Abstract
PIWI-interacting RNAs (piRNAs) of between approximately 24 and 31 nucleotides in length guide PIWI proteins to silence transposons in animal gonads, thereby ensuring fertility1. In the biogenesis of piRNAs, PIWI proteins are first loaded with 5′-monophosphorylated RNA fragments called pre-pre-piRNAs, which then undergo endonucleolytic cleavage to produce pre-piRNAs1,2. Subsequently, the 3′-ends of pre-piRNAs are trimmed by the exonuclease Trimmer (PNLDC1 in mouse)3,4,5,6 and 2′-O-methylated by the methyltransferase Hen1 (HENMT1 in mouse)7,8,9, generating mature piRNAs. It is assumed that the endonuclease Zucchini (MitoPLD in mouse) is a major enzyme catalysing the cleavage of pre-pre-piRNAs into pre-piRNAs10,11,12,13. However, direct evidence for this model is lacking, and how pre-piRNAs are generated remains unclear. Here, to analyse pre-piRNA production, we established a Trimmer-knockout silkworm cell line and derived a cell-free system that faithfully recapitulates Zucchini-mediated cleavage of PIWI-loaded pre-pre-piRNAs. We found that pre-piRNAs are generated by parallel Zucchini-dependent and -independent mechanisms. Cleavage by Zucchini occurs at previously unrecognized consensus motifs on pre-pre-piRNAs, requires the RNA helicase Armitage, and is accompanied by 2′-O-methylation of pre-piRNAs. By contrast, slicing of pre-pre-piRNAs with weak Zucchini motifs is achieved by downstream complementary piRNAs, producing pre-piRNAs without 2′-O-methylation. Regardless of the endonucleolytic mechanism, pre-piRNAs are matured by Trimmer and Hen1. Our findings highlight multiplexed processing of piRNA precursors that supports robust and flexible piRNA biogenesis.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
The sequencing data reported in this paper are publicly available in DDBJ, under the accession number DRA008549. All other data are available from the authors upon reasonable request.
Code availability
All code required for bioinformatics analysis in this paper is available at https://github.com/kshoji-nt/BmZuc_cleavage.
References
Ozata, D. M., Gainetdinov, I., Zoch, A., O’Carroll, D. & Zamore, P. D. PIWI-interacting RNAs: small RNAs with big functions. Nat. Rev. Genet. 20, 89–108 (2019).
Gainetdinov, I., Colpan, C., Arif, A., Cecchini, K. & Zamore, P. D. Single mechanism of biogenesis, initiated and directed by PIWI proteins, explains piRNA production in most animals. Mol. Cell 71, 775–790 (2018).
Izumi, N. et al. Identification and functional analysis of the pre-piRNA 3′ Trimmer in silkworms. Cell 164, 962–973 (2016).
Ding, D. et al. PNLDC1 is essential for piRNA 3′ end trimming and transposon silencing during spermatogenesis in mice. Nat. Commun. 8, 819 (2017).
Zhang, Y. et al. An essential role for PNLDC1 in piRNA 3′ end trimming and male fertility in mice. Cell Res. 27, 1392–1396 (2017).
Nishimura, T. et al. PNLDC1, mouse pre-piRNA trimmer, is required for meiotic and post-meiotic male germ cell development. EMBO Rep. 19, e44957 (2018).
Horwich, M. D. et al. The Drosophila RNA methyltransferase, DmHen1, modifies germline piRNAs and single-stranded siRNAs in RISC. Curr. Biol. 17, 1265–1272 (2007).
Kirino, Y. & Mourelatos, Z. The mouse homolog of HEN1 is a potential methylase for Piwi-interacting RNAs. RNA 13, 1397–1401 (2007).
Saito, K. et al. Pimet, the Drosophila homolog of HEN1, mediates 2′-O-methylation of Piwi-interacting RNAs at their 3′ ends. Genes Dev. 21, 1603–1608 (2007).
Ipsaro, J. J., Haase, A. D., Knott, S. R., Joshua-Tor, L. & Hannon, G. J. The structural biochemistry of Zucchini implicates it as a nuclease in piRNA biogenesis. Nature 491, 279–283 (2012).
