Abstract
The proteasome is a major proteolytic machine that regulates cellular proteostasis through selective degradation of ubiquitylated proteins1,2. A number of ubiquitin-related molecules have recently been found to be involved in the regulation of biomolecular condensates or membraneless organelles, which arise by liquid–liquid phase separation of specific biomolecules, including stress granules, nuclear speckles and autophagosomes3,4,5,6,7,8, but it remains unclear whether the proteasome also participates in such regulation. Here we reveal that proteasome-containing nuclear foci form under acute hyperosmotic stress. These foci are transient structures that contain ubiquitylated proteins, p97 (also known as valosin-containing protein (VCP)) and multiple proteasome-interacting proteins, which collectively constitute a proteolytic centre. The major substrates for degradation by these foci were ribosomal proteins that failed to properly assemble. Notably, the proteasome foci exhibited properties of liquid droplets. RAD23B, a substrate-shuttling factor for the proteasome, and ubiquitylated proteins were necessary for formation of proteasome foci. In mechanistic terms, a liquid–liquid phase separation was triggered by multivalent interactions of two ubiquitin-associated domains of RAD23B and ubiquitin chains consisting of four or more ubiquitin molecules. Collectively, our results suggest that ubiquitin-chain-dependent phase separation induces the formation of a nuclear proteolytic compartment that promotes proteasomal degradation.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
Raw cryo-ET data have been deposited to the Electron Microscopy Data Bank under accession code EMD-10494. The mass spectrometry proteomics data have been deposited to the ProteomeXchange Consortium via the PRIDE partner repository with the data set identifier PXD01637 and PXD016369. The uncropped blots and gels are provided in Supplementary Fig. 1. Source Data for Figs. 1–4 and Extended Data Figs. 1, 3–9 are provided with the paper.
Code availability
K2Align code is available at GitHub (https://github.com/dtegunov/k2align).
References
Finley, D. Recognition and processing of ubiquitin-protein conjugates by the proteasome. Annu. Rev. Biochem. 78, 477–513 (2009).
Livneh, I., Cohen-Kaplan, V., Cohen-Rosenzweig, C., Avni, N. & Ciechanover, A. The life cycle of the 26S proteasome: from birth, through regulation and function, and onto its death. Cell Res. 26, 869–885 (2016).
Shin, Y. & Brangwynne, C. P. Liquid phase condensation in cell physiology and disease. Science 357, eaaf4382 (2017).
Banani, S. F., Lee, H. O., Hyman, A. A. & Rosen, M. K. Biomolecular condensates: organizers of cellular biochemistry. Nat. Rev. Mol. Cell Biol. 18, 285–298 (2017).
Bouchard, J.J. et al. Cancer mutations of the tumor suppressor SPOP disrupt the formation of active, phase-separated compartments. Mol. Cell 72, 19–36 (2018).
Turakhiya, A. et al. ZFAND1 recruits p97 and the 26S proteasome to promote the clearance of arsenite-induced stress granules. Mol. Cell 70, 906–919 (2018).
Dao, T.P. et al. Ubiquitin modulates liquid-liquid phase separation of UBQLN2 via disruption of multivalent interactions. Mol. Cell 69, 965–978 (2018).
Sun, D., Wu, R., Zheng, J., Li, P. & Yu, L. Polyubiquitin chain-induced p62 phase separation drives autophagic cargo segregation. Cell Res. 28, 405–415 (2018).
Wójcik, C. & DeMartino, G. N. Intracellular localization of proteasomes. Int. J. Biochem. Cell Biol. 35, 579–589 (2003).
Enenkel, C. Proteasome dynamics. Biochim. Biophys. Acta 1843, 39–46 (2014).
Umpierrez, G. & Korytkowski, M. Diabetic emergencies—ketoacidosis, hyperglycaemic hyperosmolar state and hypoglycaemia. Nat. Rev. Endocrinol. 12, 222–232 (2016).
Janer, A. et al. PML clastosomes prevent nuclear accumulation of mutant ataxin-7 and other polyglutamine proteins. J. Cell Biol. 174, 65–76 (2006).
