncogenic Ras. Cell 168, 890–903.e815 (2017).
Rothhammer, V. et al. Sphingosine 1-phosphate receptor modulation suppresses pathogenic astrocyte activation and chronic progressive CNS inflammation. Proc. Natl Acad. Sci. USA 114, 2012–2017 (2017).
Yates, A. et al. Ensembl 2016. Nucleic Acids Res. 44, D710–D716 (2016).
Ovcharenko, I. et al. Mulan: multiple-sequence local alignment and visualization for studying function and evolution. Genome Res. 15, 184–194 (2005).
van Galen, P. et al. A multiplexed system for quantitative comparisons of chromatin landscapes. Mol. Cell 61, 170–180 (2016).
Untergasser, A. et al. Primer3—new capabilities and interfaces. Nucleic Acids Res. 40, e115 (2012).
Foo, L. C. Purification of rat and mouse astrocytes by immunopanning. Cold Spring Harb. Protoc. 2013, 421–432 (2013).
Esmonde-White, C. et al. Distinct function-related molecular profile of adult human A2B5+ pre-oligodendrocytes versus mature oligodendrocytes. J. Neuropathol. Exp. Neurol. 78, 468–479 (2019).
Zheng, G. X. et al. Massively parallel digital transcriptional profiling of single cells. Nat. Commun. 8, 14049 (2017).
Polman, C. H. et al. Diagnostic criteria for multiple sclerosis: 2010 revisions to the McDonald criteria. Ann. Neurol. 69, 292–302 (2011).
Dhaeze, T. et al. CD70 defines a subset of proinflammatory and CNS-pathogenic TH1/TH17 lymphocytes and is overexpressed in multiple sclerosis. Cell. Mol. Immunol. 16, 652–665 (2019).
Kuhlmann, T. et al. An updated histological classification system for multiple sclerosis lesions. Acta Neuropathol. 133, 13–24 (2017).
Moffitt, J. R. et al. High-throughput single-cell gene-expression profiling with multiplexed error-robust fluorescence in situ hybridization. Proc. Natl Acad. Sci. USA 113, 11046–11051 (2016).
Moffitt, J. R. et al. Molecular, spatial, and functional single-cell profiling of the hypothalamic preoptic region. Science 362, eaau5324 (2018).
Soumillon, M., Cacchiarelli, D., Semrau, S., van Oudenaarden, A. & Mikkelsen, T. S. Characterization of directed differentiation by high-throughput single-cell RNA-Seq. Preprint at https://www.bioRxiv.org/content/10.1101/003236v1 (2014).
Trombetta, J. J. et al. Preparation of single-cell RNA-seq libraries for next generation sequencing. Curr. Protoc. Mol. Biol. 107, 4.22.1–4.22.17 (2014).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Li, H. et al. The sequence alignment/map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
ENCODE Project Consortium. An integrated encyclopedia of DNA elements in the human genome. Nature 489, 57–74 (2012).
Zhang, Y. et al. Model-based analysis of ChIP-Seq (MACS). Genome Biol. 9, R137 (2008).
Buenrostro, J. D., Giresi, P. G., Zaba, L. C., Chang, H. Y. & Greenleaf, W. J. Transposition of native chromatin for fast and sensitive epigenomic profiling of open chromatin, DNA-binding proteins and nucleosome position. Nat. Methods 10, 1213–1218 (2013).
Buenrostro, J. D., Wu, B., Chang, H. Y. & Greenleaf, W. J. ATAC-seq: a method for assaying chromatin accessibility genome-wide. Curr. Protoc. Mol. Biol. 109, 21.29.1–21.29.9 (2015).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102, 15545–15550 (2005).
Mootha, V. K. et al. PGC-1α-responsive genes involved in oxidative phosphorylation are coordinately downregulated in human diabetes. Nat. Genet. 34, 267–273 (2003).
Chen, E. Y. et al. Enrichr: interactive and collaborative HTML5 gene list enrichment analysis tool. BMC Bioinformatics 14, 128 (2013).
Kuleshov, M. V. et al. Enrichr: a comprehensive gene set enrichment analysis web server 2016 update. Nucleic Acids Res. 44, W90–W97 (2016).
Mi, H., Muruganujan, A., Ebert, D., Huang, X. & Thomas, P. D. PANTHER version 14: more genomes, a new PANTHER GO-slim and improvements in enrichment analysis tools. Nucleic Acids Res. 47, D419–D426 (2019).
