Abstract
ATP13A2 (PARK9) is a late endolysosomal transporter that is genetically implicated in a spectrum of neurodegenerative disorders, including Kufor-Rakeb syndrome—a parkinsonism with dementia1—and early-onset Parkinson’s disease2. ATP13A2 offers protection against genetic and environmental risk factors of Parkinson’s disease, whereas loss of ATP13A2 compromises lysosomes3. However, the transport function of ATP13A2 in lysosomes remains unclear. Here we establish ATP13A2 as a lysosomal polyamine exporter that shows the highest affinity for spermine among the polyamines examined. Polyamines stimulate the activity of purified ATP13A2, whereas ATP13A2 mutants that are implicated in disease are functionally impaired to a degree that correlates with the disease phenotype. ATP13A2 promotes the cellular uptake of polyamines by endocytosis and transports them into the cytosol, highlighting a role for endolysosomes in the uptake of polyamines into cells. At high concentrations polyamines induce cell toxicity, which is exacerbated by ATP13A2 loss due to lysosomal dysfunction, lysosomal rupture and cathepsin B activation. This phenotype is recapitulated in neurons and nematodes with impaired expression of ATP13A2 or its orthologues. We present defective lysosomal polyamine export as a mechanism for lysosome-dependent cell death that may be implicated in neurodegeneration, and shed light on the molecular identity of the mammalian polyamine transport system.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
Gel source data for immunoblots and radiograms (Figs. 1, 2, Extended Data Figs. 1–3, 6) are available with the online version of the paper (Supplementary Fig. 1). All other datasets generated within this study are presented and analysed within this manuscript and are available from the corresponding author upon reasonable request.
References
Ramirez, A. et al. Hereditary parkinsonism with dementia is caused by mutations in ATP13A2, encoding a lysosomal type 5 P-type ATPase. Nat. Genet. 38, 1184–1191 (2006).
Di Fonzo, A. et al. ATP13A2 missense mutations in juvenile parkinsonism and young onset Parkinson disease. Neurology 68, 1557–1562 (2007).
Dehay, B. et al. Loss of P-type ATPase ATP13A2/PARK9 function induces general lysosomal deficiency and leads to Parkinson disease neurodegeneration. Proc. Natl Acad. Sci. USA 109, 9611–9616 (2012).
Palmgren, M. G. & Nissen, P. P-type ATPases. Annu. Rev. Biophys. 40, 243–266 (2011).
van Veen, S. et al. Cellular function and pathological role of ATP13A2 and related P-type transport ATPases in Parkinson’s disease and other neurological disorders. Front. Mol. Neurosci. 7, 48 (2014).
Narayanaswamy, N. et al. A pH-correctable, DNA-based fluorescent reporter for organellar calcium. Nat. Methods 16, 95–102 (2019).
De La Hera, D. P., Corradi, G. R., Adamo, H. P. & De Tezanos Pinto, F. Parkinson’s disease-associated human P5B-ATPase ATP13A2 increases spermidine uptake. Biochem. J. 450, 47–53 (2013).
Heinick, A. et al. Caenorhabditis elegans P5B-type ATPase CATP-5 operates in polyamine transport and is crucial for norspermidine-mediated suppression of RNA interference. FASEB J. 24, 206–217 (2010).
Holemans, T. et al. A lipid switch unlocks Parkinson's disease-associated ATP13A2. Proc. Natl Acad. Sci. USA 112, 9040–9045, (2015).
Martin, S., Holemans, T. & Vangheluwe, P. Unlocking ATP13A2/PARK9 activity. Cell Cycle 14, 3341–3342 (2015).
Martin, S. et al. Protection against mitochondrial and metal toxicity depends on functional lipid binding sites in ATP13A2. Parkinsons Dis. 2016, 9531917 (2016).
Sørensen, D. M. et al. Parkinson disease related ATP13A2 evolved early in animal evolution. PLoS ONE 13, e0193228 (2018).
Vanhoutte, R., Kahler, J. P., Martin, S., van Veen, S. & Verhelst, S. H. L. Clickable polyamine derivatives as chemical probes for the polyamine transport system. ChemBioChem 19, 907–911 (2018).