Nishimasu, H. et al. Structure and function of Zucchini endoribonuclease in piRNA biogenesis. Nature 491, 284–287 (2012).
Han, B. W., Wang, W., Li, C., Weng, Z. & Zamore, P. D. piRNA-guided transposon cleavage initiates Zucchini-dependent, phased piRNA production. Science 348, 817–821 (2015).
Mohn, F., Handler, D. & Brennecke, J. piRNA-guided slicing specifies transcripts for Zucchini-dependent, phased piRNA biogenesis. Science 348, 812–817 (2015).
Brennecke, J. et al. Discrete small RNA-generating loci as master regulators of transposon activity in Drosophila. Cell 128, 1089–1103 (2007).
Gunawardane, L. S. et al. A slicer-mediated mechanism for repeat-associated siRNA 5′ end formation in Drosophila. Science 315, 1587–1590 (2007).
De Fazio, S. et al. The endonuclease activity of Mili fuels piRNA amplification that silences LINE1 elements. Nature 480, 259–263 (2011).
Reuter, M. et al. Miwi catalysis is required for piRNA amplification-independent LINE1 transposon silencing. Nature 480, 264–267 (2011).
Aravin, A. A. et al. A piRNA pathway primed by individual transposons is linked to de novo DNA methylation in mice. Mol. Cell 31, 785–799 (2008).
Kuramochi-Miyagawa, S. et al. DNA methylation of retrotransposon genes is regulated by Piwi family members MILI and MIWI2 in murine fetal testes. Genes Dev. 22, 908–917 (2008).
Sienski, G., Dönertas, D. & Brennecke, J. Transcriptional silencing of transposons by Piwi and maelstrom and its impact on chromatin state and gene expression. Cell 151, 964–980 (2012).
Nishida, K. M. et al. Hierarchical roles of mitochondrial Papi and Zucchini in Bombyx germline piRNA biogenesis. Nature 555, 260–264 (2018).
Ge, D. T. et al. The RNA-binding ATPase, Armitage, couples piRNA amplification in Nuage to phased piRNA production on mitochondria. Mol. Cell 74, 982–995 (2019).
Hayashi, R. et al. Genetic and mechanistic diversity of piRNA 3′-end formation. Nature 539, 588–592 (2016).
Kawaoka, S., Izumi, N., Katsuma, S. & Tomari, Y. 3′ end formation of PIWI-interacting RNAs in vitro. Mol. Cell 43, 1015–1022 (2011).
Vourekas, A. et al. The RNA helicase MOV10L1 binds piRNA precursors to initiate piRNA processing. Genes Dev. 29, 617–629 (2015).
Pandey, R. R. et al. Recruitment of Armitage and Yb to a transcript triggers its phased processing into primary piRNAs in Drosophila ovaries. PLoS Genet. 13, e1006956 (2017).
Handler, D. et al. The genetic makeup of the Drosophila piRNA pathway. Mol. Cell 50, 762–777 (2013).
Muerdter, F. et al. A genome-wide RNAi screen draws a genetic framework for transposon control and primary piRNA biogenesis in Drosophila. Mol. Cell 50, 736–748 (2013).
Shiromoto, Y. et al. GPAT2, a mitochondrial outer membrane protein, in piRNA biogenesis in germline stem cells. RNA 19, 803–810 (2013).
Vagin, V. V. et al. Minotaur is critical for primary piRNA biogenesis. RNA 19, 1064–1077 (2013).
Shoji, K., Suzuki, Y., Sugano, S., Shimada, T. & Katsuma, S. Artificial “ping-pong” cascade of PIWI-interacting RNA in silkworm cells. RNA 23, 86–97 (2017).
Böttcher, R. et al. Efficient chromosomal gene modification with CRISPR/Cas9 and PCR-based homologous recombination donors in cultured Drosophila cells. Nucleic Acids Res. 42, e89 (2014).
Zhu, L., Mon, H., Xu, J., Lee, J. M. & Kusakabe, T. CRISPR–Cas9-mediated knockout of factors in non-homologous end joining pathway enhances gene targeting in silkworm cells. Sci. Rep. 5, 18103 (2015).