Cioce, M., Boulon, S., Matera, A. G. & Lamond, A. I. UV-induced fragmentation of Cajal bodies. J. Cell Biol. 175, 401–413 (2006).
Levy-Barda, A. et al. Involvement of the nuclear proteasome activator PA28γ in the cellular response to DNA double-strand breaks. Cell Cycle 10, 4300–4310 (2011).
Bohnsack, K. E. & Bohnsack, M. T. Uncovering the assembly pathway of human ribosomes and its emerging links to disease. EMBO J. 38, e100278 (2019).
Lam, Y. W., Lamond, A. I., Mann, M. & Andersen, J. S. Analysis of nucleolar protein dynamics reveals the nuclear degradation of ribosomal proteins. Curr. Biol. 17, 749–760 (2007).
Sung, M. K. et al. A conserved quality-control pathway that mediates degradation of unassembled ribosomal proteins. eLife 5, e19105 (2016).
Nguyen, A. T. et al. UBE2O remodels the proteome during terminal erythroid differentiation. Science 357, eaan0218 (2017).
Yanagitani, K., Juszkiewicz, S. & Hegde, R. S. UBE2O is a quality control factor for orphans of multiprotein complexes. Science 357, 472–475 (2017).
Kroschwald, S. et al. Promiscuous interactions and protein disaggregases determine the material state of stress-inducible RNP granules. eLife 4, e06807 (2015).
Jain, A. & Vale, R. D. RNA phase transitions in repeat expansion disorders. Nature 546, 243–247 (2017).
Jacobson, A. D., MacFadden, A., Wu, Z., Peng, J. & Liu, C. W. Autoregulation of the 26S proteasome by in situ ubiquitination. Mol. Biol. Cell 25, 1824–1835 (2014).
Yokoi, M. & Hanaoka, F. Two mammalian homologs of yeast Rad23, HR23A and HR23B, as multifunctional proteins. Gene 597, 1–9 (2017).
Walters, K. J., Lech, P. J., Goh, A. M., Wang, Q. & Howley, P. M. DNA-repair protein hHR23a alters its protein structure upon binding proteasomal subunit S5a. Proc. Natl Acad. Sci. USA 100, 12694–12699 (2003).
Nathan, J. A., Kim, H. T., Ting, L., Gygi, S. P. & Goldberg, A. L. Why do cellular proteins linked to K63-polyubiquitin chains not associate with proteasomes? EMBO J. 32, 552–565 (2013).
Kulak, N. A., Pichler, G., Paron, I., Nagaraj, N. & Mann, M. Minimal, encapsulated proteomic-sample processing applied to copy-number estimation in eukaryotic cells. Nat. Methods 11, 319–324 (2014).
Kristariyanto, Y. A. et al. K29-selective ubiquitin binding domain reveals structural basis of specificity and heterotypic nature of K29 polyubiquitin. Mol. Cell 58, 83–94 (2015).
Liu, Y., Deisenroth, C. & Zhang, Y. RP–MDM2–p53 pathway: linking ribosomal biogenesis and tumor surveillance. Trends Cancer 2, 191–204 (2016).
Dikic, I. Proteasomal and autophagic degradation systems. Annu. Rev. Biochem. 86, 193–224 (2017).
Klaips, C. L., Jayaraj, G. G. & Hartl, F. U. Pathways of cellular proteostasis in aging and disease. J. Cell Biol. 217, 51–63 (2018).
Berkers, C. R. et al. Profiling proteasome activity in tissue with fluorescent probes. Mol. Pharm. 4, 739–748 (2007).
Kroschwald, S., Maharana, S. & Simon, A. Hexanediol: a chemical probe to investigate the material properties of membrane-less compartments. Matters https://doi.org/10.19185/matters.201702000010 (2017).
D’Arcy, P. et al. Inhibition of proteasome deubiquitinating activity as a new cancer therapy. Nat. Med. 17, 1636–1640 (2011).