Acknowledgements
This work was supported by grants NS102807, NS087867, ES02530, AI126880, and AI093903 from the NIH; RSG-14-198-01-LIB from the American Cancer Society; and RG4111A1 and JF2161-A-5 from the National Multiple Sclerosis Society (to F.J.Q.) and the Canada Foundation for Innovation (CGEn and CFI-LOF 32557 to J.R.). F.J.Q., A.P. and J.P.A. received support from the International Progressive MS Alliance (PA-1604-08459). A.P. holds the T1 Canada Research Chair in Multiple Sclerosis and is funded by the Canada Institute of Health Research, the National Multiple Sclerosis Society, and Foundation of Canada. M.A.W. was supported by a training grant from the NIH and Dana-Farber Cancer Institute (T32CA207201), a postdoctoral fellowship from the NIH (F32NS101790), and a traveling neuroscience fellowship from the Program in Interdisciplinary Neuroscience at Brigham and Women’s Hospital. M.A.W. and I.C.C. received funding from the Women’s Brain Initiative at Brigham and Women’s Hospital. Sanger sequencing was carried out at the DNA Resource Core of Dana-Farber/Harvard Cancer Center (funded in part by NCI Cancer Center support grant 2P30CA006516- 48). We thank S. Jung and L. Chappell-Maor (Weizmann Institute of Science) for help in sequencing the Ribotag dataset; S. Young, T. Mason, and S. Garamszegi for assistance with coordinating deep sequencing; all members of the Quintana laboratory for helpful advice and discussions; P. Hewson for technical assistance; D. Kozoriz, A. Chicoine, and R. Krishnan for technical assistance with flow cytometry studies; Y. C. Wang for human control scRNA-seq library preparation; and the patients and their families that agreed to participate in this study.
Author information
Authors and Affiliations
Contributions
M.A.W. performed most in vitro, in vivo, and sequencing experiments. M.A.W., I.C.C., A.R., J.A.H., and S.T. assisted with and/or performed Drop-seq encapsulation experiments. M.A.W., E.C.T., and Z.L. performed bioinformatic analyses. S.E.J.Z. performed immunostaining of MS patient tissue. C.P.C. generated some human healthy control patient scRNA-seq data. B.R.W. performed multiplexed FISH experiments with assistance from M.A.W. G.S., S.A., V.R., and L.M.S. performed some in vitro, in vivo, or sequencing experiments. S.E.J.Z. and V.R. assisted with human sample isolation for scRNA-seq. J.R., D.A.W., K.P., J.R.M., B.B., J.P.A., and A.P. provided unique reagents and discussed and/or interpreted findings. M.A.W. and F.J.Q. wrote the manuscript with input from coauthors. F.J.Q. designed and supervised the study and edited the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Control analyses for scRNA-seq of B6 EAE mice.
a, Cell type marker expression in mice with EAE. n = 24,275 cells. b, Unsupervised clustering tSNE plots of CNS cells from mice with EAE. n = 6 per group, n = 4 priming, n = 3 CFA. n = 24,275 cells. c, Analysis of cluster occupation by cells across EAE time points. d, Significantly enriched genes by cell type cluster. e, PCs used in study. n = 24,275 cells. f, Cluster distribution by replicates. g, Principal componentsC used in astrocyte subclustering. n = 2,079 cells. h, Gene scatterplots of astrocyte markers in the astrocyte tSNE analysis. n = 2,079 cells. i, Astrocyte marker gene expression by time point during EAE. n = 2,079 cells. j, Correlation of NRF2 target gene expression during priming and peak EAE phases compared to in naive mice. NRF2 target genes are marked in red. In total, 88 out of 123 genes were decreased at at least one time point, whereas 40 out of 123 were decreased at both time points. n = 69 cells from cluster 4.
Extended Data Fig. 2 Sorting of TdTomatoGfap astrocytes.
The forward versus side scatter (FSC versus SSC) gating strategy, followed by exclusion of FSC and SSC doublets, and TdTomato fluorescence in the phycoerythrin (PE) channel.
Extended Data Fig. 3 Expression of Mafg and Nfe2l2 in astrocytes.
a, EAE in TdTomatoGfap mice used for scRNA-seq. n = 4 mice per time point. b, Cluster composition by replicate. c, Cluster composition by EAE time point. d, Unsupervised clustering tSNE plot of TdTomatoGfap astrocytes from mice with EAE. n = 4 per group (peak, day 20 post-induction; remission, day 42 post-induction). e, Scatterplots of astrocyte markers. f, Scatterplots of genes of interest in this study. g, Principal components used in this analysis. n = 24,963 cells for all scRNA-seq experiments.