Patil, A. A. et al. Characterization of Armitage and Yb containing granules and their relationship to nuage in ovary-derived cultured silkworm cell. Biochem. Biophys. Res. Commun. 490, 134–140 (2017).
Haley, B., Tang, G. & Zamore, P. D. In vitro analysis of RNA interference in Drosophila melanogaster. Methods 30, 330–336 (2003).
Pall, G. S. & Hamilton, A. J. Improved Northern blot method for enhanced detection of small RNA. Nat. Protoc. 3, 1077–1084 (2008).
Fu, Y., Wu, P. H., Beane, T., Zamore, P. D. & Weng, Z. Elimination of PCR duplicates in RNA-seq and small RNA-seq using unique molecular identifiers. BMC Genomics 19, 531 (2018).
Kim, H. et al. Bias-minimized quantification of microRNA reveals widespread alternative processing and 3′ end modification. Nucleic Acids Res. 47, 2630–2640 (2019).
Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet Journal. 17, 10–12 (2011).
Langmead, B., Trapnell, C., Pop, M. & Salzberg, S. L. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol. 10, R25 (2009).
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Quinlan, A. R. & Hall, I. M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26, 841–842 (2010).
Kawamoto, M. et al. High-quality genome assembly of the silkworm, Bombyx mori. Insect Biochem. Mol. Biol. 107, 53–62 (2019).
Kamminga, L. M. et al. Hen1 is required for oocyte development and piRNA stability in zebrafish. EMBO J. 29, 3688–3700 (2010).
Simon, B. et al. Recognition of 2′-O-methylated 3′-end of piRNA by the PAZ domain of a Piwi protein. Structure 19, 172–180 (2011).
Tian, Y., Simanshu, D. K., Ma, J. B. & Patel, D. J. Structural basis for piRNA 2′-O-methylated 3′-end recognition by Piwi PAZ (Piwi/Argonaute/Zwille) domains. Proc. Natl Acad. Sci. USA 108, 903–910 (2011).
Lim, S. L. et al. HENMT1 and piRNA stability are required for adult male germ cell transposon repression and to define the spermatogenic program in the mouse. PLoS Genet. 11, e1005620 (2015).
Homolka, D. et al. PIWI slicing and RNA elements in precursors instruct directional primary piRNA biogenesis. Cell Rep. 12, 418–428 (2015).
Yang, Z. et al. PIWI slicing and EXD1 drive biogenesis of nuclear piRNAs from cytosolic targets of the mouse piRNA pathway. Mol. Cell 61, 138–152 (2016).
Acknowledgements
We thank T. Kusakabe and T. Tatsuke for providing BmArmi expression vectors, K. Förstemann for providing hCas9 and sgRNA expression vectors, T. Kiuchi for sharing unpublished data and helpful discussion, and K. Kiyokawa and T. Horiuchi for technical assistance. A part of Illumina sequencing was performed in the Vincent J. Coates Genomics Sequencing Laboratory at UC Berkeley, supported by NIH S10 OD018174 Instrumentation Grant. We also thank Life Science Editors for editorial assistance, and P. Zamore and members of the Tomari laboratory for critical comments on the manuscript. This work was in part supported by a Grant-in-Aids for Scientific Research on Innovative Areas (grant 26113007 to Y.T.) from the Ministry of Education, Culture, Sports, Science and Technology in Japan and a Grant-in-Aid for Scientific Research (S) (grant 18H05271 to Y.T.), Grant-in-Aid for Scientific Research (B) (grant 16KT0064 to Y.S. and S.K.), Grant-in-Aid for Scientific Research on Innovative Areas (grant 17H06431 to S.K.), Grant-in-Aid for Young Scientists (B) (grant 17K17673 to N.I.), Grant-in-Aid for Scientific Research (C) (grant 19K06484 to N.I.), and a Grant-in-Aid for JSPS Fellows (grant 17J02408 to K.S.).