Hyer, M. L. et al. A small-molecule inhibitor of the ubiquitin activating enzyme for cancer treatment. Nat. Med. 24, 186–193 (2018).
Magnaghi, P. et al. Covalent and allosteric inhibitors of the ATPase VCP/p97 induce cancer cell death. Nat. Chem. Biol. 9, 548–556 (2013).
Sakuma, T. et al. Repeating pattern of non-RVD variations in DNA-binding modules enhances TALEN activity. Sci. Rep. 3, 3379 (2013).
Sanjana, N. E. et al. A transcription activator-like effector toolbox for genome engineering. Nat. Protoc. 7, 171–192 (2012).
Ran, F. A. et al. Genome engineering using the CRISPR–Cas9 system. Nat. Protoc. 8, 2281–2308 (2013).
Kumatori, A. et al. Abnormally high expression of proteasomes in human leukemic cells. Proc. Natl Acad. Sci. USA 87, 7071–7075 (1990).
Paul-Gilloteaux, P. et al. eC-CLEM: flexible multidimensional registration software for correlative microscopies. Nat. Methods 14, 102–103 (2017).
Tsuchiya, H. et al. In vivo ubiquitin linkage-type analysis reveals that the Cdc48-Rad23/Dsk2 axis contributes to K48-linked chain specificity of the proteasome. Mol. Cell 66, 488–502 (2017).
Fujioka, A. et al. Dynamics of the Ras/ERK MAPK cascade as monitored by fluorescent probes. J. Biol. Chem. 281, 8917–8926 (2006).
Udeshi, N. D., Mertins, P., Svinkina, T. & Carr, S. A. Large-scale identification of ubiquitination sites by mass spectrometry. Nat. Protoc. 8, 1950–1960 (2013).
Yoshida, Y. et al. A comprehensive method for detecting ubiquitinated substrates using TR-TUBE. Proc. Natl Acad. Sci. USA 112, 4630–4635 (2015).
Guo, Q. et al. In situ structure of neuronal C9orf72 Poly-GA aggregates reveals proteasome recruitment. Cell 172, 696–705 (2018).
Rigort, A. et al. Focused ion beam micromachining of eukaryotic cells for cryoelectron tomography. Proc. Natl Acad. Sci. USA 109, 4449–4454 (2012).
Mastronarde, D. N. Automated electron microscope tomography using robust prediction of specimen movements. J. Struct. Biol. 152, 36–51 (2005).
Nickell, S. et al. TOM software toolbox: acquisition and analysis for electron tomography. J. Struct. Biol. 149, 227–234 (2005).
Li, X. et al. Electron counting and beam-induced motion correction enable near-atomic-resolution single-particle cryo-EM. Nat. Methods 10, 584–590 (2013).
Kremer, J. R., Mastronarde, D. N. & McIntosh, J. R. Computer visualization of three-dimensional image data using IMOD. J. Struct. Biol. 116, 71–76 (1996).
Hrabe, T. et al. PyTom: a python-based toolbox for localization of macromolecules in cryo-electron tomograms and subtomogram analysis. J. Struct. Biol. 178, 177–188 (2012).
Bharat, T. A. & Scheres, S. H. Resolving macromolecular structures from electron cryo-tomography data using subtomogram averaging in RELION. Nat. Protoc. 11, 2054–2065 (2016).
Acknowledgements
We thank C.-G. Pack and M.-K. Jung for initial electron microscopy analysis; S. Fukai, K. Iwai, T. Yamamoto and F. Zhang for reagents; N. Noda for helpful discussion; and J. Horiuchi for critical reading of the manuscript. This research was supported by AMED under grant numbers JP18gm1110003 (S.M. and Y.S.) and JP19gm1110010 (T.I.); MEXT/JSPS KAKENHI grant numbers JP18K19352 (S.Y.), JP18K14913 (H.T.), JP26293018 (Y.S.), JP18H03993 (Y.S.), JP18H05498 (F.O. and Y.S.), JP18H03977 (T.I.), JP 19H05281 (T.I.) and JP21000012 (K.T.); the Takeda Science Foundation (Y.S., K.T. and T.I.); the Uehara Memorial Foundation (Y.S.); the European Commission, FP7 GA ERC-2012-SyG_318987-ToPAG (Q.G., W.B. and R.F.-B.); and the DFG, EXC 2067/1- 390729940 (R.F.-B.).