Demirsoy, S. et al. ATP13A2/PARK9 regulates endo-/lysosomal cargo sorting and proteostasis through a novel PI(3, 5)P2-mediated scaffolding function. Hum. Mol. Genet. 26, 1656–1669 (2017).
Seglen, P. O. & Gordon, P. B. Effects of lysosomotropic monoamines, diamines, amino alcohols, and other amino compounds on protein degradation and protein synthesis in isolated rat hepatocytes. Mol. Pharmacol. 18, 468–475 (1980).
Chatr-aryamontri, A. et al. The BioGRID interaction database: 2017 update. Nucleic Acids Res. 45, D369–D379 (2017).
Estrada-Cuzcano, A. et al. Loss-of-function mutations in the ATP13A2/PARK9 gene cause complicated hereditary spastic paraplegia (SPG78). Brain 140, 287–305 (2017).
Pegg, A. E. Functions of polyamines in mammals. J. Biol. Chem. 291, 14904–14912 (2016).
Qiao, C. et al. Atp13a2 deficiency aggravates astrocyte-mediated neuroinflammation via NLRP3 inflammasome activation. CNS Neurosci. Ther. 22, 451–460 (2016).
Aits, S. & Jäättelä, M. Lysosomal cell death at a glance. J. Cell Sci. 126, 1905–1912 (2013).
Bourdenx, M. et al. Nanoparticles restore lysosomal acidification defects: implications for Parkinson and other lysosomal-related diseases. Autophagy 12, 472–483 (2016).
Zielich, J. et al. Overlapping expression patterns and functions of three paralogous P5B ATPases in Caenorhabditis elegans. PLoS ONE 13, e0194451 (2018).
Belting, M. et al. Glypican-1 is a vehicle for polyamine uptake in mammalian cells: a pivital role for nitrosothiol-derived nitric oxide. J. Biol. Chem. 278, 47181–47189 (2003).
Uemura, T., Stringer, D. E., Blohm-Mangone, K. A. & Gerner, E. W. Polyamine transport is mediated by both endocytic and solute carrier transport mechanisms in the gastrointestinal tract. Am. J. Physiol. Gastrointest. Liver Physiol. 299, G517–G522 (2010).
Klein, C. & Westenberger, A. Genetics of Parkinson’s disease. Cold Spring Harb. Perspect. Med. 2, a008888 (2012).
Ha, H. C. et al. The natural polyamine spermine functions directly as a free radical scavenger. Proc. Natl Acad. Sci. USA 95, 11140–11145 (1998).
Madeo, F., Eisenberg, T., Pietrocola, F. & Kroemer, G. Spermidine in health and disease. Science 359, eaan2788 (2018).
Li, C. et al. Spermine synthase deficiency causes lysosomal dysfunction and oxidative stress in models of Snyder–Robinson syndrome. Nat. Commun. 8, 1257 (2017).
Lewandowski, N. M. et al. Polyamine pathway contributes to the pathogenesis of Parkinson disease. Proc. Natl Acad. Sci. USA 107, 16970–16975 (2010).
Sørensen, D. M., Buch-Pedersen, M. J. & Palmgren, M. G. Structural divergence between the two subgroups of P5 ATPases. Biochim. Biophys. Acta 1797, 846–855 (2010).
Axelsen, K. B. & Palmgren, M. G. Evolution of substrate specificities in the P-type ATPase superfamily. J. Mol. Evol. 46, 84–101 (1998).
Grunewald, A. et al. ATP13A2 mutations impair mitochondrial function in fibroblasts from patients with Kufor-Rakeb syndrome. Neurobiol Aging 33, 1843.e1–1843.e7 (2012).
Usenovic, M., Tresse, E., Mazzulli, J. R., Taylor, J. P. & Krainc, D. Deficiency of ATP13A2 leads to lysosomal dysfunction, α-synuclein accumulation, and neurotoxicity. J. Neurosci. 32, 4240–4246 (2012).
Santoro, L. et al. Novel ATP13A2 (PARK9) homozygous mutation in a family with marked phenotype variability. Neurogenetics 12, 33–39 (2011).