Author information
Authors and Affiliations
Contributions
N.I., K.S. and Y.T. conceived and designed the experiments and wrote the manuscript. N.I. performed biochemical experiments. K.S. performed bioinformatics analyses. Y.S. and S.K. supervised the bioinformatics analyses. Y.T. supervised the research. All the authors discussed the results and approved the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature thanks René Ketting and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 The current model for the piRNA biogenesis initiated by the piRNA-guided PIWI-catalysed slicing in animal germ cells.
The ping-pong cycle produces pairs of piRNAs (‘initiator’ and ‘responder’ piRNAs) that show 10-nt overlapping at their 5′ ends as well as multiple ‘trailing’ piRNAs downstream of the responder piRNAs (top)1. The ping-pong cycle is initiated by the slicing of a precursor transcript by an initiator piRNA-loaded PIWI protein. The PIWI-cleaved 5′-monophosphorylated fragment is handed over to a corresponding PIWI protein as a pre-pre-piRNA. Then, the PIWI-loaded pre-pre-piRNA is endonucleolytically cleaved at a downstream position1,2,12,13. The resultant 5′ cleavage fragment, called a pre-piRNA, is further trimmed by the 3′-to-5′ exonuclease Trimmer (PNLDC1 in mouse)3,4,5,6 to the mature length, 2′-O-methylated by the methyltransferase Hen1 (HENMT1 in mouse)7,8,9, and becomes a responder piRNA (left pathway). Hen1-mediated 2′-O-methylation protects mature piRNAs from degradation and tightens their binding to PIWI proteins7,44,45,46,47. The 3′ endonucleolytic cleavage fragment of the pre-pre-piRNA is loaded into a next PIWI protein as a new pre-pre-piRNA and endonucleolytically cleaved again at a downstream position, producing a new PIWI-loaded pre-piRNA1,2,12,13. This pre-piRNA is then processed by Trimmer and Hen1 at the 3′ end into a mature trailing piRNA (middle pathway). The 3′ cleavage fragment of the second endonucleolytic cleavage is also loaded into a next PIWI protein, serving as a new pre-pre-piRNAs. As a result, a series of trailing piRNAs are consecutively produced downstream of the responder piRNA (right pathway)1,2,12,13,48,49. These two piRNA biogenesis pathways lead to target-dependent amplification of piRNAs (via the ping-pong cycle) and expansion of piRNA sequences (via trailing piRNA production).
Extended Data Fig. 2 Generation and characterization of Trimmer-knockout cells.
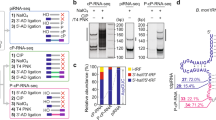
a, Schematic representation of the domain structure of Trimmer and the position of the sgRNA target site for CRISPR–Cas9. b, Genomic PCR of a region including the sgRNA target site. In addition to the main PCR product (ii), two additional PCR products (i and iii) were detected only in Tri-KO#4 cells. Detailed genome sequences are shown in c. c, Genome sequences around the sgRNA target site in naive or Tri-KO#4 BmN4 cells. Genomic sequencing revealed various mutations at the sgRNA target site, suggesting a polyploid nature of the trimmer locus and/or imperfect cell cloning. d, Western blot analysis of Trimmer in two different Tri-KO cell lines (#4 and #6). Tri-KO line #4 was used in this study. e, Western blot analysis of whole-cell lysate from naive or Tri-KO#4 BmN4 cells. f, In vitro trimming assay for Siwi-loaded 1U50 RNA using 1,000g ppt. from naive or two different Tri-KO cell lines. ppt., pellet. g, SYBR Gold staining of total RNAs from naive or three different Tri-KO cell lines (#3, #4 and #6). h, Total RNAs extracted from Tri-KO #4 cells overexpressing wild-type Trimmer (WT) or its catalytic mutant E30A (EA) were 5′ radiolabelled and detected by phosphor imaging. Mock indicates transfection of a control plasmid. Trimmer expression was analysed by western blotting (upper). i, Length distribution of small RNAs mapped to 3,236 piRNA loci in NaIO4-treated small RNA library from naive or Tri-KO BmN4 cells. j, Relative fraction of 2′-O-methylated Tri-KO small RNAs in each length. k, Peak length distribution of piRNAs mapped to 3,236 piRNA loci in the NaIO4-treated library from naive BmN4 cells. l, Changes by the depletion of BmZuc in the length distribution of Type-N (lower) or Type-E (upper) NaIO4-treated small RNAs in Tri-KO cells.