Author information
Authors and Affiliations
Contributions
S.Y., K.T. and Y.S. designed most of the experiments; S.Y., A.K. and Y.S. generated the KI/KO cell lines with assistance of S.M.; S.Y. and A.K. performed the fluorescent microscopy analyses; S.Y. processed microscopy images; H.T., A.K., N.A. and A.E. performed the immunoprecipitation and western blotting; H.T. performed the mass spectrometry proteomics analysis with assistance of A.E. and F.O.; H.T. and N.A. prepared recombinant proteins; S.Y. and Y.S. performed in vitro phase separation assay; Q.G., W.B. and R.F.-B. performed and analysed cryo-ET experiments; K.I. and T.I. performed northern blotting; S.Y., K.T. and Y.S. wrote the manuscript with input from all co-authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Characterization of KI cells stably expressing fluorescent protein-labelled proteasome subunit or ubiquitin.
a, HCT116 cells stably expressing proteasome subunits (PSMB2 or PSMD6) or ubiquitin fused with either eGFP or FusionRed. Scale bars, 10 μm. b, Proteasome assembly state of cell lines indicated in a, as determined by native PAGE. c, d, Immunoblot analysis of the KI cells in a with antibodies against ubiquitin (c), PSMD6 or PSMB2 (d). Ponceau S staining was performed as a loading control. e, Cell viability of DBeQ-treated cells was determined using the CellTiter assay kit. The MTS assay was performed in triplicate. Data are mean ± s.d. (n = 3). In a–d, representative results from two independent experiments. Gel source data for b–d are shown in Supplementary Fig. 1.
Extended Data Fig. 2 Proteasome foci formation occurs in various cell types following hyperosmotic stress.
a, Localization of endogenous proteasome in HCT116 cells, with or without 0.2 M sucrose stimulation for 30 min, was observed using PSMA1 (α6) antibodies. Scale bars, 10 μm. b, PSMB2-eGFPKI/KI cells were stimulated with sucrose, NaCl or glucose at the indicated concentrations for 5 min. Scale bars, 10 μm. c, PSMD6-eGFPKI/KI cells were stimulated with 0.2 M sucrose for 30 min. Scale bars, 10 μm. d, Tomographic slices of the nuclear region in cells not stimulated (left) or stimulated with 0.2 M sucrose (right). For the whole tomogram, proteasomes were mapped to their original positions and orientations by template matching and sub-tomogram averaging. Higher-magnification tomographic slices of a representative proteasome (arrow; coloured in yellow) detected in the tomograms are shown in the top right. In total, nine tomograms were collected from sucrose-stimulated cells, of which three exhibited proteasome clustering. Seven tomograms were collected from non-stimulated cells, none of which exhibited proteasome clustering. Three-dimensional reconstruction revealed a proteasome structure resolved by averaging 280 sub-tomograms from all tomograms obtained with and without sucrose stimulation. Scale bars, 0.2 μm. e, HCT116 cells and hTERT RPE-1 cells stably expressing PSMB2–eGFP and ES-E14TG2a cells from mouse embryos were stimulated with 0.2 M sucrose for 30 min. For ES-E14TG2a cells, endogenous proteasome activity was detected using a proteasome probe (Me4BodipyFL-Ahx3Leu3VS). Scale bars, 10 μm. Representative results from three (a) or two (b–e) independent experiments.