Extended Data Fig. 4 Nfe2l2 knockdown in astrocytes.
a, RNA-seq analysis of astrocytes following intracerebroventricular injection of IL-1β/TNF, EAE induction, or no treatment. n = 3 per group, n = 2 naive. b, siRNA-based knockdown of Nfe2l2 in primary astrocytes. n = 6 biologically independent samples per condition. Experiment repeated twice. Unpaired two-tailed t-test. c, Nfe2l2 expression determined by qPCR in flow cytometry-sorted astrocytes. n = 3 mice per group. One-sample t-test. d, Flow cytometry sorting strategy for astrocytes, microglia and pro-inflammatory monocytes. e, Quantification of astrocytes, microglia and pro-inflammatory monocytes. n = 3 mice per condition. One-way ANOVA, Tukey post-test. Data shown as mean ± s.e.m. ***P < 0.001, NS (not significant) P > 0.05.
Extended Data Fig. 5 Analysis of Nfe2l2–Mafg signature by Ribotag and scRNA-seq.
a, EAE in mice used in RibotagGfap studies. n = 3 mice per time point. b, RINs for Ribotag preparation. IP, immunoprecipitated HA-tagged ribosomes. mRNA direct, enrichment of polyadenylated mRNA using mRNA direct kit (Thermo Fisher, #61011). n = 4 biologically independent samples per condition. c, K-means clustering of RibotagGfap RNA-seq data for five CNS regions. d, ENRICHR analysis of upregulated genes in EAE (top). Analysis of gene expression associated with the altered glutathione metabolism KEGG pathway by CNS region (bottom). Number of independent mouse samples studied: n = 7 cortex, n = 8 spinal cord, n = 7 parenchyma, n = 9 cerebellum, n = 8 cranial nerves. e, Gene expression scatterplots of genes of interest in B6 EAE scRNA-seq studies. n = 24,275 cells. Data shown as mean ± s.e.m.
Extended Data Fig. 6 Mafg knockdown in astrocytes.
a, Quantification of GFAP immunoreactivity in CNS samples from naive or EAE mice. Cortex: n = 8 naive, n = 9 EAE; spinal cord: n = 13 naive, n = 9 EAE. Kolmogorov–Smirnov test. b, Immunostaining (left) and quantification (right) of MAFG+ GFAP+ astrocytes in mice targeted with sgMafg-delivering or sgScrmbl-delivering lentiviruses. n = 6 images per group from n = 3 mice. Two-tailed Mann–Whitney test. c, T cell subsets, astrocytes, microglia and pro-inflammatory monocytes in sgMafg-targeted versus sgScrmbl-targeted mice. n = 3 per condition for T cells; n = 2 for sgScrmbl IL-10+ group. Unpaired two-tailed t-test. n = 6 per condition for astrocytes, microglia, and monocytes. Experiment repeated twice. Unpaired two-tailed t-test. Data shown as mean ± s.e.m.
Extended Data Fig. 7 Mat2a and Csf2rb knockdown in astrocytes.
a, Validation of Mat2a knockdown by western blot. n = 3 biologically independent samples per group. Single sample t-test. b, Quantification of Mafg expression by scRNA-seq in Csf2rb conditional knockout mice. n = 3 per condition. Unpaired two-tailed t-test. c, Quantification of cell populations in Csf2rb conditional knockout mice. n = 3 per condition. Unpaired two-tailed t-test. d, Large area scan of down-sampled stitched multiplexed FISH images. Arrowheads indicate T cells. Representative images from three independent experiments with n = 3 mice per group. Data shown as mean ± s.e.m.
Extended Data Fig. 8 Control analyses of scRNA-seq on human samples.
a, tSNE plots of MS and control cells. b, Cluster occupation by disease state and patient. c, RINs and scRNA-seq data quality. n = 9 per analysis. Pearson’s correlation. d, Age and sex corresponding to samples analysed. n = 5 control, n = 4 MS. Unpaired two-tailed t-test (age), Fisher’s exact test (sex). e, Principal components used in this analysis. f, Expression scatterplots of genes of interest. n = 43,670 cells. g, Cell type classification based on significantly enriched genes by cluster. h, Cell type marker scatterplots. n = 43,670 cells. Data shown as mean ± s.e.m.
Extended Data Fig. 9 Validation of GM-CSF–MAFG–NRF2 transcriptional signature in multiple human scRNA-seq datasets.
a, Analysis of regionally matched cortical astrocytes derived from patients with MS and control individuals analysed by Schirmer et al.20. n = 9 controls, n = 12 patients. b, Analysis of regionally matched white matter cortical astrocytes derived from patients with MS and control individuals analysed by Jäkel et al.17. n = 5 controls, n = 4 patients. c, Analysis of regionally matched cerebellar astrocytes derived from patients with MS analysed in this study and control individuals analysed by Lake et al.40. n = 9 controls, n = 2 patients.