Extended Data Fig. 3 BmZuc is required to produce type-E pre-piRNAs for both Siwi and BmAgo3, whereas trailing piRNA production is largely restricted to Siwi.
a, Scatter plot showing normalized piRNA abundance co-immunoprecipitated with Siwi or BmAgo3 from naive BmN4 cells for each piRNA loci. Green dots, Siwi-dominant piRNA loci (n = 1,946); purple dots, BmAgo3-dominant piRNA loci (n = 1,259). b, Peak length frequency of Tri-KO small RNAs for Siwi-dominant (left) or BmAgo3-dominant (right) piRNA loci. c, Length distribution of Tri-KO small RNAs bearing the peak length of 35 or 36 nt (type-E) for Siwi-dominant (upper) or BmAgo3-dominant (lower) piRNA loci. BmZuc knockdown abolished small RNAs with the peak lengths. Zn denotes the z score at position n (c, e). d, Siwi-dominant type-E pre-piRNAs show a stronger +1U preference than BmAgo3-dominant ones. e, Siwi-dominant type-E pre-piRNAs show a greater tendency to have downstream trailing piRNAs than BmAgo3-dominant ones. f, The 5′ ends of piRNAs mapped to 20−100 nt downstream of type-N (top) or type-E (bottom) Tri-KO small RNAs were mapped on the antisense strand, separately for Siwi-dominant (left) and BmAgo3-dominant (right) piRNA loci. Type-N piRNAs have more antisense piRNAs at ~41−52 nt from the 5′ ends than type-E piRNAs, regardless of which PIWI protein they bind (two-sided Wilcoxon signed rank test, n = 12). g, Four representative type-E (piRNA-1528 and 66) and type-N (piRNA-2986 and 304) piRNA loci and their downstream genomic regions were mapped with the 5′ ends of sense (grey) and antisense (red) piRNAs. h, Distribution of type-E and type-N piRNAs mapped to a transposon called MER85. i, An example of mixed modes of pre-piRNA production. Pre-pre-piRNA-1249 contains a BmZuc cleavage site and a slicing site by an antisense piRNA-loaded PIWI protein. The ~35 nt BmZuc cleavage product, but not the 59 nt slicing product, is 2′-O-methylated. An unmethylated ~75 nt fragment, which is possibly produced by another antisense piRNA-guided slicing, locates in an unannotated genomic region and cannot be assigned.
Extended Data Fig. 4 BmZuc-mediated cleavage of Siwi-loaded pre-pre-piRNAs in vitro.
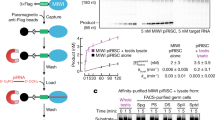
a, Detailed protocol for in vitro recapitulation of BmZuc-mediated cleavage of Siwi-loaded pre-pre-piRNAs. b, Siwi-loaded 28–80U RNA was incubated with Tri-KO 1,000g ppt. overexpressing BmZuc WT or HN, with or without ATP and the ATP-regeneration system. Mock indicates transfection of a control plasmid. c, Siwi-loaded 111750 RNA was incubated with Tri-KO 1,000g ppt. depleted of the indicated protein by RNAi. Mock indicates RNAi against Renilla luciferase (in c−e). d, Confocal images of BmN4 cells stably expressing GFP-tagged BmArmi in the presence or absence of BmGasz (scale bars, 10 μm). e, Quantitative real-time PCR analysis of BmArmi and BmGPAT1. Tri-KO cells were depleted of BmGPAT1 or BmGasz by RNAi, and the mRNA levels for BmArmi or BmGPAT1 were analysed by real-time PCR. The graph shows the average of two independent experiments. f, Siwi-loaded 1U50 RNA was incubated with Tri-KO 1,000g ppt. overexpressing BmZuc and BmArmi, or BmZuc HN. Mock indicates transfection of a control plasmid. 1U50 RNA was cleaved multiple sites within a region that is devoid of U in a manner dependent on the BmZuc activity. ppt., pellet.