Extended Data Fig. 3 Proteasome foci are distinct from known nuclear structures.
a, Colocalization of proteasome foci induced by 0.2 M sucrose, as determined by staining for markers of various nuclear bodies. Scale bar, 10 μm. Representative images from two independent experiments. b, PSMB2-eGFPKI/KI cells were transiently transfected with PML siRNA or control siRNA for 48 h, and then stimulated with 0.2 M sucrose for 30 min. The graph indicates the number of foci per cell (siControl, n = 300 cells; siPML, n = 294 cells). Data are mean ± s.d., P = 0.0541 by two-tailed unpaired Student’s t-test. Scale bars, 10 μm. c, PSMB2-eGFPKI/KI cells were stimulated with 0.2 M sucrose for 30 min and were probed with pan-ubiquitin (anti-Ub, clone FK2), K48-ubiquitin (anti-K48Ub) and K63-ubiquitin (anti-K63Ub) specific antibodies. Line profiling of representative sections of cells is indicated by white dashed lines. The mean value of the Pearson correlation coefficient in the nucleoplasm is shown in the image (n = 10 cells in two view fields). Scale bars, 10 μm. Representative images from three (anti-Ub, anti-K48) or two (anti-K63Ub) independent experiments. d, Wild-type or PSMB2-eGFPKI/KI HCT116 cells were stimulated with the E1 inhibitor MLN-7243 (1 μM) for the indicated times and immunoblotted with ubiquitin antibody. Representative result from three independent experiments. For gel source data, see Supplementary Fig. 1.
Extended Data Fig. 4 Changes in ubiquitylation levels and intranuclear structure in response to hyperosmotic stress.
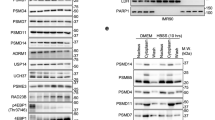
a, HCT116 cells treated with 0.2 M sucrose for the indicated times were subjected to immunoblotting with indicated antibodies. Representative results from two (anti-K48) or three (anti-Ubiquitin) independent experiments. b, Mass spectrometry screening of ubiquitylated substrates by the double-concentration method using TR-TUBE and anti-diGly antibody. PSMB2-eGFPKI/KI cells were stimulated with 0.2 M sucrose at the indicated times. The numbers of peptide spectrum matches for each identified protein are shown as a heat map. Similar results were obtained from two independent experiments. See Source Data for identified peptide list. c, Correlative light and electron microscopy (CLEM) analysis of PSMB2-FusionRedKI/KIeGFP-UbAAVS1/AVS1 cells. Scale bars, 10 μm. Right, square enlarged images on the left. The places where proteasome foci exist are circled in white. Scale bars, 0.5 μm. Representative images from two independent experiments. d, Time course images of PSMB2-eGFPKI/KI cells stimulated with 0.2 M sucrose, with or without pre-treatment with 1 μM MLN-7243 or 50 μM MG-132 for 1 h. Scale bars, 1 μm. Two independent experiments were performed with 0 min, 30 min, 240 min and MG-132, with similar results. e, Schematic of northern blot probes and positions of the pre-rRNA processing intermediates discussed in this Article. RNA was extracted from HCT116 cells stimulated with 0.2 M sucrose for the indicated times and analysed by northern hybridization using probes specific for the 5′ end and the middle of ITS1 and ITS2. Representative results from three independent experiments. Gel source data for a and e are shown in Supplementary Fig. 1.
Extended Data Fig. 5 Unassembled ribosomal proteins are present in proteasome foci.