Extended Data Fig. 10 Control analyses of human astrocytes.
a, Gene scatterplots for astrocyte-specific markers in dataset representing control individuals and patients with MS compiled using data from Schirmer et al.20, Jäkel et al.17, Lake et al.40, and this study. n = 28 controls, n = 20 patients. n = 9,673 cells. b, Principal component analysis of all cells and astrocytes in each study. Number of cells analysed: Schirmer et al.20: 48,919 (all), 5,831 (astrocytes); Jäkel et al.17: 17,799 (all), 1,422 (astrocytes); this study: 43,670 cells (all), 2,332 (astrocytes). c, Principal components used in this study. d, Fraction occupation by cluster by patient. e, tSNE plot by condition and study (n = 9,673 cells). f, Analysis of IL-1β/TNF signalling in cluster 1 astrocytes from n = 28 controls and n = 20 patients with MS from the four studies. g, Canonical correlation analysis of mouse (Fig. 1, B6 EAE) and human (Fig. 6) astrocyte clusters and IPA analysis.
Supplementary information
Supplementary Table
Supplementary Table 1: Cluster marker analysis in B6 EAE scRNA-seq. n=24,275 cells. Statistics by Benjamini-Hochberg test (adjusted p-value FDR) and MAST (p-value).
Supplementary Table
Supplementary Table 2: Differential gene expression by cluster of astrocytes during EAE in WT B6 mice by scRNA-seq. n=2,079 cells. Statistics by Benjamini-Hochberg test (adjusted p-value FDR) and MAST (p-value).
Supplementary Table
Supplementary Table 3: Differential gene expression by cluster of astrocytes during EAE in TdTomatoGfap mice by scRNA-seq. n=24,963 cells. Statistics by Benjamini-Hochberg test (adjusted p-value FDR) and MAST (p-value).
Supplementary Table
Supplementary Table 4: K-means clustering and differential gene expression of RNA-seq of RibotagGfap mice in EAE.
Supplementary Table
Supplementary Table 5: Differential gene expression of sgScrmbl- versus sgMafg-targeted astrocytes during EAE. n=3 mice per group. Statistics by Wald test (p-value).
Supplementary Table
Supplementary Table 6: Differential gene expression of Csf2rbAldh1l1-creERT2 versus Csf2rbf/f mice during EAE. n=3 mice per group. Statistics by Wald test (p-value).
Supplementary Table
Supplementary Table 7: Clinical information of samples analyzed.
Supplementary Table
Supplementary Table 8: Cluster marker analysis of human CNS samples by scRNA-seq from this study. n=43,670 cells. Statistics by Benjamini-Hochberg test (adjusted p-value FDR) and MAST (p-value).
Supplementary Table
Supplementary Table 9: Differential gene expression by cluster of cortical astrocytes in MS versus healthy control samples in the Schirmer et al. dataset. n=5,381 cells. Statistics by Benjamini-Hochberg test (adjusted p-value FDR) and MAST (p-value).
Supplementary Table
Supplementary Table 10: Differential gene expression by cluster of cortical white matter astrocytes in MS versus control samples in the Jäkel et al. dataset. n=1,422 cells. Statistics by Benjamini-Hochberg test (adjusted p-value FDR) and MAST (p-value).
Supplementary Table
Supplementary Table 11: Differential gene expression by cluster of cerebellar astrocytes in MS versus control samples in this study and Lake et al. n=823 cells. Statistics by Benjamini-Hochberg test (adjusted p-value FDR) and MAST (p-value).
Supplementary Table
Supplementary Table 12: Differential gene expression by cluster of astrocytes in MS versus control samples from Schirmer et al., Jäkel et al., Lake et al., and this study. n=9,673 cells. Statistics by Benjamini-Hochberg test (adjusted p-value FDR) and MAST (p-value).
Source data
Rights and permissions
About this article
Cite this article
Wheeler, M.A., Clark, I.C., Tjon, E.C. et al. MAFG-driven astrocytes promote CNS inflammation. Nature 578, 593–599 (2020). https://doi.org/10.1038/s41586-020-1999-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-020-1999-0
This article is cited by
-
Drug targeting in psychiatric disorders — how to overcome the loss in translation?
Nature Reviews Drug Discovery (2024)
-
A phenotypic screening platform for identifying chemical modulators of astrocyte reactivity
Nature Neuroscience (2024)
-
Potential of Nano-Engineered Stem Cells in the Treatment of Multiple Sclerosis: A Comprehensive Review
Cellular and Molecular Neurobiology (2024)
-
Disease-associated astrocyte epigenetic memory promotes CNS pathology
Nature (2024)
-
The astrocyte-produced growth factor HB-EGF limits autoimmune CNS pathology
Nature Immunology (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.