Funayama, M. et al. Rapid screening of ATP13A2 variant with high-resolution melting analysis. Mov. Disord. 25, 2434–2437 (2010).
Lin, C. H. et al. Novel ATP13A2 variant associated with Parkinson disease in Taiwan and Singapore. Neurology 71, 1727–1732 (2008).
Mao, X. Y. et al. ATP13A2 G2236A variant is rare in patients with early-onset Parkinson’s disease and familial Parkinson’s disease from mainland China. Parkinsonism Relat. Disord. 16, 235–236 (2010).
Sanjana, N. E., Shalem, O. & Zhang, F. Improved vectors and genome-wide libraries for CRISPR screening. Nat. Methods 11, 783–784 (2014).
Kleinstiver, B. P. et al. High-fidelity CRISPR–Cas9 nucleases with no detectable genome-wide off-target effects. Nature 529, 490–495 (2016).
Doench, J. G. et al. Optimized sgRNA design to maximize activity and minimize off-target effects of CRISPR–Cas9. Nat. Biotechnol. 34, 184–191 (2016).
Covy, J. P., Waxman, E. A. & Giasson, B. I. Characterization of cellular protective effects of ATP13A2/PARK9 expression and alterations resulting from pathogenic mutants. J. Neurosci. Res. 90, 2306–2316 (2012).
Gitler, A. D. et al. α-synuclein is part of a diverse and highly conserved interaction network that includes PARK9 and manganese toxicity. Nat. Genet. 41, 308–315 (2009).
Kong, S. M. et al. Parkinson’s disease-linked human PARK9/ATP13A2 maintains zinc homeostasis and promotes α-synuclein externalization via exosomes. Hum. Mol. Genet. 23, 2816–2833 (2014).
Lambie, E. J., Tieu, P. J., Lebedeva, N., Church, D. L. & Conradt, B. CATP-6, a C. elegans ortholog of ATP13A2 PARK9, positively regulates GEM-1, an SLC16A transporter. PLoS ONE 8, e77202 (2013).
Tsunemi, T. et al. Increased lysosomal exocytosis induced by lysosomal Ca2+ channel agonists protects human dopaminergic neurons from α-synuclein toxicity. J. Neurosci. 39, 5760–5772 (2019).
Gietz, R. D. & Woods, R. A. Transformation of yeast by lithium acetate/single-stranded carrier DNA/polyethylene glycol method. Methods Enzymol. 350, 87–96 (2002).
Azouaoui, H. et al. A high-yield co-expression system for the purification of an intact Drs2p-Cdc50p lipid flippase complex, critically dependent on and stabilized by phosphatidylinositol-4-phosphate. PLoS ONE 9, e112176 (2014).
Jidenko, M., Lenoir, G., Fuentes, J. M., le Maire, M. & Jaxel, C. Expression in yeast and purification of a membrane protein, SERCA1a, using a biotinylated acceptor domain. Protein Expr. Purif. 48, 32–42 (2006).
Papadopulos, A. et al. Flippase activity detected with unlabeled lipids by shape changes of giant unilamellar vesicles. J. Biol. Chem. 282, 15559–15568 (2007).
Byun, J. A. et al. Analysis of polyamines as carbamoyl derivatives in urine and serum by liquid chromatography–tandem mass spectrometry. Biomed. Chromatogr. 22, 73–80 (2008).
Stroikin, Y., Mild, H., Johansson, U., Roberg, K. & Ollinger, K. Lysosome-targeted stress reveals increased stability of lipofuscin-containing lysosomes. Age 30, 31–42 (2008).
Nilsson, C., Kagedal, K., Johansson, U. & Ollinger, K. Analysis of cytosolic and lysosomal pH in apoptotic cells by flow cytometry. Methods Cell. Sci. 25, 185–194 (2003).
Osório, L. et al. Viral vectors expressing a single microRNA-based short-hairpin RNA result in potent gene silencing in vitro and in vivo. J. Biotechnol. 169, 71–81 (2014).
Brenner, S. The genetics of Caenorhabditis elegans. Genetics 77, 71–94 (1974).