a, Colocalization of proteasome foci and endogenous ribosome components. PSMB2-eGFPKI/KI cells were stimulated with 0.2 M sucrose for 30 min, and endogenous ribosome components were detected by specific antibodies. The mean value of the Pearson correlation coefficient in the nucleoplasm is shown in the image (n = 10 cells in two fields of view). Scale bars, 10 μm. Each graph represents the normalized fluorescence distribution over white dashed lines of the cells. Representative images from two (RPL4, RPL7A, RPL35, RPS6, RPS9 and 5.8S rRNA) or three (RPS2) independent experiments. b, PSMB2-FusionRedKI/KIRPL29-eGFPAAVS1/AAVS1 cells were stimulated with 0.2 M sucrose for 30 min. Endogenous RPL29 was detected with anti-RPL29 antibody. Representative xy, xz and yz images of a single cell from two independent experiments. Scale bars, 5 μm. c, PSMB2-FusionRedKI/KIRPL29-eGFPAAVS1/AAVS1 cells were treated with the Pol I inhibitor CX-5461 (1 μM, 5 h prior), and stimulated with 0.2 M sucrose. Scale bars, 5 μm. Right graphs indicate the foci numbers per cell and their diameters. RPL29 foci number are mean ± s.d. from n = 82 cells (control) and n = 68 cells (CX-5461); ****P < 0.0001 by two-tailed unpaired t-test. RPL29 foci diameter are presented as mean ± s.d. from n = 542 foci (control) and n = 690 foci (CX-5461); ****P < 0.0001 by two-tailed Mann–Whitney U-test. PSMB2 foci number are mean ± s.d. from n = 216 cells (control) and n = 228 cells (CX-5461); **P = 0.0092 by two-tailed Mann–Whitney U-test. PSMB2 foci diameter are mean ± s.d. from n = 815 foci (control) and n = 1,149 foci (CX-5461); ****P < 0.0001 by two-tailed Mann–Whitney U-test. d, PSMB2-FusionRedKI/KI cells transiently overexpressing (white arrows) or not overexpressing RPL7A–eGFP or RPS2–eGFP were stimulated with 0.2 M sucrose for 30 min. Scale bars, 5 μm. Similar results were obtained from three independent experiments.
Extended Data Fig. 6 Microscopy analysis of proteasome-interacting proteins.
a, PSMB2-FusionRedKI/KI cells were stimulated with 0.2 M sucrose for 30 min. Endogenous p97 and ubiquitin were detected with anti-p97 and anti-ubiquitin antibodies, respectively. Representative xy, xz and yz images of a single cell from two independent experiments. Scale bars, 5 μm. b, PSMB2-FusionRedKI/KI cells were stimulated with 0.2 M sucrose for 30 min. Endogenous p97 and RAD23B were detected with anti-p97 and anti-RAD23B antibodies, respectively. Representative xy, xz and yz images of a single cell from two independent experiments. Scale bars, 5 μm. c, PSMB2-eGFPKI/KI cells were transiently transfected for 48 h with siRNA targeting RAD23A, RAD23B, UBQLN1/2/4 (mixture) or XPC, or a control siRNA, and then stimulated with 0.2 M sucrose for 30 min. Right graph indicates the number of foci per cell (siControl, n = 323 cells; siRAD23A, n = 129 cells; siRAD23B, n = 178 cells; siUBQLN1/2/4, n = 349 cells; siXPC, n = 339 cells). Data are mean ± s.d., ****P < 0.0001 by Kruskal–Wallis with Dunn’s multiple comparisons test.
Extended Data Fig. 7 Inhibition of proteasome or p97 activity results in enlargement of RPL29 condensates.
a, PSMB2-FusionRedKI/KIRPL29-eGFPAAVS1/AAVS1 cells treated with the proteasome inhibitor b-AP15 (5 μM, 10 min posterior), the p97 inhibitor NMS-873 (1 μM, 1 h prior), or the ubiquitin-activating enzyme (E1) inhibitor MLN-7243 (1 μM, 1 h prior), or lacking RAD23B, were stimulated with 0.2 M sucrose. Right graphs indicate the RPL29 foci numbers per cell and their diameters. The foci number per cell are presented as mean ± s.d. from n = 35 cells (control), n = 44 cells (b-AP15), n = 37 cells (MLN-7243), n = 50 cells (RAD23B KO) and n = 45 cells (NMS-873). P = 0.2250 (b-AP15), **P = 0.0085 (MLN-7243), **P = 0.0017 (RAD23B KO) and P = 0.9744 (NMS-873) by one-way ANOVA with Dunnett’s multiple comparisons test. The foci diameters are presented as mean ± s.d. from n = 220 foci (control), n = 210 foci (b-AP15), n = 132 foci (MLN-7243), n = 170 foci (RAD23B KO) and n = 271 foci (NMS-873). ****P < 0.0001 (b-AP15), ****P < 0.0001 (MLN-7243), ****P < 0.0001 (RAD23B KO) and ****P < 0.0001 (NMS-873) by Kruskal–Wallis with Dunn’s multiple comparisons test. Scale bars, 10 μm. b, Time-lapse images of single foci in the b-AP15 or NMS-873 treated PSMB2-FusionRedKI/KIRPL29-eGFPAAVS1/AAVS1 cells under the same conditions described in a. Scale bars, 0.5 μm. Representative images from four (control) or two (b-AP15 and NMS-873) independent experiments.