Extended Data Fig. 5 Calculation of BmZuc scores for Siwi or BmAgo3 based on the randomized sequence library analysis.
a, Peak length distribution of small RNAs derived from the randomized sequence library. b, RNA substrates used in c. The top 6 nucleotides in the BmZuc motif are shown in colour and their mutations are shown in black. c, Siwi-loaded 111750-derived RNAs were incubated with Tri-KO 1,000g ppt. overexpressing BmZuc and BmArmi, or BmZuc(HN). Each gel image was adjusted to equalize the loading signal. ppt., pellet. d, Schematic representation of the randomized sequence library analysis for Siwi- or BmAgo3-loaded pre-piRNAs cleaved by BmZuc. e, Peak length distribution of Siwi- or BmAgo3-bound 2′-O-methylated small RNAs derived from the corresponding randomized sequence library. For Siwi immunoprecipitation, the same plasmid library as in a was used. f, Nucleotide composition around the 3′ ends of mature piRNAs in naive BmN4 cells (right) or type-E pre-piRNAs in Tri-KO cells (left), separately analysed for Siwi-dominant (top) and BmAgo3-dominant (bottom) piRNA loci. The 6 nucleotides in the BmZuc motif are highlighted. g, Schematic explanation for the weighted BmZuc motif (top) and the calculation of the BmZuc score in the 17-nt sliding window analysis (bottom). h, Similarity scores with the weighted BmZuc motif (BmZuc score) for Siwi were calculated for 111750 RNA and their mutant sequences in sliding windows and plotted as in c. i, Box plots show the maximum similarity scores with the weighted BmZuc motif for Siwi or BmAgo3 within the positions of 19−45 nt of Siwi-dominant (top) or BmAgo3-dominant (bottom) type-N or type-E piRNA loci or the shuffled control sequences (a pool of 3,236 species of 27-nt scrambled sequences that have the average nucleotide composition of the silkworm genome). Type-E piRNA loci have significantly higher BmZuc scores than the shuffled control sequences for both Siwi- and BmAgo3-dominant piRNAs (Mann–Whitney U test). Centre line, median; box limits, upper and lower quartiles; whiskers, 1.5 × interquartile range; points, outliers.
Extended Data Fig. 6 Nucleotide preference around the cleavage site by mouse MitoPLD.
a, Peak length distribution of 2′-O-methylated small RNAs in wild-type (WT) or Pnldc1−/− mouse secondary spermatocytes. Data are from ref. 2. b, Nucleotide composition around the 3′ end of small RNAs in WT (left) or Pnldc1−/− mice. The 3′ ends of pre-piRNAs in Pnldc1−/−mice exhibit strong +1U bias. In addition, −9A and −3C preferences, which are similar to the BmZuc motif (Fig. 4e), are observed in fully elongated 37–42 nt pre-piRNAs in Pnldc1−/−mice (right). Data are from ref. 2.
Supplementary information
Supplementary Information
This file contains Supplementary Discussion, Supplementary Notes 1, 2, and Supplementary References.
Supplementary Figure 1
This file contains uncropped blot/gel images used in the figures and extended data figures.
Rights and permissions
About this article
Cite this article
Izumi, N., Shoji, K., Suzuki, Y. et al. Zucchini consensus motifs determine the mechanism of pre-piRNA production. Nature 578, 311–316 (2020). https://doi.org/10.1038/s41586-020-1966-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-020-1966-9
This article is cited by
-
The dual role of Spn-E in supporting heterotypic ping-pong piRNA amplification in silkworms
EMBO Reports (2024)
-
The epigenetic regulatory mechanism of PIWI/piRNAs in human cancers
Molecular Cancer (2023)
-
Emerging roles and functional mechanisms of PIWI-interacting RNAs
Nature Reviews Molecular Cell Biology (2023)
-
piRNA processing by a trimeric Schlafen-domain nuclease
Nature (2023)
-
PiRNA in Cardiovascular Disease: Focus on Cardiac Remodeling and Cardiac Protection
Journal of Cardiovascular Translational Research (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.