Casadaban, M. J., Martinez-Arias, A., Shapira, S. K. & Chou, J. β-galactosidase gene fusions for analyzing gene expression in Escherichia coli and yeast. Methods Enzymol. 100, 293–308 (1983).
The C. elegans Deletion Mutant Consortium. Large-scale screening for targeted knockouts in the Caenorhabditis elegans genome. G3 2, 1415–1425 (2012).
Mello, C. C., Kramer, J. M., Stinchcomb, D. & Ambros, V. Efficient gene transfer in C. elegans: extrachromosomal maintenance and integration of transforming sequences. EMBO J. 10, 3959–3970 (1991).
Roy, A., Kucukural, A. & Zhang, Y. I-TASSER: a unified platform for automated protein structure and function prediction. Nat. Protoc. 5, 725–738 (2010).
Zhang, Y. I-TASSER server for protein 3D structure prediction. BMC Bioinformatics 9, 40 (2008).
Omasits, U., Ahrens, C. H., Müller, S. & Wollscheid, B. Protter: interactive protein feature visualization and integration with experimental proteomic data. Bioinformatics 30, 884–886 (2014).
Acknowledgements
This work was funded by the Fonds Wetenschappelijk Onderzoek (FWO, Research Foundation Flanders) (G094219N to P.V., G092714 and G080517N to V. Baekelandt, SBO Neuro-TRAFFIC S006617N to V. Baekelandt, P.V., W.A. and J.V.S. and 1503117N to S.M.), the KU Leuven (OT/13/091; LysoCaN C16/15/073 to P.V., J.V.S., W.A. and P.A.; OT/14/120 to V. Baekelandt) and the Queen Elisabeth Medical Foundation for Neurosciences (P.V. and V. Baekelandt). S.v.V. is an aspirant FWO research fellow (11Y7518N) and J.P.K. is the recipient of a strategic basic research doctoral grant of the FWO (1S18518N). J.L. received Lundbeck postdoctoral fellowships R209-2015-2704 and R171-2014-663. Funding for E.L. and J.Z. was provided by DFG grant LA3380/2-1. Some nematode strains were provided by the Caenorhabditis Genetics Center, which is funded by NIH Office of Research Infrastructure Programs (P40 OD010440). We thank P. P. Van Veldhoven (KU Leuven) for discussions; P. Chaltin (Center for Drug Design and Discovery) for financial support for V. Benoy; and M. Schuermans, T. Arslan, A. Florizoone, J. Van Asselberghs, J. Chen and N. Hamouda for technical assistance. We also acknowledge our frequent use of the facilities and equipment of the Leuven Viral Vector Core facility (KU Leuven), Mass Spectrometry Core (R. Derua, KU Leuven), Cell and Tissue Imaging Cluster (P. Vanden Berghe, KU Leuven) and the FACS Core (KU Leuven/VIB).
Author information
Authors and Affiliations
Contributions
The study was designed by P.V., S.M., S.v.V. and J.E.; C.V.d.H. generated all stable cell lines and analysed ATP13A2 knockdown; J.-P.D. performed and analysed the CRISPR experiments; R.V. and J.P.K. synthesized BODIPY-labelled polyamine analogues; J.P.K performed and analysed acidic nanoparticle preparations; S.v.V performed and analysed the biochemical experiments (except Extended Data Fig. 2f by V. Benoy); J.L. contributed to the ATP13A2 purification protocol; S.M. performed and analysed the cell biology experiments; B.G. performed metabolomics; G.G. isolated mouse neurons; E.L. and J.Z. performed C. elegans experiments; and P.V., S.v.V. and S.M. wrote the manuscript, which was reviewed by all authors.
Corresponding author
Ethics declarations
Competing interests
Patent WO-2018002350-A1 of KU Leuven describes methods for detecting compounds with therapeutic use that target ATP13A2 or related isoforms using biological material and assays described in the current manuscript. A second patent of KU Leuven describing ATP13A2 cell models described in this manuscript has also been filed.