Extended Data Fig. 8 Characterization of RAD23B mutants in foci formation.
a, Wild-type, RAD23B-KO (RAD23BKO/KO) or UBE3A-KO (UBE3AKO/KO) HCT116 cells were stimulated with 0.2 M sucrose for 30 min and immunoblotted with the indicated antibodies. Similar results were obtained from two independent experiments. b, PSMB2-eGFPKI/KIRAD23BKO/KO cells stably expressing FusionRed-fused RAD23B(WT), UBL (L8A) or UBA (L225A–L401A) were treated with 0.2 M sucrose for 30 min. Endogenous K48-linked ubiquitin chains were detected with K48-ubiquitin antibody and Alexa Fluor 647-labelled secondary antibody. Merged images represent K48-ubiquitin antibody (red), PSMB2–eGFP (green) and DAPI (blue). Scale bars, 10 μm. Representative images from two independent experiments. c, RAD23A–FusionRed-overexpressing PSMB2-eGFPKI/KIRAD23BKO/KO cells were treated with 0.2 M sucrose for 30 min. Scale bars, 10 μm. Similar results were obtained from two independent experiments. d, Cellular abundances of RAD23A and RAD23B in HCT116 cells. See Source Data for details. e, Quantification of cell death by PI assay in wild-type or RAD23B-KO (RAD23BKO/KO) HCT116 cells stimulated with 0.2 M sucrose for 2 h, followed by 6 h recovery under normal conditions. The graph indicates percentage of PI-positive cells. n represents the number of fields of view (untreated wild-type cells, n = 12, total 2,829 cells; sucrose-treated wild-type cells, n = 11, total 2,868 cells; untreated RAD23BKO/KO cells, n = 12, total 2,865 cells; sucrose-treated RAD23BKO/KO cells, n = 13, total 2,960 cells). Data are mean ± s.d., P = 0.0029 by one-way ANOVA with Tukey’s multiple comparison test. f, Western blot analysis of wild-type or RAD23B-KO HCT116 cells under the same conditions described in e. The presence of cleaved caspase-3 was used as a marker for apoptosis. Representative results from two independent experiments. Gel source data for a, f are shown in Supplementary Fig. 1.
Extended Data Fig. 9 Liquid–liquid phase separation of ubiquitin chains and RAD23B in vitro.
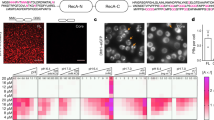
a, SDS–PAGE and fluorescent images of Cy5-labelled proteins used in the in vitro liquid–liquid phase separation assay. Cy5-labelled K48-linked ubiquitin chains (Cy5–K48Ubmix) generated by enzymatic reactions using E2-25K (left), Cy5-labelled K48-linked ubiquitin chains size-fractionated by gel filtration (middle) and Cy5-labelled K63-linked ubiquitin chains size-fractionated by gel filtration (right). Note that K63-Ub2 was omitted owing to difficulty of separation. b, SDS–PAGE and fluorescent images of Cy3-labelled RAD23 proteins. c, Effects of concentration of molecular crowding agents and NaCl. Images of the solution were obtained 90 min after mixing 20 μM Cy5–K48Ubmix and 20 μm Cy3–RAD23B(WT) with the indicated conditions. Scale bars, 5 μm. d, Average of quantification of fluorescence recovery of 20 μM Cy3–RAD23B and 60 μM Cy5–K48Ub4 (5 min after mixing in 10% PEG) and double-exponential fitting curve with s.e.m. (n = 2). e, Chain length-dependent formation of liquid droplets. Images of the solution were obtained 90 min after mixing in 20 μM size-fractionated Cy5–K48Ub, Cy5–K63Ub, or ubiquitin monomer (Cy5–Ub) and 20 μm Cy3–RAD23B(WT). All samples contained 3% PEG in 200 mM NaCl. Scale bars, 5 μm. Representative results from two independent experiments. Gel source data for a, b, are shown in Supplementary Fig. 1.