Additional information
Peer review information Nature thanks Guillaume Lenoir, Hanne Poulsen, Taiji Tsunemi and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 ATP13A2 is a polyamine transporter.
a–d, The ATPase activity of ATP13A2 was measured in solubilized microsomes (5 μg) collected from SH-SY5Y cells stably overexpressing wild-type ATP13A2 (WT-OE) in the presence of 100 nM CaCl2, MnCl2, ZnCl2 or FeCl3 and 100 μM SPD or SPM (D508N-OE as a negative control, wild type was referenced from Fig. 1b) (a) or in the presence of the indicated doses of inorganic ions and heavy metals CaCl2, MnCl2, ZnCl2 or FeCl3 (b), diamines (cadaverine, agmatine and the amino acid l-arginine) (c), monoamines (dopamine and histamine) (d) and acetylated polyamines (N1-acetylspermine, N8-acetylspermidine or N1-acetylspermidine) (e). As a reference for c–e, we plotted the dose–response curve of SPM from Fig. 1b. f, Microsomes (20 μg) collected from SH-SY5Y cells that overexpress ATP13A2 were incubated for 60 s with [γ-32P]ATP in the presence of 10 mM ornithine (ORN) or SPM (referenced from Fig. 1c). Left, a representative autoradiogram of the phosphoenzymes (EP); right, quantification. CON, control. g, The ATPase activity of purified ATP13A2 was assessed after 1 mM SPM was administered in the presence or absence of 0.25 mM orthovanadate (ORTH), a general P-type ATPase inhibitor (supplemented with 125 μM phosphatidic acid/PtdIns(3,5)P2; conditions (−) and (−)/SPM refer to Fig. 1f). h, Purified ATP13A2 was incubated with [γ-32P]ATP in the presence of 1 mM SPM, and radioactivity of the phospho-intermediate was assessed by scintillation counting. i, Comparison of the pulse (5 μM, 15 min) chase (105 min, medium) BODIPY–SPM uptake in KO/WT and KO/D508N cell lines by confocal microscopy. Cells were subsequently stained with LAMP1 and imaged with the same laser settings by confocal microscopy. DAPI was used to visualize the nuclei. Scale bar, 5 μm. j, Line intensity plots of the indicated dashed lines in Fig. 2d. k, Analysis of the Pearson’s coefficient of LAMP1 and BODIPY–SPM for the images in Fig. 2d (KO/WT, 78 images; KO/D508N, 85 images). l, Mean fluorescence intensities (MFI) of BODIPY in DAPI-positive regions of samples shown in Fig. 2d (KO/WT, 233 nuclei; KO/D508N, 243 nuclei). Data are presented as the mean ± s.e.m. or mean with individual data points shown (points represent replicates), with n = 3 independent biological experiments. Analysis was carried out using one-way ANOVA with Dunnett’s (f) or Tukey’s (a, g) corrections, or by two-tailed t-tests (unpaired, h or Welch’s, k, l). Fitted lines are semi-log lines (b) or nonlinear allosteric sigmoidal association (c–e). For gel source data, see Supplementary Fig. 1.
Extended Data Fig. 2 Streptavidin-based purification of wild-type ATP13A2 and the catalytically dead E343A mutant.
a, Coomassie staining showing the purification process for wild-type ATP13A2, starting from solubilized yeast membrane fractions, followed by streptavidin affinity chromatography and on-column thrombin cleavage to elute the protein. b, Western blot analysis of stages in the purification of ATP13A2. c, Bar graph depicting protein purity as determined by densitometry from Coomassie-stained SDS–PAGE. d, Mass spectrometry analysis of the purified ATP13A2 sample. Singly, doubly and triply charged species are indicated. e, To evaluate phosphoenzyme formation, yeast P3 membranes (20 μg) and purified ATP13A2 (1 μg) were incubated for 60 s with [γ-32P]ATP. As a positive control, microsomes collected from SH-SY5Y cells that overexpress wild-type ATP13A2 (20 μg) were used. The image is a representative radiogram depicting the ATP13A2 phosphoenzyme. f, The ATPase activity of purified ATP13A2 (0.3 μg) was measured in the presence of 2 mM SPM and the indicated concentrations of the ATP13A2 regulatory lipids phosphatidic acid (PA) and PtdIns(3,5)P2. g, Coomassie staining showing the purification process for ATP13A2(E343A). h, The ATPase activity of purified wild-type ATP13A2 or ATP13A2(E343A) (0.5 μg) was measured in the presence of the indicated concentrations of SPM with 125 μM phosphatidic acid and 125 μM PtdIns(3,5)P2. Data are expressed as mean with individual data points (points represent replicates) (c, h). The number of independent biological experiments were as follows: n = 3 (b, e–h); n = 6 (d); n = 22 (a, c). Analysis was performed using one-way ANOVA with Tukey’s post-hoc correction (h). For gel source data, see Supplementary Fig. 1.
Extended Data Fig. 3 Confirmation of CRISPR–Cas9-mediated ATP13A2 knockout and subsequent rescue with wild-type ATP13A2 or the D508N mutant.
a, The ATP13A2 knockout cell lines (KO) were generated by CRISPR–Cas9 in SH-SY5Y cells and confirmed by qPCR (top) and immunoblotting (bottom). Atp13a2 mRNA expression was normalized to hypoxanthine phosphoribosyltransferase (HPRT) and TATA-sequence-binding protein (TBP), and GAPDH was used as a loading control for the ATP13A2 protein levels. Two fragments of the same blot are depicted and separated by a dotted line. b, Rescue of ATP13A2 knockout was performed by lentiviral transduction to express wild-type ATP13A2 (KO/WT) and the catalytically dead mutant D508N, which was used as a negative control (KO/D508N). The expression of the ATP13A2 constructs was confirmed via immunoblotting. The number of biologically independent experiments were as follows: n = 1 (a, top panel); n = 3 (a, bottom panel, b). For gel source data, see Supplementary Fig. 1.
Extended Data Fig. 4 Polyamine uptake by ATP13A2 complements cytosolic polyamine synthesis.
a–c, Assessment of the cellular uptake of BODIPY-labelled polyamine analogues (a, b) or FITC–dextran (c) by flow cytometry. Uptake of BODIPY–SPD (a) or BODIPY–SPM (b) in SH-SY5Y cells overexpressing Fluc (negative control), wild-type ATP13A2 (WT-OE) or the catalytically dead mutant D508N (D508N-OE). The cells were incubated with 5 μM BODIPY–SPM or BODIPY–SPD for 2 h before analysis by flow cytometry. c, Analysis of FITC–dextran uptake (as a measure of endocytic capacity) was performed in SH-SY5Y control (CON) cells with endogenous ATP13A2 expression, ATP13A2 knockout cells (KO) and rescue cell lines with expression of wild-type ATP13A2 (KO/WT) or the D508N mutant (KO/D508N). The cells were pre-treated for 30 min with a combination (combo) of endocytosis inhibitors Dynasore (100 μM), genistein (50 μM) and Pitstop 2 (50 μM). The cells were incubated for an additional 2 h with FITC–dextran at 37 °C, followed by flow cytometry. d, Schematic representation of polyamine synthesis. SRM, spermidine synthase; SMS, spermine synthase. Specific inhibitors are indicated in red. Control, KO, KO/WT or KO/D508N cells were subjected to inhibition of polyamine synthesis by DFMO (e), 4MCHA (f) and APCHA (g) before measuring cell viability via the MUH assay. All data represent the average of two independent CRISPR–Cas9 knockout and control clones. All data are presented as mean with data points overlaid (points represent replicates) or mean ± s.e.m. The number of biologically independent experiments were as follows: n = 3 (a, b); n = 4 (c, e–g). Analysis was performed using one-way ANOVA with Dunnett’s (a, b) or Tukey’s (c) post-hoc correction or two-way ANOVA with Dunnett’s post-hoc correction (e–g). Fitted lines indicate nonlinear log(inhibitor) versus response (variable slope) (e–g).