Supplementary information
Supplementary Figures
Supplementary Figure 1: Uncropped blots and gels. The original source images of all the blots and gels with size marker in this study.
Video 1: Hyperosmotic stress induces the formation of proteasome foci.
PSMB2-EGFPKI/KI cells were stimulated with 0.2 M sucrose. Images were collected at 2 second frame intervals. Scale bar, 10 μm. Representative results from three independent experiments.
Video 2: Ribosomal proteins are degraded in proteasome foci.
PSMB2-FusionRedKI/KI RPL29-EGFPAAVS1/AAVS1 cells were stimulated with 0.2 M sucrose. Images were collected at indicated frame intervals. Scale bar, 5 μm. Representative results from five independent experiments.
Video 3: Ribosomal proteins are not degraded in proteasome foci in the presence of E1 inhibitor.
PSMB2-FusionRedKI/KI RPL29-EGFPAAVS1/AAVS1 cells were treated with the E1 inhibitor MLN-7243 (1 μM, 1 h prior), were stimulated with 0.2 M sucrose. Images were collected at indicated frame intervals. Scale bar, 5 μm. Representative results from three independent experiments.
Video 4: Proteasome foci are liquid droplets.
Foci fusion in PSMB2-FusionRedKI/KI EGFP-UbAAVS1/AAVS1 cells after stimulation with 0.2 M sucrose. Images were collected at 10 second frame intervals. Scale bar, 10 μm. Representative results from three independent experiments.
Video 5: Inhibition of proteasome activity results in enlargement RPL29 condensates.
PSMB2-FusionRedKI/KI RPL29-EGFPAAVS1/AAVS1 cells treated with the proteasome inhibitor b-AP15 (5 μM, 10 min posterior) were stimulated with 0.2 M sucrose. Images were collected at indicated frame intervals. Scale bar, 5 μm. Representative results from two independent experiments.
Video 6: Inhibition of p97 activity results in enlargement RPL29 condensates.
PSMB2-FusionRedKI/KI RPL29-EGFPAAVS1/AAVS1 cells treated with the p97 inhibitor NMS-873 (1 μM, 1 h prior) were stimulated with 0.2 M sucrose. Images were collected at indicated frame intervals. Scale bar, 5 μm. Representative results from two independent experiments.
Video 7: Polyubiquitin chains and RAD23B induces liquid-liquid phase separation in vitro.
Liquid droplets were formed by mixing Cy3-RAD23B and Cy5-K48-ubiquitin in vitro. One minute after mixing, images were collected at 7 second frame intervals. Scale bar, 5 μm. Representative results from two independent experiments.
Source data
Rights and permissions
About this article
Cite this article
Yasuda, S., Tsuchiya, H., Kaiho, A. et al. Stress- and ubiquitylation-dependent phase separation of the proteasome. Nature 578, 296–300 (2020). https://doi.org/10.1038/s41586-020-1982-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-020-1982-9
This article is cited by
-
Phase separation-mediated biomolecular condensates and their relationship to tumor
Cell Communication and Signaling (2024)
-
High-throughput and proteome-wide discovery of endogenous biomolecular condensates
Nature Chemistry (2024)
-
Elucidation of Site-Specific Ubiquitination on Chaperones in Response to Mutant Huntingtin
Cellular and Molecular Neurobiology (2024)
-
Label-free autofluorescence lifetime reveals the structural dynamics of ataxin-3 inside droplets formed via liquid–liquid phase separation
Scientific Reports (2023)
-
Biomolecular condensates in kidney physiology and disease
Nature Reviews Nephrology (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.