Extended Data Fig. 5 Catalytic and clinical mutations of ATP13A2 perturb polyamine function.
a, Sequence alignment of predicted transmembrane helices M4 (left), M6 (middle) and M8 (right). The alignment was generated using Clustal Omega. We generated mutants in M4 (A467V), M6 (D962N) and M8 (K1062A). The A467V mutation converts the protein sequence PPALP of the predicted substrate-binding site in transmembrane segment M4 into the protein sequence PPVLP that is present in ATP13A54,30. Neighbouring membrane helices also contribute to substrate coordination in P-type ATPases, which often relies on conserved and charged residues, such as D962 in M6 and K1062 in M8 of ATP13A24. b, Densitometry of the expression of catalytic mutants presented in Fig. 2e. c, Flow cytometric analysis of cellular BODIPY–SPD uptake in SH-SY5Y cells overexpressing wild-type ATP13A2, the D508N mutant or catalytic mutants. d, Quantification of ATP13A2 phosphorylation levels (EP) presented in Fig. 2g. e, Densitometry analysis of the expression of disease-related mutants presented in Fig. 2i. f, Flow cytometric analysis of cellular BODIPY–SPD uptake in SH-SY5Y cells overexpressing wild-type ATP13A2, D508N or disease mutants. g, Quantification of ATP13A2 phosphorylation levels presented in Fig. 2k. All data are depicted as mean with individual data points (points represent replicates). The number of independent biological experiments were as follows: n = 3 b, c, d (D508N (SPM), A467V (SPM), D962N (SPM), and K1062A (SPM)), e (T12M, G872R), f, g (T12M (–), T12M (SPM), T512I (SPM), G528R (SPM), A741T (SPM), and G872R (SPM)); n = 4 d (D508N (–), A467V (–), D962N (–), and K1062A (–)), e (wild-type, T512I, G528R and A741T), g (wild-type (–), T512I (–), G528R (–), A741T (–), and G872R (–)); n = 5 d (wild-type (–)); n = 6 d (wild-type (SPM)) and g (wild-type (SPM)). Analysis by one-way ANOVA with Dunnett’s (b, c, e, f) or two-way ANOVA with Sidak’s (d, g) post-hoc corrections.
Extended Data Fig. 6 The ATP sensitivity of ATP13A2(D962N) and ATP13A2(E343A) is independent of SPM.
a, Overview of rate constants of ATP13A2 phosphoenzyme decay following a chase with non-radioactive ATP with or without 1 mM SPM. b, After 30 s of incubating D962N microsomes (20 μg) with [γ-32P]ATP, the time course of dephosphorylation after an ATP chase was measured in the presence or absence of SPM. The top panel shows a representative autoradiogram of the phosphoenzymes (EP), whereas the bottom panel depicts the quantification of ATP13A2 phosphorylation levels. As a reference we plotted the wild-type curve, shown in Fig. 1d. Data are presented as the mean ± s.e.m. of n = 4 biologically independent experiments. Analysis by two-way ANOVA with Tukey’s test (b). The fitted line indicates two-phase decay (b). For gel source data, see Supplementary Fig. 1.
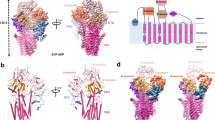
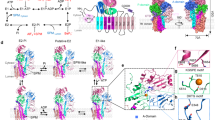
Extended Data Fig. 7 Predicted topology of ATP13A2.
a, Homology model of ATP13A2 based on the structure of Na+/K+-ATPase (ATP1A1, PDB ID: 3A3Y) as a template, generated by iTASSER (https://zhanglab.ccmb.med.umich.edu/I-TASSER/)58,59. b, Predicted membrane topology of ATP13A2 visualized by Protter60 (http://wlab.ethz.ch/protter). ATP13A2 consists of 10 transmembrane helices (M1–10) and an N-terminal membrane-associated region (Ma)9. Kufor-Rakeb syndrome (KRS)-associated mutations are highlighted in red; early-onset Parkinson’s disease (PD)-associated mutations in green; hereditary spastic paraplegia (HSP)-associated mutations in light blue; neuronal ceroid lipofuscinosis (NCL)-associated mutations in orange. Catalytic mutations and mutations in the predicted substrate-binding region are highlighted in dark blue. Residues that were subjected to mutagenesis in this study are labelled (only in a). P-type ATPase signature motifs in the cytosolic domains are indicated in pink (only in b).
Extended Data Fig. 8 Lysosomal functionality and recovery.
a–d, The impact of exogenous polyamines on cell toxicity (24 h) and lysosomal functionality (4 h) was assessed in SH-SY5Y control cells (CON) with endogenous ATP13A2, ATP13A2 knockout cells (KO) and rescue cell lines with wild-type expression (KO/WT) or expression of the catalytically dead mutant D508N (KO/D508N) on the KO background. Cytotoxicity of SPD (a), SPM (b), ORN (c) and PUT (d) were assessed via a MUH cell-viability assay. e, Death of the aforementioned cells was assessed after 4 h of SPM exposure (10 μM) by propidium iodide (PI)-based flow cytometry. f, Measurement of cathepsin D activity. g, h, Lysosomal rupture under basal (–) and SPM (10 μM) conditions was assessed via loss of FITC–dextran (FITC–DEX) punctae (g) or loss of cathepsin B (CTSB)/LAMP1 colocalization (h). Confocal images depict representative images with or without SPM exposure (4 h, DAPI staining for nuclei was included as a reference). Scale bars, 10 μm. The box and whisker plots in g depict the frequency (left) and size (right) of FITC–DEX punctae; in h the Pearson coefficient of colocalization of cathepsin B and LAMP1. i, Lysosomal pH (Fig. 3b) was evaluated using the fluorescent probe FITC–dextran and a dual-emission ratiometric technique. FITC is excited at 488 nm and emission is analysed at 530 nm (BL1) and 610 nm (BL2). A pH calibration curve was generated using FITC–dextran in cells permeabilized with 100 μM monensin and equilibrated with calibration buffers (pH 3–8). j, Representative size distribution of the acidic nanoparticles used in this study. Data are presented as the mean ± s.e.m. (a–d, i) or individual data points (representing replicates) overlaid on group means (e) or box and whisker plots (f–h, line, median; box boundaries, 25th and 75th percentiles). The number of independent biological experiments were as follows: n = 3 (e–j); n = 4 (a–d). Analysis was performed using two-way ANOVA with Dunnett’s (a, b) or Bonferroni’s (c, d) post-hoc corrections, or one-way ANOVA with Dunnett’s (e), Sidak’s (f, g (right)) or Tukey’s (g (left), h) post-hoc corrections. Fitted lines indicate nonlinear log(inhibitor) versus response (variable slope) (a, b).
Extended Data Fig. 9 Inhibition of cathepsin B activity attenuates SPM-induced neuronal death.
The effect of a cathepsin B inhibitor (CA-074, 10 μM) on SPM-induced (10 μM, 24 h) cell death in control (miR-Fluc) and Atp13a2 knockdown (miR-3 and miR-5) neurons was assayed via TUNEL-based staining. Left, representative confocal images depicting TUNEL-positive neurons; right, box and whisker plots with the quantification of the TUNEL staining. Data are presented as box and whisker plots (line, median; box boundaries, 25th and 75th percentiles) for which individual data points (representing replicates) are shown. n = 3 biologically independent experiments. Analysis by one-way ANOVA with Tukey’s post-hoc correction.
Supplementary information
41586_2020_1968_MOESM1_ESM.pdf
Supplementary Figures Supplementary Figure 1 – Uncropped blots with size marker indications and uncropped autoradiograms. The associated figure panel is indicated above each scan. Supplementary Figure 2 – Flow cytometry gating. Representative dot plot cascade and subsequent gating used for flow cytometry analysis.
Rights and permissions
About this article
Cite this article
van Veen, S., Martin, S., Van den Haute, C. et al. ATP13A2 deficiency disrupts lysosomal polyamine export. Nature 578, 419–424 (2020). https://doi.org/10.1038/s41586-020-1968-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-020-1968-7
This article is cited by
-
Antizyme inhibitor family: biological and translational research implications
Cell Communication and Signaling (2024)
-
Lysosomes as coordinators of cellular catabolism, metabolic signalling and organ physiology
Nature Reviews Molecular Cell Biology (2024)
-
Key genes and convergent pathogenic mechanisms in Parkinson disease
Nature Reviews Neuroscience (2024)
-
A novel ATP13A2 variant causing complicated hereditary spastic paraplegia
Neurological Sciences (2024)
-
Polyamines: their significance for maintaining health and contributing to diseases
Cell Communication and Signaling